Abstract
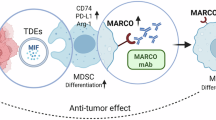
Pancreatic ductal adenocarcinoma (PDAC) is one of the deadliest cancers worldwide. The role of macrophage receptor with collagenous structure (MARCO), a scavenger receptor class-A protein expressed on macrophage surface, in PDAC progression remains unclear. Here, we identified a subset of MARCO-expressing macrophages with strong immunosuppressive signatures that were markedly increased in PDAC patients. Analysis of MARCOhi PDAC samples displayed reduced proportion of CD8⁺ T cells and NK cells, accompanied by an increased proportion of regulatory T cells (Tregs). In vitro, co-culture with multiple PDAC cell lines potently induced MARCO expression on both human and murine macrophages, driving them to a pro-tumorigenic polarization phenotype. Cell-cell interaction analyses further indicated that vascular endothelial growth factor (VEGF) selectively targets MARCO⁺ macrophages, and VEGF stimulation significantly upregulates MARCO expression in vitro. Notably, genetic ablation of Marco markedly suppressed tumor growth in a murine PDAC model, at least partly through enhanced proportion of NK and T cells. Furthermore, MARCO⁺ macrophages were also enriched across several other cancer types, suggesting a potential pan-cancer relevance. Collectively, our findings uncover a critical role of MARCO⁺ macrophages in PDAC progression and highlight MARCO as a promising therapeutic target with potential applicability across multiple malignancies.
Similar content being viewed by others
Introduction
Pancreatic ductal adenocarcinoma (PDAC), often referred to as the “king of cancers”, is one of the cancers with the worst prognosis1,2. Recent epidemiological data show that the global incidence and mortality of this disease are steadily increasing, with a growing impact on younger populations, making it a major threat to public health3. Despite advances in surgery, chemotherapy, and radiotherapy, PDAC remains highly lethal, with a 5-year survival of just 3% in metastatic cases4,5. In addition, immunotherapy against PDAC produces a limited response due to low immunogenicity, high immunosuppression and dense desmoplastic stroma in the PDAC tumor microenvironment (TME)6. By contrast, successful infiltration and activation of T cells in the TME is associated with exceptional long-term survival in patients7. Thus, exploring strategies to modulate immune cell infiltration and activation in the PDAC TME may be key to improving patient prognosis.
Macrophages represent a major component of tumor-infiltrating immune cells, and depending on activation signals, macrophages can acquire a spectrum of phenotypic states8. Tumor-associated macrophages (TAMs), key early infiltrators in the PDAC TME, critically regulate tumor progression, metastasis, immune modulation, angiogenesis, and responses to chemo- and immunotherapy9. Single-cell RNA sequencing (scRNA-seq) and spatial genomics increasingly uncover the complex heterogeneity of PDAC and its TME10,11,12,13,14,15. Despite these advances, the role of targeted macrophage immunotherapy in solid tumors, particularly PDAC, remains limited16,17. As such, there is urgent need to further understand the heterogeneity and biological functions of TAMs in PDAC and to explore new therapeutic targets.
Macrophage receptor with collagenous structure (MARCO) is a scavenger receptor on the surface of macrophages18. MARCO is highly expressed on TAMs, and blocking antibodies reprogram them into pro-inflammatory effectors, triggering antitumor immunity without reducing TAM numbers in breast, melanoma, and colon cancers19. In non-small cell lung cancer, MARCO-expressing macrophages exhibit an immunosuppressive phenotype20. Silencing cancer-cell-derived interleukin-37 or blocking its receptor IL-1R820, which also restrains NK cell antitumor activity, reverses this phenotype21. Recent studies have shown that anti-MARCO mAbs can activate NK cell-mediated killing via the TRAIL pathway18. Indeed, treatment with anti-MARCO mAbs in a mouse melanoma model did not engage CD8+ T cells and was mediated only by NK cells18. In glioblastoma, MARCOhi TAMs significantly accelerated tumor growth in vivo22. Our previous study also found that high MARCO expression indicates a poorer prognosis for PDAC23. Although these findings underscore the important role of MARCO+ macrophages in anti-tumor immunity, their pan-cancer-shared and cancer-type-specific characteristics, and especially the potential role of MARCO+ macrophages subsets underlying the progression of PDAC, remain less understood.
In this study, we identified a previously unrecognized MARCO⁺ macrophage subtype in PDAC. MARCO⁺ macrophages are markedly elevated in primary, metastatic, and post-chemotherapy PDAC and correlate with poor prognosis. scRNA-seq analysis revealed their immunosuppressive gene signatures. In vitro, co-culture with PDAC cells or treating with VEGF upregulated MARCO expression in macrophages. In vivo, Marco deficiency reduced PDAC tumor growth and increased NK cell proportion. Immunofluorescence analysis of patient samples revealed high levels of MARCO⁺ macrophage, reduced levels of CD8⁺ T and NK cells, and increased levels of Tregs. These findings suggest that targeting MARCO⁺ macrophages could be a new therapy for PDAC, and pan-cancer analysis indicates its potential applicability to other tumor types.
Results
Identification of a subpopulation of highly expressed MARCO macrophages uniquely present in primary and metastatic PDAC tissues
To identify macrophage-expressed genes driving PDAC progression and metastasis, we integrated scRNA-seq data from primary (N = 59), metastatic (N = 13), and adjacent tissues (N = 4) to generate single-cell transcriptomes covering tumor, epithelial, stromal, and immune compartments11,24,25,26,27,28,29. We used Seurat for quality control, assessing gene counts, UMIs, mitochondrial gene proportion, and HB/Ribo gene percentages per cell (Supplementary Fig. 1A). Correlations between mitochondrial gene proportion and mRNA reads, as well as mRNA quantity and reads, were analyzed (Supplementary Fig. 1B), and batch effects across cancer and paracancerous tissues were examined (Supplementary Fig. 1C). After filtering, 218,289 cells were grouped into 15 clusters, revealing 12 major cell types, including ductal, T/NK, mast, fibroblast, macrophage, plasma, cycling T, endocrine/acinar, endothelial, B, and acinar cells based on marker genes (Supplementary Figs. 2A–F; Supplementary Data 2–3). Notably, macrophages were enriched in primary and metastatic PDAC tissues (Supplementary Figs. 2G, H), suggesting a key role in PDAC development and progression.
Sub-clustering analysis classified macrophages into 14 clusters, identifying five major populations: IL1B+, C1QA+, MARCO+, HSP+, and MKI67+ macrophages (Fig. 1A–E; Supplementary Data 4, 5). IL1B+ macrophages expressed GRASP, NR4A3, THBS1, CCL3L1, IL1B, and EREG, linked to PDAC progression11. C1QA+ macrophages expressed IFI27, GPR34, C1QC, SELENOP, SLC40A1, and FOLR2, associated with phagocytosis and iron transport. HSP+ macrophages were enriched for heat-shock genes (ZFAND2A, HSPA6, HSPH1, DNAJB1, HSPA1B, BAG3), while MKI67+ macrophages expressed proliferation-related genes (ASPM, GTSE1, DLGAP5, MKI67, TOP2A, RRM2). MARCO+ macrophages expressed FLT1, MARCO, RETN, MCEMP1, SEMAC3, and SLC2A1 and were significantly elevated in primary and metastatic PDAC compared with para-cancerous tissue (Fig. 1D, F).
A The UMAP plot of macrophages from primary, metastatic, and paracancerous pancreatic cancers, including clusters 0–13; B The marker genes in each cluster. C UMAP plot of scRNA-seq data from primary, metastatic, and paracancerous pancreatic cancers, including C1QA+ macrophage, HSP+ macrophages, IL1B+ macrophages, MARCO+ macrophages, MKI67+ macrophages; D Heat map of scaled expression of the top marker genes for each cluster; E UMAP plot of macrophage subpopulation from primary (middle), metastatic (right) and paracancerous (left) pancreatic cancers; F Frequencies from scRNA-seq data of macrophage subpopulation from primary, metastatic and paracancerous pancreatic cancers; G Differences in transcriptional expression between MARCO+ macrophages and MARCO-macrophages; H Pathway analysis of MARCO+ macrophages and MARCO-macrophages; I Changes in M1 scores, M2 scores, regulation of T cell activation, MHC class II biosynthetic process scores, antigen receptor medicated scores, T cell receptor signaling pathway, and NK cell activation scores in various subpopulations of macrophages.
We further compared MARCO+ and MARCO- macrophages and found that MARCO+ cells expressed higher levels of MARCO, FLT1, CCL7, HK2, PPARG, VCAN, and CTSB, whereas MARCO- cells showed higher HLA-DPB1, HLA-DOA, THBS1, FOLR2, SELENOP, and IL1B (Fig. 1G). MARCO+ macrophages were transcriptionally enriched in hypoxia, glycolysis, interferon responses, mTORC1 signaling, EMT, angiogenesis, cholesterol homeostasis, IL2-STAT5 signaling, complement, and adipogenesis, while MARCO- macrophages displayed enhanced mitotic spindle, inflammatory response, allograft rejection, apoptosis, androgen response, G2M checkpoint, E2F targets, and TNF-α signaling via NFκB (Fig. 1H). Across five macrophage subpopulations, MARCO+ cells showed higher M2, MHC-II biosynthesis, glycolysis, oxidative phosphorylation, EMT, and MTO scores, but lower M1, T-cell activation, antigen presentation, TCR signaling, and NK cell activation (Fig. 1I, Supplementary Fig. 2I), indicating a strong immunosuppressive and tumor-promoting profile. These patterns were consistent in primary and metastatic PDAC versus paracancer tissues (Supplementary Fig. 2J), highlighting MARCO+ macrophages as an immunosuppressive, tumor-promoting subset.
