Abstract
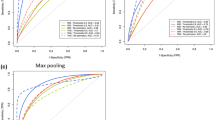
The differentiation between pathological subtypes of non-small cell lung cancer (NSCLC) is an essential step in guiding treatment options and prognosis. However, current clinical practice relies on multi-step staining and labelling processes that are time-intensive and costly, requiring highly specialised expertise. In this study, we propose a label-free methodology that facilitates autofluorescence imaging of unstained NSCLC samples and deep learning (DL) techniques to distinguish between non-cancerous tissue, adenocarcinoma (AC), squamous cell carcinoma (SqCC), and other subtypes (OS). We conducted DL-based classification and generated virtual immunohistochemical (IHC) stains, including thyroid transcription factor-1 (TTF-1) for AC and p40 for SqCC. We evaluated these methods using two types of autofluorescence imaging: intensity imaging and lifetime imaging. The results demonstrate the exceptional ability of this approach for NSCLC subtype differentiation, achieving an area under the curve above 0.981 and 0.996 for binary- and multi-class classification. Furthermore, this approach produces clinical-grade virtual IHC staining, which was blind-evaluated by three experienced thoracic pathologists. Our label-free NSCLC subtyping approach enables rapid and accurate diagnosis without the need for conventional tissue processing and staining. Both strategies can significantly accelerate diagnostic workflows and support efficient lung cancer diagnosis, without compromising clinical decision-making.
Similar content being viewed by others
Data availability
The authors declare that all data supporting the results of this study are available in the paper and the Supplementary Information section.
Code availability
The TIMM library implementing DL models for subtyping is available at https://github.com/huggingface/pytorch-image-models. The pix2pix model utilised for virtual IHC staining is available at https://github.com/phillipi/pix2pix. The Style Loss function is available at https://pytorch.org/tutorials/advanced/neural_style_tutorial.html. FLIM images were stitched using Fiji MIST stitching plugin (https://github.com/usnistgov/MIST). MATLAB® was used for affine transformation to co-register FLIM and true histology images.
References
Bray, F. et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. Ca. Cancer J. Clin. 74, 229–263 (2024).
Navada, S., Lai, P., Schwartz, A. & Kalemkerian, G. Temporal trends in small cell lung cancer: analysis of the National Surveillance, Epidemiology, and End-Results (SEER) database. J. Clin. Oncol. 24, 7082 (2006).
Sher, T., Dy, G. K. & Adjei, A. A. Small cell lung cancer. In Mayo Clinic Proceedings Vol. 83, 355–367 (Elsevier, 2008).
Kim, H. S., Mitsudomi, T., Soo, R. A. & Cho, B. C. Personalized therapy on the horizon for squamous cell carcinoma of the lung. Lung Cancer 80, 249–255 (2013).
Coudray, N. et al. Classification and mutation prediction from non–small cell lung cancer histopathology images using deep learning. Nat. Med. 24, 1559–1567 (2018).
Noorbakhsh, J. et al. Deep learning-based cross-classifications reveal conserved spatial behaviors within tumor histological images. Nat. Commun. 11, 6367 (2020).
Chen, C.-L. et al. An annotation-free whole-slide training approach to pathological classification of lung cancer types using deep learning. Nat. Commun. 12, 1193 (2021).
Yu, K.-H. et al. Classifying non-small cell lung cancer types and transcriptomic subtypes using convolutional neural networks. J. Am. Med. Inform. Assoc. 27, 757–769 (2020).
Sadhwani, A. et al. Comparative analysis of machine learning approaches to classify tumor mutation burden in lung adenocarcinoma using histopathology images. Sci. Rep. 11, 16605 (2021).
Kanavati, F. et al. A deep learning model for the classification of indeterminate lung carcinoma in biopsy whole slide images. Sci. Rep. 11, 8110 (2021).
Chen, J. et al. Automatic lung cancer subtyping using rapid on-site evaluation slides and serum biological markers. Respir. Res. 25, 1–10 (2024).
Zhang, M. et al. Decreased green autofluorescence intensity of lung parenchyma is a potential non-invasive diagnostic biomarker for lung cancer. bioRxiv 343533 (2018).
Wang, N., Liu, Y. & Li, H. An efficient and fast, noninvasive, auto-fluorescence detection method for early-stage oral cancer. IEEE Trans. Instrum. Meas. 71, 1–11 (2022).
