Abstract
Proteins are manufactured by ribosomes—macromolecular complexes of protein and RNA molecules that are assembled within major nuclear compartments called nucleoli1,2. Existing models suggest that RNA polymerases I and III (Pol I and Pol III) are the only enzymes that directly mediate the expression of the ribosomal RNA (rRNA) components of ribosomes. Here we show, however, that RNA polymerase II (Pol II) inside human nucleoli operates near genes encoding rRNAs to drive their expression. Pol II, assisted by the neurodegeneration-associated enzyme senataxin, generates a shield comprising triplex nucleic acid structures known as R-loops at intergenic spacers flanking nucleolar rRNA genes. The shield prevents Pol I from producing sense intergenic noncoding RNAs (sincRNAs) that can disrupt nucleolar organization and rRNA expression. These disruptive sincRNAs can be unleashed by Pol II inhibition, senataxin loss, Ewing sarcoma or locus-associated R-loop repression through an experimental system involving the proteins RNaseH1, eGFP and dCas9 (which we refer to as ‘red laser’). We reveal a nucleolar Pol-II-dependent mechanism that drives ribosome biogenesis, identify disease-associated disruption of nucleoli by noncoding RNAs, and establish locus-targeted R-loop modulation. Our findings revise theories of labour division between the major RNA polymerases, and identify nucleolar Pol II as a major factor in protein synthesis and nuclear organization, with potential implications for health and disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Data are in the paper, Supplementary Fig. 1 (uncropped blots) and the Source Data files related to Figs. 1–4 and Extended Data Figs. 1–3, 5–8. RNA-seq and DRIP–seq data sets have been deposited at GEO under accession codes GSE115731 and GSE68847. All data and materials are available upon reasonable request. In light of the pandemic, shipping of reagents and materials may be slightly delayed. Source data are provided with this paper.
Code availability
All scripts used to analyse data are available upon request.
References
Boisvert, F. M., van Koningsbruggen, S., Navascués, J. & Lamond, A. I. The multifunctional nucleolus. Nat. Rev. Mol. Cell Biol. 8, 574–585 (2007).
Feric, M. et al. Coexisting liquid phases underlie nucleolar subcompartments. Cell 165, 1686–1697 (2016).
Mekhail, K. & Moazed, D. The nuclear envelope in genome organization, expression and stability. Nat. Rev. Mol. Cell Biol. 11, 317–328 (2010).
Kobayashi, T. & Ganley, A. R. Recombination regulation by transcription-induced cohesin dissociation in rDNA repeats. Science 309, 1581–1584 (2005).
Kaeberlein, M., McVey, M. & Guarente, L. The SIR2/3/4 complex and SIR2 alone promote longevity in Saccharomyces cerevisiae by two different mechanisms. Genes Dev. 13, 2570–2580 (1999).
Mekhail, K. et al. Regulation of ubiquitin ligase dynamics by the nucleolus. J. Cell Biol. 170, 733–744 (2005).
Mitrea, D. M. et al. Self-interaction of NPM1 modulates multiple mechanisms of liquid-liquid phase separation. Nat. Commun. 9, 842 (2018).
Audas, T. E. et al. Adaptation to stressors by systemic protein amyloidogenesis. Dev. Cell 39, 155–168 (2016).
Wang, M. et al. Stress-induced low complexity RNA activates physiological amyloidogenesis. Cell Rep. 24, 1713–1721 (2018).
Caudron-Herger, M. et al. Alu element-containing RNAs maintain nucleolar structure and function. EMBO J. 34, 2758–2774 (2015).
Burger, K. et al. Cyclin-dependent kinase 9 links RNA polymerase II transcription to processing of ribosomal RNA. J. Biol. Chem. 288, 21173–21183 (2013).
Burger, K. et al. Chemotherapeutic drugs inhibit ribosome biogenesis at various levels. J. Biol. Chem. 285, 12416–12425 (2010).
