Abstract
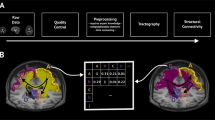
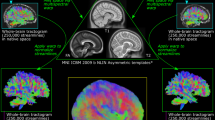
Connectional neuroanatomical maps can be generated in vivo by using diffusion-weighted magnetic resonance imaging (dMRI) data, and their representation as structural connectome (SC) atlases adopts network-based brain analysis methods. We explain the generation of high-quality SCs of brain connectivity by using recent advances for reconstructing long-range white matter connections such as local fiber orientation estimation on multi-shell dMRI data with constrained spherical deconvolution, which yields both increased sensitivity to detecting crossing fibers compared with competing methods and the ability to separate signal contributions from different macroscopic tissues, and improvements to streamline tractography such as anatomically constrained tractography and spherical-deconvolution informed filtering of tractograms, which have increased the biological accuracy of SC creation. Here, we provide step-by-step instructions to creating SCs by using these methods. In addition, intermediate steps of our procedure can be adapted for related analyses, including region of interest-based tractography and quantification of local white matter properties. The associated software MRtrix3 implements the relevant tools for easy application of the protocol, with specific processing tasks deferred to components of the FSL software. The protocol is suitable for users with expertise in dMRI and neuroscience and requires between 2 h and 13 h to complete, depending on the available computational system.
Key points
-
The procedure enables the creation of structural connectomes from diffusion-weighted magnetic resonance imaging data processed by using state-of-the-art techniques such as constrained spherical deconvolution, anatomically constrained tractography and spherical-deconvolution informed filtering of tractograms implemented by using MRtrix3 and FSL software.
-
The intermediate steps can be adapted for region-of-interest tractography and quantification of local white matter properties.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The tutorial data (raw and derived) are freely available at Open Science Framework at https://osf.io/tm5x8/ (ref. 183).
Code availability
The complementary Docker image is available on Github at https://github.com/martahedl/SC-construction-using-CSD-in-MSMT-dMRI. The relevant code to perform the present protocol’s steps is provided within this manuscript and may be used freely. More code, for example, on quality assessment, is provided in the Supplementary Information.
References
Avena-Koenigsberger, A., Misic, B. & Sporns, O. Communication dynamics in complex brain networks. Nat. Rev. Neurosci. 19, 17–33 (2018).
Bettinardi, R. G. et al. How structure sculpts function: unveiling the contribution of anatomical connectivity to the brain’s spontaneous correlation structure. Chaos 27, 047409 (2017).
Honey, C. J., Kötter, R., Breakspear, M. & Sporns, O. Network structure of cerebral cortex shapes functional connectivity on multiple time scales. Proc. Natl Acad. Sci. USA 104, 10240–10245 (2007).
Honey, C. J. et al. Predicting human resting-state functional connectivity from structural connectivity. Proc. Natl Acad. Sci. USA 106, 2035–2040 (2009).
Petersen, S. E. & Sporns, O. Brain networks and cognitive architectures. Neuron 88, 207–219 (2015).
Rubinov, M. & Sporns, O. Complex network measures of brain connectivity: uses and interpretations. Neuroimage 52, 1059–1069 (2010).
Sporns, O. From simple graphs to the connectome: networks in neuroimaging. Neuroimage 62, 881–886 (2012).
Fornito, A., Zalesky, A. & Breakspear, M. The connectomics of brain disorders. Nat. Rev. Neurosci. 16, 159–172 (2015).
Sporns, O., Tononi, G. & Kötter, R. The human connectome: a structural description of the human brain. PLoS Comput. Biol. 1, 0245–0251 (2005).
Bullmore, E. T. & Sporns, O. Complex brain networks: graph theoretical analysis of structural and functional systems. Nat. Rev. Neurosci. 10, 186–198 (2009).
Stejskal, E. O. & Tanner, J. E. Spin diffusion measurements: spin echoes in the presence of a time‐dependent field gradient. J. Chem. Phys. 42, 288–292 (1965).
Moseley, M. E. et al. Diffusion-weighted MR imaging of anisotropic water diffusion in cat central nervous system. Radiology 176, 439–445 (1990).
Dell’Acqua, F. & Tournier, J. D. Modelling white matter with spherical deconvolution: how and why? NMR Biomed. 32, e3945 (2019).
Mori, S. & van Zijl, P. C. M. Fiber tracking: principles and strategies—a technical review. NMR Biomed. 15, 468–480 (2002).
Savadjiev, P. et al. Labeling of ambiguous subvoxel fibre bundle configurations in high angular resolution diffusion MRI. Neuroimage 41, 58–68 (2008).
Thomas, C. et al. Anatomical accuracy of brain connections derived from diffusion MRI tractography is inherently limited. Proc. Natl Acad. Sci. USA 111, 16574–16579 (2014).
Tournier, J. D. et al. MRtrix3: a fast, flexible and open software framework for medical image processing and visualisation. Neuroimage 202, 116137 (2019).
