Abstract
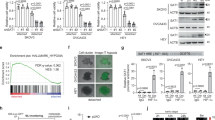
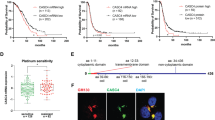
Peritoneal dissemination is the major cause of mortality in epithelial ovarian cancer (EOC) and requires tumor cells to survive in a detached state by evading anoikis. However, the molecular mechanisms supporting anchorage-independent survival remain poorly defined. Here, we identify glutamate dehydrogenase 1 (GLUD1) as a key regulator of anoikis resistance and metastatic progression in EOC. GLUD1 expression was elevated in metastatic EOC tissues and associated with unfavorable clinical outcomes. Loss of GLUD1 impaired anoikis resistance and reduced metastatic capacity of ovarian cancer cells in vitro, while markedly suppressing peritoneal dissemination and prolonging survival in vivo. Mechanistically, GLUD1 was found to interact with a key protein ARAF, the A-Raf proto-oncogene. By limiting ubiquitin–proteasome-mediated degradation of ARAF, GLUD1 exerted a non-enzymatic function that stabilized ARAF protein levels and sustained MEK/ERK signaling.
Together, these findings reveal a non-canonical role of GLUD1 in regulating protein stability and identify the GLUD1–ARAF axis as a critical mechanism supporting anchorage-independent survival during peritoneal dissemination of EOC.
Similar content being viewed by others
Data availability
All data supporting the findings of this study are presented in the article and the Supplementary Materials. Our study does not generate any new datasets. The datasets presented in this study can be found from the following links: TCGA (https://portal.gdc.cancer.gov/), GSE26712, GSE27651 GSE40595. All other raw data are available upon reasonable request from the corresponding author.
Code availability
The methods described in this manuscript have been used the public available limma package in R (v3.46.0) for differential gene expression analysis. No custom algorithm generated in this manuscript.
References
Cabasag, C. J. et al. Ovarian cancer today and tomorrow: A global assessment by world region and human development index using GLOBOCAN 2020. Int. J. Cancer 151, 1535–1541 (2022).
Lin, S.-C. et al. Periostin promotes ovarian cancer metastasis by enhancing M2 macrophages and cancer-associated fibroblasts via integrin-mediated NF-κB and TGF-β2 signaling. J. Biomed. Sci. 29, 109 (2022).
Saha, M. et al. Sustained AMPK activation and proline metabolism play critical roles in the survival of matrix-deprived transformed cells. Front. Cell Dev. Biol. 9, 771366 (2021).
Davison, C. A. et al. Antioxidant enzymes mediate survival of breast cancer cells deprived of extracellular matrix. Cancer Res. 73, 3704–3715 (2013).
Shaw, P. et al. Anoikis resistance in cancer: Mechanisms, therapeutic strategies, potential targets, and models for enhanced understanding. Cancer Lett. 624, 217750 (2025).
Endo, H. et al. Metabolic reprogramming sustains cancer cell survival following extracellular matrix detachment. Redox Biol. 36, 101643 (2020).
Zhu, C. et al. Adipose-derived stem cells promote glycolysis and peritoneal metastasis via TGF-β1/SMAD3/ANGPTL4 axis in colorectal cancer. Cell. Mol. Life Sci. 81, 189 (2024).
Li, S. et al. Obesity promotes gastric cancer metastasis via diacylglycerol acyltransferase 2–dependent lipid droplets accumulation and redox homeostasis. Redox Biol. 36, 101596 (2020).
Sullivan, L. B. et al. Supporting aspartate biosynthesis is an essential function of respiration in proliferating cells. Cell 162, 552–563 (2015).
Altman, B. J. et al. From Krebs to clinic: Glutamine metabolism to cancer therapy. Nat. Rev. Cancer 16, 749–764 (2016).
Jin, L. et al. The PLAG1–GDH1 axis promotes anoikis resistance and tumor metastasis through CamKK2–AMPK signaling in LKB1-deficient lung cancer. Mol. Cell 69, 87–99.e7 (2018).
Yang, R. et al. EGFR activates GDH1 transcription to promote glutamine metabolism through the MEK/ERK/ELK1 pathway in glioblastoma. Oncogene 39, 2975–2986 (2020).
