Abstract
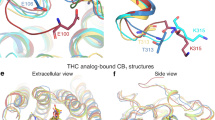
Cannabinoid receptor 1 (CB1) is the primary target of the partial agonist Δ9-tetrahydrocannabinol (Δ9-THC), the psychoactive constituent of marijuana1. Here we report two agonist-bound crystal structures of human CB1 in complex with a tetrahydrocannabinol (AM11542) and a hexahydrocannabinol (AM841). The two CB1–agonist complexes reveal important conformational changes in the overall structure relative to the antagonist-bound state2, including a 53% reduction in the volume of the ligand-binding pocket and an increase in the surface area of the G protein-binding region. Furthermore, a twin toggle switch of Phe2003.36 and Trp3566.48 (where the superscripts denote Ballesteros–Weinstein numbering3) is experimentally observed and seems to be essential for receptor activation. The structures reveal important insights into the activation mechanism of CB1 and provide a molecular basis for predicting the binding modes of Δ9-THC, and endogenous and synthetic cannabinoids. The plasticity of the binding pocket of CB1 seems to be a common feature among certain class A G protein-coupled receptors. These findings should inspire the design of chemically diverse ligands with distinct pharmacological properties.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Atomic coordinates and structures have been deposited in the PDB with accession codes 5XRA (CB1–AM11542) and 5XR8 (CB1–AM841). Source Data are provided with this paper.
References
Mechoulam, R., Hanuš, L. O., Pertwee, R. & Howlett, A. C. Early phytocannabinoid chemistry to endocannabinoids and beyond. Nat. Rev. Neurosci. 15, 757–764 (2014).
Hua, T. et al. Crystal structure of the human cannabinoid receptor CB1. Cell 167, 750–762 (2016).
Ballesteros, J. A. & Weinstein, H. in Methods in Neurosciences Vol. 25 (ed. Sealfon S. C.) 366–428 (Academic, 1995).
Lemberger, L. Potential therapeutic usefulness of marijuana. Annu. Rev. Pharmacol. Toxicol. 20, 151–172 (1980).
Li, H.-L. An archaeological and historical account of cannabis in China. Econ. Bot. 28, 437–448 (1973).
Makriyannis, A. 2012 Division of Medicinal Chemistry Award Address. Trekking the cannabinoid road: a personal perspective. J. Med. Chem. 57, 3891–3911 (2014).
Shao, Z. et al. High-resolution crystal structure of the human CB1 cannabinoid receptor. Nature 540, 602–606 (2016).
Nikas, S. P. et al. The role of halogen substitution in classical cannabinoids: a CB1 pharmacophore model. AAPS J. 6, e30 (2004).
Nikas, S. P. et al. Novel 1′,1′-chain substituted hexahydrocannabinols: 9β-hydroxy-3-(1-hexyl-cyclobut-1-yl)-hexahydrocannabinol (AM2389) a highly potent cannabinoid receptor 1 (CB1) agonist. J. Med. Chem. 53, 6996–7010 (2010).
Xie, X. Q., Melvin, L. S. & Makriyannis, A. The conformational properties of the highly selective cannabinoid receptor ligand CP-55,940. J. Biol. Chem. 271, 10640–10647 (1996).
Makriyannis, A. & Rapaka, R. S. The medicinal chemistry of cannabinoids: an overview. NIDA Res. Monogr. 79, 204–210 (1987).
Ahn, K. H., Bertalovitz, A. C., Mierke, D. F. & Kendall, D. A. Dual role of the second extracellular loop of the cannabinoid receptor 1: ligand binding and receptor localization. Mol. Pharmacol. 76, 833–842 (2009).
Feigenbaum, J. J. et al. Nonpsychotropic cannabinoid acts as a functional N-methyl-d-aspartate receptor blocker. Proc. Natl Acad. Sci. USA 86, 9584–9587 (1989).
Mechoulam, R. et al. Enantiomeric cannabinoids: stereospecificity of psychotropic activity. Experientia 44, 762–764 (1988).
Hanson, M. A. et al. Crystal structure of a lipid G protein-coupled receptor. Science 335, 851–855 (2012).
