Abstract
Breast ductal carcinoma in situ is a common non-invasive clinical finding that can progress to invasive breast cancer (IBC). Spatial proteomics can provide an additional dimension to our understanding of this disease and its capacity to progress. A subset (n = 103 patients) of a previously established cohort of primary DCIS specimens with known clinical outcomes was analyzed using a multiplexed proteomic platform (Nanostring GeoMx) for simultaneous quantitative measurement of 53 antigens. 1262 spatially distinct regions of interest (ROIs) (1226 ROIs after filtering) were collected, including inside DCIS epithelium, adjacent stroma, co-existing benign breast epithelium, and biopsy sites. We identified two predominant subgroups of DCIS, ER high/HER2 low and ER low/HER2 high. Levels of tumor associated proteins varied between benign and DCIS, between ER + and ER- patients, and between different regions within the DCIS epithelium. In addition, we identified several immune-related antigens (CD127, CD8, and PD-L2) within the DCIS epithelium that are associated with invasive progression. Comparison of antigen levels in matched ipsilateral breast events (both DCIS recurrences and IBC) demonstrates an effect of hormonal therapy on the phenotype of subsequent cancers. This study adds a spatially resolved proteomic dimension to our understanding of DCIS, its microenvironment, and its propensity to progress to IBC.
Similar content being viewed by others
Introduction
Ductal carcinoma in situ (DCIS) of the breast is a common diagnosis typically detected by its characteristic calcifications on mammography. This non-invasive epithelial proliferation does have the potential to become invasive and thus is typically removed surgically with adjuvant radiotherapy delivered to improve local control of the disease. However, many of these lesions do not progress1,2,3 and this has prompted a series of clinical trials of active monitoring, particularly for DCIS that has a low or intermediate histologic grade4,5,6. Complementary to these trials are efforts to identify DCIS that are most or least likely to progress, either with or without therapy7,8,9,10,11. We previously reported an expression signature derived from RNA sequencing of the epithelium and nearby stroma containing genes expressed from both types of cells that discriminates recurrent (either DCIS or invasive disease) from non-recurrent patients12. We have also reported on a signature derived from a mass spectrometry-based metal tagged multiplexed antibody approach (MIBI) on a small subset of samples, indicating that spatial features are important in identifying high risk DCIS13.
Spatial detection methods are rapidly advancing and becoming more widely available for investigational purposes. While single-cell RNA techniques measure a large fraction of the genome, protein detection methods are constrained by the necessity of using a series of specific affinity reagents, i.e., antibodies. In this study, we analyzed a series of patients on which we established our RNA expression signature using a commercial multiplexed spatial protein detection platform, Nanostring GeoMx, at a resolution that fills an important gap between bulk sequencing and single cell methods. Moreover, evaluation of protein expression provides data that are more proximal to tumor biology than transcriptomic technologies. The study had two aims: (1) determine how protein expression differs according to distinct spatial locations within the DCIS microenvironment and (2) determine whether DCIS protein expression is associated with disease progression.
Methods
Clinical materials
The study was performed on a subset of samples from a retrospective cohort collected through a multi-center study activated at 12 participating TBCRC (Translational Breast Cancer Consortium, TBCRC038) sites, which identified women treated for ductal carcinoma in situ (DCIS) at one of the enrolling institutions between 01/01/1998 and 02/29/2016 with follow-up to determine whether the DCIS recurred or progressed to invasive cancer. This cohort was described in detail in our prior study where we also conducted RNA expression profiling and low pass whole genome sequencing on DCIS from these patients12. The TBCRC and the Department of Defense (DOD) approved this study for the collection of archival tissues. Duke served as the initiating and central site for all data, samples, assays, and analyses. The study was approved by the Duke Health Institutional Review Board (Protocol ID: Pro00068646) as well as the IRB at each participating institution. Individual sites reviewed medical records to identify patients eligible for the study. All methods were performed in accordance with the relevant guidelines and regulations regarding human subjects’ research. Due to the retrospective nature of the study, the Duke Health Institutional Review Board waived the need of obtaining informed consent.
We selected three subsets of the patients from the TBCRC study attempting to balance these three sets only by age at diagnosis and the age of the tissue specimen: No recurrence (DCIS with No Recurrence; NR, n = 35) had no evidence of DCIS recurrence or invasive progression during the follow-up interval; DCIS recurrence (DCIS with DCIS Recurrence; DR, n = 35) experienced a recurrent DCIS in the ipsilateral breast > 1 year after the initial DCIS diagnosis. Invasive recurrence (DCIS with Invasive Recurrence, IR, n = 33) progressed to ipsilateral invasive breast cancer > 1 year after the initial DCIS diagnosis. For a subset of the DR and IR patients, we also analyzed specimens containing the ipsilateral breast event; DR, DCIS to DCIS, n = 20, and IR, DCIS to IBC, n = 18. The median follow-up time for the NR cohort was 7.83 years, and the median times to recurrence for the DR and IR cohorts were 3.83 and 4.83 years, respectively. The event times for all three cohorts are shown in Supplementary Fig. 1. All cases had clear margins after definitive surgery. The selection of the cases for these subsets was made without taking into account clinical or pathologic parameters.