Santos-Pereira, J. M. & Aguilera, A. R loops: new modulators of genome dynamics and function. Nat. Rev. Genet. 16, 583–597 (2015).
Bhatia, V. et al. BRCA2 prevents R-loop accumulation and associates with TREX-2 mRNA export factor PCID2. Nature 511, 362–365 (2014).
Sanz, L. A. et al. Prevalent, dynamic, and conserved R-loop structures associate with specific epigenomic signatures in mammals. Mol. Cell 63, 167–178 (2016).
Abraham, K. J. et al. Intersection of calorie restriction and magnesium in the suppression of genome-destabilizing RNA-DNA hybrids. Nucleic Acids Res. 44, 8870–8884 (2016).
Zentner, G. E., Saiakhova, A., Manaenkov, P., Adams, M. D. & Scacheri, P. C. Integrative genomic analysis of human ribosomal DNA. Nucleic Acids Res. 39, 4949–4960 (2011).
Shiue, C. N., Nematollahi-Mahani, A. & Wright, A. P. Myc-induced anchorage of the rDNA IGS region to nucleolar matrix modulates growth-stimulated changes in higher-order rDNA architecture. Nucleic Acids Res. 42, 5505–5517 (2014).
Groh, M., Albulescu, L. O., Cristini, A. & Gromak, N. Senataxin: genome guardian at the interface of transcription and neurodegeneration. J. Mol. Biol. 429, 3181–3195 (2017).
Skourti-Stathaki, K., Proudfoot, N. J. & Gromak, N. Human senataxin resolves RNA/DNA hybrids formed at transcriptional pause sites to promote Xrn2-dependent termination. Mol. Cell 42, 794–805 (2011).
Steinmetz, E. J. et al. Genome-wide distribution of yeast RNA polymerase II and its control by Sen1 helicase. Mol. Cell 24, 735–746 (2006).
Suraweera, A. et al. Functional role for senataxin, defective in ataxia oculomotor apraxia type 2, in transcriptional regulation. Hum. Mol. Genet. 18, 3384–3396 (2009).
Vasiljeva, L., Kim, M., Terzi, N., Soares, L. M. & Buratowski, S. Transcription termination and RNA degradation contribute to silencing of RNA polymerase II transcription within heterochromatin. Mol. Cell 29, 313–323 (2008).
Mischo, H. E. et al. Yeast Sen1 helicase protects the genome from transcription-associated instability. Mol. Cell 41, 21–32 (2011).
Stamatopoulou, V., Parisot, P., De Vleeschouwer, C. & Lafontaine, D. L. J. Use of the iNo score to discriminate normal from altered nucleolar morphology, with applications in basic cell biology and potential in human disease diagnostics. Nat. Protoc. 13, 2387–2406 (2018).
Gorthi, A. et al. EWS-FLI1 increases transcription to cause R-loops and block BRCA1 repair in Ewing sarcoma. Nature 555, 387–391 (2018); correction 559, E11 (2018).
El Hage, A., French, S. L., Beyer, A. L. & Tollervey, D. Loss of topoisomerase I leads to R-loop-mediated transcriptional blocks during ribosomal RNA synthesis. Genes Dev. 24, 1546–1558 (2010).
Kawauchi, J., Mischo, H., Braglia, P., Rondon, A. & Proudfoot, N. J. Budding yeast RNA polymerases I and II employ parallel mechanisms of transcriptional termination. Genes Dev. 22, 1082–1092 (2008).
Salvi, J. S. et al. Roles for Pbp1 and caloric restriction in genome and lifespan maintenance via suppression of RNA-DNA hybrids. Dev. Cell 30, 177–191 (2014).
Bierhoff, H. et al. Quiescence-induced LncRNAs trigger H4K20 trimethylation and transcriptional silencing. Mol. Cell 54, 675–682 (2014).
