Abstract
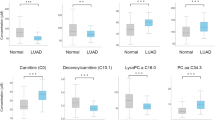
Lung adenocarcinoma (LUAD) remains a leading cause of cancer-related mortality worldwide, highlighting the urgent need for non-invasive strategies for early detection. Here, we present a machine learning-assisted metabolomics approach for the early detection of LUAD. Untargeted metabolomic profiling was performed on 199 serum samples from healthy individuals, patients with lung precancerous lesions, and those with stage I LUAD. An ensemble machine learning workflow was developed to identify metabolite panels capable of discriminating clinical status with high accuracy. We observed progressive metabolic alterations in bile acid, lipid, amino acid, and purine metabolism during LUAD initiation and stepwise progression. Notably, ensemble learning identified a six-metabolite panel, including 12-hydroxydodecanoic acid, hypoxanthine, xanthosine, cholic acid, agmatine, and paraxanthine, for accurate detection of early-stage LUAD, and a distinct four-metabolite panel, comprising 7-α,27-dihydroxycholesterol, 11-undecanedicarboxylic acid, biliverdin, and Prolyl-Valine, for precise differentiation between pre-invasive and invasive lesions. Both panels demonstrated promising diagnostic potential, with performance metrices comparing favorably to established methodologies within the current study cohort. This study delineates the evolutionary trajectory of the serum metabolome associated with early LUAD pathogenesis and provides promising biomarkers for non-invasive early detection.
Similar content being viewed by others
Data availability
All the data and materials that support the findings of this study are available within the article and Supplementary Information or available from the authors upon request.
References
Siegel, R. L. et al. Cancer statistics. CA Cancer J. Clin. 72, 7–33 (2022).
Cheng, T. Y. et al. The international epidemiology of lung cancer: latest trends, disparities, and tumor characteristics. J. Thorac. Oncol. 11, 1653–1671 (2016).
Nair, A. et al. Variable radiological lung nodule evaluation leads to divergent management recommendations. Eur. Respir. J. 52, 1801359 (2018).
Vachani, A. et al. Factors that influence physician decision making for indeterminate pulmonary nodules. Ann. Am. Thorac. Soc. 11, 1586–1591 (2014).
Crowley, E. et al. Liquid biopsy: monitoring cancer-genetics in the blood. Nat. Rev. Clin. Oncol. 10, 472–84 (2013).
Fedyuk, V. et al. Multiplexed, single-molecule, epigenetic analysis of plasma-isolated nucleosomes for cancer diagnostics. Nat. Biotechnol. 41, 212–221 (2023).
Gardner, L. et al. Nano-omics: nanotechnology-based multidimensional harvesting of the blood-circulating cancerome. Nat. Rev. Clin. Oncol. 19, 551–561 (2022).
Hu, A. et al. Cancer serum atlas-supported precise pan-targeted proteomics enable multicancer detection. Anal. Chem. 95, 862–871 (2023).
Buergel, T. et al. Metabolomic profiles predict individual multidisease outcomes. Nat. Med. 28, 2309–2320 (2022).
Chen, F. et al. Integrated analysis of the faecal metagenome and serum metabolome reveals the role of gut microbiome-associated metabolites in the detection of colorectal cancer and adenoma. Gut 71, 1315–1325 (2022).
Yi, R. et al. Multi-omic profiling of multi-biosamples reveals the role of amino acid and nucleotide metabolism in endometrial cancer. Front. Oncol. 12, 861142 (2022).
Sinclair, E. et al. Metabolomics of sebum reveals lipid dysregulation in Parkinson’s disease. Nat. Commun. 12, 1592 (2021).
Wang, Y. et al. Self-assembled hyperbranched gold nanoarrays decode serum united urine metabolic fingerprints for kidney tumor diagnosis. ACS Nano 18, 2409–2420 (2024).
Wang, G. et al. Lung cancer Scrna-Seq and lipidomics reveal aberrant lipid metabolism for early-stage diagnosis. Sci. Transl. Med. 14, eabk2756 (2022).
You, L. et al. Liquid chromatography-mass spectrometry-based tissue metabolic profiling reveals major metabolic pathway alterations and potential biomarkers of lung cancer. J. Proteome Res. 19, 3750–3760 (2020).