MARCO-expressing macrophages exhibit immunosuppressive and pro-tumorigenic signatures in post-chemotherapy PDAC patients
Next, we analyzed ScRNA-seq of freshly dissociated tumor samples from treatment-naïve (before-chemotherapy) (N = 4) or post-chemotherapy patients (N = 6) with PDAC11. We applied the same quality control (Supplementary Fig. 3A–C) and classified 47,402 cells into 11 clusters, identifying 10 major cell types, including acinar, ductal, endocrine, endothelial, fibroblasts, mast, macrophages, plasma/B, T/NK cells, and VSMCs (Supplementary Figs. 4A–F; Supplementary Data 6, 7). Notably, post-chemotherapy patients showed a higher proportion of macrophages (Supplementary Figs. 4G, H), suggesting their potential correlation with chemotherapy resistance. Sub-clustering revealed five macrophage populations: HSP+, C1QA+, IL1B+, MARCO+, and MKI67+ macrophages (Fig. 2A–E; Supplementary Data 8, 9), with MARCO+ macrophages enriched in post-chemotherapy patients (Fig. 2F). These cells exhibited elevated MARCO, VEGFA, SPP1, and VCAN, but reduced HLA-DRB1, HLA-DQA1, and HLA-DOA expression (Fig. 2G). Importantly, MARCO+ macrophages showed higher M2, glycolysis, oxidative phosphorylation, EMT, hypoxia, and angiogenesis scores, but lower M1, T-cell activation, and antigen presentation scores (Fig. 2H; Supplementary Fig. 4I), highlighting their tumor-promoting signature in post-chemotherapy PDAC patients.
A UMAP plot of macrophages from treatment naïve (before-chemothrapy) and post-chemotherapy PDAC patients, including clusters 0–13; B Marker genes in each cluster; C UMAP plot of scRNA-seq data from before-chemothrapy and post-chemotherapy PDAC patients, including C1QA+ macrophage, HSP+ macrophages, IL1B+ macrophages, MARCO+ macrophages, MKI67+ macrophages; D Heat map of scaled expression of the top marker genes for each cluster; E UMAP plot of each cell type from before-chemothrapy (right) and post-chemotherapy PDAC patients (left); F MARCO expression in all macrophages from post-chemotherapy and before-chemothrapy PDAC patients; G Differences in transcriptional expression of MARCO+ macrophages between post-chemotherapy and before-chemothrapy PDAC patients; H Changes in M1 scores, M2 scores, antigen receptor medicated scores, and regulation of T cell activation in various subpopulations of macrophages.
MARCOhi PDAC patients are positively correlated with lower immune cell proportion and higher immune checkpoint blockade (ICB) treatment resistance signatures
Using RNA-seq data from 179 PDAC cases and 332 controls in the TCGA dataset, we confirmed that MARCO expression was significantly higher in PDAC, particularly in advanced stages (IIB–IV) compared to early stages (I–IIA) (Supplementary Fig. 5A). Comparing the top 25% MARCO-high and MARCO-low PDAC tissues, we identified 961 differentially expressed genes, including 887 upregulated and 74 downregulated (Supplementary Fig. 5B; Supplementary Data 10). MARCO-high tumors also showed elevated expression of TREM1, TREM2, APOC1, CCL18, VSIG4, FCGR3A, CCL13, MRC1, ITGB2, MMPs, FN1, CXCL5, and CCL17 (Supplementary Fig. 5C). Immunophenotype analysis using Tumor Immunophenotype (TIP)30 revealed reduced T, NK, and B cell proportion in MARCO-high PDACs (Supplementary Fig. 5D), and these tumors had higher TIDE scores31, indicating poor predicted response to ICB therapy (Supplementary Fig. 5E). GSEA further showed that MARCO expression positively correlated with EMT, glycolysis, proliferation, angiogenesis, PI3K/AKT/mTOR signaling, and inflammatory pathways (Supplementary Fig. 5F). Survival analysis indicated that high MARCO expression was associated with shorter overall survival (Supplementary Fig. 5G). Together, these results suggest that MARCO expression in TAMs may serve as a biomarker promoting PDAC progression.
Cross talk analysis between MARCO+ macrophages and T/NK cells in PDAC patients
To explore how MARCO+ and MARCO- macrophages differently affect T and NK cells, we performed cellular interaction analyses. In primary PDAC, MARCO+ macrophages showed enhanced outgoing and incoming signaling via SPP1, RESISTIN, MIF, GALECTIN, ITGB2, FN1, and APP (Supplementary Fig. 6A). Notably, the SPP1/CD44 axis linked to tumor progression, chemoresistance, and cancer stemness32,was highly active between MARCO+ macrophages and T/NK subpopulations (Supplementary Fig. 6B). Other pathways enriched between MARCO+ macrophages and T/NK subpopulations, including MIF-CD74/CXCR4/CD44, FN1-CD44/ITGA4/ITGB1, and APP-CD74 (Supplementary Fig. 6B), also contributed to an immunosuppressive microenvironment13,33,34. In metastatic PDAC, MARCO+ macrophages exhibited similar signaling patterns, with THBS, SPP1, RESISTIN, MIF, GALECTIN, ICAM1, and FN1 enriched (Supplementary Fig. 6C). Corresponding receptor-ligand pairs, including SPP1/CD44/ITGA4/ITGB1, MIF-CD74/CXCR4/CD44, FN1-CD44/ITGA4/ITGB1, and APP-CD74, were significantly expressed with T/NK cells (Supplementary Fig. 6D). Comparisons between primary and metastatic PDAC showed largely consistent inhibitory interaction signatures (Supplementary Fig. 6E). Overall, MARCO+ macrophages display distinct immunosuppressive interactions with T and NK cells.
Human PDAC cell lines enhance macrophage MARCO expression and polarize macrophage to M2-like phenotype
To begin this, we first analyzed the differences between MARCO+ macrophages and MARCO- macrophages in the expression of M1 and M2 markers. Our results indicated that MARCO+ macrophages express higher MRC1, TNFSF10, ADORA2B, and NECTIN2 but not CD163 (Supplementary Fig. 7A). Conversely, MARCO+ macrophages express lower HLA-DRA, HLA-DRB5, and CD80 but not CD86 (Supplementary Fig. 7B). We next sought to investigate the mechanisms underlying the upregulation of MARCO and the associated immunosuppressive signatures in PDAC. To this end, we established an in vitro co-culture system using CD14⁺ monocyte-derived macrophages and several PDAC cell lines. Our results showed that MARCO expression was markedly induced in macrophages co-cultured with four out of five PDAC cell lines (PANC-1, BxPC-3, APSC-1/GEM, and APSC-1), to a comparable extent as that observed following IL-10 plus IL-4 stimulation (Fig. 3A, B). Phenotypically, both PDAC cell line–derived conditioned media and IL-10 plus IL-4 treatment promoted the generation of anti-inflammatory macrophages, characterized by reduced CD86 and elevated CD206 and CD163 expression, compared with LPS plus IFN-γ–stimulated proinflammatory macrophages (Fig. 3C–H). Collectively, these findings indicate that multiple human PDAC cell lines can enhance MARCO expression in macrophages and drive their polarization toward a pro-tumorigenic M2-like phenotype.
Representative MARCO expression (A), CD86 expression (C), CD163 expression (E) and CD206 expression (G) flow charts of LPS combined with IFN-γ, IL10 combined with IL4 and human pancreatic cancer cell lines (ASPC1, ASPC1/GEM, BXPC3, SW1990, PANC1) supernatant stimulated with PBMC-derived macrophages for 48 h; Quantifications of MARCO expression (B), CD86 expression (D), CD163 expression (F), and CD206 expression (H) in LPS combined with IFN-γ, IL10 combined with IL4, and human pancreatic cancer cell lines (ASPC1, ASPC1/GEM, BXPC3, SW1990, PANC1) supernatant stimulated PBMC-derived macrophages for 48 h (N = 3). I Receptors and ligands between MARCO+ macrophages or MARCO- macrophages and ductal cells (cancer cells) in primary (left) and metastatic (right) PDAC patients; J Upregulated (left) and downregulated (right) signaling in metastatic PDAC patients from ductal cells to MARCO+ macrophages; K Overall information flow of each signaling pathway between primary and metastatic PDAC; L Molecular docking results show that the ARG152 hydrogen bond of the VEGFR1 molecule is tightly bound to the MET462 hydrogen bond of the MARCO molecule; M Representative MARCO expression flow charts of LPS combined with IFN-γ, IL10 combined with IL4, human pancreatic cancer cell line PANC1 supernatant and VEGF stimulated with PBMC-derived macrophages for 48 h; N Quantifications of MARCO expression in LPS combined with IFN-γ, IL10 combined with IL4, human pancreatic cancer cell line PANC1 supernatant and VEGF stimulated with PBMC-derived macrophages for 48 h (N = 3). *P < 0.05, **P < 0.01, ***P < 0.001.