Savić, N. et al. lncRNA maturation to initiate heterochromatin formation in the nucleolus is required for exit from pluripotency in ESCs. Cell Stem Cell 15, 720–734 (2014).
Mayer, C., Schmitz, K. M., Li, J., Grummt, I. & Santoro, R. Intergenic transcripts regulate the epigenetic state of rRNA genes. Mol. Cell 22, 351–361 (2006).
Zhao, Z., Sentürk, N., Song, C. & Grummt, I. lncRNA PAPAS tethered to the rDNA enhancer recruits hypophosphorylated CHD4/NuRD to repress rRNA synthesis at elevated temperatures. Genes Dev. 32, 836–848 (2018).
Zhang, H. et al. Targeting CDK9 reactivates epigenetically silenced genes in cancer. Cell 175, 1244–1258 (2018).
Ho, E. C., Donaldson, M. E. & Saville, B. J. Detection of antisense RNA transcripts by strand-specific RT-PCR. Methods Mol. Biol. 630, 125–138 (2010).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44 (W1), W160–W165 (2016).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Agrawal, S. & Ganley, A. R. D. The conservation landscape of the human ribosomal RNA gene repeats. PLoS ONE 13, e0207531 (2018).
Acknowledgements
We thank D. Durocher and H. O. Lee for critical reading of the manuscript; A. F. Palazzo for technical assistance with phase-contrast microscopy; R. Kandel for technical tumour imaging; and F. Chédin for technical assistance with DRIP. K.J.A. is funded by a Canadian Institutes of Health Research (CIHR) Vanier Doctoral Scholarship, Ruggles Innovation Award, and Adel S. Sedra Award. N.K. is supported by a CIHR Scholarship. L.A.O. is funded by an Ontario Graduate Scholarship (OGS). R.O. is funded by a Natural Sciences and Engineering Research Council (NSERC) Doctoral Scholarship. This work was also supported by funds from the National Institutes of Health (NIH; grants K22ES012264, 1R01CA152063, 1R01CA241554), a Voelcker Fund Young Investigator Award and the Cancer Prevention and Research Institute of Texas (CPRIT; grant RP150445) to A.J.R.B.; by CPRIT (grant RP101491), a National Cancer Institute (NCI) T32 postdoctoral training grant (T32CA148724) and a National Center for Advancing Translational Sciences (NCATS) TL1 grant (TL1TR002647) to A.G.; and NCI funding (P30CA054174) to the sequencing facility. Funds were also provided to S.L. from the National Institute of General Medical Sciences (R01GM115342) and the NCI (R01CA200676) of the NIH, and the Sylvester Comprehensive Cancer Center. This work was mainly supported by grants to K.M. from the CIHR (388041, 399687), the Canada Research Chairs Program (CRC; 950-230661), and the Ontario Ministry of Research and Innovation (MRI-ERA; ER13-09-111).
Author information
Authors and Affiliations
Contributions
K.J.A. and K.M. conceived the study. K.M. together with A.J.R.B. or S.L. conceptualized the Ewing sarcoma and in vitro liquid droplet studies. K.J.A. and K.M. wrote the paper. Most DRIP and ChIP revision experiments and related data analysis were by N.K., supervised by K.J.A. General text editing was by J.N.Y.C., N.K., A.S., L.A.O., R.O., R.H. and J.F.G. The sole lead on all experiments was K.J.A. except as indicated here. Leads were: on ‘red laser’, K.J.A., N.K. and J.N.Y.C., supervised by K.M.; on ENCODE data analysis, R.S. supervised by D.D.D.C. and V.T.Y. supervised by K.M.; on Ewing sarcoma genome-wide analysis, A.G. supervised by A.J.R.B.; on SETX-knockout generation, D.Y.Z. supervised by J.F.G.; and on heat-shock, cell fractionation and in vitro work, M.W. and M.B. supervised by S.L. DRIP RNH1 overexpression controls were by P.S.P. and A.A., supervised by R.H., and DRIF controls were by V.P., supervised by M.O. and K.M. Tumour sections were by B.C.D. Contributions supervised by K.J.A. and K.M. were from: J.N.Y.C., A.S. and Y.L. for ncRNA detection by northern blotting and RT–qPCR; from E.V. for immunofluorescence microscopy; from L.A.O. for ss-RT design and experimentation; and from R.O. for FRAP design and experimentation.