Waaijer, L. et al. Detection of breast cancer precursor lesions by autofluorescence ductoscopy. Breast Cancer 28, 119–129 (2021).
Pu, Y., Wang, W., Yang, Y. & Alfano, R. R. Stokes shift spectroscopic analysis of multifluorophores for human cancer detection in breast and prostate tissues. J. Biomed. Opt. 18, 017005–017005 (2013).
Vasanthakumari, P. et al. Discrimination of cancerous from benign pigmented skin lesions based on multispectral autofluorescence lifetime imaging dermoscopy and machine learning. J. Biomed. Opt. 27, 066002–066002 (2022).
Becker, W. Fluorescence lifetime imaging–techniques and applications. J. Microsc. 247, 119–136 (2012).
Fernandes, S. et al. Fibre-based fluorescence-lifetime imaging microscopy: a real-time biopsy guidance tool for suspected lung cancer. Transl. Lung Cancer Res. 13, 355 (2024).
Adams, A. C. et al. Fibre-optic based exploration of lung cancer autofluorescence using spectral fluorescence lifetime. Biomed. Opt. Express 15, 1132–1147 (2024).
Zang, Z. et al. Fast analysis of time-domain fluorescence lifetime imaging via extreme learning machine. Sensors 22, 3758 (2022).
Sorrells, J. E. et al. Real-time pixelwise phasor analysis for video-rate two-photon fluorescence lifetime imaging microscopy. Biomed. Opt. Express 12, 4003–4019 (2021).
Luo, T., Lu, Y., Liu, S., Lin, D. & Qu, J. Phasor–FLIM as a Screening tool for the differential diagnosis of actinic keratosis, Bowen’s disease, and basal cell carcinoma. Anal. Chem. 89, 8104–8111 (2017).
Walsh, A. J. et al. Classification of T-cell activation via autofluorescence lifetime imaging. Nat. Biomed. Eng. 5, 77–88 (2021).
Hu, L., Ter Hofstede, B., Sharma, D., Zhao, F. & Walsh, A. J. Comparison of phasor analysis and biexponential decay curve fitting of autofluorescence lifetime imaging data for machine learning prediction of cellular phenotypes. Front. Bioinforma. 3, 1210157 (2023).
Alfonso-García, A. et al. Label-free identification of macrophage phenotype by fluorescence lifetime imaging microscopy. J. Biomed. Opt. 21, 046005 (2016).
Wang, Q. et al. Deep learning in ex-vivo lung cancer discrimination using fluorescence lifetime endomicroscopic images. In 2020 42nd annual international conference of the IEEE Engineering in Medicine & Biology Society (EMBC) 1891–1894 (IEEE, 2020).
Wang, Q., Vallejo, M. & Hopgood, J. Fluorescence lifetime endomicroscopic image-based ex-vivo human lung cancer differentiation using machine learning. Authorea Prepr. (2023).
Wang, Q. et al. A layer-level multi-scale architecture for lung cancer classification with fluorescence lifetime imaging endomicroscopy. Neural Comput. Appl. 34, 18881–18894 (2022).
Wang, Q. et al. Deep learning-based virtual H& E staining from label-free autofluorescence lifetime images. Npj Imaging 2, 17 (2024).
Bai, B. et al. Deep learning-enabled virtual histological staining of biological samples. Light Sci. Appl. 12, 57 (2023).
Kreiss, L. et al. Digital staining in optical microscopy using deep learning-a review. PhotoniX 4, 34 (2023).
Rivenson, Y. et al. Virtual histological staining of unlabelled tissue-autofluorescence images via deep learning. Nat. Biomed. Eng. 3, 466–477 (2019).
Zhang, Y. et al. Digital synthesis of histological stains using micro-structured and multiplexed virtual staining of label-free tissue. Light Sci. Appl. 9, 78 (2020).
Li, X. et al. Unsupervised content-preserving transformation for optical microscopy. Light Sci. Appl. 10, 44 (2021).
DoanNgan, B., Angus, D., Sung, L., & others. Label-free virtual HER2 immunohistochemical staining of breast tissue using deep learning. BME Front. (2022).
Zhang, G. et al. Image-to-images translation for multiple virtual histological staining of unlabeled human carotid atherosclerotic tissue. Mol. Imaging Biol. 1–11 (2022).