VEGF increases the expression of MARCO on macrophages in vitro
Having shown that PDAC induces macrophage MARCO expression, we next investigated which PDAC-derived factors drive this effect. Using the cellchart package in R, we analyzed receptor-ligand interactions between tumor cells and macrophages in PDAC patients. We found that both primary and metastatic PDAC cells exhibited distinct receptor–ligand profiles when interacting with MARCO⁻ and MARCO⁺ macrophages (Fig. 3I). Numerous studies have indicated that VEGF may contribute to macrophage recruitment and M2 polarization in several tumors35,36. Fibronectin 1 (FN1) is a crucial member of the extracellular matrix glycoprotein family and has been found to induce M2 polarization of macrophages in several tumors37,38. SPP1 affects the receptors of different integrins and CD44, causing tumor progression and immune escape in many tumors39. Our analyses indicated that VEGFA-VEGFR1 and FN1-ITGA5/ITGB1 are present only between primary PDAC and MARCO+ macrophages (Fig. 3I, J). While, SPP1-ITGA5/ITGB1 features both primary and metastatic PDAC and MARCO+ macrophages (Fig. 3I, J).
We next analyzed the signaling pathways from tumor cells to MARCO⁺ macrophages and found that VEGF, CXCL, and complement were primarily present in primary PDAC, whereas FN1, MIF, and ANNEXIN were primarily present in metastatic PDAC (Fig. 3K). Studies have indicated that VEGF and VEGFR1 binding activates downstream PI3K/AKT or MAPK signaling pathways40,41. We next examined whether VEGF could induce MARCO expression in macrophages. Molecular docking revealed multiple hydrogen-bonding interactions between VEGFR1 and MARCO, such as between ARG152 of VEGFR1 and MET462 of MARCO, suggesting a potential binding affinity (Fig. 3L). As expected, VEGF induced the increased expression of MARCO compared to the control (Fig. 3M, N).
Mouse PDAC cell line enhances macrophage Marco expression and polarize macrophage to M2-like phenotype and Marco -/- BMDMs reverse this process in vitro
We further co-cultured BMDMs with the mouse PDAC cell line Panc02 to examine Marco expression at both mRNA and protein levels. Marco was markedly upregulated following treatment with Panc02 supernatant (Supplementary Figs. 8A–C). Likewise, macrophages treated with Panc02-conditioned medium or IL10 plus IL4 showed reduced CD86 but increased CD206 and CD163 compared to LPS plus IFN-γ stimulated proinflammatory macrophages (Supplementary Figs. 8A, D–I). Consistently, pro-inflammatory genes (Ifn-γ, Tnf-α, Il12, and Nos2) were downregulated, whereas anti-inflammatory genes (Arg1 and Vegf) were upregulated after Panc02 supernatant treatment (Supplementary Fig. 8J).
To further assess the impact of Marco deficiency on macrophage, we performed bulk RNA-seq on BMDMs from Marco⁻/⁻ and WT mice (Supplementary Fig. 9A). A total of 631 genes were differentially expressed, including 347 upregulated and 284 downregulated in Marco⁻/⁻ BMDMs compared with WT (Supplementary Fig. 9B, Supplementary Data 12). GSVA analysis revealed enrichment in pathways related to chemokine response, cytokine-cytokine receptor interaction, antigen receptor signaling, chemokine signaling, immune receptor activity, and TNF-α signaling via NFκB, indicating enhanced immune activation upon Marco deletion (Supplementary Fig. 9C). Moreover, M1- and MHC-related genes (e.g., Cd14, Tnf, Cd86, Ccl9, Cxcl16, Tlr8, Ccr1, Ly86, H2-Aa, H2-Ab1, and CD74) were upregulated, while M2-related genes (Marco, Chil3, CD163, Arg1, Spp1, and Lsp1) were downregulated in Marco⁻/⁻ BMDMs (Supplementary Fig. 9D). These findings suggest that PDAC cells promote macrophage MARCO expression and M2-like polarization, whereas Marco deficiency reverses this process in vitro.
Genetic ablation of Marco inhibits the growth of PDAC in vivo
Previous studies have shown that KPC mice recapitulate the progression of human PDAC42. We analyzed bulk RNA-seq data from KPC tumors collected on Days 3, 8, and 15 after the tumors became palpable11. PCA and heatmap analyses revealed distinct transcriptomic profiles across these time points (Supplementary Figs. 10A–C). The expression of Marco and Folr2 decreased on Day 8 but markedly increased by Day 15, with Marco showing higher levels than Folr2 (Supplementary Fig. 10D). Similarly, Vegfa and Vegfb were upregulated at Day 15. Immune infiltration analysis showed that Marco expression negatively correlated with M1 macrophage polarization and NK cell activation, but positively correlated with M2 macrophages and resting NK cells (Supplementary Fig. 10E). Together, these data indicate that tumor progression in KPC mice is associated with elevated Marco expression and suppressed NK cell activity.
We next examined the effect of Marco deficiency on PDAC progression in vivo. Wild-type and Marco⁻/⁻ mice were subcutaneously injected with Panc02 cells. Tumor formation was observed by day 6, and tumor volumes were measured thereafter to generate growth curves (Fig. 4A). Marco⁻/⁻ mice exhibited markedly reduced tumor growth and weight compared with WT mice (Fig. 4B–D). To explore the underlying mechanisms, we performed bulk RNA-seq on tumors from WT and Marco⁻/⁻ mice. A total of 549 genes were differentially expressed (246 upregulated and 303 downregulated; Supplementary Fig. 11A, B, Supplementary Data 11). Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis revealed that upregulated pathways in Marco⁻/⁻ tumors included cytokine-cytokine receptor interaction and vascular smooth muscle contraction, while MAPK, Wnt, and HIF-1 signaling pathways were downregulated (Supplementary Fig. 11C, D). Immune profiling showed a significant increase in NK cell proportion in Marco⁻/⁻ PDAC tissues (Fig. 4E). Moreover, Marco-deficient BMDMs co-cultured with pancreatic cancer cells exhibited elevated TNF-α, IL6, and IL12 expression, but reduced Arg1 expression (Fig. 4F). Together, these findings indicate that Marco deficiency suppresses PDAC growth in vivo.
A Subcutaneous tumor formation pattern of pancreatic cancer in mice; B Representative tumor images of WT and Marco-/- mice at day 34 (N = 5); C Tumor growth curves of WT and Marco-/- mice (N = 17); D Tumor weight of WT and Marco-/- mice at day 34 (N = 17); E Detailed diferences in mMCP-counter scores between WT (N = 3) and Marco-/- mice (N = 5). Comparisons are computed using KruskalWalis tests; F qPCR analysis of the mRNA expression levels of TNF-α, IL-6, IL-12 and Arg-1 on mouse BMDMs from Marco-/- and WT mouse exposed to LPS + IFN-γ, IL-4 + IL-10, and Panc02 murine pancreatic cancer cells. *P < 0.05, **P < 0.01, ***P < 0.001.
MARCO-expressing macrophages are increased while NK cells are decreased in both primary and metastatic PDAC
To further elucidate the impact of MARCO-expressing macrophages on immune cells during pancreatic carcinogenesis, we analyzed the immune landscape of control, primary PDAC, and metastatic PDAC tissues using multiplex immunofluorescence (mIF). A total of 46 human samples were examined. The proportions of CD68⁺ macrophages and CD68⁺MARCO⁺ macrophages were markedly increased in both primary and metastatic PDAC tissues, while CD56⁺ NK cells were significantly reduced (Fig. 5A–D). Notably, tissues with a higher percentage of CD68⁺MARCO⁺ macrophages exhibited fewer proportion of CD56⁺ NK cells (Fig. 5E). We further evaluated CD4⁺ T cells, CD4⁺FoxP3⁺ regulatory T cells (Tregs), and CD8⁺ T cells. CD4⁺FoxP3⁺ Tregs were significantly enriched, whereas CD8⁺ T cells were decreased in both primary and metastatic PDAC tissues (Fig. 5F, G). MARCOhi tissues showed a higher Treg proportion and a lower CD8⁺ T cell proportion compared with MARCOlow tissues (Fig. 5H). Finally, mIF analysis revealed increased Pan-CK and Ki-67 staining in PDAC tissues compared to controls, with Ki-67 expression further elevated in MARCOhi tumors (Fig. 5I, J; Supplementary Figs. 12A, B), suggesting enhanced tumor proliferation. Collectively, these data demonstrate that PDAC tissues, particularly those with high MARCO expression, harbor more MARCO⁺ macrophages and fewer cytotoxic immune cells, indicating an immunosuppressive microenvironment.