Basser, P. J., Mattiello, J. & LeBihan, D. MR diffusion tensor spectroscopy and imaging. Biophys. J. 66, 259–267 (1994).
Jeurissen, B., Leemans, A., Tournier, J. D., Jones, D. K. & Sijbers, J. Investigating the prevalence of complex fiber configurations in white matter tissue with diffusion magnetic resonance imaging. Hum. Brain Mapp. 34, 2747–2766 (2013).
Behrens, T. E. J., Berg, H. J., Jbabdi, S., Rushworth, M. F. S. & Woolrich, M. W. Probabilistic diffusion tractography with multiple fibre orientations: what can we gain? Neuroimage 34, 144–155 (2007).
Alexander, A. L., Hasan, K. M., Lazar, M., Tsuruda, J. S. & Parker, D. L. Analysis of partial volume effects in diffusion-tensor MRI. Magn. Reson. Med. 45, 770–780 (2001).
Tuch, D. S. et al. High angular resolution diffusion imaging reveals intravoxel white matter fiber heterogeneity. Magn. Reson. Med. 48, 577–582 (2002).
Alexander, D. C. Multiple-fiber reconstruction algorithms for diffusion MRI. Ann. N. Y. Acad. Sci. 1064, 113–133 (2005).
Pierpaoli, C. et al. Water diffusion changes in Wallerian degeneration and their dependence on white matter architecture. Neuroimage 13, 1174–1185 (2001).
Jeurissen, B., Leemans, A., Jones, D. K., Tournier, J. D. & Sijbers, J. Probabilistic fiber tracking using the residual bootstrap with constrained spherical deconvolution. Hum. Brain Mapp. 32, 461–479 (2011).
Tournier, J., Yeh, C.-H., Calamante, F. & Cho, K.-H. Resolving crossing fibres using constrained spherical deconvolution: validation using diffusion-weighted imaging phantom data. Neuroimage 42, 617–625 (2008).
Farquharson, S. et al. White matter fiber tractography: why we need to move beyond DTI. J. Neurosurg. 118, 1367–1377 (2013).
Tournier, J. D., Calamante, F., Gadian, D. G. & Connelly, A. Direct estimation of the fiber orientation density function from diffusion-weighted MRI data using spherical deconvolution. Neuroimage 23, 1176–1185 (2004).
Tournier, J. D., Calamante, F. & Connelly, A. Robust determination of the fibre orientation distribution in diffusion MRI: non-negativity constrained super-resolved spherical deconvolution. Neuroimage 35, 1459–1472 (2007).
Jeurissen, B., Tournier, J. D., Dhollander, T., Connelly, A. & Sijbers, J. Multi-tissue constrained spherical deconvolution for improved analysis of multi-shell diffusion MRI data. Neuroimage 103, 411–426 (2014).
Smith, R. E., Tournier, J. D., Calamante, F. & Connelly, A. Anatomically-constrained tractography: improved diffusion MRI streamlines tractography through effective use of anatomical information. Neuroimage 62, 1924–1938 (2012).
Smith, R. E., Tournier, J. D., Calamante, F. & Connelly, A. Spherical-deconvolution informed filtering of tractograms. Neuroimage 67, 298–312 (2013).
Smith, R. E., Tournier, J. D., Calamante, F. & Connelly, A. SIFT2: enabling dense quantitative assessment of brain white matter connectivity using streamlines tractography. Neuroimage 119, 338–351 (2015).
Smith, R. E., Tournier, J. D., Calamante, F. & Connelly, A. The effects of SIFT on the reproducibility and biological accuracy of the structural connectome. Neuroimage 104, 253–265 (2015).
Yeh, C. H., Smith, R. E., Liang, X., Calamante, F. & Connelly, A. Correction for diffusion MRI fibre tracking biases: the consequences for structural connectomic metrics. Neuroimage 142, 150–162 (2016).
Smith, R. Quantitative streamlines tractography: methods and inter-subject normalisation. Apert. Neuro. 2, 1–23 (2022).
Tournier, J. D., Calamante, F. & Connelly, A. Determination of the appropriate b value and number of gradient directions for high-angular-resolution diffusion-weighted imaging. NMR Biomed. 26, 1775–1786 (2013).
Humphries, M. D. & Gurney, K. Network “small-world-ness”: a quantitative method for determining canonical network equivalence. PLoS ONE 3, e0002051 (2008).
Newman, M. E. J. & Girvan, M. Finding and evaluating community structure in networks. Phys. Rev. E Stat. Nonlin. Soft Matter Phys. 69, 026113 (2004).
Lehmann, E. L. & Romano, J. P. Generalizations of the familywise error rate. Ann. Stat. 33, 1138–1154 (2005).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B Methodol. 57, 289–300 (1995).
Genovese, C. R., Lazar, N. A. & Nichols, T. Thresholding of statistical maps in functional neuroimaging using the false discovery rate. Neuroimage 15, 870–878 (2002).