Wang, X. et al. α-Ketoglutarate-activated NF-κB signaling promotes compensatory glucose uptake and brain tumor development. Mol. Cell 76, 148–162.e7 (2019).
Yang, R. et al. GDH1-catalytic glutaminolysis feedback activates EGFR/PI3K/AKT pathway and reprograms glioblastoma metabolism. Neuro-Oncol. 26, noae222 (2024).
Kang, J. et al. EGFR-phosphorylated GDH1 harmonizes with RSK2 to drive CREB activation and tumor metastasis in EGFR-activated lung cancer. Cell Rep. 41, 111827 (2022).
Hu, Q. et al. Deacetylation of GLUD1 maintains survival of lung adenocarcinoma cells under glucose starvation by inhibiting autophagic cell death. Cell Insight 3, 100186 (2024).
Mooz, J. et al. ARAF suppresses ERBB3 expression and metastasis in a subset of lung cancers. Sci. Adv. 8, eabk1538 (2022).
Yaeger, R. & Corcoran, R. B. Targeting alterations in the RAF–MEK pathway. Cancer Discov. 9, 329–341 (2019).
Karoulia, Z. et al. New perspectives for targeting RAF kinase in human cancer. Nat. Rev. Cancer 17, 676–691 (2017).
Grbovic, O. M. et al. V600E B-RAF requires the HSP90 chaperone for stability and is degraded in response to HSP90 inhibitors. Proc. Natl. Acad. Sci. 103, 57–62 (2006).
Rebocho, A. P. & Marais, R. ARAF acts as a scaffold to stabilize BRAF–CRAF heterodimers. Oncogene 32, 3207–3212 (2013).
Venkatanarayan, A. CRAF dimerization with ARAF regulates KRAS-driven tumor growth. Cell Rep. 38, 110351 (2022).
Riegel, K. et al. RAF kinases are stabilized and required for dendritic cell differentiation and function. Cell Death Differ. 27, 1300–1315 (2020).
Kauko, O. et al. PP2A inhibition is a druggable MEK inhibitor resistance mechanism in KRAS-mutant lung cancer cells. Sci Transl Med. 10, eaaq1093 (2018).
Stelzer, G. et al. The GeneCards suite: From gene data mining to disease genome sequence analyses. Curr. Protoc. Bioinforma. 54, 1.30.1–1.30.33 (2016).
Tian, Q. et al. RICH1 inhibits breast cancer stem cell traits through activating the Hippo signaling kinase cascade. Cell Death Dis. 13, 71 (2022).
Charan, J. & Kantharia, N. D. How to calculate sample size in animal studies? J. Pharmacol. Pharmacother. 4, 303–306 (2013).
Li, Z. et al. Silencing GRHL3 promotes distant metastasis of lung squamous cell carcinoma by enhancing SOX2 stability via SIRT1. J. Pathol. 265, 302–315 (2025).
Xiong, L. et al. Glutamic-pyruvic transaminase 1 deficiency-mediated metabolic reprogramming facilitates colorectal adenoma–carcinoma progression. Sci. Transl. Med. 17, eadp9805 (2025).
Acknowledgements
This work was also supported by the Guangdong Basic and Applied Basic Research Foundation (2025A1515012496, 2022A1515010243), Guangzhou Municipal Science and Technology Project (2024A04J3475), College Students' Innovative Entrepreneurial Training Plan Program (S202510571044) and Special Funds for the Cultivation of Guangdong College Students' Scientific and Technological Innovation(No.pdjh2025bk102). We would like to express our gratitude to these funding sources for their support.
Author information
Authors and Affiliations
Contributions
L.C., H.F., and G.W. designed experiments. Z.Z. performed the bioinformatics analysis. Y.C., H.L., Y.L., S.Z., Y.L., and C.Y. performed the experiments and analyzed and interpreted the data. H.Y., S.W., and L.C. supported and supervised the research. L.C. and H.F. wrote the manuscript, and all authors read and approved the final manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Feng, H., Chen, Y., Wu, G. et al. GLUD1 supports ovarian cancer progression by counteracting anoikis via ARAF/MEK/ERK signaling. npj Precis. Onc. (2026). https://doi.org/10.1038/s41698-026-01349-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41698-026-01349-6