Borhani, N., Bower, A. J., Boppart, S. A. & Psaltis, D. Digital staining through the application of deep neural networks to multi-modal multi-photon microscopy. Biomed. Opt. Express 10, 1339–1350 (2019).
Kang, L., Li, X., Zhang, Y. & Wong, T. T. Deep learning enables ultraviolet photoacoustic microscopy based histological imaging with near real-time virtual staining. Photoacoustics 25, 100308 (2022).
Cao, R. et al. Label-free intraoperative histology of bone tissue via deep-learning-assisted ultraviolet photoacoustic microscopy. Nat. Biomed. Eng. 7, 124–134 (2023).
Levy, J. J., Jackson, C. R., Sriharan, A., Christensen, B. C. & Vaickus, L. J. Preliminary evaluation of the utility of deep generative histopathology image translation at a mid-sized NCI cancer center. BioRxiv 2020–01 (2020).
Hong, Y. et al. Deep learning-based virtual cytokeratin staining of gastric carcinomas to measure tumor–stroma ratio. Sci. Rep. 11, 19255 (2021).
Lahiani, A., Klaman, I., Navab, N., Albarqouni, S. & Klaiman, E. Seamless virtual whole slide image synthesis and validation using perceptual embedding consistency. IEEE J. Biomed. Health Inform. 25, 403–411 (2020).
Zhang, R. et al. MVFStain: multiple virtual functional stain histopathology images generation based on specific domain mapping. Med. Image Anal. 80, 102520 (2022).
Lin, Y. et al. Unpaired multi-domain stain transfer for kidney histopathological images. Proc. AAAI Conf. Artif. Intell. 36, 1630–1637 (2022).
Pati, P. et al. Accelerating histopathology workflows with generative AI-based virtually multiplexed tumour profiling. Nat. Mach. Intell. 6, 1077–1093 (2024).
Moldvay, J. et al. The role of TTF-1 in differentiating primary and metastatic lung adenocarcinomas. Pathol. Oncol. Res. 10, 85–88 (2004).
Affandi, K. A., Tizen, N. M. S., Mustangin, M. & Zin, R. R. M. R. M. p40 immunohistochemistry is an excellent marker in primary lung squamous cell carcinoma. J. Pathol. Transl. Med. 52, 283–289 (2018).
Yatabe, Y. et al. Best practices recommendations for diagnostic immunohistochemistry in lung cancer. J. Thorac. Oncol. 14, 377–407 (2019).
Goodwin, J. et al. The distinct metabolic phenotype of lung squamous cell carcinoma defines selective vulnerability to glycolytic inhibition. Nat. Commun. 8, 15503 (2017).
Song, Q. et al. Proteomic analysis reveals key differences between squamous cell carcinomas and adenocarcinomas across multiple tissues. Nat. Commun. 13, 4167 (2022).
Chattopadhay, A., Sarkar, A., Howlader, P. & Balasubramanian, V. N. Grad-cam + +: Generalized gradient-based visual explanations for deep convolutional networks. in 2018 IEEE winter conference on applications of computer vision (WACV) 839–847 (IEEE, 2018).
Cai, T. T. & Ma, R. Theoretical foundations of t-SNE for visualizing high-dimensional clustered data. J. Mach. Learn. Res. 23, 1–54 (2022).
He, K., Zhang, X., Ren, S. & Sun, J. Deep Residual Learning for Image Recognition. In 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 770–778 (2016). https://doi.org/10.1109/CVPR.2016.90.
Tan, M. EfficientNet: Rethinking model scaling for convolutional neural networks. ArXiv Prepr. ArXiv190511946 (2019).
Huang, G., Liu, Z., Van Der Maaten, L. & Weinberger, K. Q. Densely connected convolutional networks. In Proceedings of the IEEE conference on computer vision and pattern recognition 4700–4708 (2017).
WHO Classification of Tumours Editorial Board. Thoracic Tumours. WHO Classification of Tumours, 5th Edition, Volume 5. (International Agency for Research on Cancer, 2021).
Zhang, M. et al. Decreased green autofluorescence of lung parenchyma is a biomarker for lung cancer tissues. J. Biophotonics 15, e202200072 (2022).
Vorontsov, E. et al. A foundation model for clinical-grade computational pathology and rare cancers detection. Nat. Med. 30, 1–12 (2024).
Wang, X. et al. A pathology foundation model for cancer diagnosis and prognosis prediction. Nature 634, 970–978 (2024).
Vaswani, A. Attention is all you need. Adv. Neural Inf. Process. Syst. (2017).
Liu, Z. et al. A convnet for the 2020s. In Proceedings of the IEEE/CVF conference on computer vision and pattern recognition 11976–11986 (IEEE, 2022).
Bai, B. et al. Label-free virtual HER2 immunohistochemical staining of breast tissue using deep learning. BME Front 2022, 9786242 (2022).
Dalmaz, O. & Yurt, M. & Çukur, T. ResViT: Residual vision transformers for multimodal medical image synthesis. IEEE Trans. Med. Imaging 41, 2598–2614 (2022).
Zhang, Y. et al. Super-resolved virtual staining of label-free tissue using diffusion models. ArXiv Prepr. ArXiv241020073 (2024).
Kataria, T., Knudsen, B. & Elhabian, S. Y. StainDiffuser: multitask dual diffusion model for virtual staining. ArXiv Prepr. ArXiv240311340 (2024).
Preibisch, S., Saalfeld, S. & Tomancak, P. Globally optimal stitching of tiled 3D microscopic image acquisitions. Bioinformatics 25, 1463–1465 (2009).
Isola, P., Zhu, J.-Y., Zhou, T. & Efros, A. A. Image-to-image translation with conditional adversarial networks. in Proceedings of the IEEE conference on computer vision and pattern recognition 1125–1134 (IEEE, 2017).
Wightman, R. PyTorch Image Models. GitHub repository https://doi.org/10.5281/zenodo.4414861 (2019).
Wang, Z., Bovik, A. C., Sheikh, H. R. & Simoncelli, E. P. Image quality assessment: from error visibility to structural similarity. IEEE Trans. Image Process. 13, 600–612 (2004).
Gatys, L. A. A neural algorithm of artistic style. ArXiv Prepr. ArXiv150806576 (2015).
Acknowledgements
The authors acknowledge the valuable comments from Dr Marta Vallejo at the School of Mathematics and Computer Science of Heriot-Watt University. We are grateful to the staff in the Department of Pathology, NHS Lothian and the Imaging Facility at the Institute of Regeneration and Repair, The University of Edinburgh (UoE). This study was partially funded by UoE Wellcome Institutional Translational Partnership Accelerator Fund and Cancer Research Horizons Seed Fund (PIII140), UoE Medical Research Council and Harmonised Impact Accelerator Accounts awards (MRC/IAA/015 and HIAA/037), Engineering and Physical Sciences Research Council (EPSRC) Grant Ref EP/S025987/1, NVIDIA Academic Hardware Grant Program, and A.R.A. is currently supported by a UKRI Future Leaders Fellowship (MR/Y015460/1). The funders played no role in the study design, data collection, data analysis and interpretation, or this manuscript’s writing. For open access, the author has applied a Creative Commons Attribution (CC BY) licence to any Author Accepted Manuscript version arising from this submission.
Author information
Authors and Affiliations
Contributions
Q.W. conceived the research. Z.Z. and Q.W. collected and processed autofluorescence images. Z.Z. and Q.W. conducted experiments on deep classification, and Q.W. conducted experiments on virtual IHC staining. A.R.A., D.A.D., K.E.Q., and A.D.J.W. performed the clinical aspects of the study, including tissue collection and processing, IHC staining, and designing and conducting blind evaluations. J.R.H. provided expertise on signal processing and deep learning. Z.Z. and Q.W. prepared the manuscript, and all authors contributed to and approved the manuscript. Q.W. and A.R.A. supervised the research.
Corresponding authors
Ethics declarations
Competing interests
Q.W. has 2 patent applications (UK patent application numbers: GB2319396.4 and GB 2405104.7) on the methods presented in this manuscript. Q.W. is currently employed by Prothea Technologies Ltd. UK. A.R.A is a founder shareholder and consultant for Prothea Technologies Ltd. UK.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zang, Z., Dorward, D.A., Quiohilag, K.E. et al. Label-free pathological subtyping of non-small cell lung cancer using deep classification and virtual immunohistochemical staining. npj Digit. Med. (2026). https://doi.org/10.1038/s41746-026-02557-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41746-026-02557-x