We determined the ER and HER2 status for each patient as follows. For ER status, we prioritized the clinical results (102 patients). For one patient, the clinical result was unavailable and for that case, we bioinformatically inferred ER status based on RNA expression levels from RNAseq data derived from the same clinical specimen12. We assigned HER2 status (103 patients) as positive or negative based on a defined threshold of RNA expression levels that correlated closely with whole genome sequencing copy number data available on most cases.
Spatially resolved multiplexed antigen quantification
The NanoString GeoMx Digital Spatial Profiler (DSP) enables spatially resolved protein expression. Detailed methods have previously been described14. For this experiment, 5 μm serial sections derived from FFPE tissue blocks were collected. One section was stained with H&E and one unstained section was mounted on a positively charged histology slide for use in the DSP. Slides were deparaffinized followed by antigen retrieval with citrate buffer (pH6.0) at 100 °C for 15 min. Tissues were then blocked with buffer W for 1 h at RT, and primary antibody incubation overnight at 4 °C for protein core panel and protein module panels as well as morphology markers was performed. After protein incubation, tissues were fixed with 4%PFA buffer and were then stained with nuclear dye (Syto13). Slides were then loaded onto DSP for region of interest (ROI) collection or stored in 1x TBST at 4 °C for up to two weeks.
Three fluorescent markers were included in each assay in order to identify cells and cell types to facilitate ROI selection: anti-CD45 for immune cells, anti-pan-cytokeratin (PanCk) for epithelial cells, and SYTO 13 a DNA binding dye to identify all nucleated cells. DSP tissue slides were incubated with the fluorescently-conjugated antibodies to CD45, PanCk, and SMA along with a cocktail of photo-cleavable-oligo-labeled primary antibodies from Nanostring (Supplementary Table 1). The panels used were the Immune Cell Profiling Module, the Immune Activation Status Module, the Immune Cell Typing Module, and the Pan-Tumor Module. In addition, Nanostring added bar-coding to a series of 8 commercially available antibodies that were added to each run. These were: anti- Cox2 (CX-294, Agilent#M361701-2), anti-p63 (4A4 BioCare Medical#CM163B), anti-CK5/6 (D5-16B4 Leica PA0795), anti-ALDH1 (D9Q8E Cell Signaling#54135), anti-RANKL (M366 Amgen AMG161), anti-CD36 (Sigma Aldrich HPA002018), anti-E-Cadherin (4A2 Abcam ab231303), and anti-Glut1 (EPR3915, Abcam ab115730).
Sample slides were loaded onto a DSP instrument and a programmable digital micromirror device directed UV light to precisely illuminate each selected ROI and cleave photo-cleavable-oligo-labeled primary antibodies in a region-specific manner. The released indexing oligos were then collected via microcapillary aspiration and dispensed into a 96 well plates. The n-Counter Analysis System (Nanostring) was used to quantify the signal of each barcode in every well.
Slide staining and DSP profiling occurred over a 6-month period in 20 separate batches (staining and profiling) using a predetermined order that mixed the three cohorts throughout the process.
ROI selection
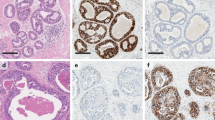
For each patient, one primary, and for some patients, one recurrent sample was collected. For each sample slide, ROIs were selected by one of the two study pathologists, both with specific expertise in breast pathology (AH, RF). Each ROI was categorized as either: DCIS epithelium, benign epithelium or DCIS adjacent stroma. The biopsy site was also sampled to serve as a baseline for non-specific inflammation. Lymphocyte clusters (identified by CD45 staining in the magenta channel, Fig. 1B) in the stroma adjacent to DCIS epithelium were specifically sampled. Further, in DCIS epithelium of sufficient size, we sampled both the peripheral (near the basement membrane) and central areas separately (Fig. 1B). Recurrent DCIS were sampled in the same manner as the primary DCIS. For invasive cancers, we sampled areas enriched in tumor epithelial cells which typically also contained stromal elements including lymphocytes and fibroblasts.
Bioinformatics and biostatistics considerations
The raw count data (Supplementary Material: Sup2_Hwang Nanostring Final Composite Data.xlsx) were downloaded from the Nanostring GeoMx Digital Spatial Profiler (DSP) platform (https://nanostring.com/products/geomx-digital-spatial-profiler/geomx-dsp-overview/). Preprocessing steps were applied to primary samples and to primary-recurrent sample pairs, including sample- and ROI-level filtering and normalization of the data. The final normalized and filtered data are provided in the supplementary material for this report (Sup3_Processed_Data.xlsx (separate tab for the primary and primary-recurrent pairs)). Data preprocessing was performed using R a statistical computing environment (version 4.2.2). Further details regarding the preprocessing and statistical analyses can be found in the supplementary materials (Sup1_Supplementary_Material.pdf). Statistical analyses (full results in Sup4_Statistical_Tests.xlsx) were performed using R statistical computing environment (version 4.2.2)15 along with extension packages from the Comprehensive R Archive Network (CRAN). Unless otherwise indicated, functions from the stats extension package were used for statistical analyses. Unless otherwise stated, changes, differences, and/or significance were claimed based on a threshold of 0.05 for the FDR-adjusted (Benjamini-Hochberg procedure) P-values.