A Multiplex immunofluorescence staining of paraffin-embedded paracancer, primary and metastatic PDAC sections for CD68 (green), MARCO (red), CD56 (saffron) and DAPI (blue). Displaying from left to right the composite image and the images for CD68, MARCO, CD56, and DAPI for representative samples defined as control, primary PDAC, and metastatic PDAC. B Quantification of percentage of CD68 in samples defined as control (N = 15), primary PDAC (N = 17), and metastatic PDAC (N = 14), C Quantification of percentage of MARCO in samples defined as control (N = 15), primary PDAC (N = 17), and metastatic PDAC (N = 18), D Quantification of percentage of CD56 in samples defined as control (N = 28), primary PDAC (N = 17), and metastatic PDAC (N = 18), as well as CD56 (E) defined as MARCOhi (N = 20) and MARCOlow (N = 26) PDAC. F Multiplex immunofluorescence staining of paraffin-embedded paracancer, primary and metastatic PDAC sections for CD8 (pink), CD4 (light blue), FoxP3 (red), and DAPI (blue). The composite image and the images for CD8, CD4, FoxP3, and DAPI for representative samples defined as control, primary PDAC, and metastatic PDAC are displayed from left to right. Quantification of percentage of CD4+, CD4+FoxP3+, and CD8+ (G) in samples defined as control (N = 15), primary PDAC (N = 17), and metastatic PDAC (N = 14), as well as CD4+, CD4+FoxP3+, and CD8+ (H) defined as MARCOhi (N = 20) and MARCOlow PDAC (N = 26). I Multiplex immunofluorescence staining of paraffin-embedded MARCOhi and MARCOlow PDAC sections for Pan-CK (red), Ki-67 (pale pink), and DAPI (blue). Displaying from left to right the composite image and the images for Pan-CK, Ki-67, and DAPI for representative samples defined as MARCOhi and MARCOlow sections. J Quantification of percentage of Ki-67 in samples defined as MARCOhi (N = 20) and MARCOlow PDAC (N = 26). *P < 0.05, **P < 0.01, ***P < 0.001.
Human MARCO+ macrophages in PDAC origins from monocytes
Next, we explored the origin of MARCO+ macrophages in human PDAC. scRNA-seq data from monocytes and macrophages in para-cancer, primary, and metastatic PDAC tissues were integrated and analyzed using Monocle 2 to reconstruct developmental trajectories. Monocytes/macrophages were categorized into seven states: the starting states (1 and 2) were mainly CD14+ and CD16+ monocytes, whereas MARCO+ macrophages were enriched in states 4–6 (Supplementary Figs. 13A–C). Trajectory analysis revealed a transition from tumor-infiltrating CD14+/CD16+ monocytes to MARCO+ macrophages, driven by key marker genes MARCO, CD68, and MND1 (Fig. 6A–C). CytoTRACE analysis indicated that CD14+/CD16+ monocytes had the highest differentiation potential, supporting their role as progenitors of MARCO+ macrophages (Fig. 6D, E). Correlation between Monocle 2 and CytoTRACE analyses further validated the trajectory (Supplementary Fig. 13D).
A Cell states are calculated by Monocle 2 and classify the cells that are on the same branch as being most similar based on their pseudotime values. B Trajectory changes of cell states across distinct macrophage subpopulations. C Modules of genes covarying across pseudotime. D UMAP plot of scRNA-seq data overlaid with CytoTRACE scores (left) and cell types (right). E Boxplot of predicted differentiation score distributions for each cell cluster. F Trajectory changes of marker gene MARCO with clinical tumor types. G Trajectory changes of marker gene MARCO with cell states. H Trajectory changes of antigen receptor mediated signaling, NK cell activation, regulation of T cell activation, and T cell receptor signaling pathway with cell states.
We next examined MARCO+ macrophage dynamics across tissue types and states. MARCO expression was highest in metastatic PDAC, intermediate in primary PDAC, and lowest in para-cancer tissues (Fig. 6F). Within states 4–6, MARCO expression peaked in state 4 and declined by state 6, whereas antigen receptor signaling, NK cell activation, and T cell activation pathways showed the opposite trend, negatively correlating with MARCO expression (Fig. 6G, H). Overall, these results suggest that MARCO+ macrophages in PDAC originate from monocytes and are associated with suppressed T and NK cell signatures.
MARCO expression by TAMs increases in 11 of 17 cancer types at single-cell analysis
TAMs play pivotal roles in shaping the TME and influencing cancer progression across diverse tumor types17,43. Therefore, a pan-cancer analysis is essential to systematically elucidate their common and cancer-specific characteristics. ScRNA-seq data from 17 cancer types (SKCM, STAD, COAD, NPC, GBM, LUAD, ESCA, PRAD, CHOL, KIRC, OS, GCTB, OV, THCA, BRCA, LIHC, and PDAC) were analyzed to identify TAMs and compare them with controls (Fig. 7A). ScRNA-seq analysis revealed that MARCO expression in TAMs was significantly higher in PDAC, SKCM, STAD, COAD, NPC, GBM, LUAD, ESCA, PRAD, CHOL, KIRC tumors compared to controls (Fig. 7B). Focusing on cancers with elevated MARCO expression, TAMs were further classified into MARCO+ and MARCO- macrophages. The proportion of MARCO+ macrophages was consistently increased across these cancer types, reflecting their elevated expression (Fig. 7C, D). Using this pan-cancer atlas, we systematically examined the homogeneity and heterogeneity of MARCO+ macrophages. Pathway analysis showed that EMT was enriched in PDAC and also upregulated in SKCM, STAD, COAD, NPC, GBM, and PRAD (Fig. 7E). Glycolysis was similarly enhanced in PDAC, STAD, COAD, NPC, GBM, and SKCM, while hypoxia was enriched in PDAC, STAD, NPC, GBM, PRAD, and SKCM. Angiogenesis was upregulated in PDAC, SKCM, NPC, GBM, and CHOL. Other pathways, including complement, pentose phosphate, IL2-STAT5 signaling, xenobiotic metabolism, and glycolysis/gluconeogenesis, were elevated in MARCO+ macrophages across specific cancer types (Fig. 7E, Table 1). These results indicate that MARCO+ macrophages share both common and cancer-specific tumor-promoting features across malignancies.
A Proportion of TAMs from SKCM, STAD, COAD, NPC, GBM, LUAD, ESCA, PRAD, CHOL, KIRC, OS, GCTB, OV, THCA, BRCA, and LIHC at the single cell level in comparison to controls; B TAMs from SKCM, STAD, COAD, NPC, PDAC,GBM, LUAD, ESCA, PRAD, CHOL, KIRC, OS, GCTB, OV, THCA, BRCA, and LIHC expressed MARCO at the single cell level in comparison to controls; C UMAP plots of macrophage subpopulations in normal (left) and tumor (right) from SKCM, STAD, COAD, NPC, GBM, LUAD, ESCA, PRAD, CHOL, and KIRC; D Frequencies from scRNA-seq data of macrophage subpopulation in normal and tumor from SKCM, STAD, COAD, NPC, GBM, LUAD, ESCA, PRAD, CHOL, and KIRC; E GSEA analysis of MARCO+ macrophages and MARCO-macrophages from SKCM, STAD, COAD, NPC, GBM, LUAD, ESCA, PRAD, CHOL, and KIRC.
Discussion
PDAC is more aggressive and carries a worse prognosis than other cancers because of its unique microenvironment, invasion, and early metastasis44. Macrophages, through their remarkable plasticity, play a pivotal role in the initiation, proliferation and development of PDAC11,45,46. We identified a distinct MARCO⁺ macrophage subpopulation in primary, metastatic, and post-chemotherapy PDAC patients, as well as in several other solid tumors. These cells display immunosuppressive characteristics, underscoring their potential as a therapeutic target.
Macrophages are a highly heterogenous population of cells that orchestrate various aspects of mammalian immunity depending on their phenotype47,48. Different macrophage subpopulations such as IL1B+ macrophages11, C1Q+ macrophages49, and SPP1+ macrophages50 have relatively specific mechanisms of action in pancreatic carcinogenesis and progression. Our previous study found that MARCO+ macrophages were positively associated with poor prognosis in PDAC patients23. In the present study, we characterized the macrophage landscape across paracancerous, primary, metastatic, and post-chemotherapy PDAC tissues using ScRNA-seq. We identified a group of MARCO+ macrophages with strong immunosuppressive signatures that were positively correlated with the progression of PDAC. Therefore, moderate MARCO expression may be crucial for the antitumor effect of macrophages.
The biological effects of VEGF-suppression of cytotoxic immunity and enhancement of glycolysis and angiogenesis in PDAC51,52,53, reflect the divergent control of gene modules in macrophages. VEGF also has a macrophage chemoattraction role, which led to the higher number of infiltrating macrophages54. MARCO expression by TAMs is associated with poor prognosis of multiple types of cancer, including lung cancer, breast cancer, gioblastoma, and so on19,20,23,55. The molecular mechanisms underlying cancer-mediated MARCO expression by macrophages remain to be elucidated. In the present study, we found that VEGF enhances the expression of MARCO by macrophages. Although our findings highlight a strong association between VEGF signaling and MARCO induction, we cannot exclude the potential contributions of other additional regulators. Further experiments, such as neutralizing VEGF or employing VEGF-deficient mouse models, will be essential to determine the specific contribution of VEGF to MARCO regulation in vivo. This limitation should be addressed in future studies.
PDAC is among the most immune-resistant tumor types6. Single-agent immune modulators have thus far proven clinically ineffective, and multi-modal therapies targeting mechanisms of immunotherapy resistance are likely needed56,57. The host immune system plays a key role within the TME, where different types of immune cells have varying impacts on PDAC progression58,59. In the present study, we demonstrated that the expression of MARCO on macrophages is associated with PDAC progression during co-culturing in vitro and in subcutaneous tumors in vivo. The expressions of SPP1/CD44/ITGA4/ITGB1 pairs, the MIF-CD74/CXCR4/CD44 pathway, FN1-CD44/ITGA4/ITGB1 pairs, and APP-CD74 pairs promotes macrophages inhibition of tumor killing by T/NK immune cells60,61,62,63. Interestingly, these receptor-ligand pairs were only expressed between MARCO+ macrophages and T and NK subpopulations in PDAC, suggesting that the unique immunosuppressive microenvironment is initiated and maintained by MARCO+ macrophages, underscoring the need for further experiments. Using PhenoCycler-Fusion (PCF), we found that MARCO+ macrophages were markedly enriched in primary and metastatic PDAC, coinciding with reduced CD8+ T and NK cell as well as increased Tregs. Notably, Marco deletion in a PDAC mouse model increased intratumoral NK cells. These findings suggest that targeting MARCO, potentially in combination with anti-TIGIT or anti-PD1 therapy, may enhance responses in patients with PDAC resistant to checkpoint inhibitors. Further preclinical and clinical studies are warranted.
