Abstract
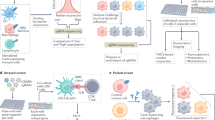
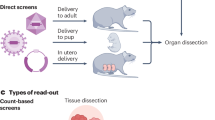
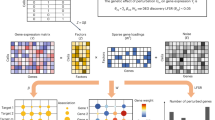
CRISPR screens are a powerful source of biological discovery, enabling the unbiased interrogation of gene function in a wide range of applications and species. In pooled CRISPR screens, various genetically encoded perturbations are introduced into pools of cells. The targeted cells proliferate under a biological challenge such as cell competition, drug treatment or viral infection. Subsequently, the perturbation-induced effects are evaluated by sequencing-based counting of the guide RNAs that specify each perturbation. The typical results of such screens are ranked lists of genes that confer sensitivity or resistance to the biological challenge of interest. Contributing to the broad utility of CRISPR screens, adaptations of the core CRISPR technology make it possible to activate, silence or otherwise manipulate the target genes. Moreover, high-content read-outs such as single-cell RNA sequencing and spatial imaging help characterize screened cells with unprecedented detail. Dedicated software tools facilitate bioinformatic analysis and enhance reproducibility. CRISPR screening has unravelled various molecular mechanisms in basic biology, medical genetics, cancer research, immunology, infectious diseases, microbiology and other fields. This Primer describes the basic and advanced concepts of CRISPR screening and its application as a flexible and reliable method for biological discovery, biomedical research and drug development — with a special emphasis on high-content methods that make it possible to obtain detailed biological insights directly as part of the screen.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 1 digital issues and online access to articles
$119.00 per year
only $119.00 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Grimm, S. The art and design of genetic screens: mammalian culture cells. Nat. Rev. Genet. 5, 179–189 (2004).
St Johnston, D. The art and design of genetic screens: Drosophila melanogaster. Nat. Rev. Genet. 3, 176–188 (2002).
Wieschaus, E. & Nüsslein-Volhard, C. The Heidelberg screen for pattern mutants of Drosophila: a personal account. Annu. Rev. Cell Dev. Biol. 32, 1–46 (2016).
Jorgensen, E. M. & Mango, S. E. The art and design of genetic screens: Caenorhabditis elegans. Nat. Rev. Genet. 3, 356–369 (2002).
Forsburg, S. L. The art and design of genetic screens: yeast. Nat. Rev. Genet. 2, 659–668 (2001).
Page, D. R. & Grossniklaus, U. The art and design of genetic screens: Arabidopsis thaliana. Nat. Rev. Genet. 3, 124–136 (2002).
Patton, E. E. & Zon, L. I. The art and design of genetic screens: zebrafish. Nat. Rev. Genet. 2, 956–966 (2001).
Boutros, M. & Ahringer, J. The art and design of genetic screens: RNA interference. Nat. Rev. Genet. 9, 554–566 (2008).
Mohr, S. E., Smith, J. A., Shamu, C. E., Neumüller, R. A. & Perrimon, N. RNAi screening comes of age: improved techniques and complementary approaches. Nat. Rev. Mol. Cell Biol. 15, 591–600 (2014).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Doench, J. G. Am I ready for CRISPR? A user’s guide to genetic screens. Nat. Rev. Genet. 19, 67–80 (2018).
Shalem, O., Sanjana, N. E. & Zhang, F. High-throughput functional genomics using CRISPR–Cas9. Nat. Rev. Genet. 16, 299–311 (2015).
Kuhn, M., Santinha, A. J. & Platt, R. J. Moving from in vitro to in vivo CRISPR screens. Gene Genome Editing 2, 100008 (2021).
Fernandes Neto, J. M. et al. Optimized Cas9 expression improves performance of large-scale CRISPR screening. Preprint at bioRxiv https://doi.org/10.1101/2021.07.13.452178v1 (2021).
Chu, V. T. et al. Efficient CRISPR-mediated mutagenesis in primary immune cells using CrispRGold and a C57BL/6 Cas9 transgenic mouse line. Proc. Natl Acad. Sci. USA 113, 12514–12519 (2016).
Platt, R. J. et al. CRISPR–Cas9 knockin mice for genome editing and cancer modeling. Cell 159, 440–455 (2014).
Gemberling, M. P. et al. Transgenic mice for in vivo epigenome editing with CRISPR-based systems. Nat. Methods 18, 965–974 (2021).
van Haasteren, J., Li, J., Scheideler, O. J., Murthy, N. & Schaffer, D. V. The delivery challenge: fulfilling the promise of therapeutic genome editing. Nat. Biotechnol. 38, 845–855 (2020).
Lawson, K. A. et al. Functional genomic landscape of cancer-intrinsic evasion of killing by T cells. Nature 586, 120–126 (2020).
Pacini, C. et al. Integrated cross-study datasets of genetic dependencies in cancer. Nat. Commun. 12, 1661 (2021).
Behan, F. M. et al. Prioritization of cancer therapeutic targets using CRISPR–Cas9 screens. Nature 568, 511–516 (2019).
Hart, T. et al. Evaluation and design of genome-wide CRISPR/SpCas9 knockout screens. G3 7, 2719–2727 (2017).
Moffat, J. et al. A lentiviral RNAi library for human and mouse genes applied to an arrayed viral high-content screen. Cell 124, 1283–1298 (2006).
Naldini, L. et al. In vivo gene delivery and stable transduction of nondividing cells by a lentiviral vector. Science 272, 263–267 (1996).
Zufferey, R., Nagy, D., Mandel, R. J., Naldini, L. & Trono, D. Multiply attenuated lentiviral vector achieves efficient gene delivery in vivo. Nat. Biotechnol. 15, 871–875 (1997).
Smits, A. H. et al. Biological plasticity rescues target activity in CRISPR knock outs. Nat. Methods 16, 1087–1093 (2019).
Esposito, R. et al. Hacking the cancer genome: profiling therapeutically actionable long non-coding RNAs using CRISPR–Cas9 screening. Cancer Cell 35, 545–557 (2019).
Shukla, A. & Huangfu, D. Decoding the noncoding genome via large-scale CRISPR screens. Curr. Opin. Genet. Dev. 52, 70–76 (2018).
Doench, J. G. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR–Cas9. Nat. Biotechnol. 34, 184–191 (2016).
Horlbeck, M. A. et al. Compact and highly active next-generation libraries for CRISPR-mediated gene repression and activation. eLife 5, e19760 (2016).
Michlits, G. et al. Multilayered VBC score predicts sgRNAs that efficiently generate loss-of-function alleles. Nat. Methods 17, 708–716 (2020).
Sanson, K. R. et al. Optimized libraries for CRISPR–Cas9 genetic screens with multiple modalities. Nat. Commun. 9, 5416 (2018).
Gonatopoulos-Pournatzis, T. et al. Genetic interaction mapping and exon-resolution functional genomics with a hybrid Cas9–Cas12a platform. Nat. Biotechnol. 38, 638–648 (2020).
Liu, J. et al. Pooled library screening with multiplexed Cpf1 library. Nat. Commun. 10, 3144 (2019).
Gonçalves, E. et al. Minimal genome-wide human CRISPR–Cas9 library. Genome Biol. 22, 40 (2021).
Aguirre, A. J. et al. Genomic copy number dictates a gene-independent cell response to CRISPR/Cas9 targeting. Cancer Discov. 6, 914–929 (2016).
Gier, R. A. et al. High-performance CRISPR–Cas12a genome editing for combinatorial genetic screening. Nat. Commun. 11, 3455 (2020).
Zhu, S. et al. Guide RNAs with embedded barcodes boost CRISPR-pooled screens. Genome Biol. 20, 20 (2019).
Gasperini, M. et al. A genome-wide framework for mapping gene regulation via cellular genetic screens. Cell 176, 377–390.e19 (2019).
Dixit, A., Kuksenko, O., Feldman, D. & Regev, A. Shuffle-Seq: en masse combinatorial encoding for n-way genetic interaction screens. Preprint at bioRxiv https://doi.org/10.1101/861443v1 (2019).
