Abstract
Terbinafine, fluconazole, and amorolfine inhibit fungal ergosterol synthesis by acting on their target enzymes at different steps in the synthetic pathway, causing the accumulation of various intermediates. We found that the effects of these three in- hibitors on yeast morphology were different. The number of morphological parameters commonly altered by these drugs was only approximately 6% of the total. Using a rational strategy to find commonly changed parameters,we focused on hidden essential similarities in the phenotypes possibly due to decreased ergosterol levels. This resulted in higher apparent morphological similarity. Improvements in morphological similarity were observed even when canonical correlation analysis was used to select biologically meaningful morphological parameters related to gene function. In addition to changes in cell morphology, we also observed differences in the synergistic effects among the three inhibitors and in their fungicidal effects against pathogenic fungi possibly due to the accumulation of different intermediates. This study provided a comprehensive understanding of the properties of inhibitors acting in the same biosynthetic pathway.
Similar content being viewed by others
Introduction
Sterols are essential components of eukaryotic cell membranes, which maintain membrane structural integrity, fluidity, and permeability, and function in the formation of lipid rafts and regulation of membrane-bound enzyme activity1. The main sterols in animals and plants are cholesterol and phytosterol, respectively, whereas ergosterol is the major sterol in fungi. The ergosterol biosynthetic pathway has been actively studied in the budding yeast Saccharomyces cerevisiae, as a model fungal organism. As ergosterol is essential for yeast growth and viability, deletions of most genes encoding its serial biosynthetic enzymes are lethal2. In addition, yeast mutants defective in the branched synthetic pathway at a late stage exhibit a wide range of phenotypes, including changes in cell morphology3, altered susceptibility to chemical treatments4, increased tolerance to high temperatures5, and reduced tolerance to other stresses, such as high ethanol, salt, and solute concentrations6. Such diverse phenotypes are observed because of the wide range of functions of various sterols in yeast cells1.
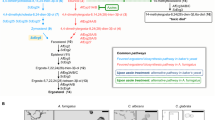
The fungal ergosterol synthetic pathway has been evaluated extensively as a target of antifungal agents7. Terbinafine (TBF), fluconazole (FCZ), and amorolfine (AMF), belonging to allylamines, azoles, and morphololines, respectively, have been shown to interfere with different steps in the serial ergosterol synthetic pathway (Fig. 1). TBF inhibits squalene epoxi- dase (Erg1), which catalyzes the conversion of squalene into squalene epoxide, the precursor of lanosterol8. FCZ inhibits a cy- tochrome P450-dependent synthetic enzyme, lanosterol-14-demethylase (Erg11), which catalyzes the conversion of lanosterol into 4,4-dimethyl cholesta-8,14,24-trienol, the precursor of 4,4-dimethyl zymosterol9,10. AMF has dual targets: 14-reductase (Erg24), which catalyzes the conversion of 4,4-dimethyl cholesta-8,14,24-trienol to 4,4-dimethyl zymosterol (the precursor of fecosterol), and C-8 sterol isomerase (Erg2), which catalyzes the conversion of fecosterol into episterol (the precursor of ergosta-5,7,24(28)-trienol and ergosterol)11. Treatments with inhibitors of each part of the ergosterol biosynthetic pathway result in a decreased level of the final product, ergosterol, resulting in common functional defects. Previous studies also showed that treatment with azole induced the accumulation of lanosterol, a substate of its target enzyme, lanosterol-14-demethylase12,13. Similarly, treatment with TBF and AMF resulted in accumulation of their intermediate compounds, squalene and fecosterol, respectively, resulting in specific effects14,15,16,17. Although the effects of ergosterol synthesis inhibitors are primarily due to marked decreases in level of the final product, ergosterol, it has also been argued that accumulation of intermediate products leads to specific perturbations in the fungal cell membrane18.
Ergosterol synthetic pathway in budding yeast. Targets of ergosterol synthesis inhibitors (AMF, FCZ, and TBF) are shown. Protein structures were obtained from AlphaFold Protein Structure Database (https://alphafold.ebi.ac.uk/).
Deletion of a gene with the same function results in a similar morphological phenotype. For example, of gene-deletion mutants of the five yeast polarisome subunits, four with obvious phenotypes (bni1, bud6, spa2, and pea2) all have round buds19. Moreover, there are numerous other examples among the subunit mutants of numerous complexes20,21. Calculating morphological similarity using over 500 morphological features increases our understanding of the close relationships between gene functions and mutant phenotypes of the subunits in many protein complexes22. Therefore, the effects of common functional defects will appear in mutants with similar morphological phenotypes. However, the addition of further effects of a specific perturbation should have different effects on the morphological phenotype.
This study revealed that the morphological features of yeast after treatment with TBF, FCZ, and AMF are quite different. Subsequently, we employed a rational approach to transform and analyze morphological profiles, aiming to understand the essential similarities induced by these chemical perturbations. Using logistic regression analysis and canonical correlation analysis (CCA), we extracted morphological phenotypic traits closely correlated with functions common to the ergosterol pathway. Based on these analyses, we propose a theory to determine rational associations with interconnected omics data, such as gene functions and mutant phenotypes.
