Abstract
Immune checkpoint inhibitors (ICIs) are utilized to treat hepatocellular carcinomas (HCC) without validated biomarkers. We sought to identify genomic signatures and immune patterns predicting treatment efficacy in 1,306 HCC patients to evaluate for potential predictive biomarkers associated with immunotherapy outcome. Programmed death ligand 1 (PD-L1) expression was high in 6.1% and low in 11.6%; deficient mismatch repair/microsatellite instability-high (dMMR/MSI-H) was detected in only 0.2%, and high tumor mutational burden (TMB-H) in 5.1% of the patients. Immuno-oncologic (IO) markers (CD80, HAVCR2, PDCD1, IFNG, CTLA4, IDO1, CD86, LAG3, PDCD1LG2) correlated with PD-L1 expression (q < 0.05). Infiltration of B cells, M1 macrophages, CD8 + T cells, and Tregs also increased with PD-L1 expression (q < 0.05). CTNNB1 mutations occurred more in PD-L1 negative (35%) than in PD-L1 high (21%) patients (p = 0.03). PD-L1 expression did not affect ICI treatment duration, but CTNNB1-high levels were associated with a shorter duration of treatment in PD-L1 negative tumors (p = 0.04). However, in IO non-exposed tumors, CTNNB1-high expression was associated with improved overall survival (HR = 0.97, 95% CI 0.96-0.98, p < 0.00001). In conclusion, our analysis indicates that expression of PD-L1, CTNNB1, and other molecular markers most associated with IO response in other gastrointestinal malignancies are not directly indicative of such responses in HCC.
Similar content being viewed by others
Introduction
Primary liver cancers in the form of hepatocellular carcinomas (HCC) present numerous challenges to patients for many reasons, including the challenge of late diagnosis and their association with other comorbid medical conditions. HCC, in general, is a recalcitrant and aggressive tumor type associated with a generally poor prognosis, carrying an overall 5-year survival rate of 18%1. Early diagnosis is rare and the vast majority are diagnosed at a late stage, making therapy particularly challenging. Although the number of available systemic therapies has increased over the past decade, they generally have limited efficacy partly because of unique features of the complex tumor immune microenvironment (TIME) comprising the hepatic parenchyma2. Until the era of targeted therapies, single or combination regimens of cytotoxic chemotherapies and other approaches failed to meaningfully improve survival and quality of life in patients with unresectable and metastatic HCC. In 2008, the SHARP trial established sorafenib, an oral multi-targeted tyrosine kinase inhibitor (TKI), as the new standard of care for patients with preserved liver function (Child-Pugh class A). Unfortunately, the mechanism of sorafenib is largely cytostatic, not likely to result in tumor regression, and benefits only a minority of patients with HCC3. Since 2017, other multi-targeted TKIs have been assessed in clinical trials and subsequently approved for use in these patients by the U.S. Food and Drug Administration (FDA), including lenvatinib4 for first-line, and regorafenib5 and cabozantinib6 as second-line treatment. The vascular endothelial growth factor (VEGF) inhibitor ramucirumab has shown efficacy following sorafenib failure in HCC, with benefit mainly limited to patients with serum alpha-fetoprotein (AFP) marker >400 ng/ml7. The sequential integration of these drugs has provided incremental improvements but yet without substantial improvements in overall survival (OS) compared to sorafenib.
The climate of integrating immunotherapies across the spectrum of solid tumor malignancies has accelerated the investigation of immune checkpoint inhibitors (ICIs) in HCC8. Nivolumab was the first FDA-approved IO agent, in 2017, as a second-line treatment based on a non-comparative open-label phase I/II trial of previously treated HCC patients, displaying an objective response rate (ORR) of 20% (95% CI 15–26)9,10. In March 2020, the FDA granted accelerated approval for the combination of nivolumab and ipilimumab as a second-line treatment based on a subgroup analysis of a phase I/II randomized trial that showed an ORR of 32% (95% CI 20–47%) and a median OS of 22.2 months11,12. In May 2020, results from a phase III trial comparing the atezolizumab/bevacizumab combination to sorafenib demonstrated a one-year OS of 67.2% compared to 54.6% with sorafenib. The combination arm had a median OS of 19.2 months compared to 13.4 months with sorafenib13. In 2022, the combination of tremelimumab and durvalumab was added to first-line systemic treatment options based on the results from a phase III trial showing an OS of 30.7% in the tremelimumab/durvalumab arm compared to 20.2% in the sorafenib arm at 36 months14. Most recently, data from a phase III trial comparing the combination of ipilimumab and nivolumab compared to either lenvatinib or sorafenib showed a median OS of 23.7 months in the ipilimumab and nivolumab arm compared to 20.6 months in the lenvatinib/sorafenib arm15. Taken together, these trials have brought immuno-oncology strategies and drugs to the forefront of care for patients suffering from HCC.
These data and the associated rise in the utilization of immunotherapies in first-line HCC highlight the importance of identifying and validating predictive biomarkers that would help tailor the selective use of these drugs in the subpopulations most likely to derive benefit. The cell surface marker programmed death ligand 1 (PD-L1) has entered routine clinical assessments for gastroesophageal and non-small cell lung cancers, among others, owing to its association with improved efficacy and survival in patients with tumors expressing this marker16,17. Other testable markers currently cited for efficacy, including deficient mismatch repair (dMMR)/high microsatellite instability (MSI-H) and high tumor mutation burden (TMB-H), have been approved using a tumor-agnostic approach across the spectrum of solid tumors and are highly investigated in multiple tumor types18. The HCC TIME is enriched with cytokines and metabolites with immunomodulatory properties. The presence of diverse immune regulatory targets in the HCC TIME is unique and enables an understanding of how HCC evades the immune system and develops resistance to novel therapies2. Confounding the question in HCC is the fact that many cases of HCC are diagnosed by standard radiologic criteria based on the distinctive appearance of contrast washout of target tumors on MRI or multi-phase CT19,20,21. Thus, tumor tissue is not required for clinical diagnosis and is not routinely available for genomic profiling.