Diehl, V. et al. Minimized combinatorial CRISPR screens identify genetic interactions in autophagy. Nucleic Acids Res. 49, 5684–5704 (2021).
Du, D. et al. Genetic interaction mapping in mammalian cells using CRISPR interference. Nat. Methods 14, 577–580 (2017).
Han, K. et al. Synergistic drug combinations for cancer identified in a CRISPR screen for pairwise genetic interactions. Nat. Biotechnol. 35, 463–474 (2017).
Horlbeck, M. A. et al. Mapping the genetic landscape of human cells. Cell 174, 953–967.e22 (2018).
Shen, J. P. et al. Combinatorial CRISPR–Cas9 screens for de novo mapping of genetic interactions. Nat. Methods 14, 573–576 (2017).
Campa, C. C., Weisbach, N. R., Santinha, A. J., Incarnato, D. & Platt, R. J. Multiplexed genome engineering by Cas12a and CRISPR arrays encoded on single transcripts. Nat. Methods 16, 887–893 (2019).
Chow, R. D. et al. In vivo profiling of metastatic double knockouts through CRISPR–Cpf1 screens. Nat. Methods 16, 405–408 (2019).
Nagy, T. & Kampmann, M. CRISPulator: a discrete simulation tool for pooled genetic screens. BMC Bioinformatics 18, 347 (2017).
Kearney, C. J. et al. Tumor immune evasion arises through loss of TNF sensitivity. Sci. Immunol. 3, eaar3451 (2018).
Patel, S. J. et al. Identification of essential genes for cancer immunotherapy. Nature 548, 537–542 (2017).
Tzelepis, K. et al. A CRISPR dropout screen identifies genetic vulnerabilities and therapeutic targets in acute myeloid leukemia. Cell Rep. 17, 1193–1205 (2016).
de Almeida, M. et al. AKIRIN2 controls the nuclear import of proteasomes in vertebrates. Nature 599, 491–496 (2021).
Bubeck, F. et al. Engineered anti-CRISPR proteins for optogenetic control of CRISPR–Cas9. Nat. Methods 15, 924–927 (2018).
Carlson-Stevermer, J. et al. CRISPRoff enables spatio-temporal control of CRISPR editing. Nat. Commun. 11, 5041 (2020).
Gilbert, L. A. et al. Genome-scale CRISPR-mediated control of gene repression and activation. Cell 159, 647–661 (2014).
Birsoy, K. et al. An essential role of the mitochondrial electron transport chain in cell proliferation is to enable aspartate synthesis. Cell 162, 540–551 (2015).
Hart, T. et al. High-resolution CRISPR screens reveal fitness genes and genotype-specific cancer liabilities. Cell 163, 1515–1526 (2015).
Jain, I. H. et al. Genetic screen for cell fitness in high or low oxygen highlights mitochondrial and lipid metabolism. Cell 181, 716–727.e11 (2020).
Wang, T., Wei, J. J., Sabatini, D. M. & Lander, E. S. Genetic screens in human cells using the CRISPR–Cas9 system. Science 343, 80–84 (2014).
Chavez, A. et al. Highly efficient Cas9-mediated transcriptional programming. Nat. Methods 12, 326–328 (2015).
Konermann, S. et al. Genome-scale transcriptional activation by an engineered CRISPR–Cas9 complex. Nature 517, 583–588 (2015).
Gao, Y. et al. Complex transcriptional modulation with orthogonal and inducible dCas9 regulators. Nat. Methods 13, 1043–1049 (2016).
Gilbert, L. A. et al. CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell 154, 442–451 (2013).
Polstein, L. R. & Gersbach, C. A. A light-inducible CRISPR–Cas9 system for control of endogenous gene activation. Nat. Chem. Biol. 11, 198–200 (2015).
Qi, L. S. et al. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152, 1173–1183 (2013).
Tanenbaum, M. E., Gilbert, L. A., Qi, L. S., Weissman, J. S. & Vale, R. D. A protein-tagging system for signal amplification in gene expression and fluorescence imaging. Cell 159, 635–646 (2014).
Zalatan, J. G. et al. Engineering complex synthetic transcriptional programs with CRISPR RNA scaffolds. Cell 160, 339–350 (2015).
Alerasool, N., Segal, D., Lee, H. & Taipale, M. An efficient KRAB domain for CRISPRi applications in human cells. Nat. Methods 17, 1093–1096 (2020).
Wang, H., La Russa, M. & Qi, L. S. CRISPR/Cas9 in genome editing and beyond. Annu. Rev. Biochem. 85, 227–264 (2016).
Xu, X. & Qi, L. S. A CRISPR–dCas toolbox for genetic engineering and synthetic biology. J. Mol. Biol. 431, 34–47 (2019).
Yeo, N. C. et al. An enhanced CRISPR repressor for targeted mammalian gene regulation. Nat. Methods 15, 611–616 (2018).
Morgens, D. W. et al. Genome-scale measurement of off-target activity using Cas9 toxicity in high-throughput screens. Nat. Commun. 8, 15178 (2017).
Rossi, A. et al. Genetic compensation induced by deleterious mutations but not gene knockdowns. Nature 524, 230–233 (2015).
Nuñez, J. K. et al. Genome-wide programmable transcriptional memory by CRISPR-based epigenome editing. Cell 184, 2503–2519.e17 (2021). This study introduces the CRISPRoff method for epigenome editing, which enables stable, mitotically heritable silencing of target genes.
Matharu, N. et al. CRISPR-mediated activation of a promoter or enhancer rescues obesity caused by haploinsufficiency. Science 363, eaau0629 (2019).
Zetsche, B. et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR–Cas system. Cell 163, 759–771 (2015).
Liu, Y. et al. Engineering cell signaling using tunable CRISPR–Cpf1-based transcription factors. Nat. Commun. 8, 2095 (2017).
Xu, X. et al. Engineered miniature CRISPR-Cas system for mammalian genome regulation and editing. Mol. Cell 81, 4333–4345.e4 (2021).
Konermann, S. et al. Transcriptome engineering with RNA-targeting type VI-D CRISPR effectors. Cell 173, 665–676.e14 (2018).
Wessels, H.-H. et al. Massively parallel Cas13 screens reveal principles for guide RNA design. Nat. Biotechnol. 38, 722–727 (2020).
Du, M., Jillette, N., Zhu, J. J., Li, S. & Cheng, A. W. CRISPR artificial splicing factors. Nat. Commun. 11, 2973 (2020).
Gapinske, M. et al. CRISPR-SKIP: programmable gene splicing with single base editors. Genome Biol. 19, 107 (2018).
Mou, H. et al. CRISPR/Cas9-mediated genome editing induces exon skipping by alternative splicing or exon deletion. Genome Biol. 18, 108 (2017).
Amabile, A. et al. Inheritable silencing of endogenous genes by hit-and-run targeted epigenetic editing. Cell 167, 219–232.e14 (2016).
Braun, S. M. G. et al. Rapid and reversible epigenome editing by endogenous chromatin regulators. Nat. Commun. 8, 560 (2017).
Hilton, I. B. et al. Epigenome editing by a CRISPR–Cas9-based acetyltransferase activates genes from promoters and enhancers. Nat. Biotechnol. 33, 510–517 (2015).
Kearns, N. A. et al. Functional annotation of native enhancers with a Cas9–histone demethylase fusion. Nat. Methods 12, 401–403 (2015).
Kim, J.-M. et al. Cooperation between SMYD3 and PC4 drives a distinct transcriptional program in cancer cells. Nucleic Acids Res. 43, 8868–8883 (2015).
Kwon, D. Y., Zhao, Y.-T., Lamonica, J. M. & Zhou, Z. Locus-specific histone deacetylation using a synthetic CRISPR–Cas9-based HDAC. Nat. Commun. 8, 15315 (2017).
O’Geen, H. et al. dCas9-based epigenome editing suggests acquisition of histone methylation is not sufficient for target gene repression. Nucleic Acids Res. 45, 9901–9916 (2017).