Rasmussen, S. G. et al. Crystal structure of the β2 adrenergic receptor–Gs protein complex. Nature 477, 549–555 (2011).
Singh, R. et al. Activation of the cannabinoid CB1 receptor may involve a W6.48/F3.36 rotamer toggle switch. J. Pept. Res. 60, 357–370 (2002).
Tiburu, E. K. et al. Structural biology of human cannabinoid receptor-2 helix 6 in membrane-mimetic environments. Biochem. Biophys. Res. Commun. 384, 243–248 (2009).
Zhang, K. et al. Structure of the human P2Y12 receptor in complex with an antithrombotic drug. Nature 509, 115–118 (2014).
Zhang, J. et al. Agonist-bound structure of the human P2Y12 receptor. Nature 509, 119–122 (2014).
Nikas, S. P. et al. A concise methodology for the synthesis of (−)-Δ9-tetrahydrocannabinol and (−)-Δ9-tetrahydrocannabivarin metabolites and their regiospecifically deuterated analogs. Tetrahedron 63, 8112–8113 (2007).
Kulkarni, S. et al. Novel C-ring-hydroxy-substituted controlled deactivation cannabinergic analogues. J. Med. Chem. 59, 6903–6919 (2016).
D’Antona, A. M., Ahn, K. H. & Kendall, D. A. Mutations of CB1 T210 produce active and inactive receptor forms: correlations with ligand affinity, receptor stability, and cellular localization. Biochemistry 45, 5606–5617 (2006).
Caffrey, M. & Cherezov, V. Crystallizing membrane proteins using lipidic mesophases. Nat. Protocols 4, 706–731 (2009).
Cherezov, V. et al. Rastering strategy for screening and centring of microcrystal samples of human membrane proteins with a sub-10 μm size X-ray synchrotron beam. J. R. Soc. Interface 6, S587–S597 (2009).
Chun, E. et al. Fusion partner toolchest for the stabilization and crystallization of G protein-coupled receptors. Structure 20, 967–976 (2012).
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Smart, O. S. et al. Exploiting structure similarity in refinement: automated NCS and target-structure restraints in BUSTER. Acta Crystallogr. D 68, 368–380 (2012).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Friesner, R. A. et al. Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J. Med. Chem. 47, 1739–1749 (2004).
Friesner, R. A. et al. Extra precision glide: docking and scoring incorporating a model of hydrophobic enclosure for protein–ligand complexes. J. Med. Chem. 49, 6177–6196 (2006).
Halgren, T. A. et al. Glide: a new approach for rapid, accurate docking and scoring. 2. Enrichment factors in database screening. J. Med. Chem. 47, 1750–1759 (2004).
Xu, T. et al. Induced-fit docking enables accurate free energy perturbation calculations in homology models. J. Chem. Theory Comput. 18, 5710–5724 (2022).
Abraham, M. J. et al. GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2, 19–25 (2015).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Sousa da Silva, A. W. & Vranken, W. F. ACPYPE—AnteChamber PYthon Parser interfacE. BMC Res. Notes 5, 367 (2012).
Berman, H. M. et al. The Protein Data Bank. Nucl. Acids Res. 28, 235–242 (2000).
Skjærven, L., Yao, X. Q., Scarabelli, G. & Grant, B. J. Integrating protein structural dynamics and evolutionary analysis with Bio3D. BMC Bioinf. 15, 399 (2014).
Acknowledgements
This work was supported by the NSF of China (grant no. 31330019 to Z.-J.L.), the MOST of China (grant nos. 2014CB910400 and 2015CB910104 to Z.-J.L.), NSF of Shanghai (grant no. 16ZR1448500 to S.Z.), the Key R&D Program of China (grant no. 2016YCF0905902 to S.Z.), the NIH (grant nos. R01DA041435 to R.C.S. and A.M., P01DA009158 to A.M. and L.M.B., and R37DA023142 to A.M.), NSF grants, the Shanghai Municipal Government, ShanghaiTech University and the GPCR Consortium. The diffraction data were collected at GM/CA@APS of Argonne National Laboratory, X06SA@SLS of the Paul Scherrer Institute, and BL41XU@Spring-8 with JASRI proposals 2015B1031 and 2016A2731. We thank M. Wang, C.-Y. Huang, V. Olieric, M. Audet and M.-Y. Lee for their help with data collection, A. Walker for critical review of the manuscript and F. Sun for high-resolution mass spectrometry analysis. We thank E. Stahl, C. Brust and V. Dang from UF Scripps for their contributions to making cell lines, mutations and generating the functional data.