Although HCC tumor specimens are relatively rare for research purposes, in this study, we accessed what is to our knowledge the largest reported dataset to date of HCC tumor tissue specimens that have undergone comprehensive genomic profiling, including next-generation sequencing (NGS) and whole exome sequencing (WES). This study aimed to identify whether this or other individual or combinations of molecular markers serve as effective biomarkers for response to immunotherapies widely used in HCC.
Results
The total prevalence of PD-L1 expression in hepatic malignancies is less than 20%
The study population consisted of 1306 patients with hepatic malignancies; 1152 had HCC, and 154 had mixed/not otherwise specified/other histology. The median age (range) was 64 (17-86) years; 74.73% of the patients were male (Table 1). The most common histologic subtype was hepatocellular (83.3%) (Table 2). The predominant location of the biopsy specimens was the liver (71.89%), followed by lymph nodes (4.67%), lungs (4.21%), and connective tissue (2.02%) (Supplementary Table 1). Of the 1306 cases, 79 (6.05%) showed PD-L1 high expression, 152 (11.64%) showed low expression, and 1075 were negative (82.31%) (Fig. 1).
A Number (%) of patients with tumors positive for PD-L1. B Pie chart showing the distribution of PD-L1 values (IHC staining value (intensity %) and percentage of patients with each value. C Table showing the distribution of patients with PD-L1 value (IHC staining (intensity | %) in each group. HCC: Hepatocellular carcinoma; PD-L1: programmed death ligand 1; IHC: immunohistochemistry. PD-L1 categories: Tumors with a value of 0 were considered Negative; values 1 ( > 1%) and 2 (1-4%) were considered Low, and 2 + ( ≥ 5%) were considered High.
Prevalence of dMMR/MSI-H and TMB-H is low–5% or less across expression levels of PD-L1
Differences in TMB-H or dMMR/MSI-H statuses were determined and stratified by PD-L1 expression status. The TMB-H prevalence, based on a cut-off of 10 mut/MB, was 3.3%, 4.8%, and 5.1% for the PD-L1 high, low, and negative groups, respectively. For each subgroup, the median TMB was 4.0 mut/MB (Table 1), and these differences were not statistically significant (Fig. 2A, B). The dMMR/MSI-H status was observed in 1.5% of cases in the PD-L1 high group, none in the PD-L1 low group, and 0.2% of the PD-L1 negative group; these differences were not statistically significant (Fig. 2A, B).
A Graph showing percentage dMMR/MSI-H or TMB-H in different PD-L1 groups. B Table showing number of dMMR/MSI-H and TMB-H in each PDL-1 group (High, Low, Negative). PD-L1: programmed death ligand 1; TMB-H: tumor mutation burden high; dMMR/MSI-H: deficient mismatch repair/microsatellite instability-high.
Clusters of alternate IO cell surface biomarkers upregulated in tandem with PD-L1
We examined the RNA expression of immune system-related genes commonly associated with possible or demonstrated responses to oncologic therapeutic agents. PD-L1 expression correlated with control CD274 expression, where CD274 shows expression at mRNA level. We divided the expression of each gene based on the extent of PD-L1 (Fig. 3A, B). CD80 (q = 1.62E-07), HAVCR2 (q = 6.57E-08), PDCD1 (q = 1.76E-07), IFNG (q = 6.69E-12), CTLA4 (q = 2.67E-08), IDO1 (q = 2.70E-09), CD86 (q = 2.32E-07), LAG3 (q = 9.71E-10), and PDCD1LG2 (q = 2.01E-10) expression tended to be higher in PD-L1 high HCC (Fig. 3A, B). CD276 (B7-H3) is another fast-emerging therapeutic target, particularly in the sphere of adoptive cell therapy development; and its expression increased with PD-L1 expression (PD-L1 high vs PD-L1 negative, p = 0.03) (Supplementary Fig. 1). Gene set enrichment analysis (GSEA) revealed that the inflammatory response, TNF-α/NF-Kappa B, IFN-γ response, IL6/JAK/STAT3, IL2/STAT5, IFN-α response, and other pathways were enriched (False Discovery Rate (FDR) q value < 0.05) (Supplementary Table 2).
A Difference in expression of immune-related genes in PD-L1 expression groups. B Table showing tumor immune-related gene counts and pairwise comparisons of PD-L1 High, Low and negative groups. TIME: tumor immune microenvironment; PD-L1: programmed death ligand 1; † CD274 is transcript of PD-L1 and serves as a control.
Immune cell infiltration in the HCC TIME stratified by PD-L1 groups
The HCC TIME is enriched in an inflammatory immune landscape that has long been believed to be the linchpin for the potential efficacy of IO approaches. We examined our dataset of HCC tumors to define the immune infiltration landscape through the identification of individual immune cell subpopulations relative to each other and to tumor cell PD-L1 expression. To this end, we performed RNA deconvolution using QuanTIseq. We examined the quantities of B cells, Tregs, M1 and M2 macrophages, and CD4+ and CD8 + T cell populations and stratified them by the extent of PD-L1 expression. Regarding the TIME, there was significantly higher infiltration of Tregs, M1 macrophages, CD4 + , and CD8 + T cells in the PD-L1 high group than in the PD-L1 low or negative groups. B cells showed higher infiltration in the PD-L1 positive (high and low) compared to the negative group. NK cells showed lower infiltration in the PD-L1 high group compared with that in the PD-L1 negative group. Neutrophil infiltration was significantly higher in the PD-L1 negative than in the PD-L1 low group (Fig. 4A, B). When examining the overall immune-related RNA signatures, the PD-L1 high group had a significantly higher T cell inflammation score and the extent of interferon expression (Fig. 5).
A Table showing tumor immune-related gene counts and pairwise comparisons of PD-L1 High, Low, and negative groups. B Infiltration of immune cell population in PD-L1 expression groups. TIME: tumor immune microenvironment; PD-L1: programmed death ligand 1.
A IFN and T cell inflamed score for PD-L1 (high, low, negative) expression group comparison. B Distribution of T cell inflamed score in PD-L1 expression (high, low, negative) groups (inflamed ≥ 80; Intermediate < 80; not inflamed ≤ 80. C Median IFN and T cell inflamed scores for high, low, negative PD-L1 expression groups with pairwise p and q values. IFN: interferon; PD-L1: programmed death ligand 1.
