Abstract
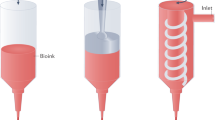
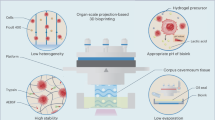
The biofabrication of complex human tissues to recapitulate organ-specific architecture and function requires a combination of engineering control and intrinsic self-assembly. Organoid bioprinting encompasses additive manufacturing approaches that can impart spatial control over the placement of organoids or organoid-forming cells to fabricate multicellular, 3D structures. In particular, bioprinting can be leveraged to control the spatial positioning of printed cells or tissues while maintaining the architecture and physiology of the constituent building blocks. In this Review, we discuss the emerging integration of bioprinting methods and tissue engineering. As bioprinting conventionally involves the patterning of a ‘material’ ink, we characterize cells and organoids as a living material and discuss how such a living material can be manipulated through biofabrication techniques. We focus on continuous and pick-and-place bioprinting methods in which spheroids, organoids or organoid-forming cells comprise the bioink. Additionally, we discuss organoid support baths into which inks are printed. Finally, we highlight how the combination of bioprinting approaches and organoid technology has the potential to improve engineered tissue models of development and disease.
Key points
-
Organoids are in vitro model systems designed to recapitulate in vivo human, tissue-specific phenotypes.
-
Organoid-forming cells, organoids, organoid suspensions and fused organoids can all be considered ‘living materials’, which indicates that they can be directly used in material-based biofabrication approaches.
-
Organoid bioprinting refers to additive manufacturing approaches that impart spatial control over the 3D arrangement of organoids or organoid-forming cells to increase the relevance, reproducibility and complexity of engineered tissues.
-
Organoid bioprinting holds potential for modelling human development and disease, standardizing drug discovery and development and biomanufacturing of human tissues for clinical use.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Rossi, G., Manfrin, A. & Lutolf, M. P. Progress and potential in organoid research. Nat. Rev. Genet. 19, 671–687 (2018).
Fatehullah, A., Tan, S. H. & Barker, N. Organoids as an in vitro model of human development and disease. Nat. Cell Biol. 18, 246–254 (2016).
Hofer, M. & Lutolf, M. P. Engineering organoids. Nat. Rev. Mater. 6, 402–420 (2021).
Lancaster, M. A. & Knoblich, J. A. Organogenesis in a dish: modeling development and disease using organoid technologies. Science 345, 1247125 (2014).
Clevers, H. Modeling development and disease with organoids. Cell 165, 1586–1597 (2016).
Neal, J. T. et al. Organoid modeling of the tumor immune microenvironment. Cell 175, 1972–1988.e16 (2018).
Sloan, S. A., Andersen, J., Pașca, A. M., Birey, F. & Pașca, S. P. Generation and assembly of human brain region-specific three-dimensional cultures. Nat. Protoc. 13, 2062–2085 (2018).
Lancaster, M. A. & Knoblich, J. A. Generation of cerebral organoids from human pluripotent stem cells. Nat. Protoc. 9, 2329–2340 (2014).
Lewis-Israeli, Y. R. et al. Self-assembling human heart organoids for the modeling of cardiac development and congenital heart disease. Nat. Commun. 12, 5142 (2021).
Guan, Y. et al. Human hepatic organoids for the analysis of human genetic diseases. JCI Insight 2, e94954 (2017).
Spence, J. R. et al. Directed differentiation of human pluripotent stem cells into intestinal tissue in vitro. Nature 470, 105–109 (2011).
Kruth, J. P., Leu, M. C. & Nakagawa, T. Progress in additive manufacturing and rapid prototyping. CIRP Ann. 47, 525–540 (1998).
Murphy, S. V. & Atala, A. 3D bioprinting of tissues and organs. Nat. Biotechnol. 32, 773–785 (2014).
Bernal, P. N. et al. Volumetric bioprinting of complex living-tissue constructs within seconds. Adv. Mater. 31, e1904209 (2019).
Loterie, D., Delrot, P. & Moser, C. High-resolution tomographic volumetric additive manufacturing. Nat. Commun. 11, 852 (2020).
Kelly, B. E. et al. Volumetric additive manufacturing via tomographic reconstruction. Science 363, 1075–1079 (2019).
Maharjan, S. et al. Advanced 3D imaging and organoid bioprinting for biomedical research and therapeutic applications. Adv. Drug Deliv. Rev. 208, 115237 (2024).
Hull, S. M., Brunel, L. G. & Heilshorn, S. C. 3D bioprinting of cell-laden hydrogels for improved biological functionality. Adv. Mater. 34, e2103691 (2022).
Sato, T. et al. Long-term expansion of epithelial organoids from human colon, adenoma, adenocarcinoma, and Barrett’s epithelium. Gastroenterology 141, 1762–1772 (2011).