Simes, R. J. An improved Bonferroni procedure for multiple tests of significance. Biometrika 73, 751–754 (1986).
Storey, J. D. A direct approach to false discovery rates. J. R. Stat. Soc. Ser. B Stat. Methodol. 64, 479–498 (2002).
Zalesky, A., Fornito, A. & Bullmore, E. T. Network-based statistic: identifying differences in brain networks. Neuroimage 53, 1197–1207 (2010).
Baggio, H. C. et al. Statistical inference in brain graphs using threshold-free network-based statistics. Hum. Brain Mapp. 39, 2289–2302 (2018).
Zalesky, A., Cocchi, L., Fornito, A., Murray, M. M. & Bullmore, E. Connectivity differences in brain networks. Neuroimage 60, 1055–1062 (2012).
Nichols, T. E. & Holmes, A. P. Nonparametric permutation tests for functional neuroimaging: a primer with examples. Hum. Brain Mapp. 15, 1–25 (2002).
Nichols, T. & Hayasaka, S. Controlling the familywise error rate in functional neuroimaging: a comparative review. Stat. Methods Med. Res. 12, 419–446 (2003).
Bassett, D. S. & Gazzaniga, M. S. Understanding complexity in the human brain. Trends Cogn. Sci. 15, 200–209 (2011).
Norman, K. A., Polyn, S. M., Detre, G. J. & Haxby, J. V. Beyond mind-reading: multi-voxel pattern analysis of fMRI data. Trends Cogn. Sci. 10, 424–430 (2006).
Burges, C. J. C. A tutorial on support vector machines for pattern recognition. Data Min. Knowl. Disc. 2, 121–167 (1998).
Dosenbach, N. U. F. et al. Prediction of individual brain maturity using fMRI. Science 329, 1358–1361 (2010).
Plis, S. M. et al. Deep learning for neuroimaging: a validation study. Front. Neurosci. 8, 229 (2014).
Mohri, M., Rostamizadeh, A. & Talwalkar, A. Foundations of Machine Learning. Edn. 2 (MIT Press, 2018).
Smith, S. M. et al. A positive-negative mode of population covariation links brain connectivity, demographics and behavior. Nat. Neurosci. 18, 1565–1567 (2015).
Krishnan, A., Williams, L. J., McIntosh, A. R. & Abdi, H. Partial Least Squares (PLS) methods for neuroimaging: a tutorial and review. Neuroimage 56, 455–475 (2011).
McIntosh, A. R., Bookstein, F. L., Haxby, J. V. & Grady, C. L. Spatial pattern analysis of functional brain images using partial least squares. Neuroimage 3, 143–157 (1996).
Drysdale, A. T. et al. Resting-state connectivity biomarkers define neurophysiological subtypes of depression. Nat. Med. 23, 28–38 (2017).
Batista-García-Ramó, K. & Fernández-Verdecia, C. I. What we know about the brain structure-function relationship. Behav. Sci. (Basel) 8, 39 (2018).
Adachi, Y. et al. Functional connectivity between anatomically unconnected areas is shaped by collective network-level effects in the macaque cortex. Cereb. Cortex 22, 1586–1592 (2012).
Biswal, B., Yetkin, F. Z., Haughton, V. M. & Hyde, J. S. Functional connectivity in the motor cortex of resting human brain using echo-planar MRI. Magn. Reson. Med. 34, 537–541 (1995).
Fox, M. D. et al. The human brain is intrinsically organized into dynamic, anticorrelated functional networks. Proc. Natl Acad. Sci. USA 102, 9673–9678 (2005).
Power, J. D. et al. Functional network organization of the human brain. Neuron 72, 665–678 (2011).
Hutchison, R. M. et al. Dynamic functional connectivity: promise, issues, and interpretations. Neuroimage 80, 360–378 (2013).
Vincent, J. L. et al. Intrinsic functional architecture in the anaesthetized monkey brain. Nature 447, 83–86 (2007).
Donnelly‐Kehoe, P. et al. Reliable local dynamics in the brain across sessions are revealed by whole‐brain modeling of resting state activity. Hum. Brain Mapp. 40, 2967–2980 (2019).
Cabral, J., Kringelbach, M. L. & Deco, G. Functional connectivity dynamically evolves on multiple time-scales over a static structural connectome: models and mechanisms. Neuroimage 160, 84–96 (2017).
Keilholz, S. D., Caballero-Gaudes, C., Bandettini, P., Deco, G. & Calhoun, V. D. Time-resolved resting state fMRI analysis: current status, challenges, and new directions. Brain Connect. 7, 465–481 (2017).
Breakspear, M. Dynamic models of large-scale brain activity. Nat. Neurosci. 20, 340–352 (2017).
Basser, P. J. & Pierpaoli, C. Microstructural and physiological features of tissues elucidated by quantitative-diffusion-tensor MRI. J. Magn. Reson. B 111, 209–219 (1996).