Author information
Authors and Affiliations
Contributions
T.H. performed crystallization, data collection, structure determination and analysis. K.V., S.P.N. and S.J. designed, synthesized and characterized the ligands. Y.W. performed the docking and molecular dynamics simulations. L.Q. and M.P. collected and processed data, and refined the structures. G.W.H. and M.A.H. contributed to structure refinement and data analysis. J.-H.H. conducted the functional studies and worked on mutations. A.K. performed the radioligand binding assays. K.D. performed structure analysis. X.L. and H.L. performed the molecular dynamics simulations. S.Z. supervised the structure and simulation analyses. L.M.B. designed and supervised the functional and kinetic studies. A.M. supervised the conceptual design, synthesis and characterization of the agonist. R.C.S. conceived the project, and supervised the data analysis. Z.-J.L. designed and supervised the experiments, and analysed the data. Z.-J.L., T.H., R.C.S., A.M., L.M.B. and S.Z. wrote the manuscript with discussions and improvements from M.A.H., K.V., S.P.N. and Y.W.
Corresponding authors
Ethics declarations
Competing interests
A.M. is a founder of MAKScientific. R.C.S. is the chief executive and a board member of Structure Therapeutics. The other authors declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Synthesis of AM841 and AM11542.
Reagents and conditions: (a) CH3I, NaH, DMF, 0 °C to room temperature, 2 h, 95%; (b) DIBAL-H, CH2Cl2, −78 °C, 0.5 h, 87%; (c) Br− P+Ph3(CH2)5OPh, (Me3Si)2NK, THF, 0–10 °C, 30 min, then addition to 3, 0 °C to room temperature, 2 h, 96%; (d) H2, 10% Pd/C, AcOEt, room temperature, 2.5 h, 89%; (e) BBr3, CH2Cl2, −78 °C to room temperature, 6 h, 85%; (f) diacetates, p-TSA, CHCl3, 0 °C to room temperature, 4 days, 64%; (g) TMSOTf, CH2Cl2/MeNO2, 0 °C to room temperature, 3 h, 71%; (h) TBDMSCl, imidazole, DMAP, DMF, room temperature, 12 h, 85%; (i) Cl− Ph3P+CH2OMe, (Me3Si)2NK, THF, 0 °C to room temperature, 1 h, then addition to 9, 0 °C to room temperature, 1.5 h, 73%; (j) Cl3CCOOH, CH2Cl2, room temperature, 50 min, 95%; (k) K2CO3, EtOH, room temperature, 3 h, 84%; (l) NaBH4, EtOH, 0 °C, 30 min, 98%; (m) TBAF, THF, −40 °C, 30 min, 96%; (n) TMG-N3, CHCl3/MeNO2, room temperature, 18 h, 84%; (o) PPh3, CS2, THF, room temperature, 10 h, 76%; (p) (+)-cis/trans-p-mentha-2,8-dien-1-ol, p-TSA, benzene, reflux 4 h, 65%.
Extended Data Fig. 2 Analytical size exclusion chromatography profile and crystals of CB1–AM11542/AM841 complex.
a, Analytical size exclusion chromatography and crystal image of the CB1–AM11542 complex. Scale bar, 70 μm. b, Analytical size exclusion chromatography and crystal image of the CB1–AM841 complex. Scale bar, 70 μm. c, The overall structures of CB1–AM11542 and CB1–AM841 complexes and crystal packing of CB1–AM11542; receptor is in orange (AM11542)/green (AM841) colour and the flavodoxin fusion protein is in purple-blue colour. The agonists AM11542 (yellow) and AM841 (pink) are shown in sticks representation. The four single mutations T2103.46A, E2735.37K, T2835.47V and R3406.32E are shown as green spheres in the CB1–AM11542 structure.