Preprocessing and all analyses were conducted with adherence to the principles of reproducible analysis. The analysis reports were generated dynamically using the rmarkdown16(version 2.19) and knitr17(version 1.41) packages. The computational software environment was containerized using Apptainer18,19.
Results
We performed a spatially resolved quantitative proteomic analysis of DCIS using the Nanostring DSP platform. We detected 53 proteins, including 8 custom antigens developed for this project and 3 housekeeping controls: RPS6, H3, and GAPDH (Supplementary Table 1). DCIS specimens from 103 unique individuals were analyzed. These included 35 DCIS patients with long-term breast cancer-free follow-up (NR), 35 patients that recurred as DCIS in the same breast (DR), and 33 patients that progressed to IBC in the same breast (IR; see Table 1). For these primary DCIS specimens, 1262 ROIs (see Methods above) were evaluated. A median of 12 ROIs per slide (mean area=31593µm2, mean cell/nuclei number = 182 per ROI) were selected, delineated, and annotated by the study pathologist using the H&E-stained section as a reference (Fig. 1A and B, representative sample). In addition, a total of 38 subsequent ipsilateral breast events (IBE) with matched primary DCIS were analyzed, including 20 DCIS recurrences and 18 subsequent invasive cancers.
Quantitative results from the Nanostring platform were based on the amount of antibody binding to antigen (tabulated primary and processed data in Sup3_Processed_Data.xlsx). We removed from subsequent analyses 36 ROIs that had fewer than 30 antigens (of the 53 total) with positive counts after normalization and background correction (mouse and rabbit IgGs). These 36 ROIs were evenly distributed among the three DCIS cohorts and ROI categories (duct, stroma) and were not associated with specific assay dates. To determine whether there were other undetected biases or structures in the data, we performed principal component analyses on the remaining ROIs (n = 1226; Fig. 1C-E). ROIs did not cluster along the first two PCs based on the batch (date of DSP profiling) or cohort but we did observe strong clustering along the PC1 axis (27.8% of the variation) based on ROI category. On PC1, benign epithelium clustered between the DCIS (ducts) and stromal clusters, whereas biopsy site ROIs clustered with the stroma.
Example of tissue ROI selection and principal components analyses of the data set from all ROIs of the primary DCIS specimens. A) Brightfield image of H&E-stained DCIS specimen. B) Parallel section used for proteomic analysis. ROIs collected from this specimen are indicated by the numbered, shaded shapes on the image. C) The first two principal components of normalized antigen expression for the 1226 filtered ROIs collected from primary DCIS specimens that were included in the final analyses, colored by DSP processing batch. D) PCA distribution by the tissue category of the ROIs. E) PCA distribution by DCIS outcome.
ER and HER2 expression differs by ROI subtype
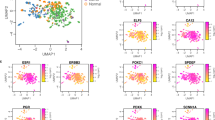
We performed unsupervised clustering of all patients and antigens separately within each of the ROI categories to examine the overall structure of the dataset (Fig. 2A and B). Notably, in the DCIS epithelium, ER positive and HER2 positive patients were in separate clusters with little overlap, suggesting that there were two dominant and distinct biologic subtypes of DCIS. In contrast, in the benign epithelium, there was no evidence of patients clustering, either related to receptor status of the co-existing DCIS or on the basis of other antigens in the panel. Stroma adjacent to DCIS did show strong clustering of a series of T cell antigens indicative of different levels of these immune cells. Dendrograms of antigen clusters indicated that the relationships between the levels of antigens were distinct and different for each tissue category (Supplementary Fig. 2).
Based on all primary DCIS samples, we identified numerous proteins that were differentially expressed between benign and DCIS ROIs (Fig. 2C and Sup4_Statistical_Tests.xlsx). Part of this was likely related to the intrinsic nature of the ROI category, where the benign epithelium had a higher ratio of basal to luminal cells (basal and basement membrane markers S100B, SMA, p63, Cytokeratin 5 (CK5), and fibronectin were higher in the benign ROIs). Notably, ALDH1A (not associated with basal cells) was also consistently higher in benign epithelium. High levels of the tumor markers, ER and HER2 were among the antigens elevated in DCIS. Other antigens that were more nonspecifically expressed (i.e., not obviously related to receptor status) included GAPDH, EpCAM, and CD127 (IL7R). Antigens with significant but less dramatic elevation in DCIS regions included the glucose transporter GLUT1 (SLC2A1), proliferation antigen Ki-67, the adhesion protein CDH1 (E-Cadherin), cancer testes antigen NY-ESO1, and CD25 (IL2RA).
Overall comparisons between the DCIS-adjacent stroma and biopsy site showed broadly increased levels of macrophage and monocyte markers (CD68, CD66B, CD11C, CD14) at the prior biopsy site (Fig. 2D and Sup4_Statistical_Tests.xlsx) consistent with a non-specific immune response. A series of epithelial markers were elevated in the adjacent stroma compared to the biopsy site possibly due to imprecise collection near the epithelium.
