Abstract
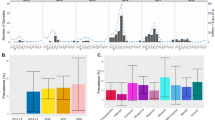
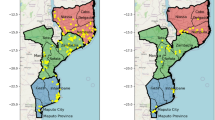
The emergence of antimalarial drug resistance is a major threat to malaria control and elimination. Using whole genome sequencing of 282 P. falciparum samples collected during the 2018 Zambia National Malaria Indicator Survey, we determined the prevalence and spatial distribution of known and candidate antimalarial drug resistance mutations in key genes, including kelch13 (Pfk13), chloroquine resistance transporter (Pfcrt), multidrug resistance protein 1 (mdr1), dihydropteroate synthase (Pfdhps), and dihydrofolate reductase (Pfdhfr) and 8 other genes. High levels of genotypic resistance were found across Zambia to pyrimethamine, > 94% (n = 266) of samples carried the Pfdhfr triple mutant (N51I, C59R, and S108N), and sulfadoxine, > 84% (n = 238) carried the Pfdhps double mutant (A437G and K540E). In northern Zambia 5.3% (n = 15) of samples additionally harbored the Pfdhps A581G mutation. While a total of 29 non-synonymous mutations were identified in Pfkelch13, these mutations were present at low frequency (< 2.5%), and only three were WHO-validated artemisinin partial resistance mutations, P441L (n = 1, 0.35%), V568M (n = 2, 0.7%) and R622T (n = 1, 0.35%). However, 91 (32%) of samples carried the E431K mutation, a candidate artemisinin resistance marker in the Pfatpase6 gene. Finally, no specimens carried any known mutations associated with chloroquine resistance in the Pfcrt gene (codons 72–76). In conclusion, P. falciparum strains circulating in Zambia appear highly resistant to sulfadoxine and pyrimethamine but remain susceptible to chloroquine and artemisinin. While this is encouraging, there are early genetic signs of artemisinin resistance developing underlining the crucial need to remain vigilant and to expand routine genomic surveillance to track these changes.
Similar content being viewed by others
Acknowledgements
The authors are grateful to the Zambian communities, particularly the volunteers and their families, for providing samples during the MIS. We would like to thank the staff of the Zambia National Malaria Elimination Centre for their generous support, especially the field researchers who conducted the nationwide survey.
Funding
This work was supported by funds to G.C. from the Purdue Department of Biological Sciences. Partial funding was provided by the Bill & Melinda Gates Foundation through a grant to PATH (OPP1134518/INV-009984 to D.J.B.). The Southern and Central Africa International Center of Excellence for Malaria Research was supported by funding from the National Institute of Allergy and Infectious Diseases (U19AI089680 to W.J.M.).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical statement
All study participants, and/or their parents or legal guardians, provided written informed consent. This study was approved by the Biomedical Research Ethics Committee of the University of Zambia (Ref 011-02-18) and the Zambian National Health Research Authority. All research procedures were conducted in accordance with the Declaration of Helsinki and relevant institutional and national guidelines and regulations.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Fola, A.A., Ciubotariu, I.I., Dorman, J. et al. National genomic profiling of Plasmodium falciparum antimalarial resistance in Zambian children participating in the 2018 malaria indicator survey. Sci Rep (2026). https://doi.org/10.1038/s41598-026-45996-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-026-45996-y