Corresponding author
Ethics declarations
Competing interests
D.D.D.C. received research funds, not related to this work, from Pfizer and Nektar Therapeutics. All other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Additional characterization of Pol I and Pol II occupancy at rDNA IGSs.
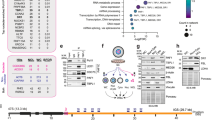
a, Organization of human rDNA repeats. At each rDNA unit, Pol I transcribes an rRNA gene encoding a 47S pre-rRNA that is processed to remove transcribed spacers, such as the 5′-ETS, and generate 18S, 5.8S and 28S rRNA molecules. The IGS constitutes the bulk of each rDNA unit. Ter, rRNA gene terminator. b, c, Specificity controls indicating that targeting Pol II for degradation with a 12-hour α-amanitin (AMN) treatment lowers anti (α)-Pol II pS2 signals in both immunofluorescence (b) and immunoblotting (c). Actin was used as a control for immunoblotting. For gel source data, see Supplementary Fig. 1. d, ChIP showing Pol II pS5 enrichment across rDNA. e, f, The enrichment of active Pol II pS2 and pS5 at rDNA IGS sites is higher than at LINE1 but lower than at β-actin sites. g–k, ChIP experiments showing the enrichment of the indicated proteins across rDNA. l, Comparison of the enrichment of RNA Pol II and Pol I across rDNA reveals the relative overrepresentation of Pol II across IGSs only. b–l, HEK293T (b–g, j–l) or IMR90 (h, i) cells were used; data shown are means ± s.d.; two-tailed t-test, n = 3 biologically independent experiments (d–l); images in b, c are representative of two independent experiments. Data in d–f, j–l and Fig. 1b were from large experimental sets sharing IgG controls. Data in h, i were from large experimental sets sharing IgG controls.
Extended Data Fig. 2 Impact of Pol I and Pol II on IGS ncRNA levels in various cell types.
a, Cell-population-based RNA pulse-chase assay used to assess pre-rRNA synthesis and processing. b, c, Confirmation of the detection of pre-rRNA synthesis (b) and processing (c) by EU–RNA pulse-chase assays, as shown in Fig. 1d, e. d, Trypan blue exclusion assay confirms that the 3-hour Pol II inhibition (iPol) regimens used in our functional assays do not compromise cell viability. e, Ponceau staining shows stable protein levels following Pol II inhibition. Veh, vehicle. f, Treatment with the fast-acting RNA Pol II inhibitor flavopiridol (FP) for 30 min is sufficient to abrogate pre-rRNA processing. g, Human IGS ncRNAs are also detected across the IGSs of diploid HeLa cells and haploid HAP1 cells. h, Pol I promotes and Pol II represses IGS ncRNAs in HeLa cells. i, Nuclear run-on assay showing de novo IGS ncRNA synthesis mediated by Pol II inhibition. j, k, Reverse-transcription experiments showing the effect of combining Pol I and Pol II inhibition on IGS ncRNAs in HEK293T cells (j) and IMR90 cells (k). l, m, Strand-specific RT–qPCR (ss-RT) showing the levels of sense and antisense intergenic ncRNAs (l) and their derived sense/antisense ratios (m) at various IGS sites. n, ss-RT shows that Pol I inhibition decreases and Pol II inhibition increases the sense/antisense ratio of the most abundant IGS ncRNAs. o, Despite the preferential enrichment of Pol II over Pol I across IGSs, Pol II is the least overrepresented relative to Pol I at IGS16 compared with all other IGSs tested. a–o, HEK293T cells were used unless otherwise indicated; data are shown as means ± s.d.; two-tailed t-test (b–d, f) or one-way ANOVA with Dunnett’s multiple comparison test (g, i, k); n = 2 biologically independent experiments (b), n = 4 biologically independent experiments (c, f), and n = 3 biologically independent experiments (d, g–o), except in the case of sense IGS18, for which n = 2 biologically independent samples (l, m); image in e is representative of two independent experiments.