Overall representations of the proteomic data from the primary DCIS specimens. A) Heat map of unsupervised clustering of DCIS and benign epithelium ROIs clustered by antigen levels. Patient ER status, HER2 status and cohort are indicated on the top rows. For the benign epithelium, the ER and HER2 status refer to the co-existing DCIS. B) Heat map of the adjacent stroma and biopsy site ROIs clustered by antigen levels. ER and HER2 statuses refer to the co-existing DCIS. C) Volcano plot of unadjusted P-values showing differences between antigen levels in the DCIS versus benign epithelium. D) Volcano plot of unadjusted P-values showing differences between the stroma adjacent to the DCIS and identified biopsy sites. In C and D, points in red indicate antigens with FDR-adjusted P-values less than 0.05. The horizontal dashed lines indicate unadjusted P-values of 0.001.
Estrogen receptor (ER) status from clinical assays and HER2 status from RNA sequencing were available for most of the patients in this study. As ER and HER2 were part of the proteomics panel, we first checked concordance between known receptor levels and those measured by the Nanostring assay (Fig. 3). We compared ER status from clinical IHC assays to the corresponding Nanostring protein data (Fig. 3A). For HER2 status categorized RNA expression levels were used in the comparison (Fig. 3B). Categorical receptor status plotted against the Nanostring data indicated a strong but not perfect concordance between the different assays. As observed with RNA expression data, most DCIS were either estrogen receptor positive or HER2 positive with very few samples negative for both of these central breast cancer drivers, therefore many aspects of these data display inverse relationships between ER and HER2 status (Fig. 3E).
The distributions of ER and HER2 levels reveal the two major categories of DCIS. A) Distribution of ER antigen levels across all primary DCIS ROIs by clinical/RNA-based ER status within tissue category. B) Distribution of HER2 antigen levels across all primary DCIS ROIs by RNA-based HER2 status within tissue category. C) Volcano plot showing differences in antigen levels in DCIS epithelium for samples from patients with ER+ versus ER- status. D) Volcano plot showing differences in antigen levels in adjacent stroma for samples from patients with ER+ versus ER- status. In C and D, points in red indicate antigens with FDR-adjusted P-values less than 0.05. The horizontal dashed lines indicate unadjusted P-values of 0.01. E) Distribution of ER and HER2 levels across all primary ROIs colored by clinical/IHC/RNA-based ER status. F) Distribution of ER levels in the DCIS epithelium and CD45 levels in the adjacent stroma. G) Distribution of ER levels in the DCIS epithelium and CD3 levels in the adjacent stroma. H) Distribution of ER levels in the DCIS epithelium and Fibronectin levels in the adjacent stroma. In F-H, plots are at the patient level, showing antigen levels averaged across all ROIs of that category for that patient.
It has previously been noted that DCIS rarely exhibits the classic basal/triple negative phenotype20 therefore the small number of these is not surprising. The small numbers of double positives and double negatives are consistent with our larger study of DCIS and dissimilar to the distribution of invasive breast cancers observed in the TCGA-BRCA cohort21(Supplementary Fig. 3).
We examined expression of other antigens related to receptor status, both in the DCIS and the surrounding stroma (Fig. 3C-D, and Sup4_Statistical_Tests.xlsx). Estrogen responsive BCL-2 and PR were both significantly elevated in ER+ DCIS along with the immune checkpoint ligand, PD-L2. In ER- DCIS, in addition to HER2, we observed significantly elevated levels of Ki-67, EpCAM, and the pan T cell marker, CD3. In the adjacent stroma (Fig. 3E), a series of antigens associated with the ER status of the DCIS were differentially expressed. For ER- DCIS, lymphocyte antigens, including CD45, CD3, CD20, and CD4, were higher, potentially indicating higher immune reactivity for this subset. ER+ DCIS exhibited increased levels of fibronectin and PanCk compared to ER- patients. Highlighting some of these antigens, scatter charts (Fig. 3F-H) displayed the inverse or direct relationships between ER status of the DCIS and expression of lymphocyte antigens (CD45, CD3) and extracellular matrix component, fibronectin.
Based on our spatial sampling, we also examined how expression levels related to the location of the ROI within the DCIS (central versus peripheral) and ER status (volcano plots, Figs. 4A-D, and Sup4_Statistical_Tests.xlsx). Antigens that were part of the basement membrane (fibronectin) or specifically expressed in the myoepithelial layer (SMA and p63) were elevated in the peripheral ROIs which were placed adjacent to the basement membrane surrounding the DCIS, as expected. Two known basal antigens, CK5 and S100B22,23, were also elevated in the peripheral ROIs compared to central regions across all patients. Notably, Ki-67 was also higher in the peripheral regions of the DCIS epithelium, indicative of differential proliferative activity based on epithelial cell location in the duct. Conversely, the glucose transporter, GLUT1, was the one antigen consistently elevated in the central compared to peripheral regions in both ER-positive and ER-negative patients, suggestive of metabolic differences of cells based on spatial location within the duct. As a comparison, we observed no difference in levels of ER or HER2 between the central and peripheral regions of the duct (Supplementary Fig. 4).
Differential antigen levels between central and peripheral areas of DCIS epithelium. A) Volcano plot showing differences in antigen levels between central and peripheral areas for all primary DCIS epithelium ROIs. B) As A, but limited to samples from ER+ patients. C) As A, but limited to samples from ER- patients. In A, B, and C, points in red indicate antigens with FDR-adjusted P-values less than 0.05. The horizontal dashed lines indicate unadjusted P-values of 0.001. D) The distributions of selected antigens across the DCIS epithelium ROIs for all samples (no fill) and also subset to samples from ER+ (red) and ER- (blue) patients separately.
