Abstract
Individuals with the Autism Susceptibility Candidate 2 (AUTS2) gene disruptions exhibit symptoms such as intellectual disability, microcephaly, growth retardation, and distinct skeletal and facial differences. The role of AUTS2 in neurodevelopment has been investigated using animal and embryonic stem cell models. However, the precise molecular mechanisms of how AUTS2 influences neurodevelopment, particularly in humans, are not thoroughly understood. Our study employed a 3D human cerebral organoid culture system, in combination with genetic, genomic, cellular, and molecular approaches, to investigate how AUTS2 impacts neurodevelopment through cellular signaling pathways. We used CRISPR/Cas9 technology to create AUTS2-deficient human embryonic stem cells and then generated cerebral organoids with these cells. Our transcriptomic analyses revealed that the absence of AUTS2 in cerebral organoids reduces the populations of cells committed to the neuronal lineage, resulting in an overabundance of cells with a transcription profile resembling that of choroid plexus (ChP) cells. Intriguingly, we found that AUTS2 negatively regulates the WNT/β-catenin signaling pathway, evidenced by its overactivation in AUTS2-deficient cerebral organoids and in luciferase reporter cells lacking AUTS2. Importantly, treating the AUTS2-deficient cerebral organoids with a WNT inhibitor reversed the overexpression of ChP genes and increased the downregulated neuronal gene expression. This study offers new insights into the role of AUTS2 in neurodevelopment and suggests potential targeted therapies for neurodevelopmental disorders.
Similar content being viewed by others
Introduction
Neurodevelopmental disorders (NDDs), including Autism Spectrum Disorders (ASD), Developmental Delay (DD), and Intellectual Disabilities (ID), arise from the impaired development of the central nervous system (CNS). Among other risk genes, Autism Susceptibility Candidate 2 (AUTS2) is frequently disrupted in various NDD patients1,2. AUTS2 disruptions manifest as neurodevelopmental issues such as microcephaly, developmental delay, and intellectual disabilities of varying degrees3,4,5. It has been established that AUTS2 plays a critical role in the development of the CNS. Various studies have highlighted its significance, with zebrafish experiments revealing that the absence of auts2 disrupts normal neurogenesis and craniofacial development5,6. Similarly, studies using mouse models have associated AUTS2 deficiency with CNS developmental issues and abnormal behaviors, including hypoplasia, imbalances in excitatory and inhibitory synaptic inputs, impaired motor skills, and deficits in cognitive and social behaviors7,8,9,10. Studies using mouse and human embryonic stem cells (ESCs) have further demonstrated that Auts2 deletion can cause differentiation anomalies in the neuronal lineage in vitro11,12,13.
AUTS2 has been identified as a part of the Polycomb Repressive Complex 1 (PRC1), an essential epigenetic regulator for suppressing gene transcription via chromatin modifications7,14. Further studies have identified additional factors, such as WDR68 and NRF1, that are crucial for the PRC1-AUTS2 complex functionality15,16. AUTS2 association with PRC1 remarkably alters the complex function, switching it from a repressor to an activator of transcription, particularly influencing gene expression in the CNS7,16. Additionally, AUTS2 influences cytoskeletal and neural development through its interaction with the Rac1 signaling pathway in the cytoplasm9. Recent research has broadened our understanding of nuclear AUTS2, revealing its role in the regulation of RNA transcription, splicing, and localization, achieved through interactions with various RNA-binding proteins17. Despite these insights into AUTS2 functions, the detailed mechanisms by which it regulates neurodevelopment, particularly at the molecular level in humans, remain a topic of ongoing investigation.
To understand the role of AUTS2 in human brain development, we utilized a cerebral organoids model system with the CRISPR/Cas9-engineered hESCs with AUTS2 deletion. Upon AUTS2 deletion, the cerebral organoids exhibited decreased size, reduced neuronal gene expression, and a defect in the neurodevelopment process. Interestingly, AUTS2 removal promotes the formation of fluid-filled compartments reminiscent of those observed in ChP organoids, suggesting a diversion from normal neuronal differentiation. Our RNA-seq analysis supported this hypothesis as AUTS2 deletion caused a downregulation in neuronal genes and an increase in ChP markers. Furthermore, our findings indicate that removing AUTS2 activates the WNT/β-catenin signaling, a pathway critical for the patterning, proliferation, and differentiation in neural development. Remarkably, introducing a WNT inhibitor reversed the decreased expression of neuronal genes and the heightened expression of ChP genes in AUTS2-deficient cerebral organoids. Our findings unveil a previously unknown function of AUTS2 in modulating neurodevelopment through negatively regulating the WNT/β-catenin signaling.
Results
Deletion of AUTS2 led to a minor growth defect in cerebral organoids
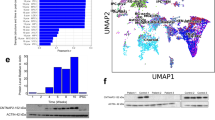
To understand the function of AUTS2 in human brain development and its contribution to NDDs, we generated a total AUTS2 knockout hESC line based on a patient-derived gene alteration (Fig. 1A and Fig. S1A). Through CRISPR/Cas9-mediated gene editing, we removed a 190 kb region containing the whole exon 6 of the AUTS2 gene and intronic promoters as evidenced by H3K4me3 enrichment in both hESC and HEK293 cells (Fig. 1A, top panel, showing the gene-editing strategy). To screen for successful knockouts, we performed genotyping PCR with two sets of primers. The first set detected a ~ 300 bp PCR product, absent in the wild-type (WT) samples but present in the successful knockouts. The second set resulted in a ~ 400 bp PCR fragment, specifically in the WT samples (Fig. S1B). Based on the genotyping PCR analysis, we identified both heterozygous (clone D3, AUTS2+/-) and homozygous AUTS2 knockout lines (clone B8, AUTS2-/-), which were further validated with Sanger sequencing and immunoblotting (Fig. 1A, bottom panel and Fig. 1B). Two main isoforms of AUTS2, the long isoform (AUTS2-L, above 150 kDa band) and short isoform (AUTS2-S, around 100 kDa band) have been either removed completely in AUTS2-/- cells or partially reduced in AUTS2+/- cells, compared with WT cells (Fig. 1B). To evaluate the role of AUTS2 in human brain development, we differentiated the WT, AUTS2+/-, and AUTS2-/- hESCs into cerebral organoids according to the protocol established by others previously18. Mature cerebral organoids were then collected at various time points for analyses (Fig. S1A). Throughout the differentiation process, we observed that AUTS2-/- cerebral organoids were slightly smaller, although WT and AUTS2+/- cerebral organoids appeared indistinguishable in size (Fig. 1C). To validate this observation further, we quantified the 21-day-old cerebral organoids from WT (n = 20), AUTS2+/- (n = 16), and AUTS2-/- (n = 22) samples under a microscope and measured the diameter of each cerebral organoid, including the fluid-filled compartments using ImageJ. Compared to WT, AUTS2+/- cerebral organoids exhibited no significant difference in size (101.1% +/- 4.0%). However, the size of AUTS2-/- cerebral organoids was significantly smaller, measuring only 76.8% +/- 7.6% of the WT (Fig. 1D), suggesting that the complete removal of AUTS2 resulted in a growth defect.