Takasato, M., Er, P. X., Chiu, H. S. & Little, M. H. Generation of kidney organoids from human pluripotent stem cells. Nat. Protoc. 11, 1681–1692 (2016).
Sampaziotis, F. et al. Reconstruction of the mouse extrahepatic biliary tree using primary human extrahepatic cholangiocyte organoids. Nat. Med. 23, 954–963 (2017).
Velasco, S. et al. Individual brain organoids reproducibly form cell diversity of the human cerebral cortex. Nature 570, 523–527 (2019).
Kadoshima, T. et al. Self-organization of axial polarity, inside-out layer pattern, and species-specific progenitor dynamics in human ES cell-derived neocortex. Proc. Natl Acad. Sci. USA 110, 20284–20289 (2013).
Qian, X. et al. Brain-region-specific organoids using mini-bioreactors for modeling ZIKV exposure. Cell 165, 1238–1254 (2016).
Ranga, A. et al. Neural tube morphogenesis in synthetic 3D microenvironments. Proc. Natl Acad. Sci. USA 113, E6831–E6839 (2016).
Abdel Fattah, A. R. et al. Actuation enhances patterning in human neural tube organoids. Nat. Commun. 12, 3192 (2021).
Lenne, P. F. et al. Roadmap for the multiscale coupling of biochemical and mechanical signals during development. Phys. Biol. 18, 041501 (2021).
Aguado, B. A., Mulyasasmita, W., Su, J., Lampe, K. J. & Heilshorn, S. C. Improving viability of stem cells during syringe needle flow through the design of hydrogel cell carriers. Tissue Eng. Pt A 18, 806–815 (2012).
Brassard, J. A., Nikolaev, M., Hübscher, T., Hofer, M. & Lutolf, M. P. Recapitulating macro-scale tissue self-organization through organoid bioprinting. Nat. Mater. 20, 22–29 (2021). This study uses extrusion bioprinting of intestinal stem cells to fabricate intestinal tube-like structures with perfusable lumens.
Lei, M. et al. The mechano-chemical circuit drives skin organoid self-organization. Proc. Natl Acad. Sci. USA 120, e2221982120 (2023).
Ehrig, S. et al. Surface tension determines tissue shape and growth kinetics. Sci. Adv. 5, eaav9394 (2019).
Riccobelli, D. & Bevilacqua, G. Surface tension controls the onset of gyrification in brain organoids. J. Mech. Phys. Solids 134, 103745 (2020).
Fernández, P. A. et al. Surface-tension-induced budding drives alveologenesis in human mammary gland organoids. Nat. Phys. 17, 1130–1136 (2021).
Roth, J. G. et al. Spatially controlled construction of assembloids using bioprinting. Nat. Commun. 14, 4346 (2023). This study develops a magnetic bioprinting approach to spatially pattern individual, specific neural organoids with minimal structural deformation.
Bernal, P. N. et al. Volumetric bioprinting of organoids and optically tuned hydrogels to build liver-like metabolic biofactories. Adv. Mater. 34, 2110054 (2022).
Shin, S. et al. Gelation of uniform interfacial diffusant in embedded 3D printing. Adv. Funct. Mater. 33, 2307435 (2023).
Kratochvil, M. J. et al. Engineered materials for organoid systems. Nat. Rev. Mater. 4, 606–622 (2019).
Hughes, C. S., Postovit, L. M. & Lajoie, G. A. Matrigel: a complex protein mixture required for optimal growth of cell culture. Proteomics 10, 1886–1890 (2010).
Aisenbrey, E. A. & Murphy, W. L. Synthetic alternatives to Matrigel. Nat. Rev. Mater. 5, 539–551 (2020).
Mulero-Russe, A. & García, A. J. Engineered synthetic matrices for human intestinal organoid culture and therapeutic delivery. Adv. Mater. 36, 2307678 (2024).
Gjorevski, N. et al. Designer matrices for intestinal stem cell and organoid culture. Nature 539, 560–564 (2016).
Hunt, D. R. et al. Engineered matrices enable the culture of human patient-derived intestinal organoids. Adv. Sci. 8, 2004705 (2021).
Cruz-Acuña, R. et al. Synthetic hydrogels for human intestinal organoid generation and colonic wound repair. Nat. Cell Biol. 19, 1326–1335 (2017).
Nerger, B. A. et al. 3D hydrogel encapsulation regulates nephrogenesis in kidney organoids. Adv. Mater. 36, e2308325 (2024).
Sorrentino, G. et al. Mechano-modulatory synthetic niches for liver organoid derivation. Nat. Commun. 11, 3416 (2020).
Hernandez-Gordillo, V. et al. Fully synthetic matrices for in vitro culture of primary human intestinal enteroids and endometrial organoids. Biomaterials 254, 120125 (2020).
Suhar, R. A., Huang, M. S., Navarro, R. S., Aviles Rodriguez, G. & Heilshorn, S. C. A library of elastin-like proteins with tunable matrix ligands for in vitro 3D neural cell culture. Biomacromolecules 24, 5926–5939 (2023).