Protein expression differs according to disease outcome
One of the main goals of this study was to determine whether the levels of specific antigens were associated with disease progression to invasive disease. Therefore, we conducted our analysis by comparing IR (progression to IBC) to the combined NR (no recurrence) plus DR (DCIS recurrence only). Antigen levels in the nearby stroma and within the DCIS epithelium were analyzed independently.
Results not adjusted for FDR are presented in (Fig. 5A and B) with volcano plots showing antigens that differed based on disease progression, including Ki-67 (higher in progressors in both the DCIS and stroma) and fibronectin (lower in progressors in the DCIS). However, these differences did not remain statistically significant after controlling for FDR (Sup4_Statistical_Tests.xlsx).
In separate analyses of disease progression based on the 93 patients for whom we sampled both benign epithelium and DCIS, we used the antigen levels in the associated benign epithelium as an interaction term in the model. Three antigens were observed to be significant in this analysis and are illustrated in spaghetti plots connecting the DCIS with its adjacent benign epithelium and box plots showing the ratio between DCIS and benign antigen levels (Fig. 5C-D and Sup4_Statistical_Tests.xlsx). This ratio was lower in progressing patients for CD127 and CD8, but higher for the checkpoint ligand PD-L2.
As receptor expression defined separate groups of DCIS as well as IBC, we also examined progression separately by ER status (Supplementary Fig. 5). Two antigens found in the primary analysis, PD-L2 and CD8, were associated with progression in ER+ patients. For ER- patients, reduced E-Cadherin and CD56 expression were significant in the model using the benign antigen level as an interaction term. Further, CD56 levels in the stroma of ER- patients was also associated with progression.
Expression differences in primary DCIS related to outcome. Volcano plots comparing DCIS with no or DCIS recurrence compared to invasive recurrence within A) the DCIS epithelium, and B) adjacent stroma. Horizontal dashed lines indicate unadjusted P-values of 0.1. Antigens above the horizontal line are titled but were not significant after adjusting for FDR. C) Spaghetti plots comparing selected antigen levels in the co-existing benign epithelium ROIs to DCIS epithelium ROIs for DCIS with no or DCIS recurrence and invasive recurrence patients. Lines in the plot are colored based on their slope. D) Box plots of the ratio of expression levels between DCIS epithelium and benign epithelium for DCIS with no or DCIS recurrence and invasive recurrence patients for selected antigens. In C and D, antigen levels were averaged across all of the ROIs of that category within each patient.
Comparison of primary DCIS and subsequent ipsilateral breast events
We previously analyzed a series of primary DCIS and subsequent ipsilateral cancers at the genetic level and found evidence for clonal relationships in the majority of cases24. This current study provides an additional level of comparison at the protein level from a series of paired ipsilateral breast events, both DCIS and IBC. For most patients, we also had information on whether endocrine therapy had been used as adjuvant therapy. Comparing ER and HER2 levels between matched primary and recurrent tissues we observed a change in receptor status for a subset of patients (Fig. 6). In patient 38 (DR), the primary DCIS was ER high and HER2 low while the recurrence was ER low and HER2 high. Patient 33 showed the reverse pattern with an ER high and HER2 low recurrence. For patients that progressed, two of the HER2 high DCIS patients (13 and 27) were observed to have much lower levels in the subsequent invasive cancer in the same breast.
We examined all antigens to determine if there was an observable effect of endocrine therapy on expression levels between the primary and recurrent tissues within the epithelial compartments (DCIS ROIs from both primary and recurrent samples for DCIS recurrence patients and DCIS ROIs from primary samples and invasive tumor region ROIs from recurrent samples for the IBC invasive cohort patients (Supplementary Fig. 6 and Sup4_Statistical_Tests.xlsx)). Two antigens (PR and BCL-2) associated with ER were observed to be lower in the subsequent IBC or DCIS in patients that received endocrine therapy compared to those that did not. Two other antigens not associated with ER also showed this same trend (CD163 and SMA). None of the four associations remained significant after controlling for FDR.
Expression differences between primary DCIS and ipsilateral breast events. A) Spaghetti plot of ER levels in primary DCIS and matched DCIS recurrence. B) Spaghetti plot of ER levels in primary DCIS and matched subsequent invasive recurrence. C) Spaghetti plot of HER2 levels in primary DCIS and matched DCIS recurrence. D) Spaghetti plot of HER2 levels in primary DCIS and matched subsequent invasive cancer. Lines are colored based on receipt of endocrine therapy for the primary DCIS. For each patient, antigen levels were averaged across all of the ROIs of that category within sample type (primary vs. recurrence) and within each patient. Each number indicates a specific patient. The counts were log2 transformed.