Generation and characterization of WT, AUTS2+/- and AUTS2-/- cerebral organoids. (A) The Integrative Genomics Viewer (IGV) snapshot of H3K4me3 ChIP-seq data from hESC and HEK293 cells shows two notable peaks in the region flanking exon 6 of AUTS2 (highlighted with a yellow box), suggesting the presence of intronic promoters. Red lines mark the sgRNA target region for creating AUTS2 knockout hESCs. The successful deletion of the targeted 190 kb region upstream of AUTS2 exon 6 is confirmed by Sanger sequencing results. (B) Immunoblot analysis confirmed the loss of AUTS2 expression in WT, AUTS2+/- , and AUTS2-/- hESC lines, with GAPDH serving as the loading control. For immunoblotting, at least three cerebral organoids for a total of three biological replicates were pooled (n = 3). (C) Microscopic examination reveals the morphology of 40-day-old cerebral organoids for each genotype. Ventricle-like structures in AUTS2-/- cerebral organoids are indicated with arrowheads. (D) The size of 21-day-old organoids was measured across genotypes: WT (n = 20), AUTS2+/- (n = 16), and AUTS2-/- (n = 22). Each bar in the graph represents the mean size, with error bars indicating the standard error. Student T-test was performed and statistical significance is marked with ** (P < 0.01).
Deletion of AUTS2 impaired neuronal differentiation in human cerebral organoids
To gain a deeper understanding of the role of AUTS2 in regulating gene expression in cerebral organoids, we performed RNA-seq analysis using 75-day-old organoids. Only a limited number of differentially expressed genes (18 upregulated genes and 43 downregulated genes) were identified between WT and AUTS2+/- cerebral organoids (Fig. 2A, Supplemental File 1). This observation indicates the deletion of one allele of AUTS2 has a minor effect on neuronal differentiation in organoid models. Meanwhile, comparing AUTS2+/- and AUTS2-/- cerebral organoids, 932 downregulated and 1045 upregulated genes were identified (Fig. 2B, Supplemental File 1). Compared with WT, 574 genes were upregulated, and 413 genes were downregulated in AUTS2-/- cerebral organoids (Fig. 2C, Supplemental File 1). Next, we conducted gene ontology (GO) analysis and revealed that downregulated genes in AUTS2-/- cerebral organoids were enriched in the functional terms related to neuronal development, such as “axonogenesis”, “peripheral nervous system development”, “dendrite development”, “dendritic spine development”, “synapse organization”, “synapse assembly”, and “positive regulation of neuron projection development” (Fig. 2D), further suggesting the essential role of AUTS2 in neurodevelopment.
Disruption of AUTS2 resulted in the downregulation of neuronal genes and the upregulation of ChP genes. (A-C). MA plots illustrate the RNA sequencing comparison between (A) WT and AUTS2+/-, (B) AUTS2+/- and AUTS2-/-, and (C) WT and AUTS2-/- 75-day-old cerebral organoids. For each genotype, three biologically replicated samples (n = 3) were processed for RNA-seq, each including 3–5 cerebral organoids. Differentially expressed genes (highlighted with blue dots) were identified using the DESeq2 R package with an adjusted p value threshold of less than 0.1. (D&E) Bar plots of the GO analysis for genes that are downregulated (D). and upregulated (E) in AUTS2-/- cerebral organoids. GO terms with an adjusted p value below 0.1 were selected and ordered by gene ratio and adjusted p values. The functional terms related to neurodevelopment are indicated by red arrows. (F) Heatmap displaying the expression of selected neuronal genes (FOXG1, POU3F2, BCL11B, RBFOX3, and MAP2) and ChP genes (TTR, CLIC6, FOLR1, and SLC4A5) based on RNA-seq data. Z-score normalization is applied for visualization.
An interesting observation is the formation of fluid-filled compartments in more than 50% of the AUTS2-/- cerebral organoids (Fig. 1C), a characteristic that has been previously reported in ChP organoids19. The ChP is a vital brain structure responsible for producing cerebrospinal fluid (CSF), a clear fluid that envelops the brain and spinal cord20,21,22. This observation led us to speculate that the deletion of AUTS2 resulted in a redirection of neuronal differentiation towards the ChP lineage. To test this hypothesis, we examined our RNA-seq results to investigate the expression profiles of neuronal and ChP genes and revealed a decrease in neuronal markers such as FOXG1, POU3F2, BCL11B, RBFOX3, and MAP2 and an increase in ChP markers, including TTR, CLIC6, FOLR1, and SLC4A5 in AUTS2-/- cerebral organoids (Fig. 2F).
AUTS2 is a key regulator of forebrain cell fate commitment
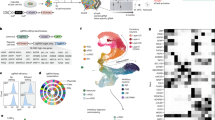
To understand how AUTS2 may regulate the cell fate commitment in cerebral organoids, we performed single-cell RNA sequencing (scRNA-seq) using Fluidigm C1TM Single-Cell Autoprep System (C1) on 75-day-old cerebral organoids. Since only a limited number of differentially expressed genes were identified between WT and AUTS2+/- cerebral organoids (Fig. 2A), we decided to compare the heterozygous and homozygous mutants. A total of 400 cells from AUTS2-/- cerebral organoids and 800 cells from AUTS2+/- cerebral organoids were sequenced. After filtering out cells with poor sequencing quality, 358 cells from AUTS2-/- cerebral organoids and 753 cells from AUTS2+/- cerebral organoids were used for further analysis. Based on the transcriptional profiles, cells from AUTS2+/- and AUTS2-/- cerebral organoids were clustered into 9 subpopulations using an unsupervised cluster algorithm23 (Fig. 3A and B). Subsequently, we identified the differentially expressed genes from each cluster and compared those with cell type-specific gene expression signatures published previously (Fig. 3C)24,25. We have successfully assigned identity to six different clusters (C0, C1, C2, C4, C6, and C8) (Fig. 3A and C). Forebrain markers, FOXG1, a master regulator for cortex development, was highly expressed in C1 cells (Fig. 3C and D). Several forebrains and neuronal markers, including TUBB3, NEUROD6, and POU3F3, also showed high expression in C1 (Fig. 3C). Building on this observation, we categorized C1 cells as forebrain neurons (Fig. 3A and C). The positive correlation between the expression of AUTS2 and neuron markers, as reflected in the high abundance of AUTS2 in the C1 cluster (Fig. 3D), further solidifies its role in neurodevelopment. C2 cells were assigned as the retinal cells, characterized by high expression of retinal cell marker genes, such as RCVRN, and NEUROD1(Fig. 3C and D). Both forebrain and proliferation markers were highly expressed in C4 cells, suggesting their identity as radial glial cells (Fig. 3C). A small number of cells were clustered in C8, exclusively expressing neuroepithelial cell marker genes (Fig. 3C). The cells from C6 cluster highly expressed TTR, CLIC6, and KRT18, markers for ChP (Fig. 3C and D). The transcription profiling of cells in C0 resembles C6 with a relatively sparse expression of these marker genes (Fig. 3C). Due to the similarity between these two clusters, we named C6 as ChP-like A and C0 as ChP-like B. The identity of C0 and C6 as ChP-like cells was further supported by calculating the enrichment of signature genes from each cluster with the list of cell identity markers from PanglaoDB, a public database that integrates published scRNA-seq data from different resources (Fig. S2A)25.