LeSavage, B. L. et al. Engineered matrices reveal stiffness-mediated chemoresistance in patient-derived pancreatic cancer organoids. Nat. Mater. 23, 1138–1149 (2024).
Ligorio, C. & Mata, A. Synthetic extracellular matrices with function-encoding peptides. Nat. Rev. Bioeng. 1, 518–536 (2023).
Roth, J. G. et al. Advancing models of neural development with biomaterials. Nat. Rev. Neurosci. 22, 593–615 (2021).
Chaudhuri, O., Cooper-White, J., Janmey, P. A., Mooney, D. J. & Shenoy, V. B. Effects of extracellular matrix viscoelasticity on cellular behaviour. Nature 584, 535–546 (2020).
Elosegui-Artola, A. et al. Matrix viscoelasticity controls spatiotemporal tissue organization. Nat. Mater. 22, 117–127 (2023).
Peng, Y. H. et al. Dynamic matrices with DNA-encoded viscoelasticity for cell and organoid culture. Nat. Nanotechnol. 18, 1463–1473 (2023).
Yavitt, F. M. et al. In situ modulation of intestinal organoid epithelial curvature through photoinduced viscoelasticity directs crypt morphogenesis. Sci. Adv. 9, eadd5668 (2023).
Haw, M. D. Jamming, two-fluid behavior, and ‘self-filtration’ in concentrated particulate suspensions. Phys. Rev. Lett. 92, 185506 (2004).
Ho, D. L. L. et al. Large-scale production of wholly cellular bioinks via the optimization of human induced pluripotent stem cell aggregate culture in automated bioreactors. Adv. Healthc. Mater. 11, e2201138 (2022).
Skylar-Scott, M. A. et al. Biomanufacturing of organ-specific tissues with high cellular density and embedded vascular channels. Sci. Adv. 5, eaaw2459 (2019). This study incorporates spheroids and organoids as the support bath, into which a sacrificial ink can be printed to yield vascular channels embedded in a high cell density fused tissue-like construct.
Steinberg, M. S. Differential adhesion in morphogenesis: a modern view. Curr. Opin. Genet. Dev. 17, 281–286 (2007).
Mironov, V. et al. Organ printing: tissue spheroids as building blocks. Biomaterials 30, 2164–2174 (2009).
Kelley, K. W. & Pașca, S. P. Human brain organogenesis: toward a cellular understanding of development and disease. Cell 185, 42–61 (2022).
Pașca, S. P. et al. A nomenclature consensus for nervous system organoids and assembloids. Nature 609, 907–910 (2022).
Bagley, J. A., Reumann, D., Bian, S., Lévi-Strauss, J. & Knoblich, J. A. Fused cerebral organoids model interactions between brain regions. Nat. Methods 14, 743–751 (2017).
Birey, F. et al. Assembly of functionally integrated human forebrain spheroids. Nature 545, 54–59 (2017).
Xiang, Y. et al. Fusion of regionally specified hPSC-derived organoids models human brain development and interneuron migration. Cell Stem Cell 21, 383–398.e7 (2017).
Sinha, S. et al. Laminin-associated integrins mediate diffuse intrinsic pontine glioma infiltration and therapy response within a neural assembloid model. Acta Neuropathol. Commun. 12, 71 (2024).
Sachs, N., Tsukamoto, Y., Kujala, P., Peters, P. J. & Clevers, H. Intestinal epithelial organoids fuse to form self-organizing tubes in floating collagen gels. Development 144, 1107–1112 (2017).
Jakab, K. et al. Relating cell and tissue mechanics: implications and applications. Dev. Dyn. 237, 2438–2449 (2008).
Fleming, P. A. et al. Fusion of uniluminal vascular spheroids: a model for assembly of blood vessels. Dev. Dyn. 239, 398–406 (2010).
Frostad, J. M., Collins, M. C. & Leal, L. G. Cantilevered-capillary force apparatus for measuring multiphase fluid interactions. Langmuir 29, 4715–4725 (2013).
Chatzigiannakis, E., Chen, Y., Bachnak, R., Dutcher, C. S. & Vermant, J. Studying coalescence at different lengthscales: from films to droplets. Rheol. Acta 61, 745–759 (2022).
Handorf, A. M., Zhou, Y., Halanski, M. A. & Li, W. J. Tissue stiffness dictates development, homeostasis, and disease progression. Organogenesis 11, 1–15 (2015).
Wolf, K. J., Weiss, J. D., Uzel, S. G. M., Skylar-Scott, M. A. & Lewis, J. A. Biomanufacturing human tissues via organ building blocks. Cell Stem Cell 29, 667–677 (2022).
Kupfer, M. E. et al. In situ expansion, differentiation, and electromechanical coupling of human cardiac muscle in a 3D bioprinted, chambered organoid. Circ. Res. 127, 207–224 (2020).