Genetic alterations in the Wnt/β-catenin pathway are more prevalent in PD-L1 negative hepatic tumors
Next, we compared PD-L1 expression in the context of molecular pathway signals known to affect the immuno-oncologic landscape and cell proliferation in HCC. Using NGS, we measured the prevalence of alterations in CTNNB1, which encodes the signaling pathway linchpin β-catenin known to affect chemokine expression and diminish the extent of immune infiltration in HCC22. CTNNB1 mutations tended to be higher in PD-L1 negative (35%) than in PD-L1 high (20%) expressing HCC tumor groups. To further explore the Wnt/β-catenin pathway, we investigated protein changes in APC, which were more common in the PD-L1 negative group than in the low or high groups (Supplementary Table 3). WNT pathway mutation analyses did not show any differences based on PD-L1 groups (Supplementary Table 4). No difference in CTNNB1 expression was found between PD-L1 groups (Supplementary Table 5), and the Wnt/catenin pathway in GSEA was not enriched (FDR q value > 0.05) (Supplementary Table 2).
Additional genetic alterations stratified by PD-L1 expression levels
Other alterations that were more prevalent in the PD-L1 high compared to the PD-L1 negative tumor group included mutations in HOXB13 (6% vs. 0%; p = 0.005), HNF1A (3% vs. 0%; p = 0.02), STK11 (5% vs. 0%; p = 0.009), and DNMT3A (4% vs. 0%; p = 0.04); as well as amplification of KRAS (4% vs. 0%; p = 0.01) and PDCD1LG2 (4% vs. 0%; p = 0.004), which were also observed in this cohort (Fig. 6). Other key genes, including mutations in TP53 (p = 0.005), ELF3 (p = 0.02), FANCC (p = 0.005), TSC2 (p = 0.02), PIK3CA (p = 0.03), and amplifications in BCL9 (p = 0.03) and CCNE1(p = 0.03), tended to be higher in the PD-L1 low versus PD-L1 negative tumor group (Fig. 6).
A Comparing alteration measured by NGS for high vs negative PD-L1 expression (all p < 0.05). B Comparing alterations measured by NGS for low vs negative PD-L1 expression (all p < 0.05). C Table describing the statistical analyses for the above comparisons. PD-L1: programmed death ligand 1; NGS: next generation sequencing; † Copy number change; § Mutation.
HER2 amplification is rare in HCC but significantly more prevalent in PD-L1 negative cases
Amplification of HER2 is rare in HCC. Using Chromogenic in situ hybridization (CISH), we detected HER2 amplification in 2.86% of the PD-L1 negative HCC tumor group, with no evidence of HER2 in PD-L1 expressing groups (either high or low). When assessing HER2 expression using copy number amplification (CNA), 0.63% of PD-L1 negative cases harbored HER2 amplification (Supplementary Fig. 2).
Survival analyses demonstrate higher survival in IO-non exposed HCC patients with CTNNB1-high expression
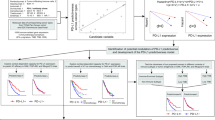
To identify associations between CTNNB1 expression and survival, we examined the time on treatment (TOT) and the time to reported death or last follow-up for unresectable HCC patients to represent overall survival (OS). In the IO-exposed cohort, when patients with CTNNB1-high (above median transcripts per million (TPM)) were compared to patients with CTNNB1-1 low tumors (below median TPM), there was no difference in OS. In the IO-non exposed cohort, CTNNB1-high cases had significantly higher survival compared to CTNNB1-low (HR = 0.97, 95% CI 0.96-0.98, p < 0.00001) (Fig. 7). In contrast, TOT was higher in the CTNNB1-high in IO-exposed group (Supplementary Fig. 3). No differences in OS or TOT were found between PD-L1 positive vs PD-L1 negative patient cases. (Supplementary Fig. 4). On further examination, we found no significant differences in either TOT or OS in patients who had received nivolumab with or without the CTLA-4 inhibitor ipilimumab when comparing PD-L1 negative patients with any level of PD-L1 expression (low or high) in HCC tumors (Supplementary Fig. 5). We conducted survival analyses based on the mRNA expression level of CD274 in PD-L1 groups, and no differences in survival were found (Supplementary Fig. 6). We investigated tyrosine kinase inhibitors (TKIs) in relation to CTNNB1 expression. Our analysis revealed no significant difference in OS or TOT between cohorts with CTNNB1-high versus CTNNB1-low expression (Supplementary Fig. 7 and 8). Furthermore, we stratified each CTNNB1 expression cohort receiving TKI treatment based on levels of PD-L1 expression (comparing PD-L1 positive to negative). Similarly, no significant differences were observed in OS and TOT (Supplementary Fig. 9 and 10). Overall, our analysis indicated some associations of low CTNNB1 expression with higher survival rates, providing signals that would need to be considered for correlative biomarker assessment in prospective therapeutic clinical trials for confirmation.
A Kaplan-Meier (KM) curves for overall survival in patients exposed to IO therapy. B Kaplan-Meier curves for overall survival in patient not exposed to ICIs. IO: Immuno-oncology. CTNNB1 gene expression is stratified by median expression within the cohorts examined.
Discussion
Herein, we provide our report on the largest dataset of HCC cases analyzed using comprehensive genomic profiling in search of potential prognostic or predictive biomarkers or genomic signatures of immunotherapy response. In the era of accelerated approval and the use of IO agents, often in combination with targeted molecular agents to treat this disease, we focused on detecting potential associations between biomarkers of IO response in other gastrointestinal malignancies and targeted therapeutic responses in HCC. We found that the prevalence of TMB-H (5.1%) and dMMR/MSI-H (0.2%) is especially rare in HCC, consistent with previous studies23,24. Overall, PD-L1 was expressed in a limited number of patients (17.7%), with only 6% showing high expression. Additionally, we found that PD-L1 expression does not have any prognostic or predictive impact on the ICIs commonly used in clinical practice for treating patients with HCC. When comparing tumors with high, low, or negative PD-L1 expression, the levels of several immuno-oncology markers, including LAG3, CTLA-4, IDO1, and others, as well as T-cell inflamed and IFN-γ scores, significantly increased with PD-L1 expression. Similar trends were observed for B cells, M1 macrophages, CD8 + T cells, and Tregs, while opposite differences were noted in NK cells. Additionally, when comparing PD-L1 high tumors to PD-L1 negative tumors, mutations in CTNNB1 tended to be lower. In contrast, the amplification of PDCD1LG2, and mutations in HNF1A, HOXB13, DNMT3A, and STK11 were found to be higher in PD-L1 high tumors than in PD-L1 negative tumors. Mutations in TP53, ELF3, TSC2, and PIK3CA, as well as amplifications in BCL9 and CCNE1, tended to be higher in PD-L1 low tumors compared to negative.