AUTS2 regulates cell fate commitment by promoting neuron differentiation and repressing ChP generation. (A) tSNE plot of 1111 cells from AUTS2+/- and AUTS2-/- 75-day-old cerebral organoids, colored by the annotated cell types (C0-C8). (B) tSNE plot of AUTS2+/- and AUTS2-/- 75-day-old cerebral organoids, colored by the genotypes. For each genotype, three biologically replicated samples (n = 3) were processed for scRNA-seq, each including 3 cerebral organoids. (C) Heatmap of selected marker genes for neurons, forebrain, proliferation, retina, neuroepithelial cells, and ChP epithelial cells. Clusters and genotypes are color-labeled on top of the heatmap. (D) Features plot of AUTS2, forebrain marker (FOXG1), retinal neurons (RCVRN), and ChP epithelial (TTR). (E) The violin plot indicates the upregulation of ChP markers (TTR, CLIC6, AQP1, HTR2C and ICFBP7) in AUTS2-/- cerebral organoids as compared to AUTS2+/- cerebral organoids. (F) The violin plot indicates the upregulation of ChP markers (TTR, CLIC6, AQP1, HTR2C and ICFBP7) in C0 and C6.
Noticeably, the clustering results showed distinct distribution of the cells from AUTS2+/- and AUTS2-/- cerebral organoids (Fig. 3B). In AUTS2+/- cerebral organoids, more than half of the cells were identified as neurons (C1) or retinal cells (C2), over 20% of cells were radial glial cells (C4) and neuroepithelial cells (C8), while a small number of cells were defined as ChP-like (C0 and C6) cells (Fig. 3C and S2B). In contrast, more than 85% of cells (308/356) from AUTS2-/- cerebral organoids were clustered in C0 (Fig. 3C). Therefore, AUTS2-/- cerebral organoids showed a biased cell composition with a low proportion of neuronal lineage (neurons, retinal cells, and radial glial cells) and an unexpectedly high proportion of ChP-like cells. This observation is consistent with the high expression of TTR, CLIC6, AQP1, HTR2C and IGFBP7 in AUTS2-/- populations as well as in C0 and C6 (Fig. 3E and F). Some of these ChP markers are shared by other lineages, including the kidney. We reason this may be why we observed the GO terms in the upregulated genes in AUTS2-/- cerebral organoids, such as “kidney development,” “connective tissue development,” “urogenital system development,” and “cartilage development” (Fig. 2E). To rule out this possibility, we analyzed marker genes for early kidney development26. As shown in Fig. S3, these markers do not show exclusive enrichment in C0 or C6 as those ChP markers. Therefore, these results strongly indicate that the AUTS2-/- cerebral organoids display a major cell population mimicking ChP cells. In summary, our results suggest that the complete deletion of AUTS2 resulted in skewed cell composition with a decreased diversity of cells committed to neuronal lineage and an overamplification of ChP-like cells.
AUTS2 regulates the transition of neural stem cells to progenitors
To better understand the impact of AUTS2 on neuronal lineage specification, we performed trajectory reconstruction analysis using our scRNA-seq data. Consistent with our clustering analysis, the pseudotime trajectory analysis clearly distinguished cells from AUTS2+/- and AUTS2-/- cerebral organoids (Fig. S4A and B). We selected neuroepithelial cells as the root for the trajectory reconstruction, simulating potential differentiation processes originating from these cells and branching into three distinct paths (Fig. S4C). The lower-left branch was enriched with cells identified as ChP-like A (C6) and ChP-like B (C0), showing a close association with neuroepithelial cells (C8) (Fig. S3C). The other two branches contained various neuron-related cells, which were predicted to emerge at later stages of differentiation based on the pseudotime analysis (Fig. S4B and C). Neurons (C1) and radial glial cells (C4) were found on the same trajectory, while retinal cells (C2) followed a separate path (Fig. S4C).
To determine the stage at which AUTS2 influences the differentiation process, we compared the proportions of various cell types in AUTS2+/- and AUTS2-/- cerebral organoids (Fig. S2B). The stacked percentage bar plot showed no significant difference in the proportions of neuroepithelial cells (C8) between the two genotypes. This suggests that the complete deletion of AUTS2 does not adversely affect the early stages of neurodevelopment, particularly the transition from embryonic stem cells to neuroepithelial cells. However, less than 5% of radial glial cells (C4), neurons (C1), and retinal cells (C2) were derived from AUTS2-/- cerebral organoids, indicating that AUTS2 is crucial for the transition of neuroepithelial cells to neuronal progenitors (Fig. S2B). The lack of neuronal lineage formation was accompanied by an overpopulation of ChP-like cells (C0) (Fig. S2B).
To further explore our hypothesis that AUTS2 regulates the transition from neuroepithelial cells to neuronal progenitors, we arranged the cells based on pseudotime and plotted the expression of selected marker genes along this timeline (Fig. S4D). We calculated the expression levels of various marker genes and AUTS2 along this trajectory. As expected, AUTS2 expression starts low at the early stage, increases during differentiation, and peaks at the end of the process (Fig. S4D). Interestingly, both MAP2 (neuronal marker) and NEUROD1 (retinal marker) exhibited expression patterns similar to AUTS2, while NES (neuronal progenitor marker) showed the highest expression in the middle stage of differentiation, coinciding with the initial increase in AUTS2 expression (Fig. S4D). Additionally, the expression of the ChP marker CLIC6 negatively correlated with AUTS2 (Fig. S4D). In summary, this pseudotime trajectory analysis strongly suggests that AUTS2 plays a crucial role in cell fate commitment beyond the neuroepithelial cell stage. Complete deletion of AUTS2 shifts neuroepithelial cells towards the ChP-like cell fate by blocking the formation of neuronal progenitors.