Results
Morphological features induced by three ergosterol synthesis inhibitors
To investigate the effects of ergosterol and cell wall inhibitors on yeast morphology (Fig. 2A), we examined the dose- dependent changes induced by the inhibitors. To exclude the secondary effects of the inhibitors, we set the highest concentra- tion of each drug to that at which only 10 % growth inhibition was observed. After 16-h of incubation of wild-type (WT) yeast cells with four or five different concentrations of the inhibitors (Supplementary Table S1), the yeast cells were harvested, fixed with formaldehyde, and used for observation of cell, actin, and nuclear DNA morphology under a fluorescence microscope. The fluorescent images were then analyzed using the CalMorph image processing system specialized for budding yeast19. To precisely estimate the correct values of drug-dependent morphological changes, we assigned the best-fit unimodal distribution model for experimental replicates in each morphological measure23. We applied generalized linear models (GLMs) with 490 morphological unimodal parameters to achieve a powerful downstream parametric approach. We extracted dose-dependent morphological changes using simple regression analysis by using them for each trait, assuming homoscedasticity, with the concentration of the drug as an explanatory variable. Significant dose-dependent morphological changes were found in 120, 80, and 57 parameters after treatment with TBF, FCZ, and AMF, respectively (FDR = 0.05, Wald-test; Fig. 2B, Supplementary Table S2). TBF mainly affected cellular and nuclear morphology, with changes in 50 cell, 25 actin, and 45 nuclear features. FCZ mainly affected nuclear morphology, including changes in 15 cell, 19 actin, and 46 nuclear features. AMF also mainly affected nuclear morphology, with changes in 15 cell, 4 actin, and 38 nuclear features. A Venn diagram showed that there were fewer common morphological defects caused by ergosterol inhibitors (12 features in Fig. 2B, only 6% of the total parame- ters altered) than by 1,3-β-glucan synthase inhibitors (226 features in Fig. 2C). The typical alterations induced by the ergosterol inhibitors included the cell axis ratio, size of the actin region, and nuclear length (Supplementary Table S2). Thus, examining morphological changes induced by TBF, FCZ, and AMF highlighted the different effects of these inhibitors on yeast morphology.
Drug effects on cell morphology. (A) Examples of the effects of ergosterol synthesis inhibitors (AMF, FCZ, and TBF) and 1,3-β-glucan synthase inhibitors (ECB and MFG) on cell morphology. Triple-stained cells are shown (red, actin; green, cell wall; blue, nucleus). Bar shows 5.0 µm. Venn diagram of morphological defects due to perturbation of ergosterol synthesis (B) and 1,3-β-glucan synthase (C) at FDR = 0.05 (Supplementary Table 2). Bar plots show number of defective parameters in actin (red), cell wall (green), and nucleus (blue). Schematic morphological defects, caused by all of the ergosterol synthesis inhibitor treatments, are also shown. Data Set 1 was used in this analysis.
Morphological similarity in the effects of terbinafine, fluconazole, and amorolfine
Morphological similarity can be estimated based on the correlation coefficient of morphological profiling in orthogonal co- ordinates. After normalizing the 490 unimodal morphological parameters using a GLM, we performed principal component analysis (PCA) using 109 replicates of WT morphological data to prepare the orthogonal coordinates. Dose-dependent morphological changes in each of the PCs were converted from the slopes estimated by simple regression analysis using the Wald test statistic, yielding morphological profiling and the correlation coefficient calculated from the principal component (PC) scores (Fig. 3A). The degrees of morphological similarity among the effects of the three ergosterol inhibitors were not very high, with a correlation coefficient of 0.61 between the highest pairs, TBF and AMF, and only of 0.48 between the lowest, TBF and FCZ (Fig. 3A, B). However, morphological similarity was high between the effects of echinocandin B (ECB) and micafungin (MFG) at 0.716 (Fig. 3A, B). As treatment with ergosterol inhibitors causes accumulation of different intermedi- ates12,13,14,15,16,17, the unexpectedly low level of morphological similarity was thought to be due to the specific effects of chemical perturbation.
Morphological similarity between chemically perturbed cells. (A) Correlation analysis among the inhibitors based on the maximum variation of quantified morphological features (CCR = 100%). Values are Pearson’s correlation coefficients (r) between ergosterol synthesis inhibitors (AMF, FCZ, and TBF) and 1,3-β-glucan synthase inhibitors (ECB and MFG). (B) Scatter plots of pairwise correlations between ergosterol synthesis inhibitors and 1,3-β-glucan synthase inhibitors. Data Set 1 was used in this analysis.
Rational selection of the morphological parameters to extract common features of chemical perturbation in the ergosterol pathway
The morphological similarity between TBF-, FCZ-, and AMF-treated cells was not high when all PCs (cumulative contribution ratio [CCR] = 100%) were used because the common chemical perturbation of each drug was masked by specific perturbations. That is, the functional defects of ergosterol production were hidden by the effects of the accumulation of intermediates. Therefore, we extracted morphological features with similar effects associated with TBF, FCZ, and AMF treatment. After selecting 40 reliable PC spaces with the lowest levels of noise (Supplementary Figure 3), we obtained a set of PCs that distinguished all drug-treated cells from untreated cells by applying logistic regression (Fig. 4A). We then selected 19 PCs as robust classifiers for the three ergosterol inhibitors after repeating 5000 times. When the morphological similarity was again calculated using these 19 selected morphological components, the correlation coefficient between TBF and AMF was increased to 0.833 (Fig. 4B, C). A similar analysis revealed that the correlation coefficients between cell wall inhibitors were still high (Fig. 4B, C). Therefore, we successfully extracted the morphological components common to the three drugs by focusing on common functional defects and concluded that the morphological similarity became noticeable when rationally selected common components were used.