Okada, M., Kanamori, M., Someya, K., Nakatsukasa, H. & Yoshimura, A. Stabilization of Foxp3 expression by CRISPR–dCas9-based epigenome editing in mouse primary T cells. Epigenetics Chromatin 10, 24 (2017).
Stepper, P. et al. Efficient targeted DNA methylation with chimeric dCas9 — Dnmt3a — Dnmt3L methyltransferase. Nucleic Acids Res. 45, 1703–1713 (2017).
Xu, X. et al. High-fidelity CRISPR/Cas9- based gene-specific hydroxymethylation rescues gene expression and attenuates renal fibrosis. Nat. Commun. 9, 3509 (2018).
Kim, J. H. et al. LADL: light-activated dynamic looping for endogenous gene expression control. Nat. Methods 16, 633–639 (2019).
Morgan, S. L. et al. Manipulation of nuclear architecture through CRISPR-mediated chromosomal looping. Nat. Commun. 8, 15993 (2017).
Wang, H. et al. CRISPR-mediated programmable 3D genome positioning and nuclear organization. Cell 175, 1405–1417.e14 (2018).
Liu, X. M., Zhou, J., Mao, Y., Ji, Q. & Qian, S. B. Programmable RNA N6-methyladenosine editing by CRISPR–Cas9 conjugates. Nat. Chem. Biol. 15, 865–871 (2019).
O’Connell, M. R. et al. Programmable RNA recognition and cleavage by CRISPR/Cas9. Nature 516, 263–266 (2014).
Rees, H. A. & Liu, D. R. Base editing: precision chemistry on the genome and transcriptome of living cells. Nat. Rev. Genet. 19, 770–788 (2018).
Cheng, L. et al. Single-nucleotide-level mapping of DNA regulatory elements that control fetal hemoglobin expression. Nat. Genet. 53, 869–880 (2021).
Gaudelli, N. M. et al. Programmable base editing of T to G C in genomic DNA without DNA cleavage. Nature 551, 464–471 (2017).
Komor, A. C. et al. Improved base excision repair inhibition and bacteriophage Mu Gam protein yields C:G-to-T:A base editors with higher efficiency and product purity. Sci. Adv. 3, eaao4774 (2017).
Molla, K. A. & Yang, Y. CRISPR/Cas-mediated base editing: technical considerations and practical applications. Trends Biotechnol. 37, 1121–1142 (2019).
Yeh, W.-H., Chiang, H., Rees, H. A., Edge, A. S. B. & Liu, D. R. In vivo base editing of post-mitotic sensory cells. Nat. Commun. 9, 2184 (2018).
Koblan, L. W. et al. Efficient C•G-to-G•C base editors developed using CRISPRi screens, target-library analysis, and machine learning. Nat. Biotechnol. 39, 1414–1425 (2021).
Arbab, M. et al. Determinants of base editing outcomes from target library analysis and machine learning. Cell 182, 463–480.e30 (2020).
Gehrke, J. M. et al. An APOBEC3a–Cas9 base editor with minimized bystander and off-target activities. Nat. Biotechnol. 36, 977 (2018).
Grünewald, J. et al. Transcriptome-wide off-target RNA editing induced by CRISPR-guided DNA base editors. Nature 569, 433–437 (2019).
Cox, D. B. T. et al. RNA editing with CRISPR–Cas13. Science 358, 1019–1027 (2017).
Abudayyeh, O. O. et al. A cytosine deaminase for programmable single-base RNA editing. Science 365, 382–386 (2019).
Shi, J. et al. Discovery of cancer drug targets by CRISPR–Cas9 screening of protein domains. Nat. Biotechnol. 33, 661–667 (2015).
Li, K., Wang, G., Andersen, T., Zhou, P. & Pu, W. T. Optimization of genome engineering approaches with the CRISPR/Cas9 system. PLoS ONE 9, e105779 (2014).
Anzalone, A. V. et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576, 149–157 (2019). This study describes CRISPR prime editing, which enables targeted introduction of short DNA sequences based on a single template.
Chen, P. J. et al. Enhanced prime editing systems by manipulating cellular determinants of editing outcomes. Cell 184, 5635–5652 (2021).
Nelson, J. W. et al. Engineered pegRNAs improve prime editing efficiency. Nat. Biotechnol. https://doi.org/10.1038/s41587-021-01039-7 (2021).
Chow, R. D., Chen, J. S., Shen, J. & Chen, S. A web tool for the design of prime-editing guide RNAs. Nat. Biomed. Eng. 5, 190–194 (2021).
Hsu, J. Y. et al. PrimeDesign software for rapid and simplified design of prime editing guide RNAs. Nat. Commun. 12, 1034 (2021).
Kim, H. K. et al. Predicting the efficiency of prime editing guide RNAs in human cells. Nat. Biotechnol. 39, 198–206 (2021).
Erwood, S. et al. Saturation variant interpretation using CRISPR prime editing. Preprint at bioRxiv https://doi.org/10.1101/2021.05.11.443710v1 (2021).
Choi, J. et al. Precise genomic deletions using paired prime editing. Nat. Biotechnol. https://doi.org/10.1038/s41587-021-01025-z (2021).
Jiang, T., Zhang, X.-O., Weng, Z. & Xue, W. Deletion and replacement of long genomic sequences using prime editing. Nat. Biotechnol. https://doi.org/10.1038/s41587-021-01026-y (2021).
Anzalone, A. V. et al. Programmable deletion, replacement, integration and inversion of large DNA sequences with twin prime editing. Nat. Biotechnol. https://doi.org/10.1038/s41587-021-01133-w (2021).
Ioannidi, E. I. et al. Drag-and-drop genome insertion without DNA cleavage with CRISPR-directed integrases. Preprint at bioRxiv https://doi.org/10.1101/2021.11.01.466786v1 (2021).
Potting, C. et al. Genome-wide CRISPR screen for PARKIN regulators reveals transcriptional repression as a determinant of mitophagy. Proc. Natl Acad. Sci. USA 115, E180–E189 (2018).
Wong, N. M. et al. Engineering digitizer circuits for chemical and genetic screens in human cells. Nat. Commun. 12, 6150 (2021).
Haney, M. S. et al. Identification of phagocytosis regulators using magnetic genome-wide CRISPR screens. Nat. Genet. 50, 1716–1727 (2018).
Mair, B. et al. High-throughput genome-wide phenotypic screening via immunomagnetic cell sorting. Nat. Biomed. Eng. 3, 796–805 (2019).
Mair, B. et al. Essential gene profiles for human pluripotent stem cells identify uncharacterized genes and substrate dependencies. Cell Rep. 27, 599–615.e12 (2019).
Shifrut, E. et al. Genome-wide CRISPR screens in primary human T cells reveal key regulators of immune function. Cell 175, 1958–1971.e15 (2018).
Dong, M. B. et al. Systematic immunotherapy target discovery using genome-scale in vivo CRISPR screens in CD8 T cells. Cell 178, 1189–1204.e23 (2019). This study highlights the feasibility of in vivo CRISPR screening in T cells and its utility for immunotherapy target discovery.
Gurusamy, D. et al. Multi-phenotype CRISPR–Cas9 screen identifies p38 kinase as a target for adoptive immunotherapies. Cancer Cell 37, 818–833.e9 (2020).
Shang, W. et al. Genome-wide CRISPR screen identifies FAM49B as a key regulator of actin dynamics and T cell activation. Proc. Natl Acad. Sci. USA 115, E4051–E4060 (2018).
Chen, Z. et al. In vivo CD8+ T cell CRISPR screening reveals control by Fli1 in infection and cancer. Cell 184, 1262–1280.e22 (2021).
Henriksson, J. et al. Genome-wide CRISPR screens in T helper cells reveal pervasive crosstalk between activation and differentiation. Cell 176, 882–896.e18 (2019).
LaFleur, M. W. et al. PTPN2 regulates the generation of exhausted CD8+ T cell subpopulations and restrains tumor immunity. Nat. Immunol. 20, 1335–1347 (2019).