AUTS2 disruption leads to hyperactivation of the WNT/β-Catenin signaling
To further investigate how AUTS2 affects important signaling pathways that may be responsible for its effect on neurodevelopment, we performed ingenuity pathway analysis (IPA) using our RNA-seq data (https://digitalinsights.qiagen.com/IPA)27. In WT cerebral organoids, we observed the activation of multiple pathways closely associated with neuronal functionality and neurogenesis, such as "CREB signaling in neurons", "Neuropathic pain signaling in dorsal horn neurons", and "Synaptic long-term depression" (Fig. 4A). Notably, we observed the activation of the WNT/β-Catenin signaling in AUTS2-/- cerebral organoids, suggesting a repressive role of AUTS2 in this pathway. Upon further analysis of the bulk RNA-seq data, we identified differentially expressed genes involved in the WNT signaling pathway (Gene Ontology term GO:0,016,055) in WT, AUTS2+/- and AUTS2-/- cerebral organoids (Fig. S5, Supplemental File 1). As demonstrated in the heatmap (Fig. S5), we observed significant upregulation of the WNT components in AUTS2-/- cerebral organoids, including some well-studied WNT target genes such as CCND1 and BMP228,29.
Disruption of AUTS2 led to the overactivation of the WNT/β-Catenin signaling. (A) The volcano plot illustrates the results of IPA, depicting the z-score of log2 fold change in gene expression between WT and AUTS2-/- cerebral organoids against the log10 p value. Pathways overactivated in WT are highlighted in red, while those overactivated in AUTS2-/- cerebral organoids are shown in blue. (B) A bar plot illustrates the outcomes of a TOP-Flash/FOP-Flash luciferase reporter assay comparing WT and AUTS2-/- HEK293T cells. Each bar represents the mean of three independent measurements, with error bars indicating the standard error. One-way ANOVA was performed using GraphPad Prism to analyze the results and statistical significance is indicated as ** (p < 0.01). (C) An immunoblotting assay showing the levels of the AUTS2 protein in both WT and AUTS2-/- HEK293T cells, with and without CHIR-99021 treatment. (D) The immunoblotting experiment reveals the protein levels of AUTS2 and MAP2 in WT, AUTS2+/-, and AUTS2-/- cerebral organoids, with and without XAV939 treatment. Beta tubulin serves as the loading control. For immunoblotting, at least three cerebral organoids for a total of three biological replicates were pooled (n = 3). (E) Heatmap of gene expression for selected ChP, neuroepithelial, and neuron markers between AUTS2-/- 75-day-old cerebral organoids with or without XAV939 treatment. Selected neuronal marker genes and ChP epithelial marker genes are annotated. (F) The RT-qPCR results indicate the rescue in the expression levels of ChP marker genes (TTR, FOLR1, and SLC4A5) and neuronal markers (FOXG1, CTIP2, and BRN2) by XAV939 in AUTS2-/- cerebral organoids. Student T-test was performed, with statistical significance indicated as * (p < 0.05) and ** (p < 0.01). For RT-qPCR and RNA-seq analyses, three biological replicates were used (n = 3), each including 3–5 organoids.
The WNT/β-Catenin signaling influences the brain's development, spanning from early patterning to adulthood. During neurogenesis, activation of this pathway leads to an increased neural precursor population30. Conversely, inhibiting the WNT/β-Catenin signaling prompts premature cell cycle exit among precursor cells, promoting their differentiation into neurons31. Moreover, disruption of the WNT/β-Catenin signaling inhibits the fate of ChP epithelial cells32. To further test the impact of AUTS2 on the WNT/β-Catenin signaling, we generated a HEK293T cell line lacking AUTS2 using the CRISPR/Cas9 gene editing (Fig. S6D), and the removal of the AUTS2 gene was confirmed by genotyping (Fig. S6B). Following this, we employed the TOP-Flash/FOP-Flash luciferase reporter system on WT and two independent AUTS2-/- clones to evaluate the influence of AUTS2 on the WNT/β-Catenin signaling (Fig. 4B). In the absence of CHIR-99021, a GSK-3β inhibitor commonly used to activate the WNT/β-Catenin signaling33,34, luciferase activity remained low. Upon the addition of CHIR-99021, luciferase activity increased, indicating WNT pathway activation. Immunoblotting showed that CHIR-99021 did not affect AUTS2 expression (Fig. 4C). Compared to WT cells, AUTS2-/- cells exhibited significantly higher luciferase activity, indicating overactivation of the WNT/β-Catenin signaling (Fig. 4B). These findings suggest that AUTS2 plays an inhibitory role in the WNT/β-Catenin signaling and its disruption may lead to the observed neuronal differentiation defects in AUTS2-/- cerebral organoids.
Inhibition of the WNT/β-Catenin signaling rescued the neuronal differentiation defect caused by AUTS2 depletion
To determine if the differentiation abnormalities observed in AUTS2-/- cerebral organoids were indeed caused by the hyperactivation of the WNT/β-Catenin signaling, we conducted a rescue experiment by treating 75-day-old AUTS2-/- cerebral organoids with XAV939, a widely used WNT inhibitor35. Our immunoblotting showed that the total protein level of MAP2, a neuronal marker, was decreased in AUTS2-/- cerebral organoids compared to WT and AUTS2+/- cerebral organoids (Fig. 4D), further supporting our transcriptome analysis indicating the downregulation of neuronal genes. When we treated AUTS2-/- cerebral organoids with XAV939, the MAP2 level was increased toward WT level, providing evidence that the observed neuronal differentiation defects were caused by hyperactivation of the WNT/β-Catenin signaling. As demonstrated in our RNA-seq results, AUTS2 deletion led to an increase in ChP gene expression in addition to the downregulation of neuronal genes (Fig. 2F). To confirm whether the deregulation of neuronal and ChP gene expression is due to increased WNT activity, we collected 75-day-old AUTS2-/- cerebral organoids treated with or without XAV939 and subjected them to RNA-seq analysis. Compared to untreated AUTS2-/- cerebral organoids, the expression of downregulated neuronal genes was increased, while the elevated ChP gene expression was inhibited in the presence of XAV939 (Fig. 4E). This rescue effect was further confirmed by RT-qPCR in 75-day-old AUTS2-/- cerebral organoids with or without XAV939 treatment (Fig. 4F). As expected, we found that expressions of forebrain neuronal marker genes, including FOXG1, CTIP2, and BRN2, were increased, and at the same time, ChP marker genes, such as TTR, FOLR1, and SLC4A5, were reduced (Fig. 4F). Overall, our studies uncover a novel mechanism through which AUTS2, a risk factor for NDD, plays a critical role in regulating cell fate choice between neuronal and ChP lineage via the WNT/β-Catenin signaling.