Morphological similarity increases with extraction of common features of chemical perturbation. (A) Selection of a set of PCs that distinguished all drug-treated cells from untreated cells using logistic regression. After logistic regression models were created, the frequency at which each PC was selected was calculated. The best thresholds for ergosterol synthesis and 1,3-β-glucan synthase inhibitors determined by the AIC were 1930 (19 PCs) and 750 (7 PCs), respectively. Red and blue bars represent treatment with ergosterol and glucan synthesis inhibitors, respectively. (B) Correlation analysis of the inhibitors after selecting common features of chemical perturbation in “A”. Values are Pearson’s correlation coefficients (r) between ergosterol synthesis inhibitors (AMF, FCZ, and TBF) and cell wall inhibitors (ECB and MFG). (C) Scatter plots of pairwise correlations between ergosterol synthesis inhibitors and 1,3-β-glucan synthase inhibitors. Data Set 1 was used in this analysis.
Selection of morphological components based on canonical correlation analysis
The morphological similarity increased when using the 19 selected morphological components affected similarly by the three drugs. As an alternative method of rational selection, we used CCA, a classic statistical method for exploring the relationships between two multivariate sets of variables: morphological components and gene functions. Using CCA, it is possible to extract biologically important morphological parameters22,23. After performing PCA and CCA, we successfully compressed all combinations of 490 morphological traits and 3568 gene ontology (GO) terms into linear combinations of morphological features (28 phenotype canonical variables [pCVs]) and gene functions (28 GO term canonical variables [gCVs]).
One advantage of using canonical variables is that we can build a rational predictor selected by combinational optimization and logistic regression. Using morphological data from WT cells treated with the ergosterol synthesis inhibitors as training data, we identified a predictor composed of the 7 pCVs for discriminating morphological phenotypes of the erg28∆ mutant defective in the synthesis of fecosterol (Fig. 1). pCV1 and pCV2 distinguished 11 replicates of erg28∆ from the WT controls (Fig. 5A). The selected predictor proved correct because its posterior probability for 9 of the 11 replicates of erg28∆ exceeded 99% (Fig. 5B). Then, the morphological similarity between TBF-, FCZ-, and AMF-treated cells was calculated using the morphological components of the 7 pCVs (Fig. 5C). When using the originally less-correlated data set (Data Set 2), the morphological similarity showed markedly high values ranging from 0.298 to 0.750 (Fig. 5A). However, using a data set with relatively strong original correlations (Data Set 1) did not result in higher degrees of morphological similarity (Supplementary Table S3), indicating that the method using CCA was not always effective. This was probably because the morphological features exhibited by erg28∆ are different from the common features caused by the three drugs. In contrast, the method using the selected 19 morphological components worked well for both data sets (Supplementary Table S3). These analyses showed that rationally selecting the common components among the three drugs was more robust than was the method using CCA.
Morphological similarity calculated by canonical variables. (A) Scatter plot of the first and second canonical variable (CV) spaces among the biologically important components. Black and yellow circles indicate 4708 non-essential gene deletion mutants and 109 wild-type (WT) replicates, respectively. Red circles indicate 11 replicates of the erg28∆ mutant as a test data set. (B) Prediction of 11 replicates of the erg28∆ mutants by logistic regression fitted with the linear predictor composed of the 7 pCVs. Red and yellow circles indicate 11 replicates of erg28∆ mutants and 109 replicates of the WT replicates, respectively. The orange curve indicates the posterior probability at each x-axis value. Gray and yellow histograms indicate 4708 non-essential gene deletion mutants and 109 replicates of WT, respectively. (C) Increase in the morphological similarity among the cells treated with ergosterol inhibitors (Data Set 2). Morphological similarities among the inhibitors were calculated by the morphological traits before (left panel) and after (right panel) CCA. Left panel: 95 PC scores of each drug were obtained by projecting Z values of the 490 morphological traits onto the 95 PC axes extracted by the 109 WT replicates at CCR = 0.99 and used for estimation of the morphological similarity by Pearson’s correlation coefficient (r). Right panel: correlation coefficients were calculated using the 7 pCV scores among the inhibitors.
Phenotypic similarity and dissimilarity caused by the ergosterol inhibitors
Phenotypic characteristics other than morphology were investigated to further examine the effects of the ergosterol inhibitors. The synergistic effects among the three inhibitors were examined after treatment with pairs of inhibitors in various combina- tions using a checkerboard assay. We found a fractional inhibitory concentration (FIC) index of 0.047 between FCZ and TBF, indicating a solid synergistic effect (Fig. 6A). However, weak synergistic effects were seen in combinations with AMF (Fig. 6A). On the other hand, ECB showed only weak synergistic effects with all ergosterol inhibitors (Supplementary Fig. 1).