Extended Data Fig. 3 Representative electron density of the CB1 agonists-bound structures and cholesterol binding sites.
a, The |Fo|− |Fc| omit maps of AM11542 and AM841 contoured at 3.0σ at 2.80 Å and 2.95 Å, respectively. b, The cholesterol binding site in the CB1–AM11542 structure (orange) with CB1–AM6538 structure (blue) superposed.
Extended Data Fig. 4 Mutations of the CB1 receptor and the effects on agonist-induced activity as assessed by the forskolin-stimulated accumulation of cAMP.
a, Primers used to generate mutations in 3×HA–CB1 and validation of cell-surface expression of wild-type and mutant CB1 in CHO-K1 cell lines quantitative flow cytometry. b, Dose response studies of agonist (AM11542, AM841 and CP55,940) activity for each mutant compared to wild type (in blue filled circles) from Fig. 3c. c, Assessment of the effect of the individual point mutations that were made to stabilize the receptor, in absence of the flavodoxin insert, on receptor activity. All experiments were repeated at least three times, and error bars denote s.e.m. of duplicate measurements (parameters are in Extended Data Table 2).
Extended Data Fig. 5 Docking poses of different cannabinoid receptor agonists and MD validation.
a–f, The r.m.s.d. values of ligand heavy atoms show that the docked poses are stable during the 1 μs molecular dynamics simulations: Δ9-THC (a), AEA (b), JWH-018 (c), HU-210 (d), 2-AG (e), WIN 55,212-2 (f). g, h, j, k, The poses of HU-210 (g), JWH-018 (h), 2-AG (j) and WIN 55,212-2 (k) are shown. i, The superimposition of HU-210 (yellow sticks) and HU-211 (blue sticks) in the binding pocket. The binding pose of HU-210 explains why HU-211, the enantiomer of HU-210, failed to stimulate CB1 because superimposed HU-211 on HU-210 shows severe clashes with H1782.65 in CB1.
Extended Data Fig. 6 Structural conformation changes of solved agonist- and antagonist-bound class A GPCRs.
a, The pattern of r.m.s.d. values of transmembrane helices between agonist- and antagonist-bound class A GPCR structures. The structures used for analysis are the same as described in Extended Data Table 3. b, Measurement of the degree of helix VI bending observed in class A GPCRs structures. All structures were superimposed onto inactive-state β2-adrenergic receptor by UCSF Chimera. The direction of helices VI were defined by vectors ηi which starts from the centre of Cα of residues 6.45–6.48 to the centre of Cα of residues 6.29-30–6.32-33. The two vectors η0 and η1 of helices VI in the inactive-state and active-state β2-adrenergic receptor were selected as reference to form a plane α. The vector ηi of helix VI of other structure was projected to the plane α as a new vector ηi’. The bending angle of each helix VI was then defined by the angle between ηi’ and η0. The structures are: ETB (PDB code 5GLH), β1-adrenergic receptor (PDB code 2Y02), P2Y12 (PDB code 4PXZ), β2-adrenergic receptor (PDB code 3PDS), FFA1 (PDB code 4PHU), 5HT2B (PDB code 4IB4), 5HT1B (PDB code 4IAR), Rho (PDB code 2HPY), A2A (PDB code 3QAK), NTS1 (PDB code 4BUO), CB1 (bound to AM11542; PDB code 5XRA), μ-opioid receptor + nanobody (NB) (PDB code 5C1M), Rho + NB (PDB code 2×72), Rho + arrestin (PDB code 4ZWJ), M2R + NB (PDB code 4MQS), β2-adrenergic receptor + NB (PDB code 4LDL), A2A + mini-Gs (PDB code 5G53), β2-adrenergic receptor + Gs (PDB code 3SN6).
Supplementary information
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hua, T., Vemuri, K., Nikas, S.P. et al. Crystal structures of agonist-bound human cannabinoid receptor CB1. Nature 646, 754–758 (2025). https://doi.org/10.1038/s41586-025-09454-5
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-09454-5