Mathé, E. A. et al. Noninvasive urinary metabolomic profiling identifies diagnostic and prognostic markers in lung cancer. Cancer Res. 74, 3259–3270 (2014).
Schult, T. A. et al. Screening human lung cancer with predictive models of serum magnetic resonance spectroscopy metabolomics. Proc. Natl. Acad. Sci. USA 118, e2110633118 (2021).
Shestakova, K. M. et al. Targeted metabolomic profiling as a tool for diagnostics of patients with non-small-cell lung cancer. Sci. Rep. 13, 11072 (2023).
Wen, T. et al. Exploratory investigation of plasma metabolomics in human lung adenocarcinoma. Mol. Biosyst. 9, 2370–2378 (2013).
Li, J. et al. Serum untargeted metabolomics reveal metabolic alteration of non-small cell lung cancer and refine disease detection. Cancer Sci. 114, 680–689 (2023).
Sun, T. et al. Lipidomics reveals new lipid-based lung adenocarcinoma early diagnosis model. EMBO Mol. Med. 16, 854–869 (2024).
Nie, M. et al. Evolutionary metabolic landscape from preneoplasia to invasive lung adenocarcinoma. Nat. Commun. 12, 6479 (2021).
Wang, L. et al. Integrative serum metabolic fingerprints based multi-modal platforms for lung adenocarcinoma early detection and pulmonary nodule classification. Adv. Sci. 9, e2203786 (2022).
Yao, Y. et al. Metabolomic differentiation of benign vs malignant pulmonary nodules with high specificity via high-resolution mass spectrometry analysis of patient sera. Nat. Commun. 14, 2339 (2023).
Huang, L. et al. Machine learning of serum metabolic patterns encodes early-stage lung adenocarcinoma. Nat. Commun. 11, 3556 (2020).
Zheng, R. et al. Machine learning-based integrated multiomics characterization of colorectal cancer reveals distinctive metabolic signatures. Anal. Chem. 96, 8772–8781 (2024).
Odenkirk, M. T. et al. Multiomic big data analysis challenges: increasing confidence in the interpretation of artificial intelligence assessments. Anal. Chem. 93, 7763–7773 (2021).
Asef, C. K. et al. Unknown metabolite identification using machine learning collision cross-section prediction and tandem mass spectrometry. Anal. Chem. 95, 1047–1056 (2023).
Malta, T. M. et al. Machine learning identifies stemness features associated with oncogenic dedifferentiation. Cell 173, 338–354.e15 (2018).
Konno, N. et al. Machine learning enables prediction of metabolic system evolution in bacteria. Sci. Adv. 9, eadc9130 (2023).
Greener, J. G. et al. A guide to machine learning for biologists. Nat. Rev. Mol. Cell. Biol. 23, 40–55 (2022).
Chen, J. et al. Machine learning aids classification and discrimination of noncanonical DNA folding motifs by an arrayed host: guest sensing system. J. Am. Chem. Soc. 143, 12791–12799 (2021).
Chen, R. J. et al. Synthetic data in machine learning for medicine and healthcare. Nat. Biomed. Eng. 5, 493–497 (2021).
Dong, X. et al. A survey on ensemble learning. Front. Comput. Sci. 14, 241–58 (2020).
Heidari, B. M. et al. Culture-free identification and metabolic profiling of microalgal single cells via ensemble learning of ramanomes. Anal. Chem. 93, 8872–8880 (2021).
Janizek, J. D. et al. Uncovering expression signatures of synergistic drug responses via ensembles of explainable machine-learning models. Nat. Biomed. Eng. 7, 811–829 (2023).
Cao, Y. et al. Ensemble deep learning in bioinformatics. Nat. Mach. Intell. 2, 500–508 (2020).
Arnaout, R. et al. An ensemble of neural networks provides expert-level prenatal detection of complex congenital heart disease. Nat. Med. 27, 882–891 (2021).
Canesin, G. et al. Heme-derived metabolic signals dictate immune responses. Front. Immunol. 11, 66 (2020).
Qian, X. et al. Integrated microbiome, metabolome, and proteome analysis identifies a novel interplay among commensal bacteria, metabolites and candidate targets in non-small cell lung cancer. Clin. Transl. Med. 12, e947 (2022).