Beaulieu, C. The basis of anisotropic water diffusion in the nervous system—a technical review. NMR Biomed. 15, 435–455 (2002).
Raffelt, D. et al. Apparent Fibre Density: a novel measure for the analysis of diffusion-weighted magnetic resonance images. Neuroimage 59, 3976–3994 (2012).
Dhollander, T., Mito, R., Raffelt, D. & Connelly, A. Improved white matter response function estimation for 3-tissue constrained spherical deconvolution. In International Society of Magnetic Resonance in Medicine 27th Annual Meeting Abstract no. 0555 (2019).
Veraart, J., Fieremans, E. & Novikov, D. S. On the scaling behavior of water diffusion in human brain white matter. Neuroimage 185, 379–387 (2019).
Dhollander, T. et al. Fixel-based analysis of diffusion MRI: methods, applications, challenges and opportunities. Neuroimage 241, 118417 (2021).
García-Gomar, M. G. et al. Long-term improvement of Parkinson disease motor symptoms derived from lesions of prelemniscal fiber tract components. Operative Neurosurg. 19, 539–550 (2020).
Le Bihan, D., Poupon, C. & Amadon, A. Artifacts and pitfalls in diffusion MRI. J. Magn. Reson. Imaging 24, 478–488 (2006).
Tax, C. M. W., Bastiani, M., Veraart, J., Garyfallidis, E. & Okan Irfanoglu, M. What’s new and what’s next in diffusion MRI preprocessing. Neuroimage 249, 118830 (2022).
Veraart, J., Fieremans, E., Jelescu, I. O., Knoll, F. & Novikov, D. S. Gibbs ringing in diffusion MRI. Magn. Reson. Med. 76, 301–314 (2016).
Anderson, A. W. & Gore, J. C. Analysis and correction of motion artifacts in diffusion weighted imaging. Magn. Reson. Med. 32, 379–387 (1994).
Ordidge, R. J., Helpern, J. A., Qing, Z. X., Knight, R. A. & Nagesh, V. Correction of motional artifacts in diffusion-weighted MR images using navigator echoes. Magn. Reson. Imaging 12, 455–460 (1994).
Ahn, C. B. & Cho, Z. H. Analysis of the eddy-current induced artifacts and the temporal compensation in nuclear magnetic resonance imaging. IEEE Trans. Med. Imaging 10, 47–52 (1991).
Spees, W. M. et al. Quantification and compensation of eddy-current-induced magnetic-field gradients. J. Magn. Reson. 212, 116–123 (2011).
Gallichan, D. et al. Addressing a systematic vibration artifact in diffusion-weighted MRI. Hum. Brain Mapp. 31, 193–202 (2010).
Andersson, J. L. R. & Graham, M. S. Incorporating outlier detection and replacement into a non-parametric framework for movement and distortion correction of diffusion MR images. Neuroimage 141, 556–572 (2016).
Haselgrove, J. C. & Moore, J. R. Correction for distortion of echo-planar images used to calculate the apparent diffusion coefficient. Magn. Reson. Med. 36, 960–964 (1996).
Shen, Y. et al. Correction of high-order eddy current induced geometric distortion in diffusion-weighted echo-planar images. Magn. Reson. Med. 52, 1184–1189 (2004).
Calamante, F., Porter, D. A., Gadian, D. G. & Connelly, A. Correction for eddy current induced Bo shifts in diffusion-weighted echo-planar imaging. Magn. Reson. Med. 41, 95–102 (1999).
Veraart, J., Fieremans, E. & Novikov, D. S. Diffusion MRI noise mapping using random matrix theory. Magn. Reson. Med. 76, 1582–1593 (2016).
Veraart, J. et al. Denoising of diffusion MRI using random matrix theory. Neuroimage 142, 394–406 (2016).
Moeller, S. et al. NOise reduction with DIstribution Corrected (NORDIC) PCA in dMRI with complex-valued parameter-free locally low-rank processing. Neuroimage 226, 117539 (2021).
Fadnavis, S., Batson, J. & Garyfallidis, E. Patch2Self: denoising diffusion MRI with self-supervised learning. In Larochelle, H., Ranzato, M., Hadsell, R., Balcan, M. F. & Lin, H. eds. Advances in Neural Information Processing Systems. Vol 33. 16293–16303 (Curran Associates, Inc., 2020).
Garyfallidis, E. et al. Dipy, a library for the analysis of diffusion MRI data. Front. Neuroinform. 8, 8 (2014).
Kellner, E., Dhital, B., Kiselev, V. G. & Reisert, M. Gibbs‐ringing artifact removal based on local subvoxel‐shifts. Magn. Reson. Med. 76, 1574–1581 (2016).
Perrone, D. et al. The effect of Gibbs ringing artifacts on measures derived from diffusion MRI. Neuroimage 120, 441–455 (2015).