Tissue-resident macrophages (RTMs) have vital roles in tissue homeostasis, inflammation, and remodeling64. During homeostasis, monocytes survey the vasculature or are recruited to continuously replenish RTMs, for example in the gut65. While, monocytes are recruited to the TME via several different chemotactic factors66,67. Furthermore, monocytes, together with VEGFA, can drive monocytes further differentiation into metastasis-associated macrophages, thus promoting tumor cell survival and persistent growth68. Tumor challenge experiments in Ccr2−/− mice, which lack the CCL2 receptor, and in wild-type mice treated with a neutralizing antibody against CSF-1 significantly impaired several cancer progressions, including PDACs11. Our analysis indicated that the origin of MARCO+ macrophages is from monocytes, which proposes a logical new paradigm of treatment, targeted monocyte recruitment combined with MARCO monotherapy. Further studies are required to confirm this.
Targeting macrophages has shown progress in hematological malignancies, but therapeutic efficacy in solid tumors remains limited, largely due to the unique TME16,17. Here, we profiled macrophage heterogeneity across cancers and identified MARCO+ macrophages enriched in over ten cancer types. Pan-cancer comparison revealed both shared and distinct macrophage states across TMEs. Previous studies showed elevated MARCO expression in pancreatic, testicular, cervical, melanoma, thyroid, brain, and renal cancers, with associations to poorer OS in BLCA, BRCA, LUSC, COAD, and PRAD, but better PFS in LGG and SKCM23,69. Using scRNA-seq, we confirmed MARCO+ macrophage enrichment in SKCM, STAD, COAD, NPC, GBM, LUAD, ESCA, PRAD, CHOL, and KIRC, consistent with prior reports in GBM55, COAD19, LUAD20,70, SKCM18, and CHOL71. Our results suggest potential roles of MARCO+ macrophages in STAD, NPC, ESCA, and PRAD, warranting further mechanistic studies. Notably, we did not observe significant MARCO+ macrophage enrichment in HCC72,73, possibly reflecting patient heterogeneity.
In conclusion, our comprehensive single-cell transcriptomic analysis reveals that MARCO+ macrophages present immunosuppressive signatures in several cancer types, especially PDAC, offering potential targets for therapeutic intervention. Marco deficiency inhibits PDAC growth, which further confirms this hypothesis. Anti-MARCO antibody is a promising novel modality for treatment of aggressive cancers, warranting further investigation.
There are several limitations of this study to consider when interpreting the results. One of the main limitations is the absence of the specific deletion of T or NK cells in the humanized PDAC mouse model to clarify the role of MARCO antibodies in adaptive immune response. To address this, we are actively planning and executing this animal study for in vivo confirmation. Another limitation is that more verification and functional experiments should be conducted in future in more cancer types. Although our data indicate an increase of Tregs and higher Ki67 expression in MARCOhi tumors, suggesting potential immunosuppressive and proliferative effects, these mechanisms were not explored in depth in the present study and require further experimental validation. While we have identified a potential therapeutic target and potential mechanisms, further translational research is required to develop and test this target in clinical settings.
Methods
Cell lines, mouse models and key resources
Human PDAC cell lines (PANC01, BCPC-3, SW1990, ASPC1, and ASPC1-GEM) were obtained from QuiCell (Shanghai QuiCell Biotechnology Co.). Mouse PDAC cell line Panc02 was also obtained from QuiCell. Cancer cell lines were maintained and sub-cultured in DMEM or RPMI 1640 medium with 10% (v/v) heat-inactivated FBS, as previously described74. C57BL/6JCya-Marcoem1/Cya mice (Cat#: S-KO-03130) were obtained from Cyagen Biotechnology Co. Wild type (WT) mice were used as a control, and both of these two types of mice were on the C57BL/6 background and maintained at the Experimental Animal Center of Zhengzhou University Animal Facility. Adult mice 8- to 12-week-old of age were used for the studies. All animal protocols are approved by the institutional animal care and use committee of Zhengzhou University Laboratory Animal Center. For euthanasia, animals were performed using an isoflurane overdose and death was confirmed by cervical dislocation. This study and included experimental procedures were approved by the Laboratory Animal Center of Zhengzhou University (ZZU-LAC20250117[21]). The collection and analysis of human samples and clinical data received approval from the Ethics Committees of the Affiliated Cancer Hospital of Zhengzhou University (2024-KY-2343-001). The key resources in this study are presented in Supplementary Data 1.
ScRNA-seq datasets collected in this study
We collected published ScRNA-seq data on human primary PDAC tissues, metastatic PDAC tissues, post-chemotherapy PDAC tissues, treatment-naïve PDAC tissues and normal adjacent tissues (GSE197177, GSE205013, GSE212966, GSE214295, GSE217845, GSE231535, GSE242230). We also collected scRNA-seq datasets from KPC mouse models (PRJNA590588). In addition to this, we collected ScRNA-seq data from 17 tumors in SKCM (GSE151177), STAD (GSE134520, GSE183904), COAD (GSE188711), NPC (GSE150430), GBM (GSE162631), LUAD (GSE139555, GSE189357), ESCA (GSE145370, GSE196756), PRAD (GSE181294), CHOL (GSE171899), KIRC (GSE139555), OS (GSE162454), GCTB (GSE168664), OV (GSE184880), THCA (GSE191288), BRCA (GSE114725, GSE180286, GSE243526), and LIHC (GSE112271, GSE179795, GSE125449, GSE134355). For these datasets we downloaded: raw fastqs, raw count matrices and normalized counts, respectively. Data were processed as follows.
ScRNA-seq processing
ScRNA-seq analysis was similar as our previously described75. Briefly, Fastq files were processed with Cell Ranger (v 4.0.0), using default parameters. Reads were aligned to reference genome mm10 for mouse samples and hg38 for human samples (version 2020-A, 10× Genomics). Only confidently mapped reads with valid barcodes and unique molecular identifiers (UMIs) were retained to compute a gene expression matrix containing the number of UMI for every cell and gene. Gene counts were imported in R environment (v 4.0.3) and processed with Seurat (v 4.0.3). When creating the Seurat object, genes expressed in less than 3 cells were removed. Putative doublets were identified and discarded using scDblFinder R package (v 1.4.0) by imputing doublet rates (dbr) equal to 0.07 for mouse samples and 0.05 for human samples. Dbr were established in agreement with the number of loaded cells and following the 10× Genomics guidelines. Cells expressing less than 1000 UMI counts were discarded. Cells expressing less than 200 genes (mouse sample) or less than 500 genes (human sample) were also excluded. Lastly, cells with a ratio of mitochondrial versus endogenous gene expression exceeding 0.25 (mouse sample) or 0.40 (human sample) were discarded. Raw expression data were normalized applying log2 transformation with the NormalizeData function, scaled using the ScaleData function, and regressed on the percentage of mitochondrial gene expression and cell cycle scores previously computed using the CellCycleScoring function. The top 3000 genes with the highest standardized variance were computed using the FindVariableFeatures function (selection.method = “vst”). Principal component analysis (PCA) was computed using the RunPCA function with default parameters.
Batch effect correction
Harmony (version 0.1.1), the top batch mixing method with good performance, was used to eliminate batch effects in the PCA space when clustering major cell lineages before any clustering analysis or cell type annotation76,77. Similarly, Harmony was employed to merge the expression profiles of all cells, eliminating batch effects arising from the heterogeneity between cells of individual patients within each dataset.
Graph-based clustering and differential gene expression analyses
A shared nearest neighbor graph was computed using the FindNeighbors function, with the first 20 PCA dimensions as input. Cell clusters were defined using the Louvain algorithm with the FindCluster function. Uniform manifold approximation and projection (UMAP) and t-distributed stochastic neighbor embedding were used for the two-dimensional visualization. Cluster-specific genes were identified using the FindAllMarkers function with option only.pos = TRUE and min.pct = 0.1, setting a cut-off of FDR < 0.01.
Differentially expressed genes analysis at the single cell level
Differentially gene expression analyses between two clusters at the single cell level were performed as described previously75,78. For each gene, the Wilcoxon rank-sum test was used to compute a z-statistic from the down-sampled UMI counts of the two populations. These z-scores were averaged by randomly sampling 100 times, then converted into p values and corrected.
Gene set enrichment analysis
Hallmarks gene sets were retrieved from msigdbr (v 7.5.1)73. For Gene Ontology (GO) biological processes gene sets and KEGG, we used org.Hs.eg.db (v 3.12.0) and org.Mm.eg.db (v 3.12.0) as genome-wide annotations for humans and mice, respectively.