Based on multiple clinical trials, PD-L1 has been well-established as a marker of ICI response in upper gastrointestinal (GI) malignancies25,26. In contrast, PD-L1 has not been widely explored as a valid predictive marker for lower GI malignancies, partly due to investigations demonstrating the higher reliability of dMMR/MSI-H27,28,29. The role of ICIs in HCC has been established based on data from several landmark trials reported over the past decade6,30,31,32. However, the utility of PD-L1 IHC as a biomarker for immunotherapy has not been established despite numerous recent clinical trials for HCC31,32. Unlike HCC, other indications for ICI use, such as upper and lower gastrointestinal cancers and non-small cell lung cancer (NSCLC), utilize biomarker testing (PD-L1 expression/MMR/MSI status) to identify the subsets of patients who would benefit from ICIs25,28,33. The use of ICIs in HCC remains independent of any available biomarkers because of challenges inherent to this cancer, including but not limited to the lack of available tissue for genomic profiling, the uniqueness of the HCC TIME, and, to some extent, the immune privilege inherent in the natural microenvironment of hepatic parenchyma. Currently, the validation of testable biomarkers predictive of efficacy is urgently needed, most prominently because immunotherapy has moved upstream from use in refractory cases to first-line treatment with atezolizumab-bevacizumab and tremelimumab-durvalumab over the past several years14,30. Based on our study, PD-L1 is not an effective marker for ICI response in HCC, as no differences in TOT or OS were found between PD-L1 negative and PD-L1 low/positive HCC patients.
Owing to the immune cell-rich microenvironment of HCC, it is crucial to recognize differences in the identification of PD-L1 expression using different clonal antibodies used for commercial and even institutional testing8,34. Studies on other disease types have shown fluctuations in the level of PD-L1 expression as the disease progresses35. Additionally, the level of PD-L1 expression between the primary tumor and metastasis (inter-tumor heterogeneity) can play a role in differing results36. Responses to ICIs can vary based on the location of metastases. Studies combining ICIs with multi-kinase inhibitors in metastatic colorectal cancer have shown better survival in patients with non-liver metastatic disease37,38,39. Studies in animal models have shown an immune response within the liver by coordinated activation of regulatory T cells (Tregs), and modulation of intratumoral CD11b+ monocytes, leading to suppression of systemic antitumor immunity, resulting in worse response rates for liver metastases40. This can further explain the lower response of HCC to ICIs and the inability to utilize IO predictive markers (PD-L1, TMB, and MMR/MSI) for ICI response.
The complex HCC TIME mixture includes but is not limited to hepatic non-parenchymal cells, tumor cells, immune cells, and cancer-associated fibroblasts. This complex cellular interplay plays a significant role in tumor immune evasion, and both adaptive and innate immune responses are blunted. Immune and stromal cell types, such as NK cells and Tregs, are abundant in HCC, and their presence is a sign of poor prognosis41,42,43,44. CTNNB1 gain-of-function mutation has been associated with a paucity of immune TIME with decreased chemokine expression and increased NK cells, Tregs, and other immune cells, which might predict resistance to ICIs or low PD-L1 expression22,45. Data from our study showed that CTNNB1 mutations tended to be higher in PD-L1 negative (35%) than in PD-L1 high (21%). We tried to explore the relationship between CTNNB1-expression and clinical outcomes in relation to IO therapy, TKIs and PD-L1 expression. However, there was no clear evidence of CTNNB-1 being a predictive biomarker of immunotherapeutic response, a conclusion that is consistent with prior analysis of KEYNOTE-240 trial data showing no association of CTNNB-1 mutation with IO related outcomes in HCC46. Our analyses of the IO non-exposed cohort showed improved survival in CTNNB1-high group compared to CTNNB-1 group; this result may be attributed to the exceptionally large sample size and would require further exploration before drawing any firm conclusions. Overall, the role of interfering with the Wnt/β catenin pathway as a potential therapeutic target would require further exploration for confirmation in future studies.
Several studies have sought to identify other markers of ICI response in this disease, focusing on the expression of cell surface markers in relation to immune signatures. In 2019, Ang et al. reported the landscape of HCC biomarkers in this setting using tumor samples from 755 patients. The prevalence of MSI-H and TMB-H was less than 1% in these cases. The authors reported clinical outcomes from a subset of 17 patients and enduring responses to nivolumab exceeding two years in a patient whose HCC was TMB-high and MSI-low HCC24. Hsu et al. reported that the expression of markers of exhausted CD8 T cells (LAG3, CD244, CCL5, CXCL9, CXCL13, MSR1, CSF3R, CYBB, and KLRK1) correlated with the efficacy of ICI in HCC47. Haber et al. showed that genes related to IFN-γ signaling (STAT1, STAT2, and IRF1), antigen presentation (B2M, HLA-DRB5, and HLA-DRA), and chemotaxis (CXCL9) were correlated with ICI efficacy48. Zhu et al. found that pre-existing immunity (high expression of CD274, T-effector signature, and intratumoral CD8+ T cell density) was associated with better clinical outcomes with combination therapy. Reduced clinical benefit was associated with a high regulatory T cell (Treg) to effector T cell (Teff) ratio and expression of oncofetal genes (GPC3, AFP)49. More recently, the retrospective analysis of 242 patients by Cowzer et al. found no molecular marker of efficacy for ICI therapy in HCC patients50. Thus, the landscape for enriching the efficacy of ICIs in response to ICI remains largely unknown and conflicted. The HCC TIME harbors a wide variety of genomic heterogeneities. Studies are underway to target a growth driver, fibroblast growth factor 19 (FGF19), and its receptor, FGFR451. Early-phase results from the selective FGFR4 inhibitor fisogatinib are promising52. However, these findings have not been validated in large, randomized trials. It is worth mentioning that a recently published study by Magen et al. analyzed 29 early-stage HCC tumors and matched uninvolved adjacent liver tissue controls after exposing them to either cemiplimab or nivolumab. They showed that the clonal expansion of CXCL13 + CH25H + IL-21 + PD-1 + CD4 + T helper cells (“CXCL13 + TH”) and Granzyme K + PD-1+ effector-like CD8 + T cells in HCC correlates with response to ICIs53. This immunologic signature needs to be explored further in larger prospective studies.