Fulco, C. P. et al. Activity-by-contact model of enhancer-promoter regulation from thousands of CRISPR perturbations. Nat. Genet. 51, 1664–1669 (2019).
Reilly, S. K. et al. Direct characterization of cis-regulatory elements and functional dissection of complex genetic associations using HCR–FlowFISH. Nat. Genet. 53, 1166–1176 (2021).
Wroblewska, A. et al. Protein barcodes enable high-dimensional single-cell CRISPR screens. Cell 175, 1141–1155.e16 (2018).
Adamson, B. et al. A multiplexed single-cell CRISPR screening platform enables systematic dissection of the unfolded protein response. Cell 167, 1867–1882.e21 (2016).
Dixit, A. et al. Perturb-seq: dissecting molecular circuits with scalable single-cell RNA profiling of pooled genetic screens. Cell 167, 1853–1866.e17 (2016).
Jaitin, D. A. et al. Dissecting immune circuits by linking CRISPR-pooled screens with single-cell RNA-seq. Cell 167, 1883–1896.e15 (2016).
Datlinger, P. et al. Pooled CRISPR screening with single-cell transcriptome readout. Nat. Methods 14, 297–301 (2017). Together with Adamson et al. (2016), Dixit et al. (2016) and Jaitin et al. (2016), these studies introduce CRISPR screens with single-cell sequencing read-out as a broadly useful method for dissecting gene regulation and cell states.
Xie, S., Duan, J., Li, B., Zhou, P. & Hon, G. C. Multiplexed engineering and analysis of combinatorial enhancer activity in single cells. Mol. Cell 66, 285–299.e5 (2017).
Jin, X. et al. In vivo Perturb-seq reveals neuronal and glial abnormalities associated with autism risk genes. Science 370, aaz6063 (2020).
Hill, A. J. et al. On the design of CRISPR-based single-cell molecular screens. Nat. Methods 15, 271–274 (2018).
Adamson, B., Norman, T. M., Jost, M. & Weissman, J. S. Approaches to maximize sgRNA-barcode coupling in Perturb-seq screens. Preprint at bioRxiv https://doi.org/10.1101/298349v1.full (2018).
Mimitou, E. P. et al. Multiplexed detection of proteins, transcriptomes, clonotypes and CRISPR perturbations in single cells. Nat. Methods 16, 409–412 (2019).
Replogle, J. M. et al. Combinatorial single-cell CRISPR screens by direct guide RNA capture and targeted sequencing. Nat. Biotechnol. 38, 954–961 (2020).
Schraivogel, D. et al. Targeted Perturb-seq enables genome-scale genetic screens in single cells. Nat. Methods 17, 629–635 (2020).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019).
Rosenberg, A. B. et al. Single-cell profiling of the developing mouse brain and spinal cord with split-pool barcoding. Science 360, 176–182 (2018).
Datlinger, P. et al. Ultra-high-throughput single-cell RNA sequencing and perturbation screening with combinatorial fluidic indexing. Nat. Methods 18, 635–642 (2021).
Bock, C., Farlik, M. & Sheffield, N. C. Multi-omics of single cells: strategies and applications. Trends Biotechnol. 34, 605–608 (2016).
Chappell, L., Russell, A. J. C. & Voet, T. Single-cell (multi)omics technologies. Annu. Rev. Genomics Hum. Genet. 19, 15–41 (2018).
Liscovitch-Brauer, N. et al. Profiling the genetic determinants of chromatin accessibility with scalable single-cell CRISPR screens. Nat. Biotechnol. 39, 1270–1277 (2021).
Pierce, S. E., Granja, J. M. & Greenleaf, W. J. High-throughput single-cell chromatin accessibility CRISPR screens enable unbiased identification of regulatory networks in cancer. Nat. Commun. 12, 2969 (2021).
Rubin, A. J. et al. Coupled single-cell CRISPR screening and epigenomic profiling reveals causal gene regulatory networks. Cell 176, 361–376.e17 (2019).
Frangieh, C. J. et al. Multimodal pooled Perturb-CITE-seq screens in patient models define mechanisms of cancer immune evasion. Nat. Genet. 53, 332–341 (2021).
Papalexi, E. et al. Characterizing the molecular regulation of inhibitory immune checkpoints with multimodal single-cell screens. Nat. Genet. 53, 322–331 (2021).
Lin, S., Schorpp, K., Rothenaigner, I. & Hadian, K. Image-based high-content screening in drug discovery. Drug Discov. Today 25, 1348–1361 (2020).
Wheeler, E. C. et al. Pooled CRISPR screens with imaging on microraft arrays reveals stress granule-regulatory factors. Nat. Methods 17, 636–642 (2020).
Hasle, N. et al. High-throughput, microscope-based sorting to dissect cellular heterogeneity. Mol. Syst. Biol. 16, e9442 (2020).
Kanfer, G. et al. Image-based pooled whole-genome CRISPRi screening for subcellular phenotypes. J. Cell Biol. 220, e202006180 (2021).
Yan, X. et al. High-content imaging-based pooled CRISPR screens in mammalian cells. J. Cell Biol. 220, e202008158 (2021).
Emanuel, G., Moffitt, J. R. & Zhuang, X. High-throughput, image-based screening of pooled genetic-variant libraries. Nat. Methods 14, 1159–1162 (2017).
Lawson, M. J. et al. In situ genotyping of a pooled strain library after characterizing complex phenotypes. Mol. Syst. Biol. 13, 947 (2017).
Camsund, D. et al. Time-resolved imaging-based CRISPRi screening. Nat. Methods 17, 86–92 (2020).
Chen, K. H., Boettiger, A. N., Moffitt, J. R., Wang, S. & Zhuang, X. RNA imaging. Spatially resolved, highly multiplexed RNA profiling in single cells. Science 348, aaa6090 (2015).
Wang, C., Lu, T., Emanuel, G., Babcock, H. P. & Zhuang, X. Imaging-based pooled CRISPR screening reveals regulators of lncRNA localization. Proc. Natl Acad. Sci. USA 116, 10842–10851 (2019).
Feldman, D. et al. Optical pooled screens in human cells. Cell 179, 787–799.e17 (2019). Together with Camsund et al. (2020) and Wang et al. (2019), these studies demonstrate pooled CRISPR screening with spatial imaging read-outs.
Ke, R. et al. In situ sequencing for RNA analysis in preserved tissue and cells. Nat. Methods 10, 857–860 (2013).
Lee, J. H. et al. Highly multiplexed subcellular RNA sequencing in situ. Science 343, 1360–1363 (2014).
Li, W. et al. MAGeCK enables robust identification of essential genes from genome-scale CRISPR/Cas9 knockout screens. Genome Biol. 15, 554 (2014).
Meyers, R. M. et al. Computational correction of copy number effect improves specificity of CRISPR–Cas9 essentiality screens in cancer cells. Nat. Genet. 49, 1779–1784 (2017).
Jeong, H.-H., Kim, S. Y., Rousseaux, M. W. C., Zoghbi, H. Y. & Liu, Z. β-Binomial modeling of CRISPR pooled screen data identifies target genes with greater sensitivity and fewer false negatives. Genome Res. 29, 999–1008 (2019).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Luecken, M. D. & Theis, F. J. Current best practices in single-cell RNA-seq analysis: a tutorial. Mol. Syst. Biol. 15, e8746 (2019).
Stuart, T. & Satija, R. Integrative single-cell analysis. Nat. Rev. Genet. 20, 257–272 (2019).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902.e21 (2019).
Wolf, F. A., Angerer, P. & Theis, F. J. ScanPy: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Caicedo, J. C. et al. Data-analysis strategies for image-based cell profiling. Nat. Methods 14, 849–863 (2017).
McQuin, C. et al. CellProfiler 3.0: next-generation image processing for biology. PLoS Biol. 16, e2005970 (2018).
Pau, G., Fuchs, F., Sklyar, O., Boutros, M. & Huber, W. EBImage — an R package for image processing with applications to cellular phenotypes. Bioinformatics 26, 979–981 (2010).