Lawlor, K. T. et al. Cellular extrusion bioprinting improves kidney organoid reproducibility and conformation. Nat. Mater. 20, 260–271 (2021). This study uses extrusion bioprinting for the reproducible and scalable generation of kidney organoids.
Groll, J. et al. Biofabrication: reappraising the definition of an evolving field. Biofabrication 8, 013001 (2016).
Jakab, K. et al. Tissue engineering by self-assembly of cells printed into topologically defined structures. Tissue Eng. Part A 14, 413–421 (2008). This study uses capillary extrusion of pre-formed spheroids to fabricate fused spheroid assemblies of defined geometries.
Norotte, C., Marga, F. S., Niklason, L. E. & Forgacs, G. Scaffold-free vascular tissue engineering using bioprinting. Biomaterials 30, 5910–5917 (2009).
Mekhileri, N. V. et al. Automated 3D bioassembly of micro-tissues for biofabrication of hybrid tissue engineered constructs. Biofabrication 10, 024103 (2018).
Mekhileri, N. V. et al. Biofabrication of modular spheroids as tumor-scale microenvironments for drug screening. Adv. Healthc. Mater. 12, e2201581 (2023).
Goulart, E. et al. 3D bioprinting of liver spheroids derived from human induced pluripotent stem cells sustain liver function and viability in vitro. Biofabrication 12, 015010 (2019).
De Moor, L. et al. Engineering microvasculature by 3D bioprinting of prevascularized spheroids in photo-crosslinkable gelatin. Biofabrication 13, 045021 (2021).
LeSavage, B. L., Suhar, R. A., Broguiere, N., Lutolf, M. P. & Heilshorn, S. C. Next-generation cancer organoids. Nat. Mater. 21, 143–159 (2022).
Cruz-Acuña, R. et al. Engineered hydrogel reveals contribution of matrix mechanics to esophageal adenocarcinoma and identifies matrix-activated therapeutic targets. J. Clin. Invest. 133, e168146 (2023).
Swaminathan, S., Hamid, Q., Sun, W. & Clyne, A. M. Bioprinting of 3D breast epithelial spheroids for human cancer models. Biofabrication 11, 025003 (2019).
Kim, E. et al. Creation of bladder assembloids mimicking tissue regeneration and cancer. Nature 588, 664–669 (2020).
Andersen, J. et al. Generation of functional human 3D cortico-motor assembloids. Cell 183, 1913–1929.e26 (2020).
Miura, Y. et al. Generation of human striatal organoids and cortico-striatal assembloids from human pluripotent stem cells. Nat. Biotechnol. 38, 1421–1430 (2020).
Schmidt, C. et al. Multi-chamber cardioids unravel human heart development and cardiac defects. Cell 186, 5587–5605.e27 (2023).
Kim, J.-I. et al. Human assembloid model of the ascending neural sensory pathway. Preprint at bioRxiv https://doi.org/10.1101/2024.03.11.584539 (2024).
Nakayama, K. in Biofabrication (eds G. Forgacs & W. Sun) 1–21 (William Andrew Publishing, 2013).
Itoh, M. et al. Scaffold-free tubular tissues created by a bio-3D printer undergo remodeling and endothelialization when implanted in rat aortae. PLoS ONE 10, e0136681 (2015). This study introduces the Kenzan method, which is a method of spheroid assembly that involves skewering pre-formed spheroids on a microneedle array.
Yamasaki, A. et al. Osteochondral regeneration using constructs of mesenchymal stem cells made by bio three-dimensional printing in mini-pigs. J. Orthop. Res. 37, 1398–1408 (2019).
Imamura, T. et al. Biofabricated structures reconstruct functional urinary bladders in radiation-injured rat bladders. Tissue Eng. Part A 24, 1574–1587 (2018).
Murata, D., Arai, K. & Nakayama, K. Scaffold-free bio-3D printing using spheroids as ‘Bio-Inks’ for tissue (re-)construction and drug response tests. Adv. Healthc. Mater. 9, 1901831 (2020).
Ayan, B. et al. Aspiration-assisted bioprinting for precise positioning of biologics. Sci. Adv. 6, eaaw5111 (2020). This study describes aspiration-assisted bioprinting for spatially positioning spheroids by using vacuum aspiration to select and deposit individual spheroids.
Heo, D. N. et al. Aspiration-assisted bioprinting of co-cultured osteogenic spheroids for bone tissue engineering. Biofabrication 13, 015013 (2020).
Daly, A. C., Davidson, M. D. & Burdick, J. A. 3D bioprinting of high cell-density heterogeneous tissue models through spheroid fusion within self-healing hydrogels. Nat. Commun. 12, 753 (2021).
Kim, W., Gwon, Y., Park, S., Kim, H. & Kim, J. Therapeutic strategies of three-dimensional stem cell spheroids and organoids for tissue repair and regeneration. Bioact. Mater. 19, 50–74 (2023).