The limitations of this study include the inherent constraints of retrospective studies, including the unavailability of some time points and incomplete baseline clinical information. The results must be interpreted with the understanding that the low occurrence of TMB-H and MSI-H increases the potential for low prevalence bias in our analyses. Additionally, future studies should assess these IO biomarkers using larger sample sizes although our cohort was a large one considering that tissue biopsy is not a requirement for routinely establishing diagnosis in this cancer type. The non-availability of treatment information by histology of hepatic malignancy is one of the major limitations. It is plausible that there is some difference in biology for patients needing a biopsy for an HCC diagnosis. Moreover, prior studies indicate a clear detection bias with antibody-based PD-L1 testing, a trend reflected in our analyses as well. The variability in PD-L1 testing for HCC is greater than that observed in melanoma or NSCLC. This disparity can be attributed to both the type of antibody employed and the spatial variability of PD-L1 expression, which varies between malignant cells, tumor-infiltrating immune cells, and non-tumorous cirrhotic tissue34.
In summary, our analysis encompassed a broad spectrum of potential molecular and other IO biomarkers utilized and validated in other forms of gastrointestinal cancers for which immunotherapy is used selectively or otherwise based on biomarker expression. Regarding PD-L1 expression, which is most commonly used for decision-making in upper gastrointestinal cancers, we determined it has no predictive or prognostic benefit in HCC. We found that tumors with PD-L1 (high, low) expression also have higher numbers of B cells, M1 macrophages, CD8 + T cells, and Tregs, but not NK cells. The most prevalent mutations were CTNNB1 (higher in negative PD-L1) and TP53 (higher in positive PD-L1). Other notable alterations were the amplification of KRAS, PDCD1LG2, BCL9, CCNE1, and mutations in HNF1A, HOXB13, STK11, ELF3, TSC2, and PIK3CA, which tended to be higher with increased PD-L1. Owing to the complexity of TIME in HCC, a combination of approaches, targets, and therapies may help improve survival. Hence, finding appropriate molecular signatures and targetable therapies is imperative for advancing HCC treatment. The data obtained from our study could not support CTNNB1-expression as a biomarker of therapeutic response. However, further exploration of the role of the Wnt/-catenin pathway in HCC and its interaction with the immune system in advanced HCC is recommended.
Methods
Subjects
A total of 1,306 Hepatocellular cancer (HCC) samples were submitted to Caris Life Sciences (Phoenix, AZ) and analyzed using Next-generation sequencing (NGS), whole exome sequencing (WES), whole transcriptome sequencing (WTS), and immune-histochemistry (IHC). Patients of all age groups and both sexes, including men and women, were involved. Besides the testing results, only basic demographic information, including age and sex, was available. This study was conducted in accordance with the guidelines of the Declaration of Helsinki, the Belmont Report, and the U.S. Common Rule. In keeping with the 45 CFR 46.101(b) (4), this study utilized retrospective, de-identified clinical data. This study was considered Institutional Review Board exempt, and no consent was required from the subject.
Next-generation Sequencing (NGS) of DNA
NGS was performed using NextSeq or NovaSeq platform (Illumina, Inc., San Diego, CA) on DNA isolated from formalin-fixed paraffin-embedded (FFPE) tumor samples. For NextSeq, a custom-designed SureSelect XT assay (Agilent Technologies, Santa Clara, CA, USA) was used to enrich 592 whole-gene targets. For NovaSeq, 720 clinically relevant genes at high coverage ( > 500x) and high read-depth were used, along with another panel designed to enrich for additional >20,000 genes at a low depth (250x). Tumor enrichment was achieved by harvesting the targeted tissue using manual microdissection. A board-certified molecular geneticist interpreted genetic variants identified. They were categorized as “pathogenic,” “likely pathogenic,” “variant of unknown significance,” “likely benign,” or “benign,” according to the American College of Medical Genetics and Genomics standards. When assessing mutation frequencies of individual genes, ‘pathogenic’ and ‘likely pathogenic’ were counted as mutations while ‘benign,’ ‘likely benign’ variants, and ‘variants of unknown significance’ were excluded. All variants were detected with greater than 99% confidence based on allele frequency and amplicon coverage, with an average sequencing depth of coverage greater than 500 and an analytic sensitivity of 5%.
mRNA expression (WTS)
Expression data were evaluated for mRNA isolated from FFPE tumor samples using the Illumina NovaSeq platform (Illumina, Inc., San Diego, CA) and Agilent SureSelect Human All Exon V7 bait panel (Agilent Technologies, Santa Clara, CA), and transcripts per million (TPM) were reported. CTNNB1 mRNA high expression was determined as above the median (TPM), whereas low expression was determined as below the median (TPM). The immune cell fraction was calculated by QuantiSeq (Finotello 2019, Genome medicine) using transcriptomic data54. Additionally, mRNA expression data were used to analyze both the interferon-gamma signature and T-cell inflamed score55,56.
Tumor mutation burden (TMB)
TMB was measured by counting all non-synonymous missense, nonsense, in-frame insertion/deletion, and frameshift mutations found per tumor that had not been previously described as germline alterations in dbSNP151, the Genome Aggregation Database (gnomAD) databases or benign variants identified by Caris’s geneticists. Samples with ≥10 mutations per MB were defined as TMB-high (TMB-H)57.
MSI/MMR status
A combination of multiple test platforms was used to determine the MSI or MMR proficiency status of the tumors, including fragment analysis (FA, Promega, Madison, WI), immunohistochemistry (IHC) (MLH1, M1 antibody; MSH2, G2191129 antibody; MSH6, 44 antibody; PMS2, EPR3947 antibody [Ventana Medical Systems, Inc., Tucson, AZ, USA]), and NGS. The three platforms generated highly concordant results, as previously reported58, and in rare cases of discordant results, the MSI or MMR status of the tumor was determined in the order of IHC, FA, and NGS.