Li, W. et al. Quality control, modeling, and visualization of CRISPR screens with MAGeCK-VISPR. Genome Biol. 16, 281 (2015).
Hart, T., Brown, K. R., Sircoulomb, F., Rottapel, R. & Moffat, J. Measuring error rates in genomic perturbation screens: gold standards for human functional genomics. Mol. Syst. Biol. 10, 733 (2014).
Daley, T. P. et al. CRISPhieRmix: a hierarchical mixture model for CRISPR pooled screens. Genome Biol. 19, 159 (2018).
Hart, T. & Moffat, J. BAGEL: a computational framework for identifying essential genes from pooled library screens. BMC Bioinformatics 17, 164 (2016).
Imkeller, K., Ambrosi, G., Boutros, M. & Huber, W. gscreend: modelling asymmetric count ratios in CRISPR screens to decrease experiment size and improve phenotype detection. Genome Biol. 21, 53 (2020).
Gonçalves, E. et al. Structural rearrangements generate cell-specific, gene-independent CRISPR–Cas9 loss of fitness effects. Genome Biol. 20, 27 (2019).
Iorio, F. et al. Unsupervised correction of gene-independent cell responses to CRISPR–Cas9 targeting. BMC Genomics 19, 604 (2018).
Bodapati, S., Daley, T. P., Lin, X., Zou, J. & Qi, L. S. A benchmark of algorithms for the analysis of pooled CRISPR screens. Genome Biol. 21, 62 (2020).
Allen, F. et al. JACKS: joint analysis of CRISPR/Cas9 knockout screens. Genome Res. 29, 464–471 (2019).
Wang, B. et al. Integrative analysis of pooled CRISPR genetic screens using MAGeCKFlute. Nat. Protoc. 14, 756–780 (2019).
Colic, M. et al. Identifying chemogenetic interactions from CRISPR screens with DrugZ. Genome Med. 11, 52 (2019).
Burkhardt, D. B. et al. Quantifying the effect of experimental perturbations at single-cell resolution. Nat. Biotechnol. 39, 619–629 (2021).
Duan, J. & Hon, G. FBA: feature barcoding analysis for single cell RNA-seq. Bioinformatics 22, 4266–4268 (2021).
Liberzon, A. et al. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Szklarczyk, D. et al. The STRING database in 2017: quality-controlled protein–protein association networks, made broadly accessible. Nucleic Acids Res. 45, D362–D368 (2017).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Sheffield, N. C. & Bock, C. LOLA: enrichment analysis for genomic region sets and regulatory elements in R and Bioconductor. Bioinformatics 32, 587–589 (2016).
Grevet, J. D. et al. Domain-focused CRISPR screen identifies HRI as a fetal hemoglobin regulator in human erythroid cells. Science 361, 285–290 (2018).
Sher, F. et al. Rational targeting of a NuRD subcomplex guided by comprehensive in situ mutagenesis. Nat. Genet. 51, 1149–1159 (2019).
Dixon, G. et al. QSER1 protects DNA methylation valleys from de novo methylation. Science 372, eabd0875 (2021).
Cheng, W. et al. CRISPR–Cas9 screens identify the RNA helicase DDX3X as a repressor of C9ORF72 (GGGGCC)n repeat-associated non-AUG translation. Neuron 104, 885–898.e8 (2019).
Ebright, R. Y. et al. Deregulation of ribosomal protein expression and translation promotes breast cancer metastasis. Science 367, 1468–1473 (2020).
Breslow, D. K. et al. A CRISPR-based screen for Hedgehog signaling provides insights into ciliary function and ciliopathies. Nat. Genet. 50, 460–471 (2018).
Pusapati, G. V. et al. CRISPR screens uncover genes that regulate target cell sensitivity to the morphogen Sonic Hedgehog. Dev. Cell 44, 113–129.e8 (2018).
Chen, S. et al. Genome-wide CRISPR screen in a mouse model of tumor growth and metastasis. Cell 160, 1246–1260 (2015).
Shalem, O. et al. Genome-scale CRISPR–Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Li, Q. V. et al. Genome-scale screens identify JNK–JUN signaling as a barrier for pluripotency exit and endoderm differentiation. Nat. Genet. 51, 999–1010 (2019).
Liu, Y. et al. CRISPR activation screens systematically identify factors that drive neuronal fate and reprogramming. Cell Stem Cell 23, 758–771.e8 (2018).
Condon, K. J. et al. Genome-wide CRISPR screens reveal multitiered mechanisms through which mTORC1 senses mitochondrial dysfunction. Proc. Natl Acad. Sci. USA 118, e2022120118 (2021).
Tian, R. et al. Genome-wide CRISPRi/a screens in human neurons link lysosomal failure to ferroptosis. Nat. Neurosci. 24, 1020–1034 (2021).
Morita, K. et al. Genome-wide CRISPR screen identifies TMEM41B as a gene required for autophagosome formation. J. Cell Biol. 217, 3817–3828 (2018).
Shoemaker, C. J. et al. CRISPR screening using an expanded toolkit of autophagy reporters identifies TMEM41B as a novel autophagy factor. PLoS Biol. 17, e2007044 (2019).
McFaline-Figueroa, J. L. et al. A pooled single-cell genetic screen identifies regulatory checkpoints in the continuum of the epithelial-to-mesenchymal transition. Nat. Genet. 51, 1389–1398 (2019).
Dede, M., McLaughlin, M., Kim, E. & Hart, T. Multiplex enCas12a screens detect functional buffering among paralogs otherwise masked in monogenic Cas9 knockout screens. Genome Biol. 21, 262 (2020).
De Kegel, B. & Ryan, C. J. Paralog buffering contributes to the variable essentiality of genes in cancer cell lines. PLoS Genet. 15, e1008466 (2019).
Wang, T. et al. Gene essentiality profiling reveals gene networks and synthetic lethal interactions with oncogenic ras. Cell 168, 890–903.e15 (2017).
Wang, T. et al. Identification and characterization of essential genes in the human genome. Science 350, 1096–1101 (2015).
Badano, J. L. & Katsanis, N. Beyond Mendel: an evolving view of human genetic disease transmission. Nat. Rev. Genet. 3, 779–789 (2002).
Gurdasani, D., Barroso, I., Zeggini, E. & Sandhu, M. S. Genomics of disease risk in globally diverse populations. Nat. Rev. Genet. 20, 520–535 (2019).
Huang, A., Garraway, L. A., Ashworth, A. & Weber, B. Synthetic lethality as an engine for cancer drug target discovery. Nat. Rev. Drug Discov. 19, 23–38 (2020).
Norman, T. M. et al. Exploring genetic interaction manifolds constructed from rich single-cell phenotypes. Science 365, 786–793 (2019).
Gupta, A. et al. Deep learning in image cytometry: a review. Cytometry A 95, 366–380 (2019).
Lotfollahi, M., Wolf, F. A. & Theis, F. J. scGen predicts single-cell perturbation responses. Nat. Methods 16, 715–721 (2019).
Shendure, J. & Fields, S. Massively parallel genetics. Genetics 203, 617–619 (2016).
Findlay, G. M. et al. Accurate classification of BRCA1 variants with saturation genome editing. Nature 562, 217–222 (2018).
Hanna, R. E. et al. Massively parallel assessment of human variants with base editor screens. Cell 184, 1064–1080.e20 (2021). This study demonstrates that CRISPR screens with base editing are feasible and useful for characterizing genetic variants.
Kweon, J. et al. A CRISPR-based base-editing screen for the functional assessment of BRCA1 variants. Oncogene 39, 30–35 (2020).
Cuella-Martin, R. et al. Functional interrogation of DNA damage response variants with base editing screens. Cell 184, 1081–1097.e19 (2021).
Canver, M. C. et al. BCL11A enhancer dissection by Cas9-mediated in situ saturating mutagenesis. Nature 527, 192–197 (2015).
Tsherniak, A. et al. Defining a cancer dependency map. Cell 170, 564–576.e16 (2017).