Extended Data Fig. 3 Characteristics of nucleoli and nucleolar proteins in the presence or absence of Pol II inhibition.
a, b, Schematic of a nucleolus, illustrating the localization of LLPS nucleolar subcompartments marked by the resident proteins NPM and UBF (a), which are highly disordered, as revealed using the various short long 2 (VSL2) predictor of natural disordered regions (PONDR) algorithm (b). c, Effects of Pol II inhibition (iPol II) on NPM localization, as shown by immunofluorescence microscopy. Examples of normal and defective phenotypes are respectively marked by magenta and white arrowheads. d, Quantification of the percentage of cells that have any NPM phase-separated body reveals that the fast-acting Pol II inhibitor FP completely disrupts nucleoli before the slower-acting Pol II inhibitor AMN can take effect. Not depicted is the percentage of cells with perturbed nucleolar architecture as evidenced by NPM1 ruffling, which increased from 0.6 ± 4.6% to 63.3 ± 5.7% following the 1-hour FP treatment. e, Pol II inhibition also disrupts NPM localization in IMR90 cells. f, Effects of Pol II inhibition on UBF localization, as shown by immunofluorescence microscopy. Examples of normal and defective phenotypes are respectively marked by magenta and white arrowheads. g, Quantification of the percentage of cells that have any punctate UBF localization confirmed that the fast-acting FP completely disrupts nucleoli before the slower-acting AMN. h, Pol II inhibition triggers various aberrant UBF localization phenotypes, as shown in representative images. i, Global nucleolar disruption following Pol II inhibition, as revealed by phase-contrast microscopy. The fraction of cells with more than three black nucleolar bodies is indicated. j, Live-cell UBF fluorescence recovery after photobleaching (FRAP). Mock control cells were continuously imaged without a photobleaching step. FRAP FP/vehicle rate-constant ratio = 2.3. k, Formerly nucleolar space became Congo red positive after Pol II inhibition. c–k, HEK293T cells were used unless otherwise indicated; data are means ± s.d.; one-way ANOVA with Dunnett’s multiple comparisons test, n = 3 biologically independent experiments (d, g) or n = 5 biologically independent experiments (i); for j, vehicle FRAP cells n = 30, vehicle control cells n = 4, FP FRAP cells n = 15, and FP control cells n = 6; images in e, h, k are representative of two independent experiments. Scale bars, 5 μm (yellow) or 1 μm (white).
Extended Data Fig. 4 Heat shock limits asincRNAs and triggers sincRNA-dependent nucleolar phase transitions.
a, Heat shock (43 °C) rapidly induces the formation of intranucleolar liquid droplets harbouring the ACM-containing VHL protein. b, Gradual amyloid-body (A-body) formation. The stress-induced, mobile and spherical liquid-like foci (yellow arrowhead) gradually transition into irregularly shaped, solid-like amyloid bodies (cyan arrowhead) in cells subjected to heat shock (43 °C)19. c, The appearance of early-stage, ACM-marked, liquid-like foci18,19 in cells subjected to a 15-min heat-shock treatment is abrogated upon siRNA-mediated knockdown of either sincRNA16 or sincRNA22. d, In a cell-free in vitro system, the low-complexity sincRNA (1 μM) forms liquid droplets when mixed with the ACM of human VHL or β-amyloid proteins (25 μM). Droplets were detected using fluorescently labelled RNA (5′FAM) and differential interference contrast (DIC). e, ss-RNA-seq reveals that sincRNA levels increase while asincRNA levels decrease across the IGS following a 30-min heat shock. Heat shock increases sincRNA levels by 607% and decreases asincRNA levels by 38%. a–e, Nucleolar-stress hyperresponsive MCF7 cells were used where applicable; images are representative of two independent experiments; scale bars, 5 μm.