Discussion
DCIS is a common pathologic diagnosis in populations undergoing screening mammography. As DCIS can progress to invasive cancer, we and others have extensively characterized these lesions to identify distinguishing features. To date, there is no single feature or set of features that consistently separate DCIS from invasive cancer other than their histologic appearance, i.e., the absence of cancer cells invading through the basement membrane. Mutations, characteristic copy number changes, and overall expression patterns broadly overlap between these two disease states and indicate a biologic continuum25,26,27,28. In this study, we used spatial proteomics to determine distribution of markers across distinct regions of tumor architecture. We identified two predominant subgroups of DCIS, ER high/HER2 low and ER low/HER2 high. Levels of tumor associated proteins varied between benign and DCIS, between ER + and ER- patients, and between different regions within the DCIS epithelium. In addition, we identified several immune-related antigens within the DCIS epithelium that are associated with invasive progression.
Features that predict the future behavior of DCIS could have substantial clinical value in guiding surveillance and treatment of the disease29. In particular, features that are associated with progression to invasive cancer are of greatest interest as they can guide treatment decisions. The current study extends our analyses12,30 of an established cohort of DCIS by applying a quantitative spatial proteomic approach measuring 53 antigens on 103 patients sampling over 1200 regions of interest defined by the study pathologist as being within the DCIS duct, stroma adjacent to the duct, or benign epithelium within the same tissue section. Patients were matched by age and year of diagnosis across three groups based on clinical outcome (i.e., no ipsilateral breast event, DCIS recurrence or IBC progression).
Our spatial sampling allowed us to investigate systematic differences across spatial location within the DCIS epithelium, comparing the luminal and basally-located epithelium, bridging an important gap between prior studies using bulk sequencing and more recent reports using single cell approaches. Proteins known to be associated with basal or myoepithelial cells (CK5, SMA, p63) and the proliferation associated Ki-67, were significantly increased in the periphery. Higher proliferation in the peripheral compared to central areas within DCIS epithelium has not been previously reported. Conversely, the glucose transporter, GLUT1, was higher in the central regions. Together with proliferative state, these findings suggest some overall metabolic differences within these relatively large DCIS epithelium regions.
Concurrent analysis of adjacent stroma allowed us to correlate antigen status within the DCIS epithelium to the milieu outside of the duct. Specifically, we found strong inverse correlations between ER in the DCIS epithelium and lymphocyte/T cell antigens CD45 and CD3 in the adjacent stroma and a positive correlation between ER and fibronectin. Higher levels of immunity were associated with ER negative and HER2 positive invasive cancers31,32 and the DCIS in this study show a similar profile. ER expression was positively correlated with fibronectin levels, likely produced by fibroblasts in the adjacent stroma. Various types of fibroblasts have been identified and disease characteristics, including receptor status and DCIS recurrence, have been associated with specific fibroblast properties, including a senescent phenotype33,34. Our result suggests that differences in fibroblast expression do manifest at the pre-invasive stage.
Based on our proteomic analysis, we made the observation that there were two distinct categories of DCIS: those with elevated ER expression, and those with elevated HER2 expression. In this cohort, there were relatively few patients that had high levels or low levels of both of these breast cancer drivers, consistent with the altered distribution of receptor-based subtypes between DCIS and IBC20,35,36. Neither ER or HER2 expression was associated with progression to IBC which continues to suggest a non-linear course of progression from DCIS to invasive cancer and that precursor lesions for receptor negative cancers are underrepresented in most DCIS cohorts.
The cohort in this study was constructed in three matched groups based on disease recurrence and progression. As in our prior work, we consider the patients that progressed to invasive cancer as the primary comparison, grouping non-recurring patients and those that recurred only as DCIS together as non-progressors. Given the heterogeneity of normal breast epithelium (24), consideration of this variability in analyzing expression within the DCIS is a reasonable approach. By using the ratio of expression in DCIS to expression in benign epithelium from the same specimen, we identified several antigens that differed between patients that progressed to IBC and those that did not, all of which were immune based. This included higher levels of the immune checkpoint ligand PD-L2 in patients that progressed to IBC and lower levels of CD127 and CD8. While stromal tumor infiltrating lymphocytes (TIL) markers were not prognostic in our study, we did find that decreased levels of two markers of TILs within the DCIS epithelium (CD45 and CD8) were associated with progression. A recent meta-analysis of 13 studies examining TILs and outcome in DCIS concluded that there was evidence indicating that high levels of stromal TILs were associated with increased risk of subsequent events37. While this conclusion may have been warranted, the studies were highly varied with respect to how TILs and outcomes were defined, and our study indicates that further evaluation of some specific immune subtypes could be an informative direction for future research.
In the subsets of the recurring and progressing patients, we also analyzed the subsequent ipsilateral recurrence (DCIS or IBC) and compared expression to the primary DCIS. Information on receipt of hormone therapy was also available for most of these patients (patient with hormonal therapy: n = 16; without n = 19; missing information n = 3), allowing us to examine antigen expression in this context. A subset of patients exhibited dramatic changes in ER and HER2 expression between the primary and secondary cancers. This was consistent with our prior study at the genetic level (copy number, mutations) indicating that approximately 20% of these secondary cancers may be unrelated to the primary DCIS24, rather than the changes being the result of receipt of hormone therapy. Changes in other antigens between primary and secondary cancers tracked with hormonal therapy, including the estrogen regulated genes PR and BCL-2.