Tournier, J. D., Mori, S. & Leemans, A. Diffusion tensor imaging and beyond. Magn. Reson. Med. 65, 1532–1556 (2011).
Oldham, S. et al. The efficacy of different preprocessing steps in reducing motion-related confounds in diffusion MRI connectomics. Neuroimage 222, 117252 (2020).
Jenkinson, M., Beckmann, C. F., Behrens, T. E. J., Woolrich, M. W. & Smith, S. M. FSL. Neuroimage 62, 782–790 (2012).
Andersson, J. L. R. & Sotiropoulos, S. N. An integrated approach to correction for off-resonance effects and subject movement in diffusion MR imaging. Neuroimage 125, 1063–1078 (2016).
Andersson, J. L. R., Skare, S. & Ashburner, J. How to correct susceptibility distortions in spin-echo echo-planar images: application to diffusion tensor imaging. Neuroimage 20, 870–888 (2003).
Graham, M. S., Drobnjak, I. & Zhang, H. Realistic simulation of artefacts in diffusion MRI for validating post-processing correction techniques. Neuroimage 125, 1079–1094 (2016).
Sairanen, V. & Andersson, J. Outliers in diffusion-weighted MRI: exploring detection models and mitigation strategies. Neuroimage 283, 120397 (2023).
Andersson, J. L. R. et al. Towards a comprehensive framework for movement and distortion correction of diffusion MR images: within volume movement. Neuroimage 152, 450–466 (2017).
Irfanoglu, M. O., Nayak, A., Jenkins, M. & Pierpaoli, C. TORTOISEv3: improvements and new features of the NIH diffusion MRI processing pipeline. In International Society of Magnetic Resonance in Medicine 25th Annual Meeting Abstract no. 3540 (2017).
Pierpaoli, C. et al. TORTOISE: an integrated software package for processing of diffusion MRI data. In International Society of Magnetic Resonance in Medicine 18th Annual Meeting Abstract no. 1597 (2010).
Christiaens, D. et al. Scattered slice SHARD reconstruction for motion correction in multi-shell diffusion MRI. Neuroimage 225, 117437 (2021).
Edwards, A. D. et al. The developing human connectome project neonatal data release. Front. Neurosci. 16, 886772 (2022).
Weisskopf, R. M. & Davis, T. L. Correcting gross distortion on echo planar images. In Proceedings of the Society for Magnetic Resonance in Medicine 11th Annual Meeting Abstract no. 4515 (1992).
Jezzard, P. & Balaban, R. S. Correction for geometric distortion in echo planar images from B0 field variations. Magn. Reson. Med. 34, 65–73 (1995).
Holland, D., Kuperman, J. M. & Dale, A. M. Efficient correction of inhomogeneous static magnetic field-induced distortion in Echo Planar Imaging. Neuroimage 50, 175–183 (2010).
Skare, S. & Bammer, R. Jacobian weighting of distortion corrected EPI data. In Proceedings of the International Society for Magnetic Resonance in Medicine Abstract no. 5063 (2010).
Sotiropoulos, S. N., Behrens, T. E. J. & Jbabdi, S. Ball and rackets: inferring fiber fanning from diffusion-weighted MRI. Neuroimage 60, 1412–1425 (2012).
Kennis, M., van Rooij, S. J. H., Kahn, R. S., Geuze, E. & Leemans, A. Choosing the polarity of the phase-encoding direction in diffusion MRI: does it matter for group analysis? Neuroimage Clin. 11, 539–547 (2016).
Hong, G. S. et al. Diffusion-weighted imaging with reverse phase-encoding polarity: the added value to the conventional diffusion-weighted imaging in differentiating acute infarctions from hyperintense brainstem artifacts. Eur. Radiol. 27, 859–867 (2017).
Irfanoglu, M. O. et al. DR-BUDDI (Diffeomorphic Registration for Blip-Up blip-Down Diffusion Imaging) method for correcting echo planar imaging distortions. Neuroimage 106, 284–299 (2015).
Setsompop, K. et al. Improving diffusion MRI using simultaneous multi-slice echo planar imaging. Neuroimage 63, 569–580 (2012).
Andersson, J. L. R. & Sotiropoulos, S. N. Non-parametric representation and prediction of single- and multi-shell diffusion-weighted MRI data using Gaussian processes. Neuroimage 122, 166–176 (2015).
Sairanen, V., Leemans, A. & Tax, C. M. W. Fast and accurate Slicewise OutLIer Detection (SOLID) with informed model estimation for diffusion MRI data. Neuroimage 181, 331–346 (2018).
Koch, A., Zhukov, A., Stöcker, T., Groeschel, S. & Schultz, T. SHORE‐based detection and imputation of dropout in diffusion MRI. Magn. Reson. Med. 82, 2286–2298 (2019).
Tustison, N. J. et al. N4ITK: improved N3 bias correction. IEEE Trans. Med. Imaging 29, 1310–1320 (2010).