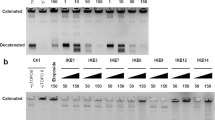
Effects of the drugs on phenotypes other than morphology. (A) Checkerboard assay performed using combinations of two of the three drugs (AMF, FCZ, and TBF) (n = 3). Green regions represent higher cell densities. The FIC index represents the effect of the combined use of the two compounds and is expressed as the mean value of three biological replicates. FIC index values were classified as follows: synergy (FIC index ≤ 0.5), weak synergy (0.5 < FIC Index < 1.0) indifference (1.0 < FIC Index < 4.0), and antagonism (4.0 < FIC Index). (B) Resazurin assay of C. albicans ATCC24433 to show fungicidal activity. An image of a cell plate showing the conversion of resazurin (blue) to resorufin (pink), with cells progressively losing their viability from right to left. (C) Venn diagram of two phenotypes: strength of synergistic effect with ergosterol inhibitors and verification of fungicidal action. Relationships between specific intermediate products accumulated by each drug treatment and the phenotype. Red or black arrows indicate increased biosynthetic product accumulation; blue arrows indicate decreased biosynthetic product accumulation.
Resazurin was then used to examine the fungicidal activity of the inhibitors against the pathogenic fungus Candida albicans. Purple resazurin converts to pink resorufin upon reduction by electron acceptance from the mitochondrial respiratory chain of living fungal cells. A high concentration of AMF yielded a purple color, indicating fungicidal activity, while FCZ and TBF yielded a pink color, indicating fungistatic activity (Fig. 6B). These studies again highlight common and specific phenotypes after treatment with the three ergosterol inhibitors (Fig. 6C).
Discussion
The ergosterol synthetic pathway inhibitors TBF, FCZ, and AMF have different effects on yeast cells. The number of morphological parameters commonly altered by these three drugs was only approximately 6% of the total altered parameters. The morphological similarity associated with these three drugs was lower than that observed with treatment with the cell wall synthesis inhibitors ECB and MCF (r = 0.716). In this study, we focused on the morphological parameters of the cell, actin, and nuclear DNA because we have previously established unimodal parameters for these components23. Unimodal parameters facilitate the application of GLMs and the estimation of true values of morphological changes. Including the morphology of the endoplasmic reticulum (ER) and vacuoles, which are significantly affected by ergosterol levels, could reveal more morphological commonalities. However, further studies are needed to determine the best-fit unimodal distribution models for ER and vacuole parameters and to accurately estimate the true extent of morphological changes. In addition to changes in yeast cell morphology, differences were observed in the synergy among these three drugs and their fungicidal effects on pathogenic fungi. As these three inhibitors cause accumulation of different intermediates of the ergosterol synthetic pathway12,13,14,15,16,17, it was assumed that they have different effects on the cells.
The three antifungal drugs have the common effect of reducing the level of ergosterol, the product of this biosynthetic pathway. Therefore, essential similarities caused by the reduction in ergosterol levels should be hidden in the phenotype. In fact, instead of calculating correlation coefficients using all morphological parameters, the apparent morphological similarity was increased by focusing on a small number of parameters that changed in sync. We also extracted biologically meaningful morphological parameters related to gene function using CCA and showed that the optimized model worked well in some cases. The high degrees of morphological similarity were logical only when they were functionally homologous. For example,the petals of Dracula simia, an epiphytic orchid found in Peru and Bolivia, resemble the face of Mandrillus sphinx. However, they are not homologous organs. On the other hand, a reasonable argument is that the wings of dragonflies, bats, and birds, which are necessary for flying, are all thin, broad, and horizontal in shape. Therefore, an approach focusing on morphology related to gene function would be necessary to select for essential morphological similarities.
Antifungal drugs with similar modes of action usually have the same effect on yeast cells.ECB and MCF, which act on Fks1 and inhibit 1,3-β-glucan synthesis, had similar morphological effects on yeast cells. The effects of Jervine and D75-4590, which act on Kre6 and Skn1 and inhibit 1,6-β-glucan synthesis, also showed significantly higher degrees of morphological similarity24. Therefore, drugs that bind to the same target protein may not show the phenomena observed with ergosterol synthesis inhibitors. Ergosterol synthesis inhibitors are used in different applications. Allylamine and morpholine are drugs primarily targeting fungal infections of the skin, nails, athlete’s foot, and ringworm, whereas azoles are most widely used to treat meningitis, oral infections, and candidiasis. Further studies are required to assess how the different modes of action of antifungal agents and their solubility and pharmacokinetics are related to their various uses.
Methods
Strains and medias
Saccharomyces cerevisiae Y8835 (MATα ura3∆0:: natMX4 can 1∆:: STE2pr-Sp_his5 lyp1∆ his3∆1 leu2∆0 met15∆0 LYS2), which is derived from S288C, was used as wild-type strain25. C. albicans ATCC24433 was obtained from Medical Mycology Research Center, Chiba University. Media and growth condition for S. cerevisiae is as described previously19. C. albicans was grown at 35°C on Sabouraud agar media (1% (w/v) Bacto-peptone (BD Biosciences, CA, USA), 4% (w/v) Glucose (Fujifilm Wako Pure Chemical Corporation, Osaka, Japan), and 1.5% (w/v) Agar). For Candida susceptibility testing, RPMI 1640 (Fujifilm Wako Pure Chemical Corporation) was adjusted to pH 6.9 by addition of 165 mM 3-(N-morpholino) propanesulfonic acid (Fujifilm Wako Pure Chemical Corporation).