Differentially expressed pathway analysis between different groups
Gene set variation analysis (GSVA) was employed to evaluate differential pathways, with activity scores for specific cell types calculated using GSVA version 1.40.1, applying a false discovery rate (FDR) threshold of less than 0.0579. Pathways were deemed significantly enriched if they exhibited an adjusted p value of less than 0.05. The GO gene sets underwent enrichment analysis via the R package clusterProfiler version 3.16.1. To address the issue of multiple comparisons, p values were adjusted using the Benjamini–Hochberg procedure. Ontologies were considered significant if the q value was less than 0.05.
Cell–cell interaction score calculation using CellphoneDB
We investigated cell–cell interactions across all cell types with CellPhoneDB (version 3.1.0) (https://www.cellphonedb.org/). To elucidate the most pertinent interactions among cell types, CellPhoneDB was employed to identify cell type-specific interactions between ligands and receptors. Only those receptors and ligands expressed in more than 10% of the cells within a specific cluster were deemed significant. Subsequently, CellPhoneDB conducted pairwise comparisons across all cell types. Ligand-receptor pairs with a P value less than 0.05 were retained for further analysis. To evaluate cell-cell interactions, the total interaction score for each cell pair was calculated by summing the scores of all ligand-receptor pairs within that cell pair. The total score indicates the presence and activity of ligand-receptor pairs in each cell pair, irrespective of whether the interactions are mediated by ligands or receptors that positively or negatively regulate antitumor immunity.
CellChat analysis
Cell–cell communication was examined utilizing the CellChat pipeline, version 1.1.3. A novel CellChat object was derived from the integrated Seurat object. The dataset of paracrine and autocrine signaling interactions from the CellChatDB served as the reference database. Subsequently, communication probabilities were computed employing a 20% truncated mean, utilizing the computeCommunProb function with the parameter type designated as “truncatedMean” and the trim parameter set to 0.2. Afterward, cell communication was inferred, and the network was aggregated with default settings. The interactions were visualized to depict the combined cell communication network and the signals transmitted from each cell cluster.
Monocle 2 analysis
Monocle 2 was used to analyze the trajectory of macrophages from human primary PDAC tissues, metastatic PDAC tissues, and normal adjacent tissues in pseudo-time based on the top 500 genes with the highest variation in expression across the macrophages comprising 7 clusters80.
CytoTRACE analysis
CytoTRACE is a computational framework that leverages single-cell gene counts, covariant gene expression, and local neighborhoods of transcriptionally similar cells to predict ordered differentiation states from scRNA-seq data as previous described81.
Murine microenvironment cell populations counter (mMCP-counter) analysis
We used TPM-normalized transcriptomic data and performed a log transformation with a pseudocount of 1 to stabilize the variance and normalize the distribution for the mMCPcounter estimate82.
The Cancer Genome Atlas (TCGA) data analysis
Using the TCGAbiolinks R package (v 2.23.2), we downloaded transcriptomic data and clinical data from the pancreatic adenocarcinoma cohort for PDAC (n = 179). We used the Limma package (version: 3.40.2) in R software to study the differential expression of mRNA. In the data from TCGA, we analyzed the adjusted P values to correct for false positive results. We defined “Adjusted P < 0.05 and log2 (fold change) > 1 or log2 (fold change) < −1” as the threshold for screening differential mRNA expression.
TIP is a system that integrates two existing methods, “ssGSEA” and “CIBERSORT,” to track and analyze the proportion of tumor-infiltrating immune cells in the cancer immunity cycle30. We collected the genes included in the corresponding pathways and then analyzed them using the GSVA package in R software, choosing the parameter method = “ssgsea” for single-sample gene set enrichment analysis (ssGSEA) as previously described83. Finally, we studied the correlation between MARCO expression and pathway scores through Spearman correlation analysis. All the above analysis methods and R packages were performed using R software version v4.1.3 (R Foundation for Statistical Computing, 2022). P < 0.05 was considered statistically significant.
Treatment of PBMC-derived macrophages with human PDAC–conditioned media
CD14+ monocytes were isolated from heathy donors’ Peripheral Blood Mononuclear Cells (PBMCs) via CD14 microbeads (Miltenyi Biotec, Cat#: 130-091-097), cultured for 3 days in Macrophage Colony-Stimulating Factor (M-CSF) (100 ng/ml), and polarized toward proinflammatory or anti-inflammatory macrophages with Lipopolysaccharide (LPS) (100 ng/ml) plus IFN-γ (20 ng/ml) or IL10 (10 ng/ml) plus IL-4 (20 ng/ml) for 48 h, respectively. Bone marrow-derived macrophages (BMDMs) were cultured as previously described84. On day 7, BMDMs were polarized toward pro-inflammatory or anti-inflammatory macrophages with LPS (100 ng/ml) and IFN-γ (20 ng/ml) or IL-4 (20 ng/ml) and IL-10 (20 ng/ml). When cancer cell lines (PANC01, BCPC-3, SW1990, ASPC1, ASPC1-GEM and Panc02) reached ∼80% confluence, cells were harvested after trypsin digestion. After extensive washing with PBS, cells were plated at 3 × 106/well in a 6-well cell culture plate. After 48 h of incubation, supernatants of cancer cell lines were collected to be used as conditioned media to treat PBMC-derived macrophages (supernatants from PANC01, BCPC-3, SW1990, ASPC1, and ASPC1-GEM cell lines) or BMDM (supernatants from Panc02) for 48 h. MARCO expression, M1 markers (CD86), and M2 markers (CD163, CD206) on macrophages were then measured using flow cytometry analysis, as previously described85,86.
Treatment of bone marrow-derived macrophages with mouse PDAC-conditioned media
BMDMs were cultured as previously described87. Briefly, mice were first taken at 8–12 weeks of age, executed by cervical dislocation after isoflurane anesthesia, placed in 75% alcohol for 2 min, and subsequently transferred to an ultra-clean bench. Then, the hind limb was clipped along the greater trochanter of the mouse leg, and the leg muscle tissue was stripped to isolate the femur and tibia. After washing with PBS, the bone marrow cavity was rinsed by drawing pre-cooled PBS with a 1 mL syringe, and the rinsing was repeated until the red color in the bone marrow cavity faded. Bone marrow cells were blown with a 1 mL pipette gun until the clumps were dispersed, and the cell suspension was passed through a 70 μm cell strainer to obtain a single-cell suspension. The single-cell suspension was centrifuged at 1400 rpm for 5 min, and 2 mL of erythrocyte lysate was added after discarding the supernatant and placed on ice for 15 min. The cells were neutralized with 6 mL of DMEM complete medium and centrifuged at 1400 rpm for 5 min, after which the supernatant was discarded and the cells were counted. The cell density was adjusted to 2 × 106/mL with macrophage induction medium containing M-CSF (20 ng/ml). Three mL of cell suspension was added to each well of a six-well plate, mixed, and placed in the incubator. After 3 days, half of the medium was replaced to continue incubation. After 5 days, the mature macrophages were adherent. The macrophage induction medium was discarded, and the adherent macrophages were divided into three groups and added to different induction media. The first group was added with M1-type induction medium containing LPS (100 ng/ml) and IFN-γ (20 ng/ml), the second group was added with M2-type induction medium containing IL-4 (20 ng/ml) and IL-10 (20 ng/ml), and the third group was added with the medium for culturing Panc02 cells for 48 h. After mixing, the cells were placed in the cell culture incubator for 48 h. Marco expression, M1 markers (Cd86), and M2 markers (Cd163, Cd206) by macrophages were then measured by flow cytometry analysis, as previously described85,86.
PhenoCycler-Fusion and image analysis
PhenoCycler-Fusion was performed in a sequential order on 5-μm-thick sections from formalin-fixed paraffin-embedded PDAC tissues (including primary PDAC tissues N = 17, and metastatic PDAC tissues N = 14) or control tissues (including normal pancreatic tissue N = 3, pancreatitis N = 4, and para-cancer N = 8). One three-marker panel and one 5-marker panel were analyzed. Macrophage and NK cell panel (CD68, MARCO, and CD56) and T cells and tumor cell panel (CD4, CD8, FoxP3, Ki-67, and PanCK) were included in the present study.
First, the paraffin section was adhered to the central area patch of the slide with an anti-debonding slide, and the slides were baked at 65 °C for 1 h, dewaxed and rehydrated, and antigenically repaired. The desired primary antibody premix was then dropped onto the slide and incubated for 3 h at room temperature without drying out.
Image analysis was as described. Briefly, images were visually evaluated to exclude areas with necrosis or staining artefacts. Then, tissue segmentation based on manual annotation of two selected regions (tumor, and stroma) was performed using the defined software algorithm. Nuclear dye was used to guide the detection of FoxP3 and Ki-67, while the cytoplasmic regions were used to evaluate the expression of MARCO, CD68, CD4, CD8, CD56, and PanCK.
Construction of a subcutaneous PDAC model in mice, tumor dissociation and Bulk-RNA-seq
WT or Marco-/- mice with 8- to 12-week-old were injected subcutaneously (s.c.) in the right flank with 2 × 106 Panc02 cells in 100 µL of PBS together with 150 µL of Matrigel (Corning; concentration 8 to 12 mg/ml). On day 6, mouse PDAC subcutaneous tumors began to appear, and subsequent measurements of mouse tumor volume were performed every 4 days and measured in a manner consistent with our previous publication74. Mice were executed at day 34 using isoflurane anesthesia, and tumor volume and weight were recorded. Then, the tumor tissues from both WT and Marco-/- mice were separated. Blood impurities were rinsed off with pre-cooled PBS twice. The tissues were dried with dust-free absorbent paper and then placed in a freezing tube, plunged into liquid nitrogen for 3 min of quick-freezing, removed, and placed at −80 °C for storage. Afterwards, total RNA from each tissue was extracted, and the concentration and integrity of the sample RNA was detected. Library construction was performed using the MGIEasy RNA Library Preparation Reagent Kit, and high-throughput sequencing was performed using the MGI DNBSEQ-T7 sequencing platform with a sequencing read length of PE150. The method of analysis was consistent with what we previously reported74.