Extended Data Fig. 5 Artificial and natural modulation of sincRNA levels.
a, In HEK293T cells treated with the Pol II inhibitor FP, introduction of ASOs targeting sincRNAs lowers IGS ncRNA levels relative to ASO control-treated cells (CTL). ASO-dependent percentage decreases in sincRNA levels are indicated for each IGS site; the average decrease in total sincRNA levels is 49%. Data are means ± s.d.; two-tailed t-test, n = 3 biologically independent experiments. b–d, In the absence of heat shock, artificial overexpression of sincRNA22 (psincRNA) in nucleolar-stress hyperresponsive MCF7 cells failed to repress rRNA biogenesis (b) or rRNA levels (c), despite the enrichment of sincRNA22 in the nucleolar fraction (d). Plasmid (pCTL), iPol I (LAD), vehicle (DMSO) and GAPDH cell fractionation controls were included. Data are means ± s.d.; n = 2 biologically independent experiments (b, d); two-tailed t-test, n = 3 biologically independent experiments (c); e, Quantification of the number of distinct NPM foci per cell in different cell types. Data are means ± s.d.; one-way ANOVA with Tukey’s multiple comparisons test, n = 5 biologically independent experiments.
Extended Data Fig. 6 Controls related to the disruption of nucleolar structure following Pol II inhibition.
a, b, The disruption of NPM phase separation following Pol II inhibition (a) coincides with time points at which the levels of IGS ncRNAs greatly increase (b; means ± s.d., n = 3 biologically independent experiments). At these time points, no reductions in the levels of the small nucleolar (sno)RNA U8 or Alu RNA were observed. c–e, Treating cells with the Pol II inhibitor FP, with various drugs that disrupt nucleolar morphology through unclear mechanisms (MG132, doxorubicin), with the LLPS/nucleolus disruptor 1,6-hexanediol, or with the global translation inhibitor cycloheximide reveals that only Pol II inhibition simultaneously disrupted NPM phase separation (c) and induced IGS ncRNA levels (d, e). Shown are representative anti-NPM immunofluorescence images (c) and two different visual representations of ncRNA levels as detected by RT–qPCR (d, e); n = 3 biologically independent experiments. In the scatter plot (e), each circle represents the value of one IGS site from one of three biological replicates. Scale bars, 5 μm.
Extended Data Fig. 7 Nucleolar R-loops and their modulation.
a, In vitro treatment with recombinant RNase H1 greatly decreases the nuclear immunofluorescence signals obtained with S9.6, an antibody against DNA–RNA hybrids. Signals remaining following RNase H1 treatment may reflect resistant hybrid structures or other nucleic acid structures. b, Immunofluorescence using S9.6, but not the anti-dsRNA antibody J2, reveals a nucleolar signal under standard cell culture conditions. c, Immunofluorescence using S9.6 with IMR90 cells also shows nucleolar signals that are repressed upon Pol II inhibition (n = 100 cells). d, In our DRIP assays, in vitro treatment with RNase H1, but not RNase III, consistently lowers DRIP signals. e, Bioinformatic analysis of the rDNA GC skew distribution and mean shows that the IGSs, but not rRNA genes, display a strongly negative GC skew; Welch’s two-tailed t-test, n = 14 (rRNA gene) and n = 30 (IGS). f, g, RNase H1 overexpression partly lowers R-loop levels (f) and increases ncRNA levels (g) at the IGS. h, Design details for the RED/dRED–LaSRR systems created to achieve inducible locus-associated R-loop repression. The zeocin resistance gene (zeoR) was used for stable cell line