Strengths of the study include the relatively large sample size of DCIS, the matched cohorts, and the availability of other detailed data on these same patients. The sampling of multiple regions and sub-regions from each patient gave us a more representative analysis and provided novel spatial information. Further, including benign epithelium and biopsy sites from many of these samples allowed us to normalize expression levels from the DCIS based on the benign epithelium and compare nonspecific wound-related expression (biopsy site) to the immune cells in the tumor microenvironment. From a proteomic perspective, simultaneous quantitative and spatial analysis of 53 selected antigens that cover tumor and microenvironmental processes provides new information and couples with genomic data from these same DCIS lesions available through the Human Tumor Atlas Network Data portal (Breast Pre-Cancer Atlas, https://humantumoratlas.org/explore). Limitations of the study include that the analysis included a select set of antigens and the limited sample size with respect to disease outcome. Further, the patients in the NR cohort were selected from among TBCRC patients who at the time of the design of this study had no reported DCIS recurrence or invasive progression following the initial DCIS diagnosis. Given that the patients in the NR cohort were selected from among TBCRC patients who were administratively censored at the time of cohort selection, the resulting biased sampling design may have been biased towards early recurrence or progression events. Finally, post-hoc analysis identified three antigens for which there was statistical evidence suggesting that the difference in expression levels between benign versus DCIS epithelia was cohort-dependent.
This analysis addresses an important gap in the current landscape between bulk analysis of tumor and single cell sequencing, allowing comparison of protein expression between geospatially distinct and clinically relevant regions of DCIS and surrounding stroma. We identified clear differences in proteomic profiles depending on spatial location of stromal and epithelial cells and discovered that a subset of these proteins, specifically those associated with the tumor immune environment, was associated with DCIS outcome. Future analyses of specific pathways using this approach will yield further insights into the mechanisms of cancer initiation and progression.
Data availability
Clinical and raw Nanostring expression data are available as part of the supplementary material of this report. The time-to-event data and the RNAseq data used to determine ER and HER2 status were made public as part of a prior publication and can be accessed through the Human Tumor Alas Network (HTAN) [https://humantumoratlas.org/explore] and subset using the patient manifest available on Mendeley [https://data.mendeley.com/datasets/tbzv5hpvw5/3]. The code to reproduce the analyses presented in this paper, including that needed to join these external data sets with the raw Nanostring expression data, along with the apptainer definition file to reproduce the computing environment used to conducted the analyses are available through [https://gitlab.oit.duke.edu/dcibioinformatics/pubs/marks-nanostring-dcis]. We note that an analysis-ready version of the data, which includes these external time-to-event data and ER and HER2 status variables, is also provided as part of the supplementary material of this report.
References
Ryser, M. D. et al. Cancer Outcomes in DCIS Patients Without Locoregional Treatment. J. Natl. Cancer Inst. 111, 952–960 (2019).
Sagara, Y. et al. Survival Benefit of Breast Surgery for Low-Grade Ductal Carcinoma In Situ: A Population-Based Cohort Study. JAMA Surg. 150, 739–745 (2015).
Maxwell, A. J. et al. Unresected screen-detected ductal carcinoma in situ: Outcomes of 311 women in the Forget-Me-Not 2 study. Breast 61, 145–155 (2022).
Hwang, E. S. et al. Active Monitoring With or Without Endocrine Therapy for Low-Risk Ductal Carcinoma In Situ: The COMET Randomized Clinical Trial. JAMA 333, 972–980 (2025).
Elshof, L. E. et al. Feasibility of a prospective, randomised, open-label, international multicentre, phase III, non-inferiority trial to assess the safety of active surveillance for low risk ductal carcinoma in situ - The LORD study. Eur. J. Cancer. 51, 1497–1510 (2015).
Francis, A. et al. Addressing overtreatment of screen detected DCIS; the LORIS trial. Eur. J. Cancer. 51, 2296–2303 (2015).
Silverstein, M. J. The University of Southern California/Van Nuys prognostic index for ductal carcinoma in situ of the breast. Am. J. Surg. 186, 337–343 (2003).
Yi, M. et al. Evaluation of a breast cancer nomogram for predicting risk of ipsilateral breast tumor recurrences in patients with ductal carcinoma in situ after local excision. J. Clin. Oncol. 30, 600–607 (2012).
Solin, L. J. et al. A multigene expression assay to predict local recurrence risk for ductal carcinoma in situ of the breast. J. Natl. Cancer Inst. 105, 701–710 (2013).
Kerlikowske, K. et al. Biomarker expression and risk of subsequent tumors after initial ductal carcinoma in situ diagnosis. J. Natl. Cancer Inst. 102, 627–637 (2010).
Debeljak, M. et al. Multimodal genome-wide survey of progressing and non-progressing breast ductal carcinoma in-situ. Breast Cancer Res. 26, 178 (2024).
Strand, S. H. et al. Molecular classification and biomarkers of clinical outcome in breast ductal carcinoma in situ: Analysis of TBCRC 038 and RAHBT cohorts. Cancer Cell. 41, 1381 (2023).
Risom, T. et al. Transition to invasive breast cancer is associated with progressive changes in the structure and composition of tumor stroma. Cell 185, 299–310e18 (2022).
Beechem, J. M. High-Plex Spatially Resolved RNA and Protein Detection Using Digital Spatial Profiling: A Technology Designed for Immuno-oncology Biomarker Discovery and Translational Research. Methods Mol. Biol. 2055, 563–583 (2020).