Statistics
All statistical analyses were performed using GraphPad Prism 9.0 (GraphPad Software, Inc.) and SPSS® 24.0. The experiments were repeated at least three times. Statistical significance was determined using Student’s t-test. The enumeration data are expressed as a percentage, and the measurement data are expressed as a mean ± standard deviation (SD) or mean ± standard error of the mean (SEM). Values of ***P < 0.001, **P < 0.01, and *P < 0.05 were considered statistically significant.
Data availability
All data generated in the study are available in the present article, supplementary information, and supplementary data.
Code availability
The code supporting this study is not publicly available but will be made available by the corresponding author upon reasonable request.
References
Siegel, R. L., Giaquinto, A. N. & Jemal, A. Cancer statistics, 2024. CA Cancer J. Clin. 74, 12–49 (2024).
Calheiros, J. et al. A first-in-class inhibitor of homologous recombination DNA repair counteracts tumour growth, metastasis and therapeutic resistance in pancreatic cancer. J. Exp. Clin. Cancer Res 44, 129 (2025).
Siegel, R. L., Miller, K. D., Wagle, N. S. & Jemal, A. Cancer statistics, 2023. CA Cancer J. Clin. 73, 17–48 (2023).
Wainberg, Z. A. et al. NALIRIFOX versus nab-paclitaxel and gemcitabine in treatment-naive patients with metastatic pancreatic ductal adenocarcinoma (NAPOLI 3): a randomised, open-label, phase 3 trial. Lancet 402, 1272–1281 (2023).
Zou, X. et al. Characterization of intratumoral tertiary lymphoid structures in pancreatic ductal adenocarcinoma: cellular properties and prognostic significance. J. Immunother. Cancer 11, e006698 (2023).
Bear, A. S., Vonderheide, R. H. & O’Hara, M. H. Challenges and opportunities for pancreatic cancer immunotherapy. Cancer Cell 38, 788–802 (2020).
Balachandran, V. P. et al. Identification of unique neoantigen qualities in long-term survivors of pancreatic cancer. Nature 551, 512–516 (2017).
Quaranta, V. et al. Macrophage-derived granulin drives resistance to immune checkpoint inhibition in metastatic pancreatic cancer. Cancer Res. 78, 4253–4269 (2018).
Zhou, J. et al. A novel role of TGFBI in macrophage polarization and macrophage-induced pancreatic cancer growth and therapeutic resistance. Cancer Lett. 578, 216457 (2023).
Bernard, V. et al. Single-cell transcriptomics of pancreatic cancer precursors demonstrates epithelial and microenvironmental heterogeneity as an early event in neoplastic progression. Clin. Cancer Res. 25, 2194–2205 (2019).
Caronni, N. et al. IL-1β+ macrophages fuel pathogenic inflammation in pancreatic cancer. Nature 623, 415–422 (2023).
Olaniru, O. E. et al. Single-cell transcriptomic and spatial landscapes of the developing human pancreas. Cell Metab. 35, 184–199.e5 (2023).
Li, W. et al. Identification and prognostic analysis of biomarkers to predict the progression of pancreatic cancer patients. Mol. Med. 28, 43 (2022).
Hosein, A. N. et al. Cellular heterogeneity during mouse pancreatic ductal adenocarcinoma progression at single-cell resolution. JCI Insight 5, e129212 (2019).
Cui Zhou, D. et al. Spatially restricted drivers and transitional cell populations cooperate with the microenvironment in untreated and chemo-resistant pancreatic cancer. Nat. Genet. 54, 1390–1405 (2022).
Li, W., Wang, F., Guo, R., Bian, Z. & Song, Y. Targeting macrophages in hematological malignancies: recent advances and future directions. J. Hematol. Oncol. 15, 110 (2022).
Liu, M., Liu, L., Song, Y., Li, W. & Xu, L. Targeting macrophages: a novel treatment strategy in solid tumors. J. Transl. Med 20, 586 (2022).
Eisinger, S. et al. Targeting a scavenger receptor on tumor-associated macrophages activates tumor cell killing by natural killer cells. Proc. Natl. Acad. Sci. USA 117, 32005–32016 (2020).
Georgoudaki, A.-M. et al. Reprogramming tumor-associated macrophages by antibody targeting inhibits cancer progression and metastasis. Cell Rep. 15, 2000–2011 (2016).
La Fleur, L. et al. Targeting MARCO and IL37R on immunosuppressive macrophages in lung cancer blocks regulatory T cells and supports cytotoxic lymphocyte function. Cancer Res. 81, 956–967 (2021).
Molgora, M. et al. IL-1R8 is a checkpoint in NK cells regulating anti-tumour and anti-viral activity. Nature 551, 110–114 (2017).
Sa, J. K. et al. Transcriptional regulatory networks of tumor-associated macrophages that drive malignancy in mesenchymal glioblastoma. Genome Biol. 21, 216 (2020).
Shi, B. et al. The scavenger receptor MARCO expressed by tumor-associated macrophages are highly associated with poor pancreatic cancer prognosis. Front Oncol. 11, 771488 (2021).
Zhang, S. et al. Single cell transcriptomic analyses implicate an immunosuppressive tumor microenvironment in pancreatic cancer liver metastasis. Nat. Commun. 14, 5123 (2023).
Werba, G. et al. Single-cell RNA sequencing reveals the effects of chemotherapy on human pancreatic adenocarcinoma and its tumor microenvironment. Nat. Commun. 14, 797 (2023).
Chen, K. et al. Immune profiling and prognostic model of pancreatic cancer using quantitative pathology and single-cell RNA sequencing. J. Transl. Med. 21, 210 (2023).
Chen, K. et al. Single-cell transcriptome profiling of primary tumors and paired organoids of pancreatobiliary cancer. Cancer Lett. 582, 216586 (2024).
Oh, K. et al. Coordinated single-cell tumor microenvironment dynamics reinforce pancreatic cancer subtype. Nat. Commun. 14, 5226 (2023).
Storrs, E. P. et al. High-dimensional deconstruction of pancreatic cancer identifies tumor microenvironmental and developmental stemness features that predict survival. NPJ Precis. Oncol. 7, 105 (2023).
Xu, L. et al. TIP: a web server for resolving tumor immunophenotype profiling. Cancer Res. 78, 6575–6580 (2018).
Jiang, P. et al. Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat. Med 24, 1550–1558 (2018).
Song, G. et al. Single-cell transcriptomic analysis suggests two molecularly subtypes of intrahepatic cholangiocarcinoma. Nat. Commun. 13, 1642 (2022).
Zhang, J. et al. Single-cell transcriptome sequencing reveals aberrantly activated inter-tumor cell signaling pathways in the development of clear cell renal cell carcinoma. J. Transl. Med. 22, 37 (2024).
Chen, G. et al. Single-cell transcriptomic analysis reveals that the APP-CD74 axis promotes immunosuppression and progression of testicular tumors. J. Pathol. 264, 250–269 (2024).
Wheeler, K. C. et al. VEGF may contribute to macrophage recruitment and M2 polarization in the decidua. PLoS One 13, e0191040 (2018).
Wang, S. et al. Targeting M2-like tumor-associated macrophages is a potential therapeutic approach to overcome antitumor drug resistance. NPJ Precis. Oncol. 8, 31 (2024).
Zhou, W.-H. et al. The overexpression of fibronectin 1 promotes cancer progression and associated with M2 macrophages polarization in head and neck squamous cell carcinoma patients. Int. J. Gen. Med. 15, 5027–5042 (2022).
Gao, Y., Dai, W., Ouyang, Z., Shen, M. & Shi, X. Dendrimer-mediated intracellular delivery of fibronectin guides macrophage polarization to alleviate acute lung injury. Biomacromolecules 24, 886–895 (2023).
Matsubara, E. et al. The significance of SPP1 in lung cancers and its impact as a marker for protumor tumor-associated macrophages. Cancers 15, 2250 (2023).
Simons, M., Gordon, E. & Claesson-Welsh, L. Mechanisms and regulation of endothelial VEGF receptor signalling. Nat. Rev. Mol. Cell Biol. 17, 611–625 (2016).
Tchaikovski, V., Fellbrich, G. & Waltenberger, J. The molecular basis of VEGFR-1 signal transduction pathways in primary human monocytes. Arterioscler. Thromb. Vasc. Biol. 28, 322–328 (2008).
Lee, J. W., Komar, C. A., Bengsch, F., Graham, K. & Beatty, G. L. Genetically engineered mouse models of pancreatic cancer: the KPC model (LSL-Kras(G12D/+);LSL-Trp53(R172H/+);Pdx-1-Cre), its variants, and their application in immuno-oncology drug discovery. Curr. Protoc. Pharm. 73, 14.39.1–14.39.20 (2016).
Cassetta, L. & Pollard, J. W. Targeting macrophages: therapeutic approaches in cancer. Nat. Rev. Drug Discov. 17, 887–904 (2018).