PDL-1 expression by IHC
IHC was performed on full-FFPE sections on glass slides. Slides were stained using automated staining techniques, per the manufacturer’s instructions, and were optimized and validated per CLIA/CAP and ISO requirements. PD-L1 IHC (SP142, Spring Biosciences) expression was evaluated in tumor cells. The staining was regarded as positive if its intensity on the membrane of the tumor cells was >=2+ (on a semiquantitative scale of 0–3: 0 for no staining, 1+ for weak staining, 2+ for moderate staining, or 3+ for strong staining) and the percentage of positively stained cells was >5% as evaluated by a board-certified pathologist.
For this study, samples were stratified by PD-L1 expression (IHC; SP142) into comparison cohorts of negative (0), low (1-2+ and 1-5%), and high ( ≥ 2+ and ≥5%).
HER2
Her-2/neu was detected either by CISH (INFORM HER-2 Dual ISH DNA Probe Cocktail) with amplification defined as Her2/chr17 ratio ≥2.0 or copy number amplification (≥6 copies).
Clinical outcomes data
Real-world overall survival (rwOS) was obtained from insurance claims and calculated from tissue collection to last contact; time-on-treatment (TOT) was calculated from the start to finish of ICI treatments. Kaplan-Meier estimates were calculated for molecularly defined patient cohorts. Statistical significance was determined at p < 0.05.
Statistical analysis
Comparative analyses among cohorts were performed using Chi-square or Fisher Exact tests. The TIME cell fractions were analyzed using the non-parametric Kruskal-Wallis test. A p value of <0.05 was considered a trending difference; p values were further corrected for multiple comparisons using the Benjamini-Hochberg method to avoid type I error, and an adjusted p-value (i.e., q value) of <0.05 was considered a significant difference.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Caris Life Sciences owns the deidentified sequencing data, which cannot be publicly shared due to the data usage agreement in place. These data will be made available to researchers for replication and verification purposes through our letter of intent process, which is generally fulfilled within 6 months. For more information on how to access this data, please contact Dr. Joanne Xiu at [jxiu@carisls.com](mailto:jxiu@carisls.com).
References
Vogel, A., Meyer, T., Sapisochin, G., Salem, R. & Saborowski, A. Hepatocellular carcinoma. Lancet 400, 1345–1362 (2022).
Chen, C., Wang, Z., Ding, Y. & Qin, Y. Tumor microenvironment-mediated immune evasion in hepatocellular carcinoma. Front. Immunol. 14, https://doi.org/10.3389/fimmu.2023.1133308 (2023).
Llovet, J. M. et al. Sorafenib in advanced hepatocellular carcinoma. N. Engl. J. Med. 359, 378–390 (2008).
Kudo, M. et al. Lenvatinib versus sorafenib in first-line treatment of patients with unresectable hepatocellular carcinoma: a randomised phase 3 non-inferiority trial. Lancet 391, 1163–1173 (2018).
Bruix, J. et al. Regorafenib for patients with hepatocellular carcinoma who progressed on sorafenib treatment (RESORCE): a randomised, double-blind, placebo-controlled, phase 3 trial. Lancet 389, 56–66 (2017).
Abou-Alfa, G. K. et al. Cabozantinib in patients with advanced and progressing hepatocellular carcinoma. N. Engl. J. Med 379, 54–63 (2018).
Zhu, A. X. et al. Ramucirumab after sorafenib in patients with advanced hepatocellular carcinoma and increased alpha-fetoprotein concentrations (REACH-2): a randomised, double-blind, placebo-controlled, phase 3 trial. Lancet Oncol. 20, 282–296 (2019).
Sangro, B., Sarobe, P., Hervas-Stubbs, S. & Melero, I. Advances in immunotherapy for hepatocellular carcinoma. Nat. Rev. Gastroenterol. Hepatol. 18, 525–543 (2021).
Yau, T. et al. Nivolumab versus sorafenib in advanced hepatocellular carcinoma (CheckMate 459): a randomised, multicentre, open-label, phase 3 trial. Lancet Oncol. 23, 77–90 (2022).
Reig, M. & Sanduzzi-Zamparelli, M. Nivolumab and sorafenib in hepatocellular carcinoma: lessons from the CheckMate 459 study. Lancet Oncol. 23, 4–6 (2022).
Yau, T. et al. Efficacy and safety of nivolumab plus ipilimumab in patients with advanced hepatocellular carcinoma previously treated with sorafenib: the CheckMate 040 randomized clinical trial. JAMA Oncol. 6, e204564 (2020).
Melero, I. et al. Nivolumab plus ipilimumab combination therapy in patients with advanced hepatocellular carcinoma previously treated with sorafenib: 5-year results from CheckMate 040☆. Ann. Oncol. 35, 537–548 (2024).
Cheng, A. L. et al. Updated efficacy and safety data from IMbrave150: atezolizumab plus bevacizumab vs. sorafenib for unresectable hepatocellular carcinoma. J. Hepatol. 76, 862–873 (2022).
Abou-Alfa, G. K. et al. Tremelimumab plus durvalumab in unresectable hepatocellular carcinoma. NEJM Evid. 1, EVIDoa2100070 (2022).
Galle, P. R. et al. Nivolumab (NIVO) plus ipilimumab (IPI) vs lenvatinib (LEN) or sorafenib (SOR) as first-line treatment for unresectable hepatocellular carcinoma (uHCC): frst results from CheckMate 9DW. J. Clin. Oncol. 42, LBA4008–LBA4008 (2024).
Fashoyin et al. FDA approval summary: pembrolizumab for recurrent locally advanced or metastatic gastric or gastroesophageal junction adenocarcinoma expressing PD-L1. Oncologist 24, 103–109 (2018).
Bodor, J. N., Boumber, Y. & Borghaei, H. Biomarkers for immune checkpoint inhibition in non–small cell lung cancer (NSCLC). Cancer 126, 260–270 (2020).
Thein, K. Z., Lemery, S. J. & Kummar, S. Tissue-agnostic drug development: a new path to drug approval. Cancer Discov. 11, 2139–2144 (2021).
