Abstract
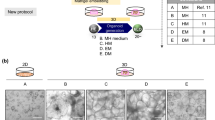
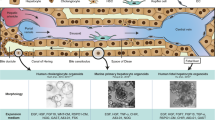
Efficient isolation and culture of liver organoids are critical for studying liver fibrosis, liver regeneration and drug toxicity and screening. However, preserving mature hepatobiliary characteristics and concurrently incorporating fibrosis-producing hepatic stellate cells (HSCs) remains a significant challenge, often hindering the large-scale production of organoids capable of replicating key liver functions. Here, we report a robust 3D organoid culture system that enables simultaneous isolation and long-term propagation of primary hepatocytes, cholangiocytes, and HSCs from a single source of mouse liver tissue. By supplementing the Hep-Med with Notch signaling inhibitor and dexamethasone, we achieved sustained organoid maturity, including stable albumin production, metabolic activity, and liver-specific gene expression, over multiple passages in culture. Quiescent HSCs within the system retained lipid droplets and could be activated into a myofibroblast-like phenotype, also called activated HSCs, via TGFβ stimulation. Activated HSCs impaired the proliferation and stemness, and induced epithelial-mesenchymal transition (EMT) of Hep-Orgs and Cho-Orgs, enabling in vitro liver fibrosis modeling. Optimized for minimal tissue input, this platform maximizes tissue utilization efficiency while preserving the liver’s heterogeneous cellular architecture. Its versatility supports diverse applications in liver disease modeling, drug discovery, and regenerative medicine.
Similar content being viewed by others
Data availability
The data underlying this article are available in GEO (Gene Expression Omnibus) at (https://www.ncbi.nlm.nih.gov/geo) and the accession number is GSE308085.
References
Asrani, S. K., Devarbhavi, H., Eaton, J. & Kamath, P. S. Burden of liver diseases in the world. J. Hepatol. 70, 151–171. https://doi.org/10.1016/j.jhep.2018.09.014 (2019).
Li, J. et al. Global burden of gallbladder and biliary diseases: A systematic analysis for the Global Burden of Disease Study 2019. J. Gastroenterol. Hepatol. 37, 1389–1399. https://doi.org/10.1111/jgh.15859 (2022).
Shao, W. et al. Advances in liver organoids: replicating hepatic complexity for toxicity assessment and disease modeling. Stem Cell. Res. Ther. 16, 27. https://doi.org/10.1186/s13287-025-04139-2 (2025).
Akbari, S. et al. Long-term culture of endoderm-derived hepatic organoids for disease modeling. Stem Cell. Rep. 13, 627–641. https://doi.org/10.1016/j.stemcr.2019.08.007 (2019). Robust.
Schutgens, F. & Clevers, H. Human organoids: tools for understanding biology and treating diseases. Annu. Rev. Pathol. 15, 211–234. https://doi.org/10.1146/annurev-pathmechdis-012419-032611 (2020).
Nuciforo, S. & Heim, M. H. Organoids to model liver disease. JHEP Rep. 3, 100198. https://doi.org/10.1016/j.jhepr.2020.100198 (2021).
Fatehullah, A., Tan, S. H. & Barker, N. Organoids as an in vitro model of human development and disease. Nat. Cell. Biol. 18, 246–254. https://doi.org/10.1038/ncb3312 (2016).
Huch, M. et al. In vitro expansion of single Lgr5 + liver stem cells induced by Wnt-driven regeneration. Nature 494, 247–250. https://doi.org/10.1038/nature11826 (2013).
Broutier, L. et al. Culture and establishment of self-renewing human and mouse adult liver and pancreas 3D organoids and their genetic manipulation. Nat. Protoc. 11, 1724–1743. https://doi.org/10.1038/nprot.2016.097 (2016).
Hu, H. et al. Long-term expansion of functional mouse and human hepatocytes as 3D organoids. Cell 175, 1591–1606. https://doi.org/10.1016/j.cell.2018.11.013 (2018).
Peng, W. C. et al. Inflammatory cytokine TNFα promotes the long-term expansion of primary hepatocytes in 3D culture. Cell 175, 1607–1619. https://doi.org/10.1016/j.cell.2018.11.012 (2018).
Hendriks, D., Artegiani, B., Hu, H., Chuva de Sousa Lopes, S. & Clevers, H. Establishment of human fetal hepatocyte organoids and CRISPR-Cas9-based gene knockin and knockout in organoid cultures from human liver. Nat. Protoc. 16, 182–217. https://doi.org/10.1038/s41596-020-00411-2 (2021).