Drug and reagents
TBF (Tokyo Chemical Industry, Tokyo, Japan), FCZ (Tokyo Chemical Industry), AMF (Tokyo Chemical Industry), MFG (Funakoshi, Tokyo, Japan) and echinocandin B (ECB, Chugai Pharmaceutical Corporation, Tokyo, Japan) were all dissolved in dimethyl sulfoxide (DMSO; Fujifilm Wako Pure Chemical Corporation). The concentrations of the drugs are described in Supplementary Table S1.
Antifungal susceptibility testing for S. cerevisiae
Antifungal susceptibility against S. cerevisiae wild-type strain Y8835 was examined using a method described previously25. The concentrations of each drug for Y8835 testing were as follows: FCZ, 0, 1.56, 3.13, 6.25, 12.5, 25, 50, and 100 μg/mL; TBF, 0, 8, 16, 32, 64, 128, 256, and 512 μg/mL; AMF, 0, 0.0313, 0.0625, 0.125, 0.25, 0.5, 1, and 2 μg/mL; ECB, 0, 0.5, 1, 2, 4, 8, 16, and 32 μg/mL. The optical density at 600 nm was measured using the SpectraMax plus 384 plate reader (Molecular Devices, San Jose, CA, USA). The half maximal inhibitory concentration was estimated using the Markov Chain Monte Carlo method (5000 iterations including the first 2000 iterations as warm-up in eight chains each) in the rstan package (https://mc-stan.org/users/interfaces/rstan) using reparameterizing of the four-parameter log-logistic equation implemented in the drc package in R26.
Checkerboard assay
The synergy of test compounds and the FIC index was examined by the checkerboard method27. Test dilutions were selected based on the minimum inhibitory concentration (MIC) of each substance. The strains were exposed to various concentrations of FCZ (0, 1.56, 3.13, 6.25, 12.5, 25, 50, and 100 μg/mL) in combination with TBF (0, 8, 16, 32, 64, 128, 256, and 512 μg/mL), AMF (0.0313, 0.0625, 0.125, 0.25, 0.5, 1, and 2 μg/mL) or ECB (0, 0.5, 1, 2, 4, 8, 16, and 32 μg/mL.). The FIC index was calculated using the following formula: FIC index = FIC index (A) + FIC index (B) = [MIC (combination) / MIC (A alone)] + [MIC (combination) / MIC (B alone)]. Synergy was defined as a FIC index of ≤ 0.5. The test was performed in 96-well microtiter plates containing yeast peptone dextrose supplemented with drugs in serial concentrations. Fungal suspensions were inoculated at a cell density of 1 × 105 cells/mL. Plates were read after incubation for 18–24 h at 30°C. Each test was performed in triplicate.
Resazurin cell viability assay
Susceptibility tests with C. albicans were performed using the Clinical and Laboratory Standards Institute document M60. Briefly, C. albicans was grown on Sabouraud glucose agar for 24 h at 35°C, washed with 0.85% saline, and diluted in RPMI 1640 to 5 × 103 cells/mL. Diluted cells and 3% DMSO (with/without drugs) were added to 96-well round-bottom microplates and incubated at 35 °C in a static incubator. The drug concentration was used as same as checkerboard assay. The fungal cell killing was determined by the resazurin cell viability assay as described previously28. After incubation of C. albicans ATCC24433 with drugs at various concentrations for 24 h, 10 μL (2.1 mg/mL) resazurin was added to each well and incubated for 24 h. The color of each well was determined visually; a blue or purple color was interpreted as the absence of metabolic activity and dead cells, whereas pink indicated the presence of living fungal cells.
Fluorescence staining
Fluorescence staining was performed as described previously19. Briefly, cells of yeast strains were cultivated until the early log phase (< 5.0 × 106 cells/mL) and fixed in medium containing 3.7% (w/v) formaldehyde (Fujifilm Wako Pure Chemical Corporation). The fixed cells were triple-stained with fluorescein isothiocyanate-conjugated concanavalin A (Sigma-Aldrich Co. LLC, St. Louis, MO, USA) for the cell wall, rhodamine–phalloidin (Invitrogen, Carlsbad, CA, USA) for the actin cytoskeleton, and 4,6-diamidino-2-phenylindole (Sigma-Aldrich Co. LLC) for nuclear DNA.
Microscopy and image acquisition
Fluorescence image of the cells were acquired using a microscope (Axioplan 2, Carl Zeiss, Oberkochen, Germany) equipped with 6100 Ecplan-Neofluar (Carl Zeiss), a cooled-charge-coupled device camera (CoolSNAP HQ; Roper Scientific Photometrics, Tucson, AZ, USA), and AxioVision software (Carl Zeiss).
Image quantification and analysis
CalMorph (version 1.2) was used for quantitative analysis of the images. Only those experiments containing at least 123 cells were considered for subsequent analysis. CalMorph generates 501 morphological parameters related to the cell cycle phase, actin cytoskeleton, cell wall, and nuclear DNA (see descriptions for each parameter at http://www.yeast.ib.k.u-tokyo.ac.jp/CalMorph/index.html). The appropriate probability distributions were reported previously for only unimodal parameters (490) using the the UNImodal MOrphological [UNIMO23] data pipeline. Briefly, CalMorph parameters were first categorized according to their data type (including the non-negative, ratio, coefficient of variation, and proportion parameters). Then, the best probability distribution was defined for each parameter using the Akaike information criterion (AIC). In this analysis, we used these 490 morphological unimodal parameters because the optimal probability distributions have been defined for them, allowing for highly accurate statistical analysis.