Xu, C. et al. β-catenin signaling in hepatocellular carcinoma. J. Clin. Investig 132, e154515 https://www.jci.org/articles/view/154515(2022).
Wang, B., Tian, T., Kalland, K.-H., Ke, X. & Qu, Y. Targeting Wnt/β-Catenin Signaling for Cancer Immunotherapy. Trends Pharmacol. Sci. 39, 648–658 (2018).
Spranger, S., Bao, R. & Gajewski, T. F. Melanoma-intrinsic β-catenin signalling prevents anti-tumour immunity. Nature 523, 231–235 (2015).
Xiao, X., Mo, H. & Tu, K. CTNNB1 mutation suppresses infiltration of immune cells in hepatocellular carcinoma through miRNA-mediated regulation of chemokine expression. Int. Immunopharmacol. 89, 107043 (2020).
Wong, M. et al. Evaluation of tumor mutational burden in small early hepatocellular carcinoma and progressed hepatocellular carcinoma. Hepatic Oncol. 8, HEP39 (2021).
Ang, C. et al. Prevalence of established and emerging biomarkers of immune checkpoint inhibitor response in advanced hepatocellular carcinoma. Oncotarget 10, 4018–4025 (2019).
Sun, J.-M. et al. Pembrolizumab plus chemotherapy versus chemotherapy alone for first-line treatment of advanced oesophageal cancer (KEYNOTE-590): a randomised, placebo-controlled, phase 3 study. Lancet 398, 759–771 (2021).
Doki, Y. et al. Nivolumab combination therapy in advanced esophageal squamous-cell carcinoma. N. Engl. J. Med. 386, 449–462 (2022).
Dienstmann, R. et al. Consensus molecular subtypes and the evolution of precision medicine in colorectal cancer. Nat. Rev. Cancer 17, 79–92 (2017).
Diaz, L. A. Jr et al. Pembrolizumab versus chemotherapy for microsatellite instability-high or mismatch repair-deficient metastatic colorectal cancer (KEYNOTE-177): final analysis of a randomised, open-label, phase 3 study. Lancet Oncol. 23, 659–670 (2022).
Ho, H.-L. et al. PD-L1 is a double-edged sword in colorectal cancer: the prognostic value of PD-L1 depends on the cell type expressing PD-L1. J. Cancer Res. Clin. Oncol. 145, 1785–1794 (2019).
Finn, R. S. et al. Atezolizumab plus bevacizumab in unresectable hepatocellular carcinoma. N. Engl. J. Med. 382, 1894–1905 (2020).
El-Khoueiry, A. B. et al. Nivolumab in patients with advanced hepatocellular carcinoma (CheckMate 040): an open-label, non-comparative, phase 1/2 dose escalation and expansion trial. Lancet 389, 2492–2502 (2017).
Zhu, A. X. et al. Pembrolizumab in patients with advanced hepatocellular carcinoma previously treated with sorafenib (KEYNOTE-224): a non-randomised, open-label phase 2 trial. Lancet Oncol. 19, 940–952 (2018).
Reck, M. et al. Pembrolizumab versus chemotherapy for PD-L1-positive non-small-cell lung cancer. N. Engl. J. Med. 375, 1823–1833 (2016).
Pinato, D. J. et al. Clinical implications of heterogeneity in PD-L1 immunohistochemical detection in hepatocellular carcinoma: the Blueprint-HCC study. Br. J. Cancer 120, 1033–1036 (2019).
Vilain, R. E. et al. Dynamic changes in PD-L1 expression and immune infiltrates early during treatment predict response to PD-1 blockade in melanoma. Clin. Cancer Res. 23, 5024–5033 (2017).
Lu, L. C. et al. Increased expression of programmed death-ligand 1 in infiltrating immune cells in hepatocellular carcinoma tissues after sorafenib treatment. Liver Cancer 8, 110–120 (2019).
Barzi, A. et al. Phase I/II study of regorafenib (rego) and pembrolizumab (pembro) in refractory microsatellite stable colorectal cancer (MSSCRC). J. Clin. Oncol. 40, 15–15 (2022).
Kim, R. D. et al. A phase I/Ib study of regorafenib and nivolumab in mismatch repair proficient advanced refractory colorectal cancer. Eur. J. Cancer 169, 93–102 (2022).
Martinelli, E. et al. Clinical outcome and molecular characterisation of chemorefractory metastatic colorectal cancer patients with long-term efficacy of regorafenib treatment. ESMO Open 2, e000177 (2017).
Lee, J. C. et al. Regulatory T cell control of systemic immunity and immunotherapy response in liver metastasis. Sci. Immunol. 5, eaba0759 (2020).
Fu, J. et al. Increased regulatory T cells correlate with CD8 T-cell impairment and poor survival in hepatocellular carcinoma patients. Gastroenterology 132, 2328–2339 (2007).
Gao, Q. et al. Intratumoral balance of regulatory and cytotoxic T cells is associated with prognosis of hepatocellular carcinoma after resection. J. Clin. Oncol. 25, 2586–2593 (2007).
Sun, C., Sun, H., Zhang, C. & Tian, Z. NK cell receptor imbalance and NK cell dysfunction in HBV infection and hepatocellular carcinoma. Cell. Mol. Immunol. 12, 292–302 (2015).
Sun, H. et al. Accumulation of tumor-infiltrating CD49a+ NK cells correlates with poor prognosis for human hepatocellular carcinoma. Cancer Immunol. Res. 7, 1535–1546 (2019).
Lachenmayer, A. et al. Wnt-pathway activation in two molecular classes of hepatocellular carcinoma and experimental modulation by sorafenib. Clin. Cancer Res. 18, 4997–5007 (2012).
Kudo, M. et al. Association between beta-catenin (CTNNB1) mutations and clinical outcomes of pembrolizumab in advanced hepatocellularcarcinoma (aHCC): Exploratory analyses from KEYNOTE-240. J Clin Oncol 42, 4109–4109 https://ascopubs.org/doi/10.1200/JCO.2024.42.16_suppl.4109 (American Society of Clinical Oncology, 2024).