Zhou, B. et al. Notch signaling pathway: architecture, disease, and therapeutics. Signal. Transduct. Target. Ther. 7, 95. https://doi.org/10.1038/s41392-022-00934-y (2022).
Felszeghy, S., Suomalainen, M. & Thesleff, I. Notch signalling is required for the survival of epithelial stem cells in the continuously growing mouse incisor. Differentiation 80, 241–248. https://doi.org/10.1016/j.diff.2010.06.004 (2010).
Ke, Z. et al. Dynamic expression characteristics of Notch signal in bone marrow-derived mesenchymal stem cells during the process of differentiation into hepatocytes. Tissue Cell. 45, 95–100. https://doi.org/10.1016/j.tice.2012.09.008 (2013).
Benne, C. et al. Notch increases T/NK potential of human hematopoietic progenitors and inhibits B cell differentiation at a pro-B stage. Stem Cells. 27, 1676–1685. https://doi.org/10.1002/stem.94 (2009).
Li, H. J., Kapoor, A., Giel-Moloney, M., Rindi, G. & Leiter, A. B. Notch signaling differentially regulates the cell fate of early endocrine precursor cells and their maturing descendants in the mouse pancreas and intestine. Dev. Biol. 371, 156–169. https://doi.org/10.1016/j.ydbio.2012.08.023 (2012).
Aguirre, A., Rubio, M. E. & Gallo, V. Notch and EGFR pathway interaction regulates neural stem cell number and self-renewal. Nature 467, 323–327. https://doi.org/10.1038/nature09347 (2010).
Egger, B., Gold, K. S. & Brand, A. H. Notch regulates the switch from symmetric to asymmetric neural stem cell division in the Drosophila optic lobe. Development 137, 2981–2987. https://doi.org/10.1242/dev.051250 (2010).
Wang, T. et al. Notch inhibition promotes fetal liver stem/progenitor cells differentiation into hepatocytes via the inhibition of HNF-1β. Cell. Tissue Res. 357, 173–184. https://doi.org/10.1007/s00441-014-1825-9 (2014).
Wang, X. et al. S-Nitrosylation of NOTCH1 Regulates Mesenchymal Stem Cells Differentiation Into Hepatocyte-Like Cells by Inhibiting Notch Signalling Pathway. J. Cell. Mol. Med. 28, e70274. https://doi.org/10.1111/jcmm.70274 (2024).
Lambiotte, M., Vorbrodt, A. & Benedetti, E. L. Expression of differentiation of rat foetal hepatocytes in cellular culture under the action of glucocorticoids: appearance of bile canaliculi. Cell. Differ. 2, 43–53 (1973).
Laishes, B. A. & Williams, G. M. Conditions affecting primary cell cultures of functional adult rat hepatocytes. II. Dexamethasone enhanced longevity and maintenance of morphology. Vitro 12, 821–832 (1976).
Jefferson, D. M., Reid, L. M., Giambrone, M. A., Shafritz, D. A. & Zern, M. A. Effects of dexamethasone on albumin and collagen gene expression in primary cultures of adult rat hepatocytes. Hepatology 5, 14–20 (1985).
Oh, H. Y. et al. Dexamethasone protects primary cultured hepatocytes from death receptor-mediated apoptosis by upregulation of cFLIP. Cell. Death Differ. 13, 512–523 (2006).
Kim, H. M., Kim, Y., Kim, Y., Kim, Y. J. & Ko, K. S. Organoid establishment of long-term culture using primary mouse hepatocytes and evaluation of liver function. Prev. Nutr. Food Sci. 28, 360–369. https://doi.org/10.3746/pnf.2023.28.3.360 (2023).
Ben-Moshe, S. & Itzkovitz, S. Spatial heterogeneity in the mammalian liver. Nat. Rev. Gastroenterol. Hepatol. 16, 395–410. https://doi.org/10.1038/s41575-019-0134-x (2019).
Reza, H. A. et al. Multi-zonal liver organoids from human pluripotent stem cells. Nature 641, 1258–1267. https://doi.org/10.1038/s41586-025-08850-1 (2025).
He, L. & Deng, C. Recent advances in organotypic tissue slice cultures for anticancer drug development. Int. J. Biol. Sci. 18, 5885–5896. https://doi.org/10.7150/ijbs.78997 (2022).
He, L. et al. CDK4/6 Inhibition Induces CD8 + T cell antitumor immunity via MIF-induced functional orchestration of tumor-associated macrophages. Adv. Sci. (Weinh). https://doi.org/10.1002/advs.202511330 (2025).