Avants, B. B. et al. The Insight ToolKit image registration framework. Front. Neuroinform. 8, 44 (2014).
Raffelt, D. et al. Bias field correction and intensity normalisation for quantitative analysis of apparent fibre density. In International Society of Magnetic Resonance in Medicine 25th Annual Meeting Abstract no. 3541 (2017).
Dhollander, T. et al. Multi-tissue log-domain intensity and inhomogeneity normalisation for quantitative apparent fibre density. In International Society of Magnetic Resonance in Medicine 29th Annual Meeting. Abstract no. 2472 (2021).
Wedeen, V. J. et al. Diffusion spectrum magnetic resonance imaging (DSI) tractography of crossing fibers. Neuroimage 41, 1267–1277 (2008).
Wedeen, V. et al. Mapping fiber orientation spectra in cerebral white matter with Fourier-transform diffusion MRI. Proc. Int. Soc. Magn. Reson. Med. 8, 5627 (2000).
Tuch, D. S. Q-ball imaging. Magn. Reson. Med. 52, 1358–1372 (2004).
Descoteaux, M., Angelino, E., Fitzgibbons, S. & Deriche, R. Regularized, fast, and robust analytical Q-ball imaging. Magn. Reson. Med. 58, 497–510 (2007).
Panagiotaki, E. et al. Compartment models of the diffusion MR signal in brain white matter: a taxonomy and comparison. Neuroimage 59, 2241–2254 (2012).
Dell’Acqua, F. et al. A model-based deconvolution approach to solve fiber crossing in diffusion-weighted MR imaging. IEEE Trans. Biomed. Eng. 54, 462–472 (2007).
Dell’Acqua, F. et al. A modified damped Richardson–Lucy algorithm to reduce isotropic background effects in spherical deconvolution. Neuroimage 49, 1446–1458 (2010).
Nigel, I., Lawes, C. & Clark, C. A. Anatomical validation of DTI and tractography. In Diffusion MRI: Theory, Methods and Applications (ed. Jones, D. K.) 439–448 (Oxford University Press, 2010).
Jeurissen, B., Descoteaux, M., Mori, S. & Leemans, A. Diffusion MRI fiber tractography of the brain. NMR Biomed. 32, e3785 (2019).
Sarwar, T., Ramamohanarao, K. & Zalesky, A. Mapping connectomes with diffusion MRI: deterministic or probabilistic tractography? Magn. Reson. Med. 81, 1368–1384 (2019).
Tournier, J. D., Calamante, F. & Connelly, A. MRtrix: diffusion tractography in crossing fiber regions. Int. J. Imaging Syst. Technol. 22, 53–66 (2012).
Guevara, P. et al. Accurate tractography propagation mask using T1-weighted data rather than FA. In International Society for Magnetic Resonance in Medicine 19th Scientific Meeting Abstract no. 2018 (2011).
Catani, M. & Thiebaut de Schotten, M. Atlas of Human Brain Connections (Oxford University Press, 2015).
Reisert, M. et al. Global fiber reconstruction becomes practical. Neuroimage 54, 955–962 (2011).
Kreher, B. W., Mader, I. & Kiselev, V. G. Gibbs tracking: a novel approach for the reconstruction of neuronal pathways. Magn. Reson. Med. 60, 953–963 (2008).
Jbabdi, S., Woolrich, M. W., Andersson, J. L. R. & Behrens, T. E. J. A Bayesian framework for global tractography. Neuroimage 37, 116–129 (2007).
Mangin, J. ‐F. et al. A framework based on spin glass models for the inference of anatomical connectivity from diffusion‐weighted MR data—a technical review. NMR Biomed. 15, 481–492 (2002).
Reisert, M., Kiselev, V. G., Dihtal, B., Kellner, E. & Novikov, D. S. MesoFT: unifying diffusion modelling and fiber tracking. In Medical Image Computing and Computer-Assisted Intervention – MICCAI 2014 (eds. Golland, P., Hata, N., Barillot, C., Hornegger, J., and Howe, R.) 201–208 (Springer, Cham, Switzerland, 2014).
Christiaens, D. et al. Global tractography of multi-shell diffusion-weighted imaging data using a multi-tissue model. Neuroimage 123, 89–101 (2015).
Sherbondy, A. J., Rowe, M. C. & Alexander, D. C. MicroTrack: an algorithm for concurrent projectome and microstructure estimation. Med. Image Comput. Comput. Assist. Interv. 13, 183–190 (2010).
Sherbondy, A. J., Dougherty, R. F., Ananthanarayanan, R., Modha, D. S. & Wandell, B. A. Think global, act local; projectome estimation with BlueMatter. Med. Image Comput. Comput. Assist. Interv. 12, 861–868 (2009).
Pestilli, F., Yeatman, J. D., Rokem, A., Kay, K. N. & Wandell, B. A. Evaluation and statistical inference for human connectomes. Nat. Methods 11, 1058–1063 (2014).