Statistical modeling to assess the effects of drugs
Two datasets were used to analyze the morphological information of cells treated with AMF, FCZ, and TBF. Data Set 1 has relatively high correlation because it is data from 10 repeated experiments. Data Set 2, on the other hand, has relatively low correlation because it is data from five repeated experiments. All statistical analyses were performed using R (http://www.r-project.org). To assess the effects of each drug on cell morphology, a regression GLM29 was built. The cell morphological traits were compared with the corresponding WT distribution (i.e., null distribution) in each model. The Wald test (two-sided) was used to transform each morphological parameter to a Z value using the summary function in the gamlss package30. The false discovery rate (FDR) was finally estimated using the qvalue package31.
Projection of morphological data
To normalize the morphological data, PCA was performed based on the Z values (Wald test) of 109 WT replicates23 in 490 unimodal parameters using the prcomp function in the stats package (Supplementary Fig. 2A). Then, to obtain orthogonal morphological data, the Z values of the unimodal parameters of 4708 non-essential mutants (Supplementary Fig. 2B)23, ergosterol synthesis inhibitors (Supplementary Fig. 2C), and glucan synthase inhibitors (Supplementary Fig. 2D) were projected by PCA of the 109 WT replicates. Orthogonal PC space was used to check the similarity of morphological profiles induced by the inhibitors by calculating the Pearson’s correlation coefficients (r) of the PC scores.
Selection of important parameters
To select important parameters, reliable PC spaces were first defined. For this purpose, a ratio of variance of 4708 non-essential mutants to 109 WT replicates was used to extract biologically meaningful signals (Supplementary Fig. 3A). A locally estimated scatterplot smoothing (LOESS) regression was fitted to the ratio values (Supplementary Fig. 3B). The best smooth span for the regression model was defined as follows30: gamlss (y~lo(~x, span = f), data = data, family = NO), where y is the ratio of variance of 4708 non-essential mutants to 109 WT replicates, x is the ordinal number of PC spaces, and f is the smooth span of the LOESS regression (ranging from 0.10 to 0.99). The best fit was chosen using the AIC. Then, the slope of the LOESS regression was calculated between successive PC spaces. Finally, the first 40 PCs (CCR = 83.64%) were considered reliable PC spaces in which the slope of the regression line was not constantly positive (Supplementary Fig. 3C). That is, the variance of 4708 non-essential mutants was not always higher than that of WT between PC1 and PC40, was always higher than that of WT from PC41 onward. Morphological similarity in fact increased after removal of noise (Supplementary Fig. 4). Next, a set of PC spaces that best distinguished drug-treated cells from WT cells were extracted. To avoid overfitting, each observation was considered as an independent entry (n = 130 and 70 for ergosterol synthesis inhibitors and 1,3-β-glucan synthase inhibitors, respectively), and Z values were calculated for each, as described above in the Statistical modeling, to assess effects of drugs subsection and projection of morphological data.
To determine the importance of each PC space to differentiate drug-treated cells from WT cells, two logistic regression models (one for ergosterol synthesis inhibitors and one for 1,3-β-glucan synthase inhibitors) were built using the brglm2 package32 using 60% of the data to train the model and 40% to test the model. The process was repeated 5000 times with random sampling in each model, and the best model was selected using the AIC in a stepwise algorithm (step function in R) in each iteration. The frequency of each PC space (Fig. 4A) and accuracy of the model (Supplementary Fig. 5) were calculated in these 5000 final models. Finally, the best frequency was chosen by defining various thresholds (t) and fitting a logistic model with the PCs that were observed at least t times. The best threshold was selected based on the AIC of fitted logistic regression. This threshold was 1930 times for ergosterol synthesis inhibitors including 19 PCs (CCR = 41.51%; Supplementary Fig. 6A) and 750 times for 1,3-β-glucan synthase inhibitors including 7 PCs (CCR = 30.91%; Supplementary Fig. 6B).
Canonical correlation analysis
CCA was used to explore the relationships between two multivariate sets of variables (morphological profile and functional profile) to find linear combinations of phenotypic traits and gene function features, as described previously22,23. CCA also helps to avoid possible overfitting that may occur by PC space selection.
Morphological profile
To examine common defects in the ergosterol pathway without bias against specific drug effects, we chose erg28∆. First, 10 replicates of erg28∆ (ERGosterol biosynthesis gene) and 10 replicates of WT cells were stained (see Fluorescence staining). Then, microscopy images (see Microscopy and image acquisition) were quantified to extract morphological traits (see Image quantification and analysis). Z values of these replicates were estimated as described in Statistical modeling to assess drug effects section. Finally, the morphological profile of 3917 non-essential mutants (see below) were subjected to PCA to provide phenotype principal components [pPCs] and Z values of 10 replicates of erg28∆ and WT replicates were projected using the PCA of the non-essential mutants. The non-essential data set also contains one replicate of erg28∆, thus, in total, 11 replicates of erg28∆ mutants were used.