Ho, W. J., Jaffee, E. M. & Zheng, L. The tumour microenvironment in pancreatic cancer—clinical challenges and opportunities. Nat. Rev. Clin. Oncol. 17, 527–540 (2020).
Griesmann, H. et al. Pharmacological macrophage inhibition decreases metastasis formation in a genetic model of pancreatic cancer. Gut 66, 1278–1285 (2017).
Nielsen, S. R. et al. Macrophage-secreted granulin supports pancreatic cancer metastasis by inducing liver fibrosis. Nat. Cell Biol. 18, 549–560 (2016).
Mass, E., Nimmerjahn, F., Kierdorf, K. & Schlitzer, A. Tissue-specific macrophages: how they develop and choreograph tissue biology. Nat. Rev. Immunol. 23, 563–579 (2023).
Nasir, I. et al. Tumor macrophage functional heterogeneity can inform the development of novel cancer therapies. Trends Immunol. 44, 971–985 (2023).
Revel, M., Sautès-Fridman, C., Fridman, W.-H. & Roumenina, L. T. C1q+ macrophages: passengers or drivers of cancer progression. Trends Cancer 8, 517–526 (2022).
Chen, K. et al. Single cell RNA-seq identifies immune-related prognostic model and key signature-SPP1 in pancreatic ductal adenocarcinoma. Genes 13, 1760 (2022).
Yang, J. et al. The enhancement of glycolysis regulates pancreatic cancer metastasis. Cell Mol. Life Sci. 77, 305–321 (2020).
Liu, Z.-L., Chen, H.-H., Zheng, L.-L., Sun, L.-P. & Shi, L. Angiogenic signaling pathways and anti-angiogenic therapy for cancer. Signal Transduct. Target. Ther. 8, 198 (2023).
Curtarello, M. et al. VEGF-targeted therapy stably modulates the glycolytic phenotype of tumor cells. Cancer Res 75, 120–133 (2015).
Lorestani, P. et al. The complex role of macrophages in pancreatic cancer tumor microenvironment: a review on cancer progression and potential therapeutic targets. Discov. Oncol. 15, 369 (2024).
Chen, A. X. et al. Single-cell characterization of macrophages in glioblastoma reveals MARCO as a mesenchymal pro-tumor marker. Genome Med. 13, 88 (2021).
Farhangnia, P., Khorramdelazad, H., Nickho, H. & Delbandi, A.-A. Current and future immunotherapeutic approaches in pancreatic cancer treatment. J. Hematol. Oncol. 17, 40 (2024).
Luo, W., Wen, T. & Qu, X. Tumor immune microenvironment-based therapies in pancreatic ductal adenocarcinoma: time to update the concept. J. Exp. Clin. Cancer Res. 43, 8 (2024).
Joseph, A. M., Al Aiyan, A., Al-Ramadi, B., Singh, S. K. & Kishore, U. Innate and adaptive immune-directed tumour microenvironment in pancreatic ductal adenocarcinoma. Front. Immunol. 15, 1323198 (2024).
Karamitopoulou, E. Tumour microenvironment of pancreatic cancer: immune landscape is dictated by molecular and histopathological features. Br. J. Cancer 121, 5–14 (2019).
Yang, D. et al. Comprehensive analysis of scRNA-Seq and bulk RNA-Seq data reveals dynamic changes in tumor-associated neutrophils in the tumor microenvironment of hepatocellular carcinoma and leads to the establishment of a neutrophil-related prognostic model. Cancer Immunol. Immunother. 72, 4323–4335 (2023).
Wang, Y. et al. Identification of potential immune-related mechanisms related to the development of multiple myeloma. Chin. Med. J. 137, 1603–1613 (2024).
Huo, X. et al. Unravelling the role of immune cells and FN1 in the recurrence and therapeutic process of skull base chordoma. Clin. Transl. Med. 13, e1429 (2023).
Sun, Y. et al. Immunometabolic changes and potential biomarkers in CFS peripheral immune cells revealed by single-cell RNA sequencing. J. Transl. Med. 22, 925 (2024).
Ginhoux, F. & Jung, S. Monocytes and macrophages: developmental pathways and tissue homeostasis. Nat. Rev. Immunol. 14, 392–404 (2014).
Liu, Z. et al. Fate mapping via Ms4a3-expression history traces monocyte-derived cells. Cell 178, 1509–1525.e19 (2019).
Mantovani, A. et al. Chemokines in the recruitment and shaping of the leukocyte infiltrate of tumors. Semin Cancer Biol. 14, 155–160 (2004).
Robinson, A., Han, C. Z., Glass, C. K. & Pollard, J. W. Monocyte regulation in homeostasis and malignancy. Trends Immunol. 42, 104–119 (2021).
Qian, B.-Z. et al. FLT1 signaling in metastasis-associated macrophages activates an inflammatory signature that promotes breast cancer metastasis. J. Exp. Med. 212, 1433–1448 (2015).
Dong, Q. et al. MARCO is a potential prognostic and immunotherapy biomarker. Int Immunopharmacol. 116, 109783 (2023).
La Fleur, L. et al. Expression of scavenger receptor MARCO defines a targetable tumor-associated macrophage subset in non-small cell lung cancer. Int J. Cancer 143, 1741–1752 (2018).
Fan, G. et al. The co-location of MARCO+ tumor-associated macrophages and CTSE+ tumor cells determined the poor prognosis in intrahepatic cholangiocarcinoma. Hepatology 82, 25–41 (2025).
Ding, L. et al. Blocking MARCO+ tumor-associated macrophages improves anti-PD-L1 therapy of hepatocellular carcinoma by promoting the activation of STING-IFN type I pathway. Cancer Lett. 582, 216568 (2024).
Sun, H. et al. Association of decreased expression of the macrophage scavenger receptor MARCO with tumor progression and poor prognosis in human hepatocellular carcinoma. J. Gastroenterol. Hepatol. 32, 1107–1114 (2017).
Zhang, B. et al. Development and evaluation of a human CD47/HER2 bispecific antibody for Trastuzumab-resistant breast cancer immunotherapy. Drug Resist. Updates 74, 101068 (2024).
Li, W. et al. Identification of potential resistance mechanisms and therapeutic targets for the relapse of BCMA CAR-T therapy in relapsed/refractory multiple myeloma through single-cell sequencing. Exp. Hematol. Oncol. 12, 44 (2023).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Tran, H. T. N. et al. A benchmark of batch-effect correction methods for single-cell RNA sequencing data. Genome Biol. 21, 12 (2020).
Wang, Z. et al. An immune cell atlas reveals the dynamics of human macrophage specification during prenatal development. Cell 186, 4454–4471.e19 (2023).
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinform. 14, 7 (2013).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Gulati, G. S. et al. Single-cell transcriptional diversity is a hallmark of developmental potential. Science 367, 405–411 (2020).
Petitprez, F. et al. The murine Microenvironment Cell Population counter method to estimate abundance of tissue-infiltrating immune and stromal cell populations in murine samples using gene expression. Genome Med. 12, 86 (2020).
Xiao, Z., Dai, Z. & Locasale, J. W. Metabolic landscape of the tumor microenvironment at single cell resolution. Nat. Commun. 10, 3763 (2019).
Auger, J.-P. et al. Metabolic rewiring promotes anti-inflammatory effects of glucocorticoids. Nature 629, 184–192 (2024).
Li, W. et al. Identification and transcriptome analysis of erythroblastic island macrophages. Blood 134, 480–491 (2019).
Wang, Y. et al. Impairment of human terminal erythroid differentiation by histone deacetylase 5 deficiency. Blood 138, 1615–1627 (2021).
Zhang, Y. et al. Glucocorticoids increase adiposity by stimulating Krüppel-like factor 9 expression in macrophages. Nat. Commun. 15, 1190 (2024).
Acknowledgements
We would like to thank all our authors listed in this manuscript. This work was supported by Natural Science Foundation of China (82400233 (B.Z.), 82270149 (W.L.), 82370144 (Y.S.)); Henan Province Medical Science and Technology Research Project (SBGJ202002087(H.S.), LHGL20240207 (B.Z.)); The Natural Science Foundation of Henan Province (242300421080 (W.L.)); Leading talent project of Henan Province (LJRC2023004, (L.X.)); Postdoctoral Innovative Talent Support Program (GZC20232429(B.Z.)); Henan Provincial Key R&D Program (251111312100 (H.S.)). Funding for Scientific Research and Innovation Team of The First Affiliated Hospital of Zhengzhou University.
Author information
Authors and Affiliations
Contributions
H.S., M.G., Z.L., and Z.Z. performed experiments, and analyzed the data. B.Z. and S.Z. drafted manuscript. Z.L., Z.Z., T.L., and J.S. analyzed the ScRNA-seq data and bulk-RNA-seq data. H.H., X.L., J.Y., M.S., M.L., Y.A., S.Y., Y.L., Z.H., Y.H., Y.L., C.L., M.L., and M.Y. performed experiments. J.C., Y.S., J.M., M.L., Z.B., and W.L. read and edited the manuscript. L.X., S.Z., B.Z., and G.W. designed and supervised the study and edited the manuscript. All authors approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sun, H., Gao, M., Liu, Z. et al. Identification and characterization of MARCO-expressing tumor-associated macrophages in pancreatic ductal adenocarcinoma with pan-cancer relevance. npj Precis. Onc. 10, 78 (2026). https://doi.org/10.1038/s41698-026-01293-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41698-026-01293-5