Discussion
The application of cell lines and animal models in biomedical research has been successful across various fields. Nonetheless, recent research has shown that certain biological processes unique to the human body, such as brain development, cannot be precisely replicated in other animals36. The use of human cerebral organoids—mini-brain structures grown in a lab—has become a valuable tool to tackle the intrinsic complexity of human brain development. These organoids closely mimic the cell composition and functionality of neurons in the developing human brain18,37. In our study, we used this advanced technique and found that the absence of AUTS2 severely disrupts neuronal development. Specifically, we noticed a reduction in genes related to neuronal lineage specification as well as the increased expression of ChP genes, suggesting a critical role of AUTS2 in regulating cell fate choice by promoting the development of neurons and preventing the development of ChP epithelial cells from the early stage of brain development. This hypothesis is further supported by scRNA-seq analysis, which revealed a shift in cell fate commitment from neuronal cell types to ChP-like cell types upon AUTS2 deletion. The overproduction of ChP-like cells resulting from AUTS2 deletion is novel, as it has not been previously reported in cell lines or animal models. Conversely, the reduction in the neuronal cell population is consistent with previous findings, where a significant decrease in neuronal cells was observed in auts2 morphant zebrafish and hypoplasia in both the dentate gyrus and the cerebellum in mice lacking AUTS217,38,39,40,41.
Previous research has noted that patients with ASD or those at high risk often have increased CSF42,43,44,45. CSF, produced by the ChP epithelial cells, is vital for delivering key growth factors and signaling molecules, and an imbalance in CSF production can significantly impact brain development46,47. Our research suggests that eliminating AUTS2 in cerebral organoids reduces their size and diminishes neuronal gene expression. Concurrently, AUTS2-/- cerebral organoids form vesicle-like structures similar to ChP organoids, with elevated ChP gene expression. A recent study using cerebral organoids derived from a patient with an AUTS2 missense mutation in exon 9 displayed similar defects to those we observed, such as reduced growth and a decrease in neural progenitor cell (NPC) proliferation48. However, unlike our findings, they did not report an increase in ChP gene expression. This discrepancy could be due to the different pathological alleles triggering distinct signaling pathways, leading to varying phenotypes. Although this AUTS2 mutation leads to the loss of interaction with P300, potentially impairing its role in activating neuronal gene transcription, it still preserves most of the AUTS2 protein structure. Therefore, we cannot rule out the possibility that this variant might acquire additional functions compared to its original form. Conversely, our model, involving the complete deletion of both AUTS2 isoforms, will likely have a more pronounced impact on AUTS2 normal functions. Future research is essential to understand how different pathological phenotypes observed in various patients correlate with distinct AUTS2 alleles. Nonetheless, our finding connects a key risk gene to the brain anomalies and dysfunctions often seen in neurodevelopmental disorders. Further investigation into this area promises to yield significant insights regarding the role of CSF in these disorders.
Mechanistically, our study revealed that AUTS2 is instrumental in regulating the balance between neuronal genes and ChP genes during brain development by modulating the WNT/β-Catenin signaling. The significance of WNT/β-Catenin signaling in the temporal and spatial precision of lineage specification and cell fate determination in brain development has been well-established49,50,51. Particularly, the activation of the WNT/β-Catenin signaling is essential for ChP morphogenesis19,32. Recent findings suggest that combining a WNT agonist with BMP4 can promote ChP structure formation and CSF production in brain organoids19. In our present study, neutralizing WNT overactivation in AUTS2-/- cerebral organoids led to the inhibition of ChP genes and reinstate neuronal genes to levels approaching those of WT, confirming that the skewed neuronal differentiation observed after AUTS2 deletion is mediated through the WNT/β-Catenin signaling. Notably, the recovery of forebrain neuron-specific genes in AUTS2-/- cerebral organoids upon WNT inhibition was only partial, indicating the involvement of additional mechanisms in AUTS2's regulation of this complex cell fate determination process. One such mechanism is BMP signaling, previously shown to mediate the action of AUTS2 in the differentiation of mouse embryonic stem cells toward neuronal progenitors11. Future research will be key in understanding how WNT, BMP, and other mechanisms collaboratively regulate cell fate determination in various lineages during human brain development using organoid models.
While organoids are a powerful model system, there is variability between different batches of organoids due to differences in starting materials, such as human embryonic stem cells or induced pluripotent stem cells (iPSCs), as well as culture conditions and handling procedures. Another study that utilized cerebral organoid culture to study AUTS2 function was generated from human iPSCs, whereas in this study, we generated the cerebral organoids from hESCs48. Although the cerebral organoid generation methods are similar, variability might arise due to the different starting materials. Even within the same batch, individual organoids can exhibit considerable heterogeneity. This intra-batch variability can arise from slight differences in the microenvironment, nutrient distribution, and local signaling cues during organoid formation and growth36,52,53. For our scRNA-seq analysis, multiple cerebral organoids from the same batch were combined for dissociation into single cells and hence, our scRNA-seq data represents an average of multiple cerebral organoids to minimize variability.
Our transcriptomic analyses revealed an increase in ChP-like cells in AUTS2-/- cerebral organoids, accompanied by the upregulation of ChP genes. Many of the ChP markers we used, such as TTR, CLIC6, FOLR1, and SLC4A5, are also expressed in other tissues because they are involved in functions that are not unique to a single tissue or organ. For example, TTR is expressed in the eyes, pancreas, and various other tissues, where it acts as a transporter54. Similarly, CLIC6, part of the chloride channel family, plays a crucial role in ion transport and cellular homeostasis55. ChP epithelial cells are characterized by features such as polarity, tight junctions, microvilli, and unique protein expression and localization, which mature over developmental time56. Comprehensive analyses of the AUTS2-/- cerebral organoids, incorporating histological staining and structural analysis via electron microscopy, would significantly enhance our study. These methods would enable us to elucidate the developmental trajectory and pinpoint the stages at which differences arise, providing valuable insights into the development of ChP cells in the absence of AUTS2.