Functional profile
The basic GO files were first downloaded from the GO Consortium and gene annotations were downloaded from the Saccharomyces Genome Database (SGD). Then, a Boolean matrix of GO terms was generated with a value of TRUE if a gene was annotated by a GO, otherwise the value was FALSE. Next, a GO slimmer process was performed as follows: Removal of global GO terms (i.e., annotated for more than 200 genes); removal of GO terms with identical sets of annotated ORFs; removal of unique GO terms (i.e., annotated for less than two genes); and exclusion of genes with no annotations. Finally, 3,568 GO terms of unique annotations for 3,917 genes were obtained. Next, PCA was performed to reduce the dimensionality of the functional profiles33, with some modifications to provide GO term principal components. This method preserved the structure of the functional relationships among the genes while reducing the dimensionality33. Finally, a zero matrix of the WT replicates was projected by the obtained PCA. CCA was finally performed to compress phenotype principal components (CCR = 95%) and GO term principal components (CCR = 99%) of the 3,917 genes into linear combinations using the cancor function of the R stats package. Bartlett’s Chi-squared test was used to check the significance of the canonical correlation coefficients. Ultimately, 28 morphological features (pCVs) and 28 gene function features (gCVs) were obtained (P < 0.05; Supplementary Fig. 7).
To obtain the best set of CVs to differentiate 11 replicates of erg28∆ mutants from WT replicates and other non-essential mutants, a logistic regression model was built using the brglm2 package32 as mentioned above. This was conducted using the morphological data of the ergosterol synthesis inhibitors as training data and 6000 times random sampling from the inhibitor data as a training data set and applying a stepwise algorithm (step function in R) in each iteration to select the best set of CVs. Finally, the best set of 7 CVs were chosen by the AIC (Supplementary Fig. 8). The precision was 0.905 with 11 replicates of erg28∆ and 10 replicates of WT as a test data set (Supplementary Fig. 8). For visual presentation, pCV1 and pCV2 among the 7 CVs were selected as having clear separation of erg28∆ replicates in this seven-dimensional space (Fig. 5A).
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Jordá, T. & Puig, S. Regulation of ergosterol biosynthesis in Saccharomyces cerevisiae. Genes 11, 795. https://doi.org/10.3390/genes11070795 (2020).
Johnston, E. J., Moses, T. & Rosser, S. J. The wide-ranging phenotypes of ergosterol biosynthesis mutants, and implications for microbial cell factories. Yeast 37, 27–44. https://doi.org/10.1002/yea.3452 (2020).
OMeara, T. R. et al. Global analysis of fungal morphology exposes mechanisms of host cell escape. Nat. Commun. 6, 6741. https://doi.org/10.1038/ncomms7741 (2015).
Mukhopadhyay, K. et al. Membrane sphingolipid-ergosterol interactions are important determinants of multidrug re-sistance in Candida albicans. Antimicrob. Agents Chemother. 48, 1778–1787. https://doi.org/10.1128/aac.48.5.1778-1787.2004 (2004).
Liu, G., Chen, Y., Færgeman, N. J. & Nielsen, J. Elimination of the last reactions in ergosterol biosynthesis alters the resistance of Saccharomyces cerevisiae to multiple stresses. FEMS Yeast Res. 17, fox063. https://doi.org/10.1093/femsyr/fox063 (2017).
Henderson, C. M. & Block, D. E. Examining the role of membrane lipid composition in determining the ethanol tolerance of Saccharomyces cerevisiae. Appl. Enviro. Microbiol. 80, 2966–2972. https://doi.org/10.1128/AEM.04151-13 (2014).
Bell, A. Comprehensive medicinal chemistry ii. 7, 445–468 (2007).
Nowosielski, M. et al. Detailed mechanism of squalene epoxidase inhibition by terbinafine. J. Chem. Inf. Model. 51, 455–462. https://doi.org/10.1021/ci100403b (2011).
Sagatova, A. A., Keniya, M. V., Wilson, R. K., Monk, B. C. & Tyndall, J. D. Structural insights into binding of the antifungal drug fluconazole to Saccharomyces cerevisiae lanosterol 14α-demethylase. Antimicrob. Agents Chemother. 59, 4982–4989. https://doi.org/10.1128/aac.00925-15 (2015).
Sheng, C. et al. Structure-based design, synthesis, and antifungal activity of new triazole derivatives. Chem. Biol. Drug Des. 78, 309–313. https://doi.org/10.1111/j.1747-0285.2011.01138.x (2011).
Polak, A. Mode of action of morpholine derivatives. Annals New York Acad. Sci. 544, 221–228. https://doi.org/10.1111/j.1749-6632.1988.tb40406.x (1988).
Van den Bossche, H. et al. In vitro and in vivo effects of the antimycotic drug ketoconazole on sterol synthesis. Antimicrob. Agents Chemother. 17, 922–928. https://doi.org/10.1128/aac.17.6.922 (1980).
Wang, T. et al. Jumonji histone demethylase inhibitor JIB-04 as a broad-spectrum antifungal agent. ACS Infect Dis. 8, 1316–1323. https://doi.org/10.1021/acsinfecdis.2c00096 (2022).
Ryder, N. S. Specific inhibition of fungal sterol biosynthesis by sf 86–327, a new allylamine antimycotic agent. Antimicrob. Agents Chemother. 27, 252–256. https://doi.org/10.1128/aac.27.2.252 (1985).