Dempster, J. M. et al. Agreement between two large pan-cancer CRISPR–Cas9 gene dependency data sets. Nat. Commun. 10, 5817 (2019). This study shows that CRISPR screens in cancer cell lines can yield consistent and reproducible results across different centres.
Aregger, M. et al. Systematic mapping of genetic interactions for de novo fatty acid synthesis identifies C12orf49 as a regulator of lipid metabolism. Nat. Metab. 2, 499–513 (2020).
Zamanighomi, M. et al. GEMINI: a variational Bayesian approach to identify genetic interactions from combinatorial CRISPR screens. Genome Biol. 20, 137 (2019).
Bryant, H. E. et al. Specific killing of BRCA2-deficient tumours with inhibitors of poly(ADP-ribose) polymerase. Nature 434, 913–917 (2005).
Farmer, H. et al. Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature 434, 917–921 (2005).
Chan, E. M. et al. WRN helicase is a synthetic lethal target in microsatellite unstable cancers. Nature 568, 551–556 (2019).
Kategaya, L., Perumal, S. K., Hager, J. H. & Belmont, L. D. Werner syndrome helicase is required for the survival of cancer cells with microsatellite instability. iScience 13, 488–497 (2019).
Lieb, S. et al. Werner syndrome helicase is a selective vulnerability of microsatellite instability-high tumor cells. eLife 8, e43333 (2019).
Lopes, R. et al. Systematic dissection of transcriptional regulatory networks by genome-scale and single-cell CRISPR screens. Sci. Adv. 7, eabf5733 (2021).
Panganiban, R. A. et al. Genome-wide CRISPR screen identifies suppressors of endoplasmic reticulum stress-induced apoptosis. Proc. Natl Acad. Sci. USA. 116, 13384–13393 (2019).
Kamber, R. A. et al. Inter-cellular CRISPR screens reveal regulators of cancer cell phagocytosis. Nature 597, 549–554 (2021).
Sheffer, M. et al. Genome-scale screens identify factors regulating tumor cell responses to natural killer cells. Nat. Genet. 53, 1196–1206 (2021).
Hussmann, J. A. et al. Mapping the genetic landscape of DNA double-strand break repair. Cell 184, 5653–5669 (2021).
Olivieri, M. et al. A genetic map of the response to DNA damage in human cells. Cell 182, 481–496.e21 (2020).
Manguso, R. T. et al. In vivo CRISPR screening identifies Ptpn2 as a cancer immunotherapy target. Nature 547, 413–418 (2017).
Artegiani, B. et al. Fast and efficient generation of knock-in human organoids using homology-independent CRISPR–Cas9 precision genome editing. Nat. Cell Biol. 22, 321–331 (2020).
Han, K. et al. CRISPR screens in cancer spheroids identify 3D growth-specific vulnerabilities. Nature 580, 136–141 (2020).
Pan, D. et al. A major chromatin regulator determines resistance of tumor cells to T cell-mediated killing. Science 359, 770–775 (2018).
Hess, G. T. et al. Directed evolution using dCas9-targeted somatic hypermutation in mammalian cells. Nat. Methods 13, 1036–1042 (2016).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
Schumann, K. et al. Functional CRISPR dissection of gene networks controlling human regulatory T cell identity. Nat. Immunol. 21, 1456–1466 (2020).
Wei, J. et al. Targeting REGNASE-1 programs long-lived effector T cells for cancer therapy. Nature 576, 471–476 (2019).
Ye, L. et al. In vivo CRISPR screening in CD8 T cells with AAV–Sleeping Beauty hybrid vectors identifies membrane targets for improving immunotherapy for glioblastoma. Nat. Biotechnol. 37, 1302–1313 (2019).
Parnas, O. et al. A genome-wide CRISPR screen in primary immune cells to dissect regulatory networks. Cell 162, 675–686 (2015).
Covarrubias, S. et al. High-throughput CRISPR screening identifies genes involved in macrophage viability and inflammatory pathways. Cell Rep. 33, 108541 (2020).
Schmid-Burgk, J. L. et al. A genome-wide CRISPR (clustered regularly interspaced short palindromic repeats) screen identifies NEK7 as an essential component of NLRP3 inflammasome activation. J. Biol. Chem. 291, 103–109 (2016).
Baggen, J. et al. Genome-wide CRISPR screening identifies TMEM106B as a proviral host factor for SARS-CoV-2. Nat. Genet. 53, 435–444 (2021).
Daniloski, Z. et al. Identification of required host factors for SARS-CoV-2 infection in human cells. Cell 184, 92–105.e16 (2021).
Flynn, R. A. et al. Discovery and functional interrogation of SARS-CoV-2 RNA–host protein interactions. Cell 184, 2394–2411.e16 (2021).
Hoffmann, M., Kleine-Weber, H. & Pöhlmann, S. A multibasic cleavage site in the spike protein of SARS-CoV-2 is essential for infection of human lung cells. Mol. Cell 78, 779–784.e5 (2020).
Hoffmann, H.-H. et al. TMEM41B is a pan-flavivirus host factor. Cell 184, 133–148.e20 (2021).
Schneider, W. M. et al. Genome-scale identification of SARS-CoV-2 and pan-coronavirus host factor networks. Cell 184, 120–132.e14 (2021).
Wang, R. et al. Genetic screens identify host factors for SARS-CoV-2 and common cold coronaviruses. Cell 184, 106–119.e14 (2021).
Wei, J. et al. Genome-wide CRISPR screens reveal host factors critical for SARS-CoV-2 infection. Cell 184, 76–91.e13 (2021).
Zhu, Y. et al. A genome-wide CRISPR screen identifies host factors that regulate SARS-CoV-2 entry. Nat. Commun. 12, 961 (2021).
Theisen, D. J. et al. WDFY4 is required for cross-presentation in response to viral and tumor antigens. Science 362, 694–699 (2018).
Wei, S. C., Duffy, C. R. & Allison, J. P. Fundamental mechanisms of immune checkpoint blockade therapy. Cancer Discov. 8, 1069–1086 (2018).
Huang, H. et al. In vivo CRISPR screening reveals nutrient signaling processes underpinning CD8+ T cell fate decisions. Cell 184, 1245–1261.e21 (2021).
Wang, X. et al. In vivo CRISPR screens identify the E3 ligase Cop1 as a modulator of macrophage infiltration and cancer immunotherapy target. Cell 184, 5357–5374.e22 (2021).
Guil-Luna, S., Sedlik, C. & Piaggio, E. Humanized mouse models to evaluate cancer immunotherapeutics. Annu. Rev. Cancer Biol. 5, 119–136 (2021).
Masopust, D. & Soerens, A. G. Tissue-resident T cells and other resident leukocytes. Annu. Rev. Immunol. 37, 521–546 (2019).
Krausgruber, T. et al. Structural cells are key regulators of organ-specific immune responses. Nature 583, 296–302 (2020).
Heng, T. S. P. & Painter, M. W. & Immunological Genome Project Consortium. The Immunological Genome Project: networks of gene expression in immune cells. Nat. Immunol. 9, 1091–1094 (2008).
Regev, A. et al. The Human Cell Atlas. eLife 6, e27041 (2017).
Adams, D. et al. BLUEPRINT to decode the epigenetic signature written in blood. Nat. Biotechnol. 30, 224–226 (2012).
Shapiro, R. S., Chavez, A. & Collins, J. J. CRISPR-based genomic tools for the manipulation of genetically intractable microorganisms. Nat. Rev. Microbiol. 16, 333–339 (2018).
Todor, H., Silvis, M. R., Osadnik, H. & Gross, C. A. Bacterial CRISPR screens for gene function. Curr. Opin. Microbiol. 59, 102–109 (2021).
Wang, T. et al. Pooled CRISPR interference screening enables genome-scale functional genomics study in bacteria with superior performance. Nat. Commun. 9, 2475 (2018).
Rousset, F. et al. Genome-wide CRISPR–dCas9 screens in E. coli identify essential genes and phage host factors. PLoS Genet. 14, e1007749 (2018).