generation, and the blasticidin-resistance gene (blast) for selection of the tetracycline repressor (TetR). i, j, Validation of noninducible and tetracycline-inducible RED and dRED protein expression using immunoblotting (i) and microscopy (j). For gel source data, see Supplementary Fig. 1. k, Using RED together with sgIGS28 decreases R-loop levels at IGS18. l, m, Using RED together with sgIGS38 fails to alter R-loop (l) or ncRNA levels (m) at IGS18. n, Using RED together with sgIGS28 does not alter Pol II enrichments across the IGS. o, The fusion-protein system can be used to preferentially enrich the dRED fusion protein at the 5′ pause site of the ACTIN locus. p, Use of the nonoverlapping sgRNAs targeting IGS28, individually instead of as a pool, failed to significantly repress R-loop levels at IGS18, arguing against nonspecific effects related to the RNase H1 moiety of RED or any of the gRNAs used. a–p, HEK293T cells were used unless otherwise indicated. Data are means ± s.d.; one-way ANOVA with Dunnett’s multiple comparisons test (p; n = 3 biologically independent experiments) or two-tailed t-test (d, f–g, k, l–n; n = 3 biologically independent experiments); n = 3 biologically independent experiments (o); images in a, b, i–j are representative of two independent experiments. Scale bars, 5 μm.
Extended Data Fig. 8 Nucleolar and IGS features of wild-type and SETX-knockout cells.
a, ChIP showing SETX enrichment at the IGS. b, SETX has a nucleolar/nucleoplasmic localization. c, Bioinformatic analysis of ENCODE-K562 data, showing coenrichment of epigenetic marks consistent with transcriptional activation near IGS28. d, Immunoblot showing CRISPR/Cas9-mediated SETX knockout (KO). e, ChIP showing Pol II enrichment across rDNA in wild-type and SETX-KO cells. f, ChIP reveals that SETX KO, in two clones, enriches RNA Pol I at the IGSs. g, h, SETX KO induces IGS ncRNA synthesis (g) and decreases Pol I enrichment at the rRNA gene (5′-ETS region) (h). i, j, siRNA-mediated knockdown of TIF1A lowers Pol-I-dependent pre-rRNA levels but fails to induce IGS ncRNAs. Because of differences in experimental design, FP/vehicle data (j) were from a different experiment (Extended Data Fig. 6d) but are shown here for better visual comparison. k, Northern blotting reveals that Pol II or SETX disruption does not induce rRNA gene read-through transcripts. A probe for the 5′-ETS of pre-rRNA was used. l, m, SETX KO disrupts nucleolar organization as indicated by NPM immunofluorescence (l), and decreases pre-rRNA processing in pulse-chase assays (m). n, ASO-mediated knockdown of sincRNAs increases rRNA biogenesis, as indicated by single-cell rRNA biogenesis assays. Shown are nucleolar fibrillar-centre-associated RNA rings revealed by single-cell FU-RNA pulse-chase immunofluorescence. Quantification shown in Fig. 4e. o, ChIP showing H3K9me2 enrichment across rDNA in wild-type and SETX-KO cells. a–o, HEK293T cells were used unless otherwise indicated. Data in e, o were from large experimental sets sharing IgG controls. Data are means ± s.d.; two-tailed t-test, n = 3 biologically independent experiments (a, j), n = 6 biologically independent experiments (e), and n = 4 biologically independent experiments (f, o); one-way ANOVA with Dunnett’s multiple comparisons test, n = 3 biologically independent experiments (g, h) and n = 4 biologically independent experiments (m); images in b–d, k are representative of two independent experiments. Scale bars, 5 μm. For gel source data (d, i, k), see Supplementary Fig. 1.