Liebl, M. et al. Quantifying isoprenoids in the ergosterol biosynthesis by gas chromatography-mass spectrometry. J. Fungi (Basel) 9, 768. https://doi.org/10.3390/jof9070768 (2023).
Polak-Wyss, A., Lengsfeld, H. & Oesterhelt, G. Effect of oxiconazole and ro 14–4767/002 on sterol pattern in Candida albicans. Sabouraudia 23, 433–441. https://doi.org/10.1080/00362178585380631 (1985).
Krauß, J. et al. Synthesis, biological evaluation, and structure-activity relationships of 4-aminopiperidines as novel antifungal agents targeting ergosterol biosynthesis. Molecules 26, 7208. https://doi.org/10.3390/molecules26237208 (2021).
Kontoyiannis, D. P. & Lewis, R. E. Antifungal drug resistance of pathogenic fungi. The Lancet 359, 1135–1144. https://doi.org/10.1016/S0140-6736(02)08162-X (2002).
Ohya, Y. et al. High-dimensional and large-scale phenotyping of yeast mutants. Proc. Natl. Acad. Sci. 102, 19015–19020. https://doi.org/10.1073/pnas.0509436102 (2005).
Sing, T. L. et al. The budding yeast rsc complex maintains ploidy by promoting spindle pole body insertion. J. Cell Biol. 217, 2445–2462. https://doi.org/10.1083/jcb.201709009 (2018).
Suzuki, G. et al. Global study of holistic morphological effectors in the budding yeast Saccharomyces cerevisiae. BMC Genomics 19, 1–14. https://doi.org/10.1186/s12864-018-4526-z (2018).
Ohnuki, S. & Ohya, Y. High-dimensional single-cell phenotyping reveals extensive haploinsufficiency. PLoS Biol. 16, e2005130. https://doi.org/10.1371/journal.pbio.2005130 (2018).
Ghanegolmohammadi, F., Ohnuki, S. & Ohya, Y. Assignment of unimodal probability distribution models for quantitative morphological phenotyping. BMC Biol. 20, 81. https://doi.org/10.1186/s12915-022-01283-6 (2022).
Kubo, K. et al. Jerveratrum-type steroidal alkaloids inhibit β-1, 6-glucan biosynthesis in fungal cell walls. Microbiol. Spectr. 10, e00873-e921. https://doi.org/10.1128/spectrum.00873-21 (2022).
Ohnuki, S. et al. High-throughput platform for yeast morphological profiling predicts the targets of bioactive compounds. Npj Syst. Biol. Appl. 8, 3. https://doi.org/10.1038/s41540-022-00212-1 (2022).
Ritz, C., Baty, F., Streibig, J. C. & Gerhard, D. Dose-response analysis using r. PloS One 10, e0146021. https://doi.org/10.1371/journal.pone.0146021 (2015).
Bidaud, A.-L., Schwarz, P., Herbreteau, G. & Dannaoui, E. Techniques for the assessment of in vitro and in vivo antifungal combinations. J. Fungi. 7, 113. https://doi.org/10.3390/jof7020113 (2021).
Zhang, X. et al. Development of lipo-γ-aa peptides as potent antifungal agents. J. Med. Chem. 65, 8029–8039. https://doi.org/10.1021/acs.jmedchem.2c00595 (2022).
Nelder, J. A. & Wedderburn, R. W. Generalized linear models. J. Royal Stat. Soc. Ser. A Stat. Soc. 135, 370–384. https://doi.org/10.2307/2344614 (1972).
Stasinopoulos, D. M. & Rigby, R. A. Generalized additive models for location scale and shape (GAMLSS) in R. J. Stat. Softw. 23, 1–46 (2008).
John, D. S., Andrew, J. B., Alan, D. & Robinson, D. qvalue: Q-value estimation for false discovery rate control. R Package version 2.6.0. https://bioconductor.org/packages/release/bioc/html/qvalue.html (2015).
Kosmidis, I., Kenne Pagui, E. C. & Sartori, N. Mean and median bias reduction in generalized linear models. Stat. Comput. 30, 43–59. https://doi.org/10.1007/s11222-019-09860-6 (2020).
De Haan, J. R. et al. Integrating gene expression and GO classification for PCA by preclustering. BMC Bioinform. 11, 1–10. https://doi.org/10.1186/1471-2105-11-158 (2010).
Acknowledgements
We thank Seiko Morinaga for obtaining morphological data using high-throughput microscopy, and Kuninori Suzuki and other members of the Laboratory of Signal Transduction for their helpful discussions. This work was supported by JSPS KAKENHI Grant Numbers 19H03205 and 23K23483 (YO) and the GAP fund of The University of Tokyo (YO).
Author information
Authors and Affiliations
Contributions
Conceptualization, Y.O.; Experiments, T.X., Y.L., and K.I.-N.; Analysis, S.O. and F.G.; Methodology, S.O. and F.G.; Writing Draft Preparation, Y.O; Writing Review and Editing, W.L. and Y.O.; Visualization, S.O., K.I.-N., and F.G.; Supervision, Y.O.; Funding, Y.O.; Project Administration, Y.O.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ghanegolmohammadi, F., Liu, W., Xu, T. et al. Rational selection of morphological phenotypic traits to extract essential similarities in chemical perturbation in the ergosterol pathway. Sci Rep 14, 17093 (2024). https://doi.org/10.1038/s41598-024-67634-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-67634-1