Peters, J. M. et al. A comprehensive, CRISPR-based functional analysis of essential genes in bacteria. Cell 165, 1493–1506 (2016).
Peters, J. M. et al. Enabling genetic analysis of diverse bacteria with mobile-CRISPRi. Nat. Microbiol. 4, 244–250 (2019).
Rousset, F. et al. The impact of genetic diversity on gene essentiality within the Escherichia coli species. Nat. Microbiol. 6, 301–312 (2021).
Wet, T. Jde et al. Arrayed CRISPRi and quantitative imaging describe the morphotypic landscape of essential mycobacterial genes. eLife 9, e60083 (2020).
Lee, H. H. et al. Functional genomics of the rapidly replicating bacterium Vibrio natriegens by CRISPRi. Nat. Microbiol. 4, 1105–1113 (2019).
Strich, J. R. & Chertow, D. S. CRISPR–Cas biology and its application to infectious diseases. J. Clin. Microbiol. 57, e01307–e01318 (2019).
Liu, X. et al. Exploration of bacterial bottlenecks and Streptococcus pneumoniae pathogenesis by CRISPRi-seq. Cell Host Microbe 29, 107–120.e6 (2021).
Morio, F., Lombardi, L. & Butler, G. The CRISPR toolbox in medical mycology: state of the art and perspectives. PLoS Pathog. 16, e1008201 (2020).
Bryant, J. M., Baumgarten, S., Glover, L., Hutchinson, S. & Rachidi, N. CRISPR in parasitology: not exactly cut and dried! Trends Parasitol. 35, 409–422 (2019).
Sidik, S. M. et al. A genome-wide CRISPR screen in Toxoplasma identifies essential apicomplexan genes. Cell 166, 1423–1435.e12 (2016).
Young, J. et al. A CRISPR platform for targeted in vivo screens identifies Toxoplasma gondii virulence factors in mice. Nat. Commun. 10, 3963 (2019).
Rosiana, S. et al. Comprehensive genetic analysis of adhesin proteins and their role in virulence of Candida albicans. Genetics 217, iyab003 (2021).
Shapiro, R. S. et al. A CRISPR–Cas9-based gene drive platform for genetic interaction analysis in Candida albicans. Nat. Microbiol. 3, 73–82 (2018).
Li, Z. & Kim, K. S. RELATe enables genome-scale engineering in fungal genomics. Sci. Adv. 6, eabb8783 (2020).
Caro, F., Place, N. M. & Mekalanos, J. J. Analysis of lipoprotein transport depletion in Vibrio cholerae using CRISPRi. Proc. Natl Acad. Sci. USA 116, 17013–17022 (2019).
Liu, X. et al. High-throughput CRISPRi phenotyping identifies new essential genes in Streptococcus pneumoniae. Mol. Syst. Biol. 13, 931 (2017).
Shields, R. C. et al. Repurposing the Streptococcus mutans CRISPR–Cas9 system to understand essential gene function. PLoS Pathog. 16, e1008344 (2020).
Jaffe, M. et al. Improved discovery of genetic interactions using CRISPRiSeq across multiple environments. Genome Res. 29, 668–681 (2019).
Jiang, W., Oikonomou, P. & Tavazoie, S. Comprehensive genome-wide perturbations via CRISPR adaptation reveal complex genetics of antibiotic sensitivity. Cell 180, 1002–1017.e31 (2020).
Halder, V., McDonnell, B., Uthayakumar, D., Usher, J. & Shapiro, R. S. Genetic interaction analysis in microbial pathogens: unravelling networks of pathogenesis, antimicrobial susceptibility and host interactions. FEMS Microbiol. Rev. 45, fuaa055 (2020).
Garst, A. D. et al. Genome-wide mapping of mutations at single-nucleotide resolution for protein, metabolic and genome engineering. Nat. Biotechnol. 35, 48–55 (2017).
Mutalik, V. K. et al. High-throughput mapping of the phage resistance landscape in E. coli. PLoS Biol. 18, e3000877 (2020).
Hein, M. Y. & Weissman, J. S. Functional single-cell genomics of human cytomegalovirus infection. Nat. Biotechnol. https://doi.org/10.1038/s41587-021-01059-3 (2021).
Bao, Z. et al. Genome-scale engineering of Saccharomyces cerevisiae with single-nucleotide precision. Nat. Biotechnol. 36, 505–508 (2018).
Donati, S. et al. Multi-omics analysis of CRISPRi-knockdowns identifies mechanisms that buffer decreases of enzymes in E. coli metabolism. Cell Syst. 12, 56–67.e6 (2021).
Li, S. et al. Genome-wide CRISPRi-based identification of targets for decoupling growth from production. ACS Synth. Biol. 9, 1030–1040 (2020).
Yao, L. et al. Pooled CRISPRi screening of the cyanobacterium Synechocystis sp PCC 6803 for enhanced industrial phenotypes. Nat. Commun. 11, 1666 (2020).
Bowman, E. K. et al. Bidirectional titration of yeast gene expression using a pooled CRISPR guide RNA approach. Proc. Natl Acad. Sci. USA 117, 18424–18430 (2020).
Hawkins, J. S. et al. Mismatch-CRISPRi reveals the co-varying expression–fitness relationships of essential genes in Escherichia coli and Bacillus subtilis. Cell Syst. 11, 523–535 (2020).
Beuter, D. et al. Selective enrichment of slow-growing bacteria in a metabolism-wide CRISPRi library with a TIMER protein. ACS Synth. Biol. 7, 2775–2782 (2018).
Guo, X. et al. High-throughput creation and functional profiling of DNA sequence variant libraries using CRISPR–Cas9 in yeast. Nat. Biotechnol. 36, 540–546 (2018).
Després, P. C., Dubé, A. K., Seki, M., Yachie, N. & Landry, C. R. Perturbing proteomes at single residue resolution using base editing. Nat. Commun. 11, 1871 (2020).
Köster, J. & Rahmann, S. Snakemake — a scalable bioinformatics workflow engine. Bioinformatics 28, 2520–2522 (2012).
Cui, Y. et al. CRISP-view: a database of functional genetic screens spanning multiple phenotypes. Nucleic Acids Res. 49, D848–D854 (2021).
Oughtred, R. et al. The BioGRID interaction database: 2019 update. Nucleic Acids Res. 47, D529–D541 (2019).
Liu, H. et al. CRISPR-ERA: a comprehensive design tool for CRISPR-mediated gene editing, repression and activation. Bioinformatics 31, 3676–3678 (2015).
Galonska, C. et al. Genome-wide tracking of dCas9–methyltransferase footprints. Nat. Commun. 9, 597 (2018).
Michlits, G. et al. CRISPR-UMI: single-cell lineage tracing of pooled CRISPR–Cas9 screens. Nat. Methods 14, 1191–1197 (2017).
Schmierer, B. et al. CRISPR/Cas9 screening using unique molecular identifiers. Mol. Syst. Biol. 13, 945 (2017).
Esk, C. et al. A human tissue screen identifies a regulator of ER secretion as a brain-size determinant. Science 370, 935–941 (2020).
Michels, B. E. et al. Pooled in vitro and in vivo CRISPR–Cas9 screening identifies tumor suppressors in human colon organoids. Cell Stem Cell 26, 782–792.e7 (2020).
Ringel, T. et al. Genome-scale CRISPR screening in human intestinal organoids identifies drivers of TGF-β resistance. Cell Stem Cell 26, 431–440.e8 (2020).
Allen, T. M. et al. Humanized immune system mouse models: progress, challenges and opportunities. Nat. Immunol. 20, 770–774 (2019).
Walsh, N. C. et al. Humanized mouse models of clinical disease. Annu. Rev. Pathol. 12, 187–215 (2017).
Kim, E. & Hart, T. Improved analysis of CRISPR fitness screens and reduced off-target effects with the BAGEL2 gene essentiality classifier. Genome Med. 13, 2 (2021).
Hsu, J. Y. et al. CRISPR-SURF: discovering regulatory elements by deconvolution of CRISPR tiling screen data. Nat. Methods 15, 992–993 (2018).