Materials and methods
Cell culture and transfection
All hESC lines were cultured in StemFlex basal medium (Gibco, Cat# A3349401) supplemented with non-essential amino acids and sodium pyruvate on Matrigel (Gibco, Cat# A14133)-coated plates. HEK293T cells were cultured in a standard DMEM medium supplemented with 10% FBS (Atlanta Biologicals, Cat# S11050), L-glutamine, and penicillin–streptomycin. Plasmids were transfected into HEK293T cells using Lipofectamine 2000 (Invitrogen, Cat# 11668027).
CRISPR/Cas9-mediated gene editing
The deletion of AUTS2 in H9 cells was generated as previously described57. Two small guide RNAs, sgRNA1 (5’-GTGCACTGAGAATGTTAACC-3’) and sgRNA2 (5’-GTGGCTTCTAGAGTCAAGGC-3’) were designed and synthesized by GenScript to target the 190 kb genomic region upstream of AUTS2 exon 6 (chr7: 70,545,519–70,735,445, hg38). These sgRNAs were mixed with recombinant Cas9 to form a ribonucleoprotein complex and then introduced into H9 cells via nucleofection using the Human Stem Cell Nucleofector Kit (Lonza, Cat# VPH-5002). Two days post-nucleofection, the cells were dissociated using Accutase (Gibco, Cat# A11105) and sorted at a density of one cell per well in a 96-well plate to facilitate expansion. Genomic DNA from the resulting individual clones was isolated, and the edited region was screened via PCR and confirmed with Sanger sequencing. The primers used for this process were as follows: geno-A2S-p-F (forward, outside): 5’-GCATGCTGTAAGGACAGAGG-3’, geno-A2S-p-R1’ (reverse, outside): 5’-GCATTCTTCGGAACATCCGC-3’, geno-A2S-p-R2 (reverse, inside): 5’-CCTTCCAGAAGAAAAGGCTTG-3’.
For CRISPR editing in HEK293T cells, sgRNAs were designed to target exon 7 and its surrounding region. Oligonucleotides corresponding to the candidate sgRNA sequences were obtained from IDT and cloned into the pX458 vector using methods previously described58. The resulting plasmids were then transfected into the cells using Lipofectamine 2000, following the manufacturer’s instructions. The sgRNA sequences used in this study are as follows: 5’-AGAGCAAGACATCTTGCGAC-3’; 5’-GGGGCCACCAGAGATCGAAT-3’. After two days of incubation, the transfected cells were sorted for GFP fluorescence and seeded into a 96-well plate. Single-cell clones were isolated and genotyped using PCR analysis. Clones that were positively identified were further confirmed through immunoblotting.
Cerebral organoid culture condition
The Cerebral Organoid kit from Stem Cell Technologies (Cat# 08570) was utilized, following the manufacturer’s instructions for generating cerebral organoids. Briefly, on day 0, embryoid bodies (EBs) were formed using 9,000 H9 cells (WT, AUTS2+/- and AUTS2-/-) per well in 96-well U-bottom suspension plates, using EB Formation media supplemented with 5 mM Y-27632 ROCK inhibitor (Stem Cell Technologies, Cat# 100–1044). Fresh EB Formation media was added bi-daily. On day 5, the EBs were transferred to a 24-well ultra-low attachment plate containing Induction media. After two days, the EBs were embedded in 15 µL of Matrigel and then cultured in a 6-well ultra-low adherent plate with Expansion medium. On day 10, the EBs were replaced with Maturation media, moved to an incubator set at 37 °C, and placed on an orbital shaker. The cerebral organoids were subsequently fed every 3–4 days with Maturation media. For the WNT inhibitor treatment, cerebral organoids were cultured in Expansion media supplemented with 10 mM XAV939 from day 7 to day 10 (Stem Cell Technologies, Cat# 72672).
Immunoblotting
The protein samples were obtained by resuspending the cell pellets in lysis buffer consisting 1/3 volume of buffer A (10 mM Tris–HCl, pH 7.9, 1.5 mM MgCl2, 10 mM KCl, 0.5 mM DTT, 0.2 mM PMSF, 1 µg/ml Pepstatin A, 1 µg/ml Leupeptin, 1 µg/ml Aprotinin), 2/3 volume of buffer C (20 mM Tris–HCl, pH 7.9, 25% glycerol, 420 mM NaCl, 1.5 mM MgCl2, 0.2 mM EDTA, 0.5 mM DTT, 0.2 mM PMSF, 1 µg/ml Pepstatin A, 1 µg/ml Leupeptin, 1 µg/ml Aprotinin) and 10% Triton X-100. The protein samples were resolved by SDS-PAGE, transferred to nitrocellulose, and probed with antibodies as follows: anti-AUTS2 (in-house), anti-MAP2 (Abcam, Cat# ab5622), anti-OCT3/4 (Santa Cruz Biotechnology, Cat# sc-365509), anti-GAPDH (Invitrogen, Cat# MA5-15738) and anti-β-tubulin (Abcam, Cat# ab6046).
Quantitative RT-PCR
Total RNA was extracted using TriPure reagent (Roche, Cat# 11667165001) followed by cDNA synthesis with the SuperScript III system (Invitrogen, Cat# 18080–044). The synthesized cDNA was combined with Brilliant III Ultra-Fast SYBR QPCR master mix (Agilent, Cat# 600883) and Random Hexamer Primer (Thermo Scientific, Cat# SO142). This mixture was subsequently processed in a Bio-Rad CFX Connect real-time PCR detection system for quantitative PCR analysis. The sequences of primers used were listed in Table S1.