Daducci, A., Dal Palù, A., Lemkaddem, A. & Thiran, J. P. COMMIT: convex optimization modeling for microstructure informed tractography. IEEE Trans. Med. Imaging 34, 246–257 (2015).
Sarwar, T. et al. Evaluation of tractogram filtering methods using human-like connectome phantoms. Neuroimage 281, 120376 (2023).
Yeh, C. H., Jones, D. K., Liang, X., Descoteaux, M. & Connelly, A. Mapping structural connectivity using diffusion MRI: Challenges and opportunities. J. Magn. Reson. Imaging 53, 1666–1682 (2020).
Watson, D. M. & Andrews, T. J. Mapping the functional and structural connectivity of the scene network. Hum. Brain Mapp. 45, e26628 (2024).
García-Gomar, M. G., Singh, K., Cauzzo, S. & Bianciardi, M. In vivo structural connectome of arousal and motor brainstem nuclei by 7 Tesla and 3 Tesla MRI. Hum. Brain Mapp. 43, 4397–4421 (2022).
Favaretto, C. et al. Subcortical-cortical dynamical states of the human brain and their breakdown in stroke. Nat. Commun. 13, 5069 (2022).
Alves, P. N., Forkel, S. J., Corbetta, M. & Thiebaut de Schotten, M. The subcortical and neurochemical organization of the ventral and dorsal attention networks. Commun. Biol. 5, 1343 (2022).
Yeh, C. H., Smith, R. E., Dhollander, T., Calamante, F. & Connelly, A. Connectomes from streamlines tractography: assigning streamlines to brain parcellations is not trivial but highly consequential. Neuroimage 199, 160–171 (2019).
Fischl, B. FreeSurfer. Neuroimage 62, 774–781 (2012).
Smith, R. & Connelly, A. MRtrix3_connectome: a BIDS application for quantitative structural connectome construction. In Organization for Human Brain Mapping W610 (2019).
Vinokur, L., Zalesky, A., Raffelt, D., Smith, R. E. & Connelly, A. A novel threshold-free network-based statistical method: demonstration and parameter optimisation using in vivo simulated pathology. In International Society of Magnetic Resonance in Medicine 23rd Annual Meeting Abstract no. 2846 (2015).
Maier-Hein, K. H. et al. The challenge of mapping the human connectome based on diffusion tractography. Nat. Commun. 8, 1349 (2017).
Wasserthal, J., Neher, P. & Maier-Hein, K. H. TractSeg—Fast and accurate white matter tract segmentation. Neuroimage 183, 239–253 (2018).
Neher, P. F., Côté, M. A., Houde, J. C., Descoteaux, M. & Maier-Hein, K. H. Fiber tractography using machine learning. Neuroimage 158, 417–429 (2017).
Théberge, A., Desrosiers, C., Descoteaux, M. & Jodoin, P. M. Track-to-Learn: a general framework for tractography with deep reinforcement learning. Med. Image Anal. 72, 102093 (2021).
Sarwar, T., Seguin, C., Ramamohanarao, K. & Zalesky, A. Towards deep learning for connectome mapping: a block decomposition framework. Neuroimage 212, 116654 (2020).
Benou, I. & Riklin Raviv, T. DeepTract: A probabilistic deep learning framework for white matter fiber tractography. In Medical Image Computing and Computer-Assisted Intervention – MICCAI 2019 (eds. Shen, D. et al.) 626–635 (Springer, Cham, Switzerland, 2019).
Poulin, P., Jörgens, D., Jodoin, P. M. & Descoteaux, M. Tractography and machine learning: current state and open challenges. Magn. Reson. Imaging 64, 37–48 (2019).
Smith, R. E., Skoch, A., Bajada, C., Caspers, S. & Connelly, A. Hybrid surface-volume segmentation for improved anatomically-constrained tractography. In Organization of Human Brain Mapping 1–5 (2020).
Yeh, C. H., Smith, R. E., Dhollander, T. & Connelly, A. Mesh-based anatomically-constrained tractography for effective tracking termination and structural connectome construction. In International Society of Magnetic Resonance in Medicine 25th Annual Meeting Abstract no. 0058 (2017).
Tournier, J. D., Calamante, F. & Connelly, A. Improved probabilistic streamlines tractography by 2nd order integration over fibre orientation distributions. In International Society for Magnetic Resonance in Medicine-European Society for Magnetic Resonance in Medicine and Biology Joint Annual Meeting Abstract no. 1670 (2010).
Szczepankiewicz, F., Hoge, S. & Westin, C. F. Linear, planar and spherical tensor-valued diffusion MRI data by free waveform encoding in healthy brain, water, oil and liquid crystals. Data Brief. 25, 104208 (2019).
Wu, D., Martin, L. J., Northington, F. J. & Zhang, J. Oscillating gradient diffusion MRI reveals unique microstructural information in normal and hypoxia-ischemia injured mouse brains. Magn. Reson. Med. 72, 1366–1374 (2014).