Brunel, L. G., Hull, S. M. & Heilshorn, S. C. Engineered assistive materials for 3D bioprinting: support baths and sacrificial inks. Biofabrication 14, 032001 (2022).
Becker, M., Gurian, M., Schot, M. & Leijten, J. Aqueous two-phase enabled low viscosity 3D (LoV3D) bioprinting of living matter. Adv. Sci. 10, e2204609 (2023).
Sakai, S. & Morita, T. One-step FRESH bioprinting of low-viscosity silk fibroin inks. ACS Biomater. Sci. Eng. 8, 2589–2597 (2022).
Tan, E. Y. S., Suntornnond, R. & Yeong, W. Y. High-resolution novel indirect bioprinting of low-viscosity cell-laden hydrogels via model-support bioink interaction. 3D Print. Addit. Manuf. 8, 69–78 (2021).
Brunel, L. G. et al. Embedded 3D bioprinting of collagen inks into microgel baths to control hydrogel microstructure and cell spreading. Adv. Healthc. Mater. 13, 2303325 (2024).
Jakab, K., Damon, B., Neagu, A., Kachurin, A. & Forgacs, G. Three-dimensional tissue constructs built by bioprinting. Biorheology 43, 509–513 (2006).
Wu, W., DeConinck, A. & Lewis, J. A. Omnidirectional printing of 3D microvascular networks. Adv. Mater. 23, H178–H183 (2011).
Bhattacharjee, T. et al. Writing in the granular gel medium. Sci. Adv. 1, e1500655 (2015).
Highley, C. B., Rodell, C. B. & Burdick, J. A. Direct 3D printing of shear-thinning hydrogels into self-healing hydrogels. Adv. Mater. 27, 5075–5079 (2015).
Hinton, T. J. et al. Three-dimensional printing of complex biological structures by freeform reversible embedding of suspended hydrogels. Sci. Adv. 1, e1500758 (2015).
Lee, A. et al. 3D bioprinting of collagen to rebuild components of the human heart. Science 365, 482–487 (2019).
Noor, N. et al. 3D printing of personalized thick and perfusable cardiac patches and hearts. Adv. Sci. 6, 1900344 (2019).
Moxon, S. R. et al. Suspended manufacture of biological structures. Adv. Mater. 29, 1605594 (2017).
Compaan, A. M., Song, K. & Huang, Y. Gellan fluid gel as a versatile support bath material for fluid extrusion bioprinting. ACS Appl. Mater. Interfaces 11, 5714–5726 (2019).
Jin, Y., Compaan, A., Chai, W. & Huang, Y. Functional nanoclay suspension for printing-then-solidification of liquid materials. ACS Appl. Mater. Interfaces 9, 20057–20066 (2017).
Morley, C. D. et al. Quantitative characterization of 3D bioprinted structural elements under cell generated forces. Nat. Commun. 10, 3029 (2019).
Shin, S. & Hyun, J. Matrix-assisted three-dimensional printing of cellulose nanofibers for paper microfluidics. ACS Appl. Mater. Interfaces 9, 26438–26446 (2017).
Bertassoni, L. E. et al. Direct-write bioprinting of cell-laden methacrylated gelatin hydrogels. Biofabrication 6, 024105 (2014).
Bertassoni, L. E. et al. Hydrogel bioprinted microchannel networks for vascularization of tissue engineering constructs. Lab Chip 14, 2202–2211 (2014).
Hammer, J., Han, L. H., Tong, X. & Yang, F. A facile method to fabricate hydrogels with microchannel-like porosity for tissue engineering. Tissue Eng. Part C Methods 20, 169–176 (2014).
Wang, X. Y. et al. Engineering interconnected 3D vascular networks in hydrogels using molded sodium alginate lattice as the sacrificial template. Lab Chip 14, 2709–2716 (2014).
Visser, J. et al. Biofabrication of multi-material anatomically shaped tissue constructs. Biofabrication 5, 035007 (2013).
Pimentel, C. R. et al. Three-dimensional fabrication of thick and densely populated soft constructs with complex and actively perfused channel network. Acta Biomater. 65, 174–184 (2018).
Miller, J. S. et al. Rapid casting of patterned vascular networks for perfusable engineered three-dimensional tissues. Nat. Mater. 11, 768–774 (2012).
Ozbolat, V., Dey, M., Ayan, B. & Ozbolat, I. T. Extrusion-based printing of sacrificial Carbopol ink for fabrication of microfluidic devices. Biofabrication 11, 034101 (2019).
Seymour, A. J., Shin, S. & Heilshorn, S. C. 3D printing of microgel scaffolds with tunable void fraction to promote cell infiltration. Adv. Healthc. Mater. 10, e2100644 (2021).
Ying, G. L. et al. Aqueous two-phase emulsion bioink-enabled 3D bioprinting of porous hydrogels. Adv. Mater. 30, e1805460 (2018).