RNA-seq and analysis
After RNA extraction, cDNA libraries were prepared at the Genome Sciences Facility of PSU College of Medicine. The libraries were then loaded onto a TruSeq Rapid flow cell and sequenced on an Illumina HiSeq 2500 system, housed at the Genomic Sciences Facility at Penn State University College of Medicine. The sequencing was performed over 50 cycles using a single-end recipe as specified by the manufacturer. The resulting raw Fastq files were mapped to the reference human genome (GRCh38/hg38). The number of reads were as follows: WT replicates (n = 3)—24,576,263, 17,855,110, and 16,661,181; AUTS2+/- replicates (n = 3)—18,001,301, 19,045,923, and 24,470,584; AUTS2-/- replicates (n = 3)—23,666,812, 18,828,165, and 6,958,366. Gene abundances were estimated using Kallisto59 for raw read counts and relative gene expression (TPM). The gene expression matrix was imported into the R programming environment, and differentially expressed genes were identified using the DESeq2 R package60, applying an adjusted p value threshold of less than 0.1. The list of differentially expressed genes (DEGs) was then curated, and Gene Ontology (GO) analysis was carried out using the clusterProfiler R package61,62. GO terms with an adjusted p value below 0.1 were selected and ordered by gene ratio and adjusted p values.
scRNA-seq and analysis
Cerebral organoids were harvested on day 75 for single-cell RNA sequencing (scRNA-seq). The cerebral organoids were then treated with Papain (Worthington, Cat# LK003178) at 37 °C for 30 min to dissociate them into single cells. These cells were loaded onto a Fluidigm C1 Single-Cell Autoprep System (C1) for automated capture and subsequent sequencing-ready library preparation. The DNA libraries were sequenced on an Illumina NovaSeq 6000 system, with the resulting data undergoing demultiplexing and readout trimming. For aligning the sequence data to the reference genome (GRCh38/hg38), Bowtie63 was employed, while HTseq64 was used to compile the gene abundance matrix for each cell. The raw count matrix was then processed in R using the Seurat R package65. Initially, genes with low or no expression across all cells were filtered out, along with cells exhibiting poor sequencing quality. After normalizing the gene count matrix, high-variance genes were identified. Principal Component Analysis (PCA) was conducted based on these genes, with the JackStraw66 plot assisting in determining the appropriate number of dimensions. Cell clustering was performed using dimensions 1 through 12 at a resolution of 0.3, and clusters were visualized via tSNE67. The FindAllMarkers function (with default parameters) was used to identify marker genes for each cluster. Cell identities were assigned based on the expression of exclusive high-expression genes within each cluster, drawing on marker information from the literature and public databases. Trajectory analysis was performed using the R package monocle368. First, the gene raw count matrix was imported to monocle3 and filtered out the low expression gene and cells with low sequencing quality, using the same criteria as Seurat. Dimension reduction and visualization were performed using the program's default method, UMAP. Crossing the clustering results and cell identity we acquired from Seurat, we set neuroepithelial cells as the root for pseudotime analysis.
Ingenuity pathway analysis
Pathway analysis was performed using QIAGEN IPA (https://digitalinsights.qiagen.com/IPA)27. The results of DEG analysis, containing gene name, fold change, pvalue and adjusted p value were imported into IPA. The pathway analysis was conducted using 39 default parameters. The results of canonical pathways were exported for downstream analysis and plotting.
Luciferase reporter assay
To assess the impact of AUTS2 on the WNT activity, M50 Super 8 × TOP-Flash and pRL-SV40 Renilla luciferase (Promega, Cat# E2231) were co-transfected into WT and AUTS2-/- HEK293T cells using Lipofectamine 2000. A mutant TOP-Flash reporter, M51 Super 8 × FOP-Flash, served as a negative control. Twenty-four hours post-transfection, the cells were treated with 10 µM CHIR-99021 (Cayman Chemical, Cat# 13122). After 18–20 h of treatment, the cells were harvested and analyzed using the Dual-Luciferase Reporter Assay System (Promega, Cat# E1910), following the manufacturer's instructions.
Statistical analyses
For RT-qPCR and RNA-seq analyses, three biological replicates were used, each including 3–5 organoids. For scRNA-seq, we combined three biological replicates, each containing 3 organoids. For immunoblotting, at least three cerebral organoids for a total of three biological replicates were pooled. Student T-test was applied to identify the statistical significance of organoids size and qRT-PCT. For RNA-seq analysis, DEGs were identified using the generalized linear models from DESeq2 R package60. Seurat R package was applied for dimension reduction, clustering, and differential gene analysis for scRNA-seq23. For the luciferase assay, biological triplicates were used for each strain, and one-way ANOVA was performed using GraphPad Prism to analyze the results.
Data availability
All data generated or analyzed during this study are included in this published article. Bulky RNA-seq data that supports the findings of this study have been deposited in the Gene Expression Omnibus (GEO) under accession code GSE255185. All other data supporting the findings of this study are available from the corresponding author upon request.
Reference:s
Hori, K., Shimaoka, K. & Hoshino, M. AUTS2 gene: keys to understanding the pathogenesis of neurodevelopmental disorders. Cells 11(1), 11 (2021).
Biel, A. et al. AUTS2 syndrome: molecular mechanisms and model systems. Front. Mol. Neurosci. 15, 858582 (2022).
Beunders, G. et al. A detailed clinical analysis of 13 patients with AUTS2 syndrome further delineates the phenotypic spectrum and underscores the behavioural phenotype. J. Med. Genet. 53(8), 523–532 (2016).
Talkowski, M. E. et al. Sequencing chromosomal abnormalities reveals neurodevelopmental loci that confer risk across diagnostic boundaries. Cell 149(3), 525–537 (2012).
Oksenberg, N. & Ahituv, N. The role of AUTS2 in neurodevelopment and human evolution. Trends Genet. 29(10), 600–608 (2013).
Jha, U., Kondrychyn, I., Korzh, V., & Thirumalai, V. (2021). High behavioral variability mediated by altered neuronal excitability in auts2 mutant zebrafish. ENeuro, 8(5).
Gao, Z. et al. An AUTS2-Polycomb complex activates gene expression in the CNS. Nature 516(7531), 349–354 (2014).
Hori, K. et al. AUTS2 regulation of synapses for proper synaptic inputs and social communication. Science 23(6), 101183 (2020).
Hori, K. et al. Cytoskeletal regulation by AUTS2 in neuronal migration and neuritogenesis. Cell Rep. 9(6), 2166–2179 (2014).
Hori, K. et al. Heterozygous disruption of Autism susceptibility candidate 2 causes impaired emotional control and cognitive memory. PLoS ONE 10(12), e0145979 (2015).
Geng, Z. et al. AUTS2 controls neuronal lineage choice through a novel PRC1-independent complex and BMP inhibition. Stem Cell Rev. Rep. 19(2), 531–549 (2023).
Monderer-Rothkoff, G. et al. AUTS2 isoforms control neuronal differentiation. Mol. Psychiatr 26(2), 666–681 (2019).
Russo, D. et al. Glycosphingolipid metabolic reprogramming drives neural differentiation. The EMBO J. 37(7), e97674 (2018).
Gao, Z. et al. PCGF homologs, CBX proteins, and RYBP define functionally distinct PRC1 family complexes. Mol. Cell 45(3), 344–356 (2012).
Wang, Q. et al. WDR68 is essential for the transcriptional activation of the PRC1-AUTS2 complex and neuronal differentiation of mouse embryonic stem cells. Stem Cell Res. 33, 206–214 (2018).
