Abstract
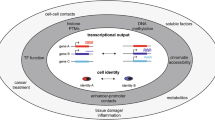
The concept of cellular neighborhoods, defined as recurring structures within the tissue with characteristic cell compositions and interactions, has transformed our understanding of the complexity and dynamics of tumor ecosystems. Recent advances in spatial omics and computational modeling have enabled high-resolution mapping of these neighborhoods, providing unprecedented insights into their roles in shaping tumor heterogeneity, evolution and therapeutic responses. Despite these advances, a unified framework for interpreting cellular neighborhoods remains lacking. This Perspective synthesizes emerging concepts and insights, focusing on the definition and classification of cellular neighborhoods in cancer, computational methods for identifying and comparing them, and their clinical relevance.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Wilson, A. & Trumpp, A. Bone-marrow haematopoietic-stem-cell niches. Nat. Rev. Immunol. 6, 93–106 (2006).
Brittan, M. & Wright, N. A. Gastrointestinal stem cells. J. Pathol. 197, 492–509 (2002).
Bressan, D., Battistoni, G. & Hannon, G. J. The dawn of spatial omics. Science 381, eabq4964 (2023).
Lewis, S. M. et al. Spatial omics and multiplexed imaging to explore cancer biology. Nat. Methods 18, 997–1012 (2021).
Park, J. et al. Spatial omics technologies at multimodal and single cell/subcellular level. Genome Biol. 23, 256 (2022).
Kang, L., Zhang, Q., Qian, F., Liang, J. & Wu, X. Benchmarking computational methods for detecting spatial domains and domain-specific spatially variable genes from spatial transcriptomics data. Nucleic Acids Res. 53, gkaf303 (2025).
Chen, C., Kim, H. J. & Yang, P. Evaluating spatially variable gene detection methods for spatial transcriptomics data. Genome Biol. 25, 18 (2024).
Keren, L. et al. A structured tumor–immune microenvironment in triple negative breast cancer revealed by multiplexed ion beam imaging. Cell 174, 1373–1387 (2018).
Schürch, C. M. et al. Coordinated cellular neighborhoods orchestrate antitumoral immunity at the colorectal cancer invasive front. Cell 182, 1341–1359 (2020).
Shiao, S. L. et al. Single-cell and spatial profiling identify three response trajectories to pembrolizumab and radiation therapy in triple negative breast cancer. Cancer Cell 42, 70–84 (2024).
Pentimalli, T. M. et al. High-resolution molecular atlas of a lung tumor in 3D. Preprint at bioRxiv https://doi.org/10.1101/2023.05.10.539644 (2023).
Bandyopadhyay, S. et al. Mapping the cellular biogeography of human bone marrow niches using single-cell transcriptomics and proteomic imaging. Cell 187, 3120–3140 (2024).
Barkley, D. et al. Cancer cell states recur across tumor types and form specific interactions with the tumor microenvironment. Nat. Genet. 54, 1192–1201 (2022).
Pelka, K. et al. Spatially organized multicellular immune hubs in human colorectal cancer. Cell 184, 4734–4752 (2021).
Gavish, A. et al. Hallmarks of transcriptional intratumour heterogeneity across a thousand tumours. Nature 618, 598–606 (2023).
Liu, M. et al. Tumor cell villages define the co-dependency of tumor and microenvironment in liver cancer. Preprint at bioRxiv https://doi.org/10.1101/2025.03.07.642107 (2025).
Yeh, C. Y. et al. Mapping spatial organization and genetic cell-state regulators to target immune evasion in ovarian cancer. Nat. Immunol. 25, 1943–1958 (2024).
Ren, Y. et al. Spatial transcriptomics reveals niche-specific enrichment and vulnerabilities of radial glial stem-like cells in malignant gliomas. Nat. Commun. 14, 1028 (2023).
Greenwald, A. C. et al. Integrative spatial analysis reveals a multi-layered organization of glioblastoma. Cell 187, 2485–2501 (2024).
Ravi, V. M. et al. Spatially resolved multi-omics deciphers bidirectional tumor–host interdependence in glioblastoma. Cancer Cell 40, 639–655 (2022).
Schumacher, T. N. & Thommen, D. S. Tertiary lymphoid structures in cancer. Science 375, eabf9419 (2022).
Sautès-Fridman, C., Petitprez, F., Calderaro, J. & Fridman, W. H. Tertiary lymphoid structures in the era of cancer immunotherapy. Nat. Rev. Cancer 19, 307–325 (2019).
Gaglia, G. et al. Lymphocyte networks are dynamic cellular communities in the immunoregulatory landscape of lung adenocarcinoma. Cancer Cell 41, 871–886 (2023).
Sorin, M. et al. Single-cell spatial landscapes of the lung tumour immune microenvironment. Nature 614, 548–554 (2023).
Cheng, S. et al. A pan-cancer single-cell transcriptional atlas of tumor infiltrating myeloid cells. Cell 184, 792–809 (2021).
Ma, R.-Y., Black, A. & Qian, B.-Z. Macrophage diversity in cancer revisited in the era of single-cell omics. Trends Immunol. 43, 546–563 (2022).
Karimi, E. et al. Single-cell spatial immune landscapes of primary and metastatic brain tumours. Nature 614, 555–563 (2023).
Wattenberg, M. M. et al. Intratumoral cell neighborhoods coordinate outcomes in pancreatic ductal adenocarcinoma. Gastroenterology 166, 1114–1129 (2024).
Ruf, B. et al. Tumor-associated macrophages trigger MAIT cell dysfunction at the HCC invasive margin. Cell 186, 3686–3705 (2023).
Xue, R. et al. Liver tumour immune microenvironment subtypes and neutrophil heterogeneity. Nature 612, 141–147 (2022).
Quail, D. F. & Joyce, J. A. Microenvironmental regulation of tumor progression and metastasis. Nat. Med. 19, 1423–1437 (2013).
Lodyga, M. & Hinz, B. TGF-β1—a truly transforming growth factor in fibrosis and immunity. Semin. Cell Dev. Biol. 101, 123–139 (2020).
Feng, Y. et al. Spatially organized tumor–stroma boundary determines the efficacy of immunotherapy in colorectal cancer patients. Nat. Commun. 15, 10259 (2024).
Cremasco, V. et al. FAP delineates heterogeneous and functionally divergent stromal cells in immune-excluded breast tumors. Cancer Immunol. Res. 6, 1472–1485 (2018).
Lavie, D., Ben-Shmuel, A., Erez, N. & Scherz-Shouval, R. Cancer-associated fibroblasts in the single-cell era. Nat. Cancer 3, 793–807 (2022).
Liu, Y. et al. Conserved spatial subtypes and cellular neighborhoods of cancer-associated fibroblasts revealed by single-cell spatial multi-omics. Cancer Cell 43, 905–924 (2025).
Xie, Y. et al. FGF/FGFR signaling in health and disease. Signal Transduct. Target. Ther. 5, 181 (2020).
Calon, A., Tauriello, D. V. F. & Batlle, E. TGF-β in CAF-mediated tumor growth and metastasis. Semin. Cancer Biol. 25, 15–22 (2014).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Sussman, J. H. et al. Multiplexed imaging mass cytometry analysis characterizes the vascular niche in pancreatic cancer. Cancer Res. 84, 2364–2376 (2024).
Gaglia, G. et al. Temporal and spatial topography of cell proliferation in cancer. Nat. Cell Biol. 24, 316–326 (2022).
Majmundar, A. J., Wong, W. J. & Simon, M. C. Hypoxia-inducible factors and the response to hypoxic stress. Mol. Cell 40, 294–309 (2010).
Qiu, G.-Z. et al. Reprogramming of the tumor in the hypoxic niche: the emerging concept and associated therapeutic strategies. Trends Pharmacol. Sci. 38, 669–686 (2017).
Smith, E. A. & Hodges, H. C. The spatial and genomic hierarchy of tumor ecosystems revealed by single-cell technologies. Trends Cancer 5, 411–425 (2019).
Kathagen, A. et al. Hypoxia and oxygenation induce a metabolic switch between pentose phosphate pathway and glycolysis in glioma stem-like cells. Acta Neuropathol. 126, 763–780 (2013).
Seim, J. et al. Hypoxia-induced irreversible S-phase arrest involves down-regulation of cyclin A. Cell Prolif. 36, 321–332 (2003).
Nirmal, A. J. et al. The spatial landscape of progression and immunoediting in primary melanoma at single-cell resolution. Cancer Discov. 12, 1518–1541 (2022).
Zhao, Y. et al. Stromal cells in the tumor microenvironment: accomplices of tumor progression? Cell Death Dis. 14, 587 (2023).
Ma, L., Li, C. C. & Wang, X. W. Roles of cellular neighborhoods in hepatocellular carcinoma pathogenesis. Annu. Rev. Pathol. 20, 169–192 (2025).
Wu, L. et al. An invasive zone in human liver cancer identified by Stereo-seq promotes hepatocyte–tumor cell crosstalk, local immunosuppression and tumor progression. Cell Res. 33, 585–603 (2023).
Lemaitre, L. et al. Spatial analysis reveals targetable macrophage-mediated mechanisms of immune evasion in hepatocellular carcinoma minimal residual disease. Nat. Cancer 5, 1534–1556 (2024).
Plaks, V., Kong, N. & Werb, Z. The cancer stem cell niche: how essential is the niche in regulating stemness of tumor cells? Cell Stem Cell 16, 225–238 (2015).
Batlle, E. & Clevers, H. Cancer stem cells revisited. Nat. Med. 23, 1124–1134 (2017).
Prager, B. C., Xie, Q., Bao, S. & Rich, J. N. Cancer stem cells: the architects of the tumor ecosystem. Cell Stem Cell 24, 41–53 (2019).
Wang, X. et al. Reciprocal signaling between glioblastoma stem cells and differentiated tumor cells promotes malignant progression. Cell Stem Cell 22, 514–528 (2018).
Su, S. et al. CD10+GPR77+ cancer-associated fibroblasts promote cancer formation and chemoresistance by sustaining cancer stemness. Cell 172, 841–856 (2018).
Wan, S. et al. Tumor-associated macrophages produce interleukin 6 and signal via STAT3 to promote expansion of human hepatocellular carcinoma stem cells. Gastroenterology 147, 1393–1404 (2014).
Lu, H. et al. A breast cancer stem cell niche supported by juxtacrine signalling from monocytes and macrophages. Nat. Cell Biol. 16, 1105–1117 (2014).
Forte, D. et al. Bone marrow mesenchymal stem cells support acute myeloid leukemia bioenergetics and enhance antioxidant defense and escape from chemotherapy. Cell Metab. 32, 829–843 (2020).
Semenza, G. L. Tumor metabolism: cancer cells give and take lactate. J. Clin. Invest. 118, 3835–3837 (2008).
Lyssiotis, C. A. & Kimmelman, A. C. Metabolic interactions in the tumor microenvironment. Trends Cell Biol. 27, 863–875 (2017).
Kennedy, K. M. & Dewhirst, M. W. Tumor metabolism of lactate: the influence and therapeutic potential for MCT and CD147 regulation. Future Oncol. 6, 127–148 (2010).
Sun, C. et al. Spatially resolved metabolomics to discover tumor-associated metabolic alterations. Proc. Natl Acad. Sci. USA 116, 52–57 (2019).
Smith, B. et al. Addiction to coupling of the Warburg effect with glutamine catabolism in cancer cells. Cell Rep. 17, 821–836 (2016).
Khaliq, A. M. et al. Spatial transcriptomic analysis of primary and metastatic pancreatic cancers highlights tumor microenvironmental heterogeneity. Nat. Genet. 56, 2455–2465 (2024).
Wilde, L. et al. Metabolic coupling and the Reverse Warburg Effect in cancer: implications for novel biomarker and anticancer agent development. Semin. Oncol. 44, 198–203 (2017).
Wang, Y. et al. Single-cell analysis of pancreatic ductal adenocarcinoma identifies a novel fibroblast subtype associated with poor prognosis but better immunotherapy response. Cell Discov. 7, 36 (2021).
Kloosterman, D. J. et al. Macrophage-mediated myelin recycling fuels brain cancer malignancy. Cell 187, 5336–5356 (2024).
Ikeda, H. et al. Immune evasion through mitochondrial transfer in the tumour microenvironment. Nature 638, 225–236 (2025).
Hioki, K. A. et al. The mosquito effect: regulatory and effector T cells acquire cytoplasmic material from tumor cells through intercellular transfer. Front. Immunol. 14, 1272918 (2023).
Mo, C.-K. et al. Tumour evolution and microenvironment interactions in 2D and 3D space. Nature 634, 1178–1186 (2024).
Wu, R. et al. Comprehensive analysis of spatial architecture in primary liver cancer. Sci. Adv. 7, eabg3750 (2021).
Matusiak, M. et al. Spatially segregated macrophage populations predict distinct outcomes in colon cancer. Cancer Discov. 14, 1418–1439 (2024).
Shi, Q. et al. Cross-tissue multicellular coordination and its rewiring in cancer. Nature 643, 529–538 (2025).
Jahanban-Esfahlan, R., de la Guardia, M., Ahmadi, D. & Yousefi, B. Modulating tumor hypoxia by nanomedicine for effective cancer therapy. J. Cell. Physiol. 233, 2019–2031 (2018).
Liu, Y. et al. Identification of a tumour immune barrier in the HCC microenvironment that determines the efficacy of immunotherapy. J. Hepatol. 78, 770–782 (2023).
Sathe, A. et al. Colorectal cancer metastases in the liver establish immunosuppressive spatial networking between tumor-associated SPP1+ macrophages and fibroblasts. Clin. Cancer Res. 29, 244–260 (2023).
Christofori, G. New signals from the invasive front. Nature 441, 444–450 (2006).
Ding, G.-Y. et al. Distribution and density of tertiary lymphoid structures predict clinical outcome in intrahepatic cholangiocarcinoma. J. Hepatol. 76, 608–618 (2022).
Black, S. et al. CODEX multiplexed tissue imaging with DNA-conjugated antibodies. Nat. Protoc. 16, 3802–3835 (2021).
Chen, A. et al. Spatiotemporal transcriptomic atlas of mouse organogenesis using DNA nanoball-patterned arrays. Cell 185, 1777–1792 (2022).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Chen, H. et al. Integrative spatial analysis reveals tumor heterogeneity and immune colony niche related to clinical outcomes in small cell lung cancer. Cancer Cell 43, 519–536 (2025).
Kim, J. et al. Unsupervised discovery of tissue architecture in multiplexed imaging. Nat. Methods 19, 1653–1661 (2022).
Singhal, V. et al. BANKSY unifies cell typing and tissue domain segmentation for scalable spatial omics data analysis. Nat. Genet. 56, 431–441 (2024).
Ma, Y. & Zhou, X. Accurate and efficient integrative reference-informed spatial domain detection for spatial transcriptomics. Nat. Methods 21, 1231–1244 (2024).
Haviv, D. et al. The covariance environment defines cellular niches for spatial inference. Nat. Biotechnol. 43, 269–280 (2025).
Blei, D. M., Ng, A. Y. & Jordan, M. I. Latent Dirichlet allocation. J. Mach. Learn. Res. 3, 993–1022 (2003).
Chen, Z., Soifer, I., Hilton, H., Keren, L. & Jojic, V. Modeling multiplexed images with Spatial-LDA reveals novel tissue microenvironments. J. Comput. Biol. 27, 1204–1218 (2020).
Zhao, E. et al. Spatial transcriptomics at subspot resolution with BayesSpace. Nat. Biotechnol. 39, 1375–1384 (2021).
Li, Z. & Zhou, X. BASS: multi-scale and multi-sample analysis enables accurate cell type clustering and spatial domain detection in spatial transcriptomic studies. Genome Biol. 23, 168 (2022).
Hu, J. et al. SpaGCN: integrating gene expression, spatial location and histology to identify spatial domains and spatially variable genes by graph convolutional network. Nat. Methods 18, 1342–1351 (2021).
Hu, Y. et al. Unsupervised and supervised discovery of tissue cellular neighborhoods from cell phenotypes. Nat. Methods 21, 267–278 (2024).
Long, Y. et al. Spatially informed clustering, integration, and deconvolution of spatial transcriptomics with GraphST. Nat. Commun. 14, 1155 (2023).
Varrone, M., Tavernari, D., Santamaria-Martínez, A., Walsh, L. A. & Ciriello, G. CellCharter reveals spatial cell niches associated with tissue remodeling and cell plasticity. Nat. Genet. 56, 74–84 (2024).
Li, Y., Zhang, J., Gao, X. & Zhang, Q. C. Tissue module discovery in single-cell-resolution spatial transcriptomics data via cell–cell interaction-aware cell embedding. Cell Syst. 15, 578–592 (2024).
Chitra, U. et al. Mapping the topography of spatial gene expression with interpretable deep learning. Nat. Methods 22, 298–309 (2025).
Birk, S. et al. Quantitative characterization of cell niches in spatially resolved omics data. Nat. Genet. 57, 897–909 (2025).
Hickey, J. W. et al. T cell-mediated curation and restructuring of tumor tissue coordinates an effective immune response. Cell Rep. 42, 113494 (2023).
Samadi, Z., Hao, K. & Askary, A. SMORE: spatial motifs reveal patterns in cellular architecture of complex tissues. Genome Biol. 26, 3 (2025).
Chen, J. H. et al. Human lung cancer harbors spatially organized stem-immunity hubs associated with response to immunotherapy. Nat. Immunol. 25, 644–658 (2024).
Salié, H. et al. Spatial single-cell profiling and neighbourhood analysis reveal the determinants of immune architecture connected to checkpoint inhibitor therapy outcome in hepatocellular carcinoma. Gut 74, 451–466 (2025).
Mascharak, S. et al. Desmoplastic stromal signatures predict patient outcomes in pancreatic ductal adenocarcinoma. Cell Rep. Med. 4, 101248 (2023).
Qi, J. et al. Single-cell and spatial analysis reveal interaction of FAP+ fibroblasts and SPP1+ macrophages in colorectal cancer. Nat. Commun. 13, 1742 (2022).
Maestri, E. et al. Spatial proximity of tumor–immune interactions predicts patient outcome in hepatocellular carcinoma. Hepatology 79, 768–779 (2024).
Peyraud, F. et al. Spatially resolved transcriptomics reveal the determinants of primary resistance to immunotherapy in NSCLC with mature tertiary lymphoid structures. Cell Rep. Med. 6, 101934 (2025).
Magen, A. et al. Intratumoral dendritic cell–CD4+ T helper cell niches enable CD8+ T cell differentiation following PD-1 blockade in hepatocellular carcinoma. Nat. Med. 29, 1389–1399 (2023).
Wang, X. Q. et al. Spatial predictors of immunotherapy response in triple-negative breast cancer. Nature 621, 868–876 (2023).
Jia, G. et al. Spatial immune scoring system predicts hepatocellular carcinoma recurrence. Nature 640, 1031–1041 (2025).
Ali, H. R. et al. Imaging mass cytometry and multiplatform genomics define the phenogenomic landscape of breast cancer. Nat. Cancer 1, 163–175 (2020).
Chen, J., Larsson, L., Swarbrick, A. & Lundeberg, J. Spatial landscapes of cancers: insights and opportunities. Nat. Rev. Clin. Oncol. 21, 660–674 (2024).
Väyrynen, S. A. et al. Composition, spatial characteristics, and prognostic significance of myeloid cell infiltration in pancreatic cancer. Clin. Cancer Res. 27, 1069–1081 (2021).
Zheng, Y., Carrillo-Perez, F., Pizurica, M., Heiland, D. H. & Gevaert, O. Spatial cellular architecture predicts prognosis in glioblastoma. Nat. Commun. 14, 4122 (2023).
Phillips, D. et al. Immune cell topography predicts response to PD-1 blockade in cutaneous T cell lymphoma. Nat. Commun. 12, 6726 (2021).
Niazi, M. K. K., Parwani, A. V. & Gurcan, M. N. Digital pathology and artificial intelligence. Lancet Oncol. 20, e253–e261 (2019).
Wang, X. et al. A pathology foundation model for cancer diagnosis and prognosis prediction. Nature 634, 970–978 (2024).
Huang, Z., Bianchi, F., Yuksekgonul, M., Montine, T. J. & Zou, J. A visual–language foundation model for pathology image analysis using medical twitter. Nat. Med. 29, 2307–2316 (2023).
Lu, M. Y. et al. A visual–language foundation model for computational pathology. Nat. Med. 30, 863–874 (2024).
Yang, Z. et al. A foundation model for generalizable cancer diagnosis and survival prediction from histopathological images. Nat. Commun. 16, 2366 (2025).
Hua, S., Yan, F., Shen, T., Ma, L. & Zhang, X. PathoDuet: foundation models for pathological slide analysis of H&E and IHC stains. Med. Image Anal. 97, 103289 (2024).
Cui, H. et al. scGPT: toward building a foundation model for single-cell multi-omics using generative AI. Nat. Methods 21, 1470–1480 (2024).
Hao, M. et al. Large-scale foundation model on single-cell transcriptomics. Nat. Methods 21, 1481–1491 (2024).
Rood, J. E. et al. The Human Cell Atlas from a cell census to a unified foundation model. Nature 637, 1065–1071 (2025).
Boiarsky, R. et al. Deeper evaluation of a single-cell foundation model. Nat. Mach. Intell. 6, 1443–1446 (2024).
Wang, H. et al. SpatialAgent: an autonomous AI agent for spatial biology. Preprint at bioRxiv https://doi.org/10.1101/2025.04.03.646459 (2025).
Tejada-Lapuerta, A. et al. Nicheformer: a foundation model for single-cell and spatial omics. Nat. Methods 22, 2525–2538 (2025).
Marks, M. et al. CellSAM: a foundation model for cell segmentation. Nat. Methods 22, 2585–2593 (2025).
Wang, C. et al. scGPT-spatial: continual pretraining of single-cell foundation model for spatial transcriptomics. Preprint at bioRxiv https://doi.org/10.1101/2025.02.05.636714 (2025).
Xu, Z. et al. STOmicsDB: a comprehensive database for spatial transcriptomics data sharing, analysis and visualization. Nucleic Acids Res. 52, D1053–D1061 (2024).
Yuan, Z. et al. SODB facilitates comprehensive exploration of spatial omics data. Nat. Methods 20, 387–399 (2023).
Zheng, Y., Chen, Y., Ding, X., Wong, K. H. & Cheung, E. Aquila: a spatial omics database and analysis platform. Nucleic Acids Res. 51, D827–D834 (2023).
Dhainaut, M. et al. Spatial CRISPR genomics identifies regulators of the tumor microenvironment. Cell 185, 1223–1239 (2022).
Binan, L. et al. Simultaneous CRISPR screening and spatial transcriptomics reveals intracellular, intercellular, and functional transcriptional circuits. Preprint at bioRxiv https://doi.org/10.1101/2023.11.30.569494 (2023).
Gu, J. et al. Mapping multimodal phenotypes to perturbations in cells and tissue with CRISPRmap. Nat. Biotechnol. 43, 1101–1115 (2025).
Chuprin, J. et al. Humanized mouse models for immuno-oncology research. Nat. Rev. Clin. Oncol. 20, 192–206 (2023).
Connolly, K. A., Fitzgerald, B., Damo, M. & Joshi, N. S. Novel mouse models for cancer immunology. Annu. Rev. Cancer Biol. 6, 269–291 (2022).
Ingber, D. E. Human organs-on-chips for disease modelling, drug development and personalized medicine. Nat. Rev. Genet. 23, 467–491 (2022).
Leung, C. M. et al. A guide to the organ-on-a-chip. Nat. Rev. Methods Primers 2, 33 (2022).
Ståhl, P. L. et al. Visualization and analysis of gene expression in tissue sections by spatial transcriptomics. Science 353, 78–82 (2016).
Chen, K. H., Boettiger, A. N., Moffitt, J. R., Wang, S. & Zhuang, X. Spatially resolved, highly multiplexed RNA profiling in single cells. Science 348, aaa6090 (2015).
Janesick, A. et al. High resolution mapping of the tumor microenvironment using integrated single-cell, spatial and in situ analysis. Nat. Commun. 14, 8353 (2023).
Merritt, C. R. et al. Multiplex digital spatial profiling of proteins and RNA in fixed tissue. Nat. Biotechnol. 38, 586–599 (2020).
Stickels, R. R. et al. Highly sensitive spatial transcriptomics at near-cellular resolution with Slide-seqV2. Nat. Biotechnol. 39, 313–319 (2021).
Wang, X. et al. Three-dimensional intact-tissue sequencing of single-cell transcriptional states. Science 361, eaat5691 (2018).
Schott, M. et al. Open-ST: high-resolution spatial transcriptomics in 3D. Cell 187, 3953–3972 (2024).
He, S. et al. High-plex imaging of RNA and proteins at subcellular resolution in fixed tissue by spatial molecular imaging. Nat. Biotechnol. 40, 1794–1806 (2022).
Goltsev, Y. et al. Deep profiling of mouse splenic architecture with CODEX multiplexed imaging. Cell 174, 968–981 (2018).
Keren, L. et al. MIBI-TOF: a multiplexed imaging platform relates cellular phenotypes and tissue structure. Sci. Adv. 5, eaax5851 (2019).
Giesen, C. et al. Highly multiplexed imaging of tumor tissues with subcellular resolution by mass cytometry. Nat. Methods 11, 417–422 (2014).
Niehaus, M., Soltwisch, J., Belov, M. E. & Dreisewerd, K. Transmission-mode MALDI-2 mass spectrometry imaging of cells and tissues at subcellular resolution. Nat. Methods 16, 925–931 (2019).
Acknowledgements
This work was supported by grants from the National Institutes of Health (NIH) under award numbers U2C CA233285 and U54HL165442 (K.T.). L.M. was supported by grants (ZIA BC 012079 and ZIA BC 012083) from the Intramural Research Program of the Center for Cancer Research, US National Cancer Institute. B.X. was supported by NIH grant F30CA298606. K.T. holds the Richard and Sheila Sanford Endowed Chair at CHOP. This research was supported in part by the Intramural Research Program of the NIH. The contributions of the NIH authors were made as part of their official duties as NIH federal employees, are in compliance with agency policy requirements and are considered works of the US government. However, the findings and conclusions presented in this paper are those of the authors and do not necessarily reflect the views of the NIH or the US Department of Health and Human Services.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cancer thanks Chenfei Wang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ma, L., Xiong, B., Liu, M. et al. Cellular neighborhoods in cancer. Nat Cancer 7, 16–28 (2026). https://doi.org/10.1038/s43018-025-01107-w
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s43018-025-01107-w