Team, R. C. R: A language and environment for statistical computingR Foundation for Statistical Computing, Vienna, Austria,. (2022).
Xie, Y. A. & Grolemund, J. J. G. R Markdown: The Definitive Guide (Chapman and Hall/CRC, 2018).
Xie, Y. Dynamic Documents with R and knitr 2nd edn, Vol. 294 (Chapman and Hall/CRC, 2015).
Developers, S. Singularity https://doi.org/10.5281/zenodo.1310023 (2021).
Kurtzer, G. M., Sochat, V., Bauer, M. W. & Singularity Scientific containers for mobility of compute. PLoS One. 12, e0177459 (2017).
Clark, S. E. et al. Molecular subtyping of DCIS: heterogeneity of breast cancer reflected in pre-invasive disease. Br. J. Cancer. 104, 120–127 (2011).
Cancer Genome Atlas, N. Comprehensive molecular portraits of human breast tumours. Nature 490, 61–70 (2012).
Abd El-Rehim, D. M. et al. Expression of luminal and basal cytokeratins in human breast carcinoma. J. Pathol. 203, 661–671 (2004).
Ichihara, S., Koshikawa, T., Nakamura, S., Yatabe, Y. & Kato, K. Epithelial hyperplasia of usual type expresses both S100-alpha and S100-beta in a heterogeneous pattern but ductal carcinoma in situ can express only S100-alpha in a monotonous pattern. Histopathology 30, 533–541 (1997).
Lips, E. H. et al. Genomic analysis defines clonal relationships of ductal carcinoma in situ and recurrent invasive breast cancer. Nat. Genet. 54, 850–860 (2022).
Pareja, F. et al. Whole-Exome Sequencing Analysis of the Progression from Non-Low-Grade Ductal Carcinoma In Situ to Invasive Ductal Carcinoma. Clin. Cancer Res. 26, 3682–3693 (2020).
Lin, C. Y. et al. Genomic landscape of ductal carcinoma in situ and association with progression. Breast Cancer Res. Treat. 178, 307–316 (2019).
Pang, J. B. et al. Breast ductal carcinoma in situ carry mutational driver events representative of invasive breast cancer. Mod. Pathol. 30, 952–963 (2017).
Abba, M. C. et al. A Molecular Portrait of High-Grade Ductal Carcinoma In Situ. Cancer Res. 75, 3980–3990 (2015).
Hwang, E. S. & Malek, V. Estimating the magnitude of clinical benefit of local therapy in patients with DCIS. Breast 48 (Suppl 1), S34–S38 (2019).
Strand, S. H. et al. Analysis of ductal carcinoma in situ by self-reported race reveals molecular differences related to outcome. Breast Cancer Res. 26, 127 (2024).
Denkert, C. et al. Tumour-infiltrating lymphocytes and prognosis in different subtypes of breast cancer: a pooled analysis of 3771 patients treated with neoadjuvant therapy. Lancet Oncol. 19, 40–50 (2018).
Almekinders, M. M. et al. Comprehensive multiplexed immune profiling of the ductal carcinoma in situ immune microenvironment regarding subsequent ipsilateral invasive breast cancer risk. Br. J. Cancer. 127, 1201–1213 (2022).
Reid, S. E. et al. Cancer-associated fibroblasts rewire the estrogen receptor response in luminal breast cancer, enabling estrogen independence. Oncogene 43, 1113–1126 (2024).
Ye, J. et al. Senescent CAFs Mediate Immunosuppression and Drive Breast Cancer Progression. Cancer Discov. 14, 1302–1323 (2024).
Bergholtz, H. et al. Contrasting DCIS and invasive breast cancer by subtype suggests basal-like DCIS as distinct lesions. NPJ Breast Cancer. 6, 26 (2020).
Horimoto, Y. et al. Estrogen Receptor-positive Ductal Carcinoma In Situ Frequently Overexpresses HER2 Protein Without Gene Amplification. Am. J. Surg. Pathol. 43, 1221–1228 (2019).
Wu, S. L., Yu, X., Mao, X. & Jin, F. Prognostic value of tumor-infiltrating lymphocytes in DCIS: a meta-analysis. BMC Cancer. 22, 782 (2022).
Funding
Research reported in this publication was supported by National Cancer Institute of the National Institutes of Health under award number R01 CA185138-01 (ESH); U2C CA-17-035 Pre-Cancer Atlas (PCA) Research Centers (ESH); UO1 CA214183 (JRM); DOD BC132057 (ESH); BCRF 19–074 (ESH); P30CA014236 (YD, ABS, KO) and Duke Cancer Institute. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
J.M. and S.H. conceived and were responsible for the overall study design. J.M., L.K., L.S., A.H., R.F., and J.N. conducted the experimental portion of the study and primary data collection. Y.D., A.S., and K.W. were responsible for data analysis. Y.D. and A.S. prepared the figures. S.N. provided input on interpretation of the data. J.M., Y.D., A.S., K.W. and S.H. wrote the main manuscript text. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Marks, J.R., Dai, Y., King, L.M. et al. Spatial proteomics of breast ductal carcinoma in situ reveal distinct regional differences. Sci Rep 16, 13350 (2026). https://doi.org/10.1038/s41598-026-43486-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-43486-9