Hull, S. M. et al. 3D bioprinting using UNIversal Orthogonal Network (UNION) Bioinks. Adv. Funct. Mater. 31, 2007983 (2021).
Jeon, O. et al. Individual cell-only bioink and photocurable supporting medium for 3D printing and generation of engineered tissues with complex geometries. Mater. Horiz. 6, 1625–1631 (2019).
Gao, T. & Hu, H. H. Deformation of elastic particles in viscous shear flow. J. Comput. Phys. 228, 2132–2151 (2009).
Chen, Y.-L. Inertia- and deformation-driven migration of a soft particle in confined shear and Poiseuille flow. RSC Adv. 4, 17908–17916 (2014).
Dubbin, K., Hori, Y., Lewis, K. K. & Heilshorn, S. C. Dual-stage crosslinking of a gel-phase bioink improves cell viability and homogeneity for 3D bioprinting. Adv. Healthc. Mater. 5, 2488–2492 (2016).
Flégeau, K., Puiggali-Jou, A. & Zenobi-Wong, M. Cartilage tissue engineering by extrusion bioprinting utilizing porous hyaluronic acid microgel bioinks. Biofabrication 14, 034105 (2022).
Park, J. Y. et al. Development of a functional airway-on-a-chip by 3D cell printing. Biofabrication 11, 015002 (2018).
Kolesky, D. B., Homan, K. A., Skylar-Scott, M. A. & Lewis, J. A. Three-dimensional bioprinting of thick vascularized tissues. Proc. Natl Acad. Sci. USA 113, 3179–3184 (2016).
Gao, G., Park, J. Y., Kim, B. S., Jang, J. & Cho, D. W. Coaxial cell printing of freestanding, perfusable, and functional in vitro vascular models for recapitulation of native vascular endothelium pathophysiology. Adv. Healthc. Mater. 7, e1801102 (2018).
Hunziker, E. B., Quinn, T. M. & Häuselmann, H. J. Quantitative structural organization of normal adult human articular cartilage. Osteoarthr. Cartil. 10, 564–572 (2002).
Quinn, T. M., Häuselmann, H. J., Shintani, N. & Hunziker, E. B. Cell and matrix morphology in articular cartilage from adult human knee and ankle joints suggests depth-associated adaptations to biomechanical and anatomical roles. Osteoarthr. Cartil. 21, 1904–1912 (2013).
Blouin, A., Bolender, R. P. & Weibel, E. R. Distribution of organelles and membranes between hepatocytes and nonhepatocytes in the rat liver parenchyma. A stereological study. J. Cell Biol. 72, 441–455 (1977).
Gullino, P. M., Grantham, F. H. & Smith, S. H. The interstitial water space of tumors. Cancer Res. 25, 727–731 (1965).
Donahue, K. M. et al. Dynamic Gd-DTPA enhanced MRI measurement of tissue cell volume fraction. Magn. Reson. Med. 34, 423–432 (1995).
Thulborn, K. et al. Quantitative sodium MRI of the human brain at 9.4 T provides assessment of tissue sodium concentration and cell volume fraction during normal aging. NMR Biomed. 29, 137–143 (2016).
Thulborn, K. R. Gender differences in cell volume fraction (CVF): a structural parameter reflecting the energy efficiency of maintaining the resting membrane potential. NMR Biomed. 35, e4693 (2022).
Stankey, P. P. et al. Embedding biomimetic vascular networks via coaxial sacrificial writing into functional tissue. Adv. Mater. 36, e2401528 (2024).
Skylar-Scott, M. A. et al. Orthogonally induced differentiation of stem cells for the programmatic patterning of vascularized organoids and bioprinted tissues. Nat. Biomed. Eng. 6, 449–462 (2022).
Ritzau-Reid, K. I. et al. Microfibrous scaffolds guide stem cell lumenogenesis and brain organoid engineering. Adv. Mater. 35, e2300305 (2023).
Trentesaux, C., Yamada, T., Klein, O. D. & Lim, W. A. Harnessing synthetic biology to engineer organoids and tissues. Cell Stem Cell 30, 10–19 (2023).
Cederquist, G. Y. et al. Specification of positional identity in forebrain organoids. Nat. Biotechnol. 37, 436–444 (2019).
Toda, S., Blauch, L. R., Tang, S. K. Y., Morsut, L. & Lim, W. A. Programming self-organizing multicellular structures with synthetic cell–cell signaling. Science 361, 156–162 (2018).
Bao, M. et al. Stem cell-derived synthetic embryos self-assemble by exploiting cadherin codes and cortical tension. Nat. Cell Biol. 24, 1341–1349 (2022).
Cakir, B. et al. Engineering of human brain organoids with a functional vascular-like system. Nat. Methods 16, 1169–1175 (2019).
Ribezzi, D. et al. Shaping synthetic multicellular and complex multimaterial tissues via embedded extrusion-volumetric printing of microgels. Adv. Mater. 35, e2301673 (2023).