Cory, D. G., Miller, J. B. & Garroway, A. N. Time-suspension multiple-pulse sequences: applications to solid-state imaging. J. Magn. Reson. 90, 205–213 (1990).
Shemesh, N. et al. Conventions and nomenclature for double diffusion encoding NMR and MRI. Magn. Reson. Med. 75, 82–87 (2016).
Caruyer, E., Lenglet, C., Sapiro, G. & Deriche, R. Design of multishell sampling schemes with uniform coverage in diffusion MRI. Magn. Reson. Med. 69, 1534–1540 (2013).
Dhollander, T., Raffelt, D. & Connelly, A. Unsupervised 3-tissue response function estimation from single-shell or multi-shell diffusion MR data without a co-registered T1 image. In Conference: ISMRM Workshop on Breaking the Barriers of Diffusion MRI. 5 (2016).
Dhollander, T. & Connelly, A. A novel iterative approach to reap the benefits of multi-tissue CSD from just single-shell (+b=0) diffusion MRI data. In Proceedings of the International Society of Magnetic Resonance in Medicine 24th Annual Meeting Abstract no. 3010 (2016).
Zeng, R. et al. FOD-Net: a deep learning method for fiber orientation distribution angular super resolution. Med. Image Anal. 79, 102431 (2022).
Jones, D. K., Horsfield, M. A. & Simmons, A. Optimal strategies for measuring diffusion in anisotropic systems by magnetic resonance imaging. Magn. Reson. Med. 42, 515–525 (1999).
Bastiani, M. et al. Automated quality control for within and between studies diffusion MRI data using a non-parametric framework for movement and distortion correction. Neuroimage 184, 801–812 (2019).
Smith, R. E., Connelly, A. & Calamante, F. Diffusion MRI fiber tractography. In Advances in Magnetic Resonance Technology and Applications (eds. Seiberlich, N. et al.) 533–569 (Academic Press, 2020).
Desikan, R. S. et al. An automated labeling system for subdividing the human cerebral cortex on MRI scans into gyral based regions of interest. Neuroimage 31, 968–980 (2006).
Tzourio-Mazoyer, N. et al. Automated anatomical labeling of activations in SPM using a macroscopic anatomical parcellation of the MNI MRI single-subject brain. Neuroimage 15, 273–289 (2002).
Schilling, K. G. et al. Synthesized b0 for diffusion distortion correction (Synb0-DisCo). Magn. Reson. Imaging 64, 62–70 (2019).
Greve, D. N. & Fischl, B. Accurate and robust brain image alignment using boundary-based registration. Neuroimage 48, 63–72 (2009).
Tahedl, M., Tournier, J.-D. & Smith, R. SC-construction-using-MSMT-CSD. Open Science Framework https://osf.io/tm5x8 (2024).
Acknowledgements
This work was supported by funding from the National Health and Medical Research Council of Australia. R.E.S. is supported by fellowship funding from the National Imaging Facility (NIF), an Australian Government National Collaborative Research Infrastructure Strategy (NCRIS) capability. J.-D.T. was supported with funding from the European Research Council under the European Union’s Seventh Framework Programme (FP7/20072013), ERC grant agreement no. 319456 (developing Human Connectome Project) and MRC strategic funds MR/K006355/1. J.-D.T. was also supported by the Wellcome/EPSRC Centre for Medical Engineering at King’s College London (WT 203148/Z/16/Z) and by the National Institute for Health Research (NIHR) Biomedical Research Centre at Guy’s and St Thomas’ NHS Foundation Trust and King’s College London. The views expressed are those of the authors and not necessarily those of the NHS, the NIHR or the Department of Health. M.T. is supported by a Walter-Benjamin Postdoc Stipend from the Deutsche Forschungsgemeinschaft (DFG, TA 1902/1-1).
Author information
Authors and Affiliations
Contributions
All authors contributed extensively to the work presented in this paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks Linden Parkes and Fan Zhang for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Key references
Tournier, J.-D. et al. Neuroimage 202, 116137 (2019): https://doi.org/10.1016/j.neuroimage.2019.116137
Smith, R. E. et al. Neuroimage 62, 1924–1938 (2012): https://doi.org/10.1016/j.neuroimage.2012.06.005
Smith, R. E. et al. Neuroimage 104, 253–265 (2015): https://doi.org/10.1016/j.neuroimage.2014.10.004
Smith, R. E. et al. Neuroimage 119, 338–351 (2015): https://doi.org/10.1016/j.neuroimage.2015.06.092
Supplementary information
Supplementary Information
Supplementary material providing additional instructions for quality assessment and visual inspection of the main protocol.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tahedl, M., Tournier, JD. & Smith, R.E. Structural connectome construction using constrained spherical deconvolution in multi-shell diffusion-weighted magnetic resonance imaging. Nat Protoc 20, 2652–2684 (2025). https://doi.org/10.1038/s41596-024-01129-1
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41596-024-01129-1