Extended Data Fig. 9 Additional nucleolar organization and sequencing analyses related to Ewing sarcoma.
a, Representative tissue sections of human Ewing sarcoma and osteosarcoma (haematoxylin and eosin staining; magnification ×400). Materials were obtained following Institutional Research Ethics Board approval (Sinai Heath Systems, 17-0103-E). The percentages of cells with one or two distinct nucleoli per nucleus are shown. Data are means ± s.d.; per cancer type, n = 5 cases (100 cells each); two-tailed t-test P = 0.0019. b, Ewing sarcoma cells (EWS502 cells) and U2OS cells with siEWSR1 display disrupted nucleoli, as indicated by the nucleolin protein, compared to their respective control IMR90 and U2OS siControl (siCTL) cells. Scale bar, 5 μm. c, Ewing sarcoma (EWS502 and TC32) cells showed increased R-loop levels across IGSs in DRIP–seq. d, Genome-wide view of sequence read alignments for DRIP–seq and RNA-seq. Chr., chromosome. e, IMR90, EWS502 and TC32 cells can exhibit similarities and differences at non-rDNA loci in sequencing read alignments from RNA-seq. f, ASOs targeting sincRNAs ameliorate nucleolar organization. Shown are representative images related to the quantifications in Fig. 4h. Images are representative of two independent experiments. Scale bar, 5 μm.
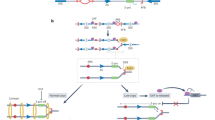
Extended Data Fig. 10 Detailed model illustrating how nucleolar Pol-II-dependent R-loops shield the IGS from sincRNA synthesis by Pol I.
Top and centre, Pol II at rDNA intergenic spacers (IGSs) synthesizes antisense intergenic ncRNAs (asincRNAs) that constitutively engage in R-loops containing DNA–RNA hybrids (orange). Centre, nucleolar Pol II function is promoted by the neurodegeneration-linked SETX protein (purple). Within rRNA genes, the formation of R-loops usually inhibits the function of Pol I, which is subject to Pol II-independent termination. However, disruption of nucleolar Pol II or its R-loops enables the recruitment of Pol I to IGSs. There, Pol I synthesizes sense intergenic ncRNAs (sincRNAs; green) that mimic environmental stress, disrupting nucleolar liquid–liquid phase separation and triggering an aberrant nucleolar liquid-to-solid phase transition. This unscheduled activation of nucleolar stress responses compromises the natural organization of nucleoli, leading to defects in pre-rRNA biogenesis, especially at the processing level. Nucleolar sincRNA levels are naturally elevated in Ewing sarcoma cells, explaining the indistinct nucleoli often seen in this cancer. In the context of Pol II inhibition, SETX loss or Ewing sarcoma, sincRNA repression ameliorates nucleolar organization and rRNA biogenesis.
Supplementary information
Supplementary Information
This file contains Supplementary Fig. 1 - the uncropped gels, and Supplementary Tables 1-4.
Source data
Rights and permissions
About this article
Cite this article
Abraham, K.J., Khosraviani, N., Chan, J.N.Y. et al. Nucleolar RNA polymerase II drives ribosome biogenesis. Nature 585, 298–302 (2020). https://doi.org/10.1038/s41586-020-2497-0
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-020-2497-0
This article is cited by
-
Atypical R-loops in cancer: decoding molecular chaos for therapeutic gain
Journal of Translational Medicine (2025)
-
Dynamic O-GlcNAcylation and phosphorylation attract and expel proteins from RNA polymerase II to regulate mRNA maturation
Journal of Biomedical Science (2025)
-
The cancer-testis lncRNA LINC01940 promotes gastric cancer malignant progression and chemoresistance by enhancing ribosome biogenesis via TAF15-mediated NOL11 SUMOylation
Cellular & Molecular Biology Letters (2025)
-
CDK9 is a dependency in GATA-3 driven and MCL-1 independent T-cell Lymphomas
Blood Cancer Journal (2025)
-
Nuclear and genome dynamics underlying DNA double-strand break repair
Nature Reviews Molecular Cell Biology (2025)