Tringides, C. M. et al. Tunable conductive hydrogel scaffolds for neural cell differentiation. Adv. Healthc. Mater. 12, 2202221 (2023).
Paunović, N. et al. 4D printing of biodegradable elastomers with tailorable thermal response at physiological temperature. J. Control. Release 361, 417–426 (2023).
Wirtz, D. Particle-tracking microrheology of living cells: principles and applications. Annu. Rev. Biophys. 38, 301–326 (2009).
Squires, T. M. & Mason, T. G. Fluid mechanics of microrheology. Annu. Rev. Fluid Mech. 42, 413–438 (2010).
McGlynn, J. A., Wu, N. & Schultz, K. M. Multiple particle tracking microrheological characterization: fundamentals, emerging techniques and applications. J. Appl. Phys. 127, 201101 (2020).
Schultz, K. M. & Furst, E. M. Microrheology of biomaterial hydrogelators. Soft Matter 8, 6198–6205 (2012).
Woolley, L., Burbidge, A., Vermant, J. & Christakopoulos, F. A microrheological examination of insulin-secreting β-cells in healthy and diabetic-like conditions. Soft Matter 20, 3464–3472 (2024).
Mason, T. G. Estimating the viscoelastic moduli of complex fluids using the generalized Stokes–Einstein equation. Rheol. Acta 39, 371–378 (2000).
Guo, M. et al. Probing the stochastic, motor-driven properties of the cytoplasm using force spectrum microscopy. Cell 158, 822–832 (2014).
Sriram, I., Furst, E. M., DePuit, R. J. & Squires, T. M. Small amplitude active oscillatory microrheology of a colloidal suspension. J. Rheol. 53, 357–381 (2009).
Efremov, Y. M., Okajima, T. & Raman, A. Measuring viscoelasticity of soft biological samples using atomic force microscopy. Soft Matter 16, 64–81 (2020).
Mattice, J. M., Lau, A. G., Oyen, M. L. & Kent, R. W. Spherical indentation load-relaxation of soft biological tissues. J. Mater. Res. 21, 2003–2010 (2006).
Alcaraz, J. et al. Laminin and biomimetic extracellular elasticity enhance functional differentiation in mammary epithelia. EMBO J. 27, 2829–2838 (2008).
Lyu, Q. et al. A soft and ultrasensitive force sensing diaphragm for probing cardiac organoids instantaneously and wirelessly. Nat. Commun. 13, 7259 (2022).
Vyas, V., Solomon, M., D’Souza, G. G. M. & Huey, B. D. Nanomechanical analysis of extracellular matrix and cells in multicellular spheroids. Cell Mol. Bioeng. 12, 203–214 (2019).
Kwon, E.-Y., Kim, Y.-T. & Kim, D.-E. Investigation of penetration force of living cell using an atomic force microscope. J. Mech. Sci. Technol. 23, 1932–1938 (2009).
Hochmuth, R. M. Micropipette aspiration of living cells. J. Biomech. 33, 15–22 (2000).
Barnes, H. A. Rheology: principles, measurements and applications. Powder Technol. 86, 313 (1996).
Ewoldt, R. H., Johnston, M. T. & Caretta, L. M. in Complex Fluids in Biological Systems: Experiment, Theory, and Computation (ed. Spagnolie, S. E.) 207–241 (Springer New York, 2015).
Du, J. et al. A visual, in-expensive, and wireless capillary rheometer for characterizing wholly-cellular bioinks. Small 20, e2304778 (2024).
Tomasina, C., Bodet, T., Mota, C., Moroni, L. & Camarero-Espinosa, S. Bioprinting vasculature: materials, cells and emergent techniques. Materials 12, 2701 (2019).
Chen, E. P., Toksoy, Z., Davis, B. A. & Geibel, J. P. 3D bioprinting of vascularized tissues for in vitro and in vivo applications. Front. Bioeng. Biotechnol. 9, 664188 (2021).
Carmeliet, P. & Jain, R. K. Angiogenesis in cancer and other diseases. Nature 407, 249–257 (2000).
Jain, R. K., Au, P., Tam, J., Duda, D. G. & Fukumura, D. Engineering vascularized tissue. Nat. Biotechnol. 23, 821–823 (2005).
Datta, P., Ayan, B. & Ozbolat, I. T. Bioprinting for vascular and vascularized tissue biofabrication. Acta Biomater. 51, 1–20 (2017).
Zhang, Y., Yu, Y., Chen, H. & Ozbolat, I. T. Characterization of printable cellular micro-fluidic channels for tissue engineering. Biofabrication 5, 025004 (2013).
Miri, A. K. et al. Multiscale bioprinting of vascularized models. Biomaterials 198, 204–216 (2019).
Zhang, Y. et al. Recent advances in 3D bioprinting of vascularized tissues. Mater. Des. 199, 109398 (2021).