Lv, M. et al. Plasma lipidomics profiling to identify the biomarkers of diagnosis and radiotherapy response for advanced non-small-cell lung cancer patients. J. Lipids 2024, 6730504 (2024).
Chen, L. et al. 25-Hydroxycholesterol promotes migration and invasion of lung adenocarcinoma cells. Biochem. Biophys. Res. Commun. 484, 857–863 (2017).
Musial, C. et al. Induction of 2-hydroxycatecholestrogens O-methylation: a missing puzzle piece in diagnostics and treatment of lung cancer. Redox. Biol. 55, 102395 (2022).
Chen, X. et al. Whole-lesion computed tomography-based entropy parameters for the differentiation of minimally invasive and invasive adenocarcinomas appearing as pulmonary subsolid nodules. J. Comput. Assist. Tomogr. 43, 817–824 (2019).
Zhang, J. et al. Why do pathological stage IA lung adenocarcinomas vary from prognosis?: a clinicopathologic study of 176 patients with pathological stage IA lung adenocarcinoma based on the IASLC/ATS/ERS classification. J. Thorac. Oncol. 8, 1196–202 (2013).
Altorki, N. K. et al. Sublobar resection is equivalent to lobectomy for clinical stage 1A lung cancer in solid nodules. J. Thorac. Cardiovasc. Surg. 147, 754–762 (2014).
Huang, Z. et al. From purines to purinergic signalling: molecular functions and human diseases. Signal. Transduct. Target. Ther. 6, 162 (2021).
Ma, C. et al. Gut microbiome-mediated bile acid metabolism regulates liver cancer via NKT cells. Science 360, eaan5931 (2018).
Martin-Perez, M. et al. The role of lipids in cancer progression and metastasis. Cell Metab. 34, 1675–1699 (2022).
Arndt, M. A. et al. The arginine metabolite agmatine protects mitochondrial function and confers resistance to cellular apoptosis. Am. J. Physiol. Cell Physiol. 296, C1411–9 (2009).
Dong, B. et al. Plasma proteometabolome in lung cancer: exploring biomarkers through bidirectional mendelian randomization and colocalization analysis. Hum. Mol. Genet. 33, 1688–1696 (2024).
Wang, C. et al. Multi-omics analyses reveal biological and clinical insights in recurrent stage I non-small cell lung cancer. Nat. Commun. 16, 1477 (2025).
Zhang, Y. et al. Evolutionary proteogenomic landscape from pre-invasive to invasive lung adenocarcinoma. Cell Rep. Med. 5, 101358 (2024).
Nicholson, A. G. et al. The 2021 WHO classification of lung tumors: impact of advances since 2015. J. Thorac. Oncol. 17, 362–387 (2022).
Li, R. et al. Deep learning applications in computed tomography images for pulmonary nodule detection and diagnosis: a review. Diagnostics 12, 298 (2022).
Wang, S. et al. Tumor evolutionary trajectories during the acquisition of invasiveness in early stage lung adenocarcinoma. Nat. Commun. 11, 6083 (2020).
Chen, Y. C. et al. Multiomics analysis reveals molecular changes during early progression of precancerous lesions to lung adenocarcinoma in never-smokers. Cancer Res. 85, 602–617 (2025).
Acknowledgements
This study was supported by the National Natural Science Foundation of China (82404096), the Science and Technology Commission of Shanghai Municipality (24Y12800300), and the National Key Clinical Specialty Discipline Construction Program of China: Establishment and Application of a Precision Diagnosis and Treatment System for Chest Tumors.
Author information
Authors and Affiliations
Contributions
C.C. and S.R. designed the study. Data collection was carried out by C.C., W.X., and L.W. Statistical analysis and graph organization were made by C.C. and W.X. The initial manuscript was written by C.C., W.X., L.W., S.Y., and J.Y. A manuscript review was made by C.C., W.X., L.W., and S.R.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Cai, C., Xu, W., Yang, S. et al. Ensemble learning on serum metabolic fingerprints for early detection of lung adenocarcinoma. npj Precis. Onc. (2026). https://doi.org/10.1038/s41698-026-01342-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41698-026-01342-z