Dempster, J. M. et al. Chronos: a cell population dynamics model of CRISPR experiments that improves inference of gene fitness effects. Genome Biol. 22, 343 (2021).
Vinceti, A. et al. CoRe: a robustly benchmarked R package for identifying core-fitness genes in genome-wide pooled CRISPR-Cas9 screens. BMC Genomics 17, 22 (2021).
Ward, H. N. et al. Analysis of combinatorial CRISPR screens with the Orthrus scoring pipeline. Nat. Protoc. 16, 4766–4798 (2021).
Duan, B. et al. Model-based understanding of single-cell CRISPR screening. Nat. Commun. 10, 2233 (2019).
Wang, L. Single-cell normalization and association testing unifying CRISPR screen and gene co-expression analyses with Normalisr. Nat. Commun. 12, 6395 (2021).
Yang, L. et al. scMAGeCK links genotypes with multiple phenotypes in single-cell CRISPR screens. Genome Biol. 21, 19 (2020).
Beneke, T. et al. Genetic dissection of a Leishmania flagellar proteome demonstrates requirement for directional motility in sand fly infections. PLoS Pathog. 15, e1007828 (2019).
Acknowledgements
The authors thank D. Seruggia for critical reading of the manuscript. C.B. is supported by a European Research Council (ERC) Starting Grant (no. 679146) and Consolidator Grant (no. 101001971) of the European Union’s Horizon 2020 Research and Innovation Programme. G.S. and L.S.Q. are supported by the Li Ka Shing Foundation, the US NIH Common Fund 4D Nucleome Program (U01 EB021240) and a US National Science Foundation CAREER award (award no. 2046650; L.S.Q.). R.S.S. is supported by a Canadian Institutes of Health Research (CIHR) Project Grant (PJT 162195). J.S., J.S.W. and X.Z. are Howard Hughes Medical Institute investigators.
Author information
Authors and Affiliations
Contributions
Introduction (C.B.); Experimentation (C.B., P.D., F.C., M.A.C., M.B.D., K.A.L., T.L., T.M.N., G.S., S.C., M.G., J.M., L.S.Q., J.S., J.S.W. and X.Z.); Results (C.B., B.S., G.S., W.L. and L.S.Q.); Applications (C.B., F.C., M.A.C., M.B.D., K.A.L., L.M., T.M.N., S.C., M.G., J.M., R.S.S., J.S. and J.S.W.); Reproducibility and data deposition (C.B., B.S., G.S., W.L. and L.S.Q.); Limitations and optimizations (C.B.); Outlook (C.B.); Figures (C.B., P.D., L.M., B.S., W.L. and R.S.S.); Overview of the Primer (C.B.); Editing (all authors).
Corresponding author
Ethics declarations
Competing interests
C.B. is a co-founder and scientific advisor of Aelian Biotechnology and Neurolentech. S.C. is a co-founder of EvolveImmune Therapeutics and Cellinfinity Bio. M.G. has performed consultancy for Sanofi, receives research funding from AstraZeneca and GlaxoSmithKline, and is a co-founder of Mosaic Therapeutics. J.M. is a shareholder of Northern Biologics and Pionyr Immunotherapeutics, and a scientific advisor and shareholder of Century Therapeutics and Aelian Biotechnology. L.S.Q. is a co-founder and scientific advisor of Epicrispr Biotechnologies and Refuge Biotechnologies. J.S. is a scientific advisor of Maze Therapeutics, Camp4 Therapeutics, Cajal Biosciences, Adaptive Biotechnologies and Guardant Health, and a co-founder of Scale Bio and Phase Genomics. J.S.W. consults for and holds equity in KSQ Therapeutics, Maze Therapeutics and Tenaya Therapeutics, is a venture partner at 5AM Ventures and is a member of the Amgen Scientific Advisory Board. X.Z. is a co-founder and consultant of Vizgen. The other authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Methods Primers thanks Sabrina D’Agosto, Francesco Iorio and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Addgene: https://www.addgene.org/
DepMap: https://depmap.org/
EBI ArrayExpress: https://www.ebi.ac.uk/arrayexpress/
EBI European Genome-phenome Archive (EGA): https://ega-archive.org/
International Nucleotide Sequence Database Collaboration: https://www.insdc.org/
NCBI database of Genotypes and Phenotypes (dbGAP): https://www.ncbi.nlm.nih.gov/gap/
NCBI Gene Expression Omnibus (GEO): https://www.ncbi.nlm.nih.gov/geo/
Zenodo: https://zenodo.org/
Glossary
- Forward genetics
-
Screening approach in which genes involved in the phenotype of interest are identified by screening genetically perturbed cells.
- Pooled CRISPR screen
-
A technique in which genetically encoded perturbations are introduced in bulk and read out with sequencing or imaging technology.
- Arrayed CRISPR screens
-
A technique in which perturbations are introduced in individual reaction compartments and remain physically separated.
- High-content CRISPR screens
-
Screens combining complex models, perturbations and stimuli with data-rich read-outs.
- Biological replicates
-
Separately conducted repetitions of the same CRISPR screen using cells from different individuals or different passages of a cell line
- Coverage
-
The average number of cells per guide RNA (gRNA) in a CRISPR screen.
- Multiplicity of infection
-
(MOI). The average number of virions (and, by extension, guide RNAs (gRNAs)) delivered per cell during infection.
- Genetic interactions
-
The combined effect of the simultaneous perturbation of several genes, which may deviate from the sum of the individual effects.
- Positive selection screens
-
Also known as enrichment screens. Cells with the phenotype of interest are selected (enriched) in the screens; other cells are depleted.
- Negative selection screens
-
Also known as dropout screens. Cells with the phenotype of interest are depleted in the screen; other cells are maintained.
- Effective dose
-
The intensity of a perturbation (such as a drug or virus) that causes an effect (such as cell death) in a specified percentage of cells.
- Screening hits
-
Target genes identified as being associated with the phenotype of interest in a CRISPR screen.
- scCRISPR-seq
-
An umbrella term for a group of methods that combine pooled CRISPR sequencing with a single-cell sequencing read-out.
- Manifold learning
-
A set of machine learning methods that seek to uncover hidden structures in the data through dimensionality reduction.
- Variants of uncertain significance
-
Genetic variants for which there is insufficient genetic evidence to support or exclude a causal phenotypic effect.
- Saturation genome editing
-
Introduction of many genome edits into a gene or regulatory element, with the goal of comprehensively assessing their phenotypic impact.
- Synthetic lethality
-
Special case of a genetic interaction in which knockouts of two genes are individually tolerated, but their combination is lethal to cells.
- False positives
-
Putative CRISPR screening hits that do not validate, which can be caused by technical biases.
- False negatives
-
Potential CRISPR screening hits that would have validated but were missed, which can be caused by suboptimal screening conditions.
- Population bottlenecks
-
Reductions in the genetic diversity among a pool of cells owing to external events.
- Genetic drift
-
Genetic changes over time in a pool of cells, caused by random and uncontrolled events.
- Unique molecular identifiers
-
Short barcodes that uniquely identify individual DNA or RNA molecules.
Rights and permissions
About this article
Cite this article
Bock, C., Datlinger, P., Chardon, F. et al. High-content CRISPR screening. Nat Rev Methods Primers 2, 8 (2022). https://doi.org/10.1038/s43586-021-00093-4
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s43586-021-00093-4
This article is cited by
-
Parallel CRISPR screens reveal pathways controlling the cell surface levels of the attractant receptor FPR1
Communications Biology (2026)
-
Progress and new challenges in image-based profiling
Molecular Systems Biology (2026)
-
Multidimensional analysis of host–virus interactions using the virus-encoded CRISPR-based direct readout system (VECOS)
Nature Protocols (2026)
-
Elucidating the coordination of RNA processing using short-read and long-read RNA-sequencing methods
Nature Reviews Molecular Cell Biology (2026)
-
A Comprehensive Review on CRISPR-Based Screening and Its Applications
Molecular Biotechnology (2026)