Wei, Z. et al. Hydrogels with tunable mechanical plasticity regulate endothelial cell outgrowth in vasculogenesis and angiogenesis. Nat. Commun. 14, 8307 (2023).
Luttun, A. et al. Revascularization of ischemic tissues by PlGF treatment, and inhibition of tumor angiogenesis, arthritis and atherosclerosis by anti-Flt1. Nat. Med. 8, 831–840 (2002).
Richardson, T. P., Peters, M. C., Ennett, A. B. & Mooney, D. J. Polymeric system for dual growth factor delivery. Nat. Biotechnol. 19, 1029–1034 (2001).
Jain, R. K. Molecular regulation of vessel maturation. Nat. Med. 9, 685–693 (2003).
Macklin, B. L. et al. Intrinsic epigenetic control of angiogenesis in induced pluripotent stem cell-derived endothelium regulates vascular regeneration. npj Regen. Med. 7, 28 (2022).
Inglis, S., Kanczler, J. M. & Oreffo, R. O. C. 3D human bone marrow stromal and endothelial cell spheres promote bone healing in an osteogenic niche. FASEB J. 33, 3279–3290 (2019).
Salmon, I. et al. Engineering neurovascular organoids with 3D printed microfluidic chips. Lab Chip 22, 1615–1629 (2022).
Stresser, D. M. et al. Towards in vitro models for reducing or replacing the use of animals in drug testing. Nat. Biomed. Eng. 8, 930–935 (2023).
Mariani, J. et al. FOXG1-dependent dysregulation of GABA/glutamate neuron differentiation in autism spectrum disorders. Cell 162, 375–390 (2015).
Vlachogiannis, G. et al. Patient-derived organoids model treatment response of metastatic gastrointestinal cancers. Science 359, 920–926 (2018).
Sachs, N. et al. A living biobank of breast cancer organoids captures disease heterogeneity. Cell 172, 373–386.e10 (2018).
LeBlanc, V. G. et al. Single-cell landscapes of primary glioblastomas and matched explants and cell lines show variable retention of inter- and intratumor heterogeneity. Cancer Cell 40, 379–392.e9 (2022).
Co, J. Y., Klein, J. A., Kang, S. & Homan, K. A. Toward inclusivity in preclinical drug development: a proposition to start with intestinal organoids. Adv. Biol. 7, e2200333 (2023).
Vandana, J. J., Manrique, C., Lacko, L. A. & Chen, S. Human pluripotent-stem-cell-derived organoids for drug discovery and evaluation. Cell Stem Cell 30, 571–591 (2023).
Chen, X. et al. Antisense oligonucleotide therapeutic approach for Timothy syndrome. Nature 628, 818–825 (2024).
Acknowledgements
The authors thank S. Pasca for helpful conversations about organoid biology and editing of the manuscript. This work was funded in part by a Stanford Cancer Institute Innovation Seed Grant, a Stanford Maternal and Child Health Research Institute Uytengsu-Hamilton 22q11 Neuropsychiatry Seed Grant, the National Institutes of Health (NIH) (R01 HL142718, R01 DK129309 and R01 EB027171), the Advanced Research Projects Agency for Health (ARPA-H) under award number AY1AX000002 and the National Science Foundation (NSF) (DMR 2103812, CBET 2033302). M.S.H. was supported by a NIH F31 Pre-doctoral Fellowship (NS132505) and a Sarafan ChEM-H O’Leary-Thiry Fellowship. F.C. was supported by a Swiss National Science Foundation Postdoc Mobility Fellowship (PN210723).
Author information
Authors and Affiliations
Contributions
M.S.H., F.C. and S.C.H. conceptualized the review. M.S.H., F.C. and J.G.R. researched data for the article and contributed to discussion of content and writing. All authors reviewed and edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
S.C.H. and J.G.R. are listed as inventors on a patent application (US 2023-0357685 A1) that covers some of the technologies described in this manuscript. The other authors declare no competing interests.
Peer review
Peer review information
Nature Reviews Bioengineering thanks Ibrahim Ozbolat and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Huang, M.S., Christakopoulos, F., Roth, J.G. et al. Organoid bioprinting: from cells to functional tissues. Nat Rev Bioeng 3, 126–142 (2025). https://doi.org/10.1038/s44222-024-00268-0
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s44222-024-00268-0
This article is cited by
-
Biomaterials with droplet microfluidics
Nature Reviews Bioengineering (2026)
-
Organoid analytical toolkits
Nature Reviews Bioengineering (2026)
-
Spatial omics in 3D culture model systems: decoding cellular positioning mechanisms and microenvironmental dynamics
Journal of Translational Medicine (2025)
-
Engineering vascularized brain tumor organoids: bridging the gap between models and reality
Biomedical Microdevices (2025)
-
Regenerative Engineering: Evolution and Its Modern Significance
Regenerative Engineering and Translational Medicine (2025)