Dowbaj, A. M. et al. Mouse liver assembloids model periportal architecture and biliary fibrosis. Nature 644, 473–482. https://doi.org/10.1038/s41586-025-09183-9 (2025).
Friedman, S. L. Mechanisms of hepatic fibrogenesis. Gastroenterology 134, 1655–1669. https://doi.org/10.1053/j.gastro.2008.03.003 (2008).
Hernandez-Gea, V. & Friedman, S. L. Pathogenesis of liver fibrosis. Annu. Rev. Pathol. 6, 425–456. https://doi.org/10.1146/annurev-pathol-011110-130246 (2011).
Tsuchida, T. & Friedman, S. L. Mechanisms of hepatic stellate cell activation. Nat. Rev. Gastroenterol. Hepatol. 14, 397–411. https://doi.org/10.1038/nrgastro.2017.38 (2017).
Mederacke, I. et al. Fate tracing reveals hepatic stellate cells as dominant contributors to liver fibrosis independent of its aetiology. Nat. Commun. 4, 2823. https://doi.org/10.1038/ncomms3823 (2013).
Mederacke, I., Dapito, D. H., Affò, S., Uchinami, H. & Schwabe, R. F. High-yield and high-purity isolation of hepatic stellate cells from normal and fibrotic mouse livers. Nat. Protoc. 10, 305–315. https://doi.org/10.1038/nprot.2015.017 (2015).
Friedman, S. L. Liver fibrosis -- from bench to bedside. J. Hepatol. 38 (Suppl 1), S38–S53 (2003).
Marrone, G., Shah, V. H. & Gracia-Sancho, J. Sinusoidal communication in liver fibrosis and regeneration. J. Hepatol. 65, 608–617. https://doi.org/10.1016/j.jhep.2016.04.018 (2016).
Kisseleva, T. & Brenner, D. Molecular and cellular mechanisms of liver fibrosis and its regression. Nat. Rev. Gastroenterol. Hepatol. 18, 151–166. https://doi.org/10.1038/s41575-020-00372-7 (2020).
Iwaisako, K. et al. Origin of myofibroblasts in the fibrotic liver in mice. Proc. Natl. Acad. Sci. U S A. 111, E3297–E3305. https://doi.org/10.1073/pnas.1400062111 (2014).
Ebrahimkhani, M. R. et al. Stimulating healthy tissue regeneration by targeting the 5-HT₂B receptor in chronic liver disease. Nat. Med. 17, 1668–1673. https://doi.org/10.1038/nm.2490 (2011).
Wallace, K., Burt, A. D. & Wright, M. C. Liver fibrosis. Biochem. J. 411, 1–18. https://doi.org/10.1042/BJ20071570 (2008).
Friedman, S. L. Hepatic stellate cells: protean, multifunctional, and enigmatic cells of the liver. Physiol. Rev. 88, 125–172. https://doi.org/10.1152/physrev.00013.2007 (2008).
Babu, R. O. et al. Beta-amyloid deposition around hepatic bile ducts is a novel pathobiological and diagnostic feature of biliary atresia. J. Hepatol. 73, 1391–1403. https://doi.org/10.1016/j.jhep.2020.06.012 (2020).
Acknowledgements
We are sincerely grateful to Professor Ye Hua from the Oxford Suzhou Centre for contributing to our major revision on the expertise of organoid-related technologies.
Funding
This work was supported by Macau Science and Technology Development Fund (FDCT No. 0086/2022/A, 0097/2022/A2 and 0011/2023/AKP).
Author information
Authors and Affiliations
Contributions
Conceptualization, Yingyu Liang, Yongqin Ye, Yan Chen and Paul Tam; Data curation, Yingyu Liang; Funding acquisition, Yan Chen and Paul Tam; Methodology, Yingyu Liang, Yongqin Ye, Hua Xie, Yan Chen and Paul Tam; Project administration, Vincent Lui, Yan Chen and Paul Tam; Software, Yingyu Liang and Yongqin Ye; Validation, Yingyu Liang and Yongqin Ye; Writing–original draft, Yingyu Liang and Yongqin Ye; Writing–review & editing, Yan Chen and Paul Tam.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liang, Y., Ye, Y., Xie, H. et al. A robust mouse liver organoid platform enables sustained multicellular maturation and fibrosis modeling from a single tissue sample. Sci Rep (2026). https://doi.org/10.1038/s41598-026-42990-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-026-42990-2