Hsu, C.-L. et al. Exploring markers of exhausted CD8 T cells to predict response to immune checkpoint inhibitor therapy for hepatocellular carcinoma. Liver Cancer 10, 346–359 (2021).
Haber, P. K. et al. Molecular markers of response to anti-PD1 therapy in advanced hepatocellular carcinoma. Gastroenterology 164, 72–88.e18 (2023).
Zhu, A. X. et al. Molecular correlates of clinical response and resistance to atezolizumab in combination with bevacizumab in advanced hepatocellular carcinoma. Nat. Med. 28, 1599–1611 (2022).
Cowzer, D. et al. Clinicogenomic predictors of outcomes in patients with hepatocellular carcinoma treated with immunotherapy. Oncologist. https://doi.org/10.1093/oncolo/oyae110 (2024).
Kanzaki, H. et al. The impact of FGF19/FGFR4 signaling inhibition in antitumor activity of multi-kinase inhibitors in hepatocellular carcinoma. Sci. Rep. 11, 5303 (2021).
Kim, R. D. et al. First-in-human phase i study of fisogatinib (BLU-554) validates aberrant FGF19 signaling as a driver event in hepatocellular carcinoma. Cancer Discov. 9, 1696–1707 (2019).
Magen, A. et al. Intratumoral dendritic cell–CD4+ T helper cell niches enable CD8+ T cell differentiation following PD-1 blockade in hepatocellular carcinoma. Nat. Med. 29, 1389–1399 (2023).
Finotello, F. et al. Molecular and pharmacological modulators of the tumor immune contexture revealed by deconvolution of RNA-seq data. Genome Med. 11, 1–20 (2019).
Ayers, M. et al. IFN-γ–related mRNA profile predicts clinical response to PD-1 blockade. J. Clin. Investig. 127, 2930–2940 (2017).
Bao, R., Stapor, D. & Luke, J. J. Molecular correlates and therapeutic targets in T cell-inflamed versus non-T cell-inflamed tumors across cancer types. Genome Med. 12, 1–19 (2020).
Merino, D. M. et al. Establishing guidelines to harmonize tumor mutational burden (TMB): in silico assessment of variation in TMB quantification across diagnostic platforms: phase I of the friends of cancer research TMB harmonization project. J. Immunother. Cancer 8 https://doi.org/10.1136/jitc-2019-000147 (2020).
Vanderwalde, A., Spetzler, D., Xiao, N., Gatalica, Z. & Marshall, J. Microsatellite instability status determined by next-generation sequencing and compared with PD-L1 and tumor mutational burden in 11,348 patients. Cancer Med. 7, 746–756 (2018).
Acknowledgements
EL thanks the following groups for donations in support of cancer research: Friends and family of Gayle Huntington, the Mu Sigma Chapter of the Phi Gamma Delta Fraternity, University of Minnesota (FIJI), the Litman Family Fund for Cancer Research, Dick and Lynnae Koats, Ms. Patricia Johnson, and Love Like Laurie Legacy. This study has no funding support. However, ESA is partially supported by NCI Cancer Center Support Grant P30 CA077598 and DOD Grant W81XWH-22-2-0025. EL reports research grants from the American Cancer Society (RSG-22-022-01-CDP) 2022-2026, and Minnesota Ovarian Cancer Alliance in 2019, 2021, and 2022.
Author information
Authors and Affiliations
Contributions
E.L. conceived the study design. G.S., Y.B., and E.L. performed initial analysis of data. G.S. and E.L. drafted the manuscript. J.H., E.S.A., A.S., S.G., A.P., B.A.W., A.S., and V.K.C. reviewed and edited the study and made major revisions. All authors read the manuscript and provided critical analysis and edits. All authors read and approved the final version for submission.
Corresponding author
Ethics declarations
Competing interests
EL disclosed the following financial relationships from the past two years: equipment for laboratory-based research 2018-present, Novocure, Ltd; honorarium for panel discussion organized by Antidote Education for a CME module on diagnostics and treatment of HER2+ gastric and colorectal cancers, funded by Daiichi-Sankyo, 2021 (honorarium donated to lab); compensation for scientific review of proposed printed content; Elsevier Publishing and Johns Hopkins Press; consultant, Nomocan Pharmaceuticals (no financial compensation); Institutional Principal Investigator for clinical trials sponsored by Celgene, Novocure, Intima Bioscience, Inc., the National Cancer Institute, and the University of Minnesota membership in the Caris Life Sciences Precision Oncology Alliance (no financial compensation). ESA has served as a paid consultant/advisor for Sanofi, Dendreon, Janssen Biotech, Merck, AstraZeneca, Clovis Oncology, Lilly, Bayer, and has received honoraria from Sanofi, Dendreon, Janssen Biotech, Astellas Pharma, Merck, AstraZeneca, and Clovis Oncology; has received research funding from Janssen Biotech, Johnson & Johnson, Sanofi, Dendreon, Genentech, Novartis, Astellas Pharma, Merck, AstraZeneca, Clovis Oncology, and Constellation Pharmaceuticals, a well as travel accommodations from Sanofi, and Dendreon; He is a co-inventor of a biomarker technology licensed to Qiagen. AS reports a leadership role with Autem therapeutics, Exelixis, KAHR medical and Bristol-Myers Squibb; consulting or advisory board role with AstraZeneca, Bristol-Myers Squibb, Merck, Exelixis, Pfizer, Xilio therapeutics, Taiho, Amgen, Autem therapeutics, KAHR medical, and Daiichi Sankyo; institutional research funding from AstraZeneca, Bristol-Myers Squibb, Merck, Clovis, Exelixis, Actuate therapeutics, Incyte Corporation, Daiichi Sankyo, Five prime therapeutics, Amgen, Innovent biologics, Dragonfly therapeutics, Oxford Biotherapeutics, Arcus therapeutics, and KAHR medical; and participation as a data safety monitoring board chair for Arcus therapeutics. YB, CN, and JX are employees of Caris Life Sciences. The authors declare no other financial or non-financial interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sharma, G., Baca, Y., Goel, S. et al. Evaluation of immuno-oncologic biomarkers and β-catenin expression in response of hepatocellular carcinomas to immunotherapy. npj Precis. Onc. 10, 86 (2026). https://doi.org/10.1038/s41698-026-01275-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41698-026-01275-7