Liu, S. et al. NRF1 association with AUTS2-Polycomb mediates specific gene activation in the brain. Mol. Cell 81(22), 4663-4676.e8 (2021).
Castanza, A. S. et al. AUTS2 regulates RNA metabolism and dentate gyrus development in mice. Cerebral Cortex 31(10), 4808–4824 (2021).
Lancaster, M. A. et al. Cerebral organoids model human brain development and microcephaly. Nature 501(7467), 373–379 (2013).
Pellegrini, L. et al. Human CNS barrier-forming organoids with cerebrospinal fluid production. Science 369(6500), eaaz5626 (2020).
Liddelow, S. A. (2015). Development of the choroid plexus and blood-CSF barrier. In Frontiers in Neuroscience (Vol. 9, Issue MAR, p. 32). Frontiers Media S.A.
Fame, R. M. & Lehtinen, M. K. Emergence and developmental roles of the cerebrospinal fluid system. Dev. Cell 52(3), 261–275 (2020).
Saunders, N. R., Dziegielewska, K. M., Fame, R. M., Lehtinen, M. K. & Liddelow, S. A. The choroid plexus: a missing link in our understanding of brain development and function. Physiol. Rev. 103(1), 919 (2023).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177(7), 1888-1902.e21 (2019).
Quadrato, G. et al. Cell diversity and network dynamics in photosensitive human brain organoids. Nature 545(7652), 48–53 (2017).
Franzén, O., Gan, L. M. & Björkegren, J. L. M. PanglaoDB: A web server for exploration of mouse and human single-cell RNA sequencing data. Database J. Biol. Databases Curat. 2019(1), 46 (2019).
Brunskill, E. W. et al. Single cell dissection of early kidney development: Multilineage priming. Dev. (Cambridge, England) 141(15), 3093–3101 (2014).
Krämer, A., Green, J., Pollard, J. & Tugendreich, S. Causal analysis approaches in ingenuity pathway analysis. Bioinformatics 30(4), 523–530 (2014).
Shtutman, M. et al. The cyclin D1 gene is a target of the β-catenin/LEF-1 pathway. Proc. Natl. Acad. Sci. United States Am. 96(10), 5522 (1999).
Hill, T. P., Taketo, M. M., Birchmeier, W. & Hartmann, C. Multiple roles of mesenchymal β-catenin during murine limb patterning. Development 133(7), 1219–1229 (2006).
Chenn, A. & Walsh, C. A. Regulation of cerebral cortical size by control of cell cycle exit in neural precursors. Science 297(5580), 365–369 (2002).
Woodhead, G. J., Mutch, C. A., Olson, E. C. & Chenn, A. Cell-autonomous β-Catenin signaling regulates cortical precursor proliferation. J. Neurosci. 26(48), 12620–12630 (2006).
Parichha, A. et al. Constitutive activation of canonical Wnt signaling disrupts choroid plexus epithelial fate. Nat. Commun. 13(1), 1–20 (2022).
Naujok, O., Lentes, J., Diekmann, U., Davenport, C. & Lenzen, S. Cytotoxicity and activation of the Wnt/beta-catenin pathway in mouse embryonic stem cells treated with four GSK3 inhibitors. BMC Res. Notes 7(1), 273 (2014).
Law, S. M. & Zheng, J. J. Premise and peril of Wnt signaling activation through GSK-3β inhibition. IScience 25(4), 104159 (2022).
Huang, S. M. A. et al. Tankyrase inhibition stabilizes axin and antagonizes Wnt signalling. Nature 461(7264), 614–620 (2009).
Kim, J., Koo, B. K. & Knoblich, J. A. Human organoids: Model systems for human biology and medicine. Nat. Rev. Mol. Cell Biol. 21(10), 571–584 (2020).
Eichmüller, O. L. & Knoblich, J. A. Human cerebral organoids—A new tool for clinical neurology research. Nat. Rev. Neurol. 18(11), 661–680 (2022).
Oksenberg, N., Stevison, L., Wall, J. D. & Ahituv, N. Function and regulation of AUTS2, a gene implicated in autism and human evolution. PLoS Genet. 9(1), e1003221 (2013).
Beunders, G. et al. Exonic deletions in AUTS2 cause a syndromic form of intellectual disability and suggest a critical role for the C terminus. Am. J. Human Genet. 92(2), 210–220 (2013).
Yamashiro, K. et al. AUTS2 governs cerebellar development, purkinje cell maturation, motor function and social communication. IScience 23(12), 101820 (2020).
Weisner, P. A. et al. A mouse mutation that dysregulates neighboring Galnt17 and Auts2 genes is associated with phenotypes related to the human AUTS2 syndrome. G3 Bethesda Md 9(11), 3891–3906 (2019).
Runge, K. et al. Cerebrospinal fluid findings of 36 adult patients with autism spectrum disorder. Brain Sci. 10(6), 1–18 (2020).
Shen, M. D. et al. Extra-axial cerebrospinal fluid in high-risk and normal-risk children with autism aged 2–4 years: A case-control study. Lancet. Psychiatr. 5(11), 895 (2018).
McAlonan, G. M. et al. Mapping the brain in autism. A voxel-based MRI study of volumetric differences and intercorrelations in autism. Brain 128(2), 268–276 (2005).
Hallahan, B. et al. Brain morphometry volume in autistic spectrum disorder: A magnetic resonance imaging study of adults. Psychol. Med. 39(2), 337–346 (2009).
Mashayekhi, F. et al. Deficient cortical development in the hydrocephalic Texas (H-Tx) rat: a role for CSF. Brain 125(8), 1859–1874 (2002).
Johanson, C. E. et al. Multiplicity of cerebrospinal fluid functions: New challenges in health and disease. Cerebrospinal Fluid Res. 5(1), 1–32 (2008).
Fair, S. R. et al. Cerebral organoids containing an AUTS2 missense variant model microcephaly. Brain. 146(1), 387–404 (2022).
Brafman, D. & Willert, K. Wnt/β-catenin signaling during early vertebrate neural development. Dev. Neurobiol. 77(11), 1239 (2017).
Kim, J. Y. et al. Wnt signal activation induces midbrain specification through direct binding of the beta-catenin/TCF4 complex to the EN1 promoter in human pluripotent stem cells. Exp. Mol. Med. 50(4), 1–13 (2018).
Harrison-Uy, S. J. & Pleasure, S. J. Wnt signaling and forebrain development. Cold Spring Harbor Perspect. Biol. 4(7), 1–11 (2012).
Andrews, M. G. & Kriegstein, A. R. Challenges of organoid research. Ann. