Abstract
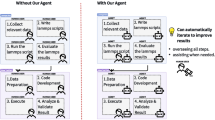
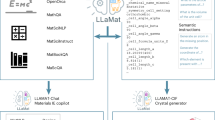
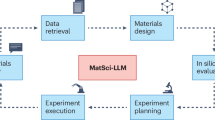
The integration of large language models (LLMs) with domain-specific computational tools provides a pathway to streamline and enhance materials science workflows. This paper introduces MatSciAgent, a multi-agent framework supporting tasks such as materials data retrieval, continuum simulation, crystal structure generation, and molecular dynamics simulation. At its core is a master agent that interprets user queries, identifies the task type, and delegates to task-specific agent(s) equipped with tools. Leveraging databases such as Materials Project and MatWeb, the framework retrieves and summarizes materials data with grounded, factual responses, addressing limitations of vanilla LLMs. When a target material is absent from databases, a generative agent can propose plausible crystal structures. For simulations, specialized agents extract parameters to perform continuum and molecular dynamics simulations using existing software or custom code. MatSciAgent demonstrates stability, with parameter extraction achieving 100% success across five runs and materials extraction consistent in 9 of 10 runs. Its modular design ensures seamless extensibility to evolve as new capabilities are integrated.

Similar content being viewed by others
Data availability
The case study results and reliability study results are provided as supplementary data.
Code availability
The necessary code used in this study can be accessed at https://github.com/cakshat/MatSci-LLM-Agents.
References
Pollice, R. et al. Data-driven strategies for accelerated materials design. Acc. Chem. Res. 54, 849–860 (2021).
Kalidindi, S. R. & De Graef, M. Materials data science: current status and future outlook. Annu. Rev. Mater. Res. 45, 171–193 (2015).
Jain, A. et al. The materials project: a materials genome approach to accelerating materials innovation. APL Mater. 1, 011002 (2013).
LLC, M. Matweb: Online Materials Information Resource. http://www.matweb.com (2020).
Kirklin, S. et al. The open quantum materials database (oqmd): assessing the accuracy of DFT formation energies. npj Comput. Mater. 1, 1–15 (2015).
Hellenbrandt, M. The inorganic crystal structure database (ICSD)-present and future. Crystallogr. Rev. 10, 17–22 (2004).
Tanaka, I., Rajan, K. & Wolverton, C. Data-centric science for materials innovation. MRS Bull. 43, 659–663 (2018).
Antunes, L. M., Butler, K. T. & Grau-Crespo, R. Crystal structure generation with autoregressive large language modeling. Nat. Commun. 15, 10570 (2024).
Xie, T., Fu, X., Ganea, O.-E., Barzilay, R. & Jaakkola, T. Crystal diffusion variational autoencoder for periodic material generation. arXiv https://arxiv.org/abs/2110.06197 (2022).
Zeni, C. et al. A generative model for inorganic materials design. Nature 639, 624–632 (2025).
Himanen, L., Geurts, A., Foster, A. S. & Rinke, P. Data-driven materials science: status, challenges, and perspectives. Adv. Sci. 6, 1900808 (2019).
Vaswani, A. et al. Attention is all you need. In Guyon, I.et al. (eds.) Advances in Neural Information Processing Systems, vol. 30 (Curran Associates, Inc., 2017).
Brown, T. et al. Language models are few-shot learners. Adv. neural Inf. Process. Syst. 33, 1877–1901 (2020).
Achiam, J. et al. Gpt-4 Technical Report. https://cdn.openai.com/papers/gpt-4.pdf (2023).
Touvron, H. et al. Llama: Open and efficient foundation language models. arXiv https://arxiv.org/abs/2302.13971 (2024).
Anthropic. Claude 3.7 Sonnet. https://www.anthropic.com (2025).
Tshitoyan, V. et al. Unsupervised word embeddings capture latent knowledge from materials science literature. Nature 571, 95–98 (2019).
Ock, J., Guntuboina, C. & Barati Farimani, A. Catalyst energy prediction with CatBERTA: unveiling feature exploration strategies through large language models. ACS Catal. 13, 16032–16044 (2023).
Ock, J., Badrinarayanan, S., Magar, R., Antony, A. & Barati Farimani, A. Multimodal language and graph learning of adsorption configuration in catalysis. Nat. Mach. Intell. 6, 1501–1511 (2024).
Cao, Z., Magar, R., Wang, Y. & Barati Farimani, A. Moformer: Self-supervised transformer model for metal-organic framework property prediction. J. Am. Chem. Soc. 145, 2958–2967 (2023).
Xu, C., Wang, Y. & Barati Farimani, A. Transpolymer: a transformer-based language model for polymer property predictions. npj Comput. Mater. 9, 64 (2023).
Kim, S., Mollaei, P., Antony, A., Magar, R. & Barati Farimani, A. Gpcr-bert: Interpreting sequential design of g protein-coupled receptors using protein language models. J. Chem. Inf. Modeling 64, 1134–1144 (2024).
Mollaei, P., Sadasivam, D., Guntuboina, C. & Barati Farimani, A. Idp-bert: Predicting properties of intrinsically disordered proteins using large language models. J. Phys. Chem. B 128, 12030–12037 (2024).
Badrinarayanan, S., Guntuboina, C., Mollaei, P. & Barati Farimani, A. Multi-peptide: Multimodality leveraged language-graph learning of peptide properties. J. Chem. Inf. Model. 65, 83–91 (2025).
Meda, R. S. & Farimani, A. B. Bapulm: Binding affinity prediction using language models. arXiv https://arxiv.org/abs/2411.04150 (2024).
Chaudhari, A., Guntuboina, C., Huang, H. & Farimani, A. B. Alloybert: Alloy property prediction with large language models. Comput. Mater. Sci. 244, 113256 (2024).
Yeh, Y.-T., Ock, J. & Farimani, A. B. Text to band gap: Pre-trained language models as encoders for semiconductor band gap prediction. arXiv https://arxiv.org/abs/2501.03456 (2025).
Pak, P. & Farimani, A. B. Additivellm: Large language models predict defects in additive manufacturing. arXiv https://arxiv.org/abs/2501.17784 (2025).
Lu, C. et al. The ai scientist: Towards fully automated open-ended scientific discovery. arXiv https://arxiv.org/abs/2408.06292 (2024).
Ock, J., Vinchurkar, T., Jadhav, Y. & Farimani, A. B. Adsorb-agent: Autonomous identification of stable adsorption configurations via large language model agent. arXiv https://arxiv.org/abs/2410.16658 (2024).
Boiko, D. A., MacKnight, R., Kline, B. & Gomes, G. Autonomous chemical research with large language models. Nature 624, 570–578 (2023).
Bran, A. M. et al. Augmenting large language models with chemistry tools. Nat. Mach. Intell. 6, 525–535 (2024).
Jadhav, Y., Pak, P. & Farimani, A. B. LLM-3D print: Large language models to monitor and control 3d printing. arXiv https://arxiv.org/abs/2408.14307 (2024).
Jadhav, Y. & Farimani, A. B. Large language model agent as a mechanical designer. arXiv https://arxiv.org/abs/2404.17525 (2024).
Huang, L. et al. A survey on hallucination in large language models: Principles, taxonomy, challenges, and open questions. arXiv https://doi.org/10.1145/3703155 (2024).
Perković, G., Drobnjak, A. & Botički, I. Hallucinations in llms: Understanding and addressing challenges. In 2024 47th MIPRO ICT and Electronics Convention (MIPRO) 2084–2088 (IEEE, 2024).
Liu, F. et al. Exploring and evaluating hallucinations in LLM-powered code generation. arXiv https://arxiv.org/abs/2404.00971 (2024).
Birhane, A., Kasirzadeh, A., Leslie, D. & Wachter, S. Science in the age of large language models. Nat. Rev. Phys. 5, 277–280 (2023).
Johnson, S. & Hyland-Wood, D. A primer on large language models and their limitations. arXiv https://arxiv.org/abs/2412.04503 (2024).
Chiang, Y., Hsieh, E., Chou, C.-H. & Riebesell, J. Llamp: Large language model made powerful for high-fidelity materials knowledge retrieval and distillation. arXiv https://arxiv.org/abs/2401.17244 (2024).
Zhang, H., Song, Y., Hou, Z., Miret, S. & Liu, B. Honeycomb: A flexible LLM-based agent system for materials science. arXiv https://arxiv.org/abs/2409.00135 (2024).
Ghafarollahi, A. & Buehler, M. J. Atomagents: Alloy design and discovery through physics-aware multi-modal multi-agent artificial intelligence. arXiv https://arxiv.org/abs/2407.10022 (2024).
Radford, A. & Narasimhan, K. Improving Language Understanding by Generative Pre-training. https://api.semanticscholar.org/CorpusID:49313245 (2018).
Vaswani, A. Attention is all you need. In 31st Conference on Neural Information Processing Systems. (NIPS, 2017).
Chase, H. Langchain. https://github.com/langchain-ai/langchain (2022).
Ling, C. et al. Domain specialization as the key to make large language models disruptive: a comprehensive survey. arXiv https://arxiv.org/abs/2305.18703 (2024).
Li, H. et al. Blade: Enhancing black-box large language models with small domain-specific models arXiv https://arxiv.org/abs/2403.18365 (2024).
Mittelstadt, B., Wachter, S. & Russell, C. To protect science, we must use LLMs as zero-shot translators. Nat. Hum. Behav. 7, 1830–1832 (2023).
Fanfoni, M. & Tomellini, M. The Johnson-Mehl-Avrami-Khokhlov model: a brief review. Il Nuovo Cim. D. 20, 1171–1182 (1998).
Kinoshita, M., Okamoto, Y. & Hirata, F. First-principle determination of peptide conformations in solvents: Combination of Monte Carlo simulated annealing and RISM theory. J. Am. Chem. Soc. 120, 1855–1863 (1998).
Wolfram, S. Statistical mechanics of cellular automata. Rev. Mod. Phys. 55, 601 (1983).
Wolfram, S. Cellular automata as models of complexity. Nature 311, 419–424 (1984).
Metropolis, N. & Ulam, S. The Monte Carlo method. J. Am. Stat. Assoc. 44, 335–341 (1949).
Saito, Y. & Enomoto, M. Monte Carlo simulation of grain growth. ISIJ Int. 32, 267–274 (1992).
Radford, A. et al. Language Models are Unsupervised Multitask Learners. https://api.semanticscholar.org/CorpusID:160025533 (2019).
Karpathy, A. NanoGPT. https://github.com/karpathy/nanoGPT (2022).
Nomad: A distributed web-based platform for managing materials science research data. J. Open Source Softw. 8, 5388 (2023).
Mavromaras, A. et al. Computational materials engineering: capabilities of atomic-scale prediction of mechanical, thermal, and electrical properties of microelectronic materials. In 2010 11th International Thermal, Mechanical & Multi-Physics Simulation, and Experiments in Microelectronics and Microsystems (EuroSimE), Bordeaux, 1–10 (IEEE, 2010).
Eyert, V. et al. Atomistic simulations of microelectronic materials: prediction of mechanical, thermal, and electrical properties. In Molecular Modeling and Multiscaling Issues for Electronic Material Applications. (eds. Iwamoto, N., Yuen, M., Fan, H.) (Springer, 2012).
Larsen, A. H. et al. The atomic simulation environment-a Python library for working with atoms. J. Phys. Condens. Matter 29, 273002 (2017).
Foiles, S., Baskes, M. & Daw, M. S. Embedded-atom-method functions for the fcc metals Cu, Ag, Au, Ni, Pd, Pt, and their alloys. Phys. Rev. B 33, 7983 (1986).
Lennard-Jones, J. E. On the forces between atoms and ions. Proc. A 109, 584–597 (1925).
Choy, T. C. Effective Medium Theory: Principles and Applications 2nd edn, Vol. 256 (Oxford University Press, 2015).
Acknowledgements
The authors gratefully acknowledge support from the H. Robert Sharbaugh Presidential Fellowship.
Author information
Authors and Affiliations
Contributions
A.C. conceived the study, developed the methodology, implemented the software, curated data, performed analysis, prepared visualizations, and wrote the original draft. J.O. contributed to methodology, validation, investigation, data curation, and writing and editing. A.B.F. supervised the project, contributed to conceptualization, project administration, funding acquisition, and writing and editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Materials thanks Federico Ottomano and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chaudhari, A., Ock, J. & Barati Farimani, A. Modular large language model agents for multi-task computational materials science. Commun Mater (2026). https://doi.org/10.1038/s43246-025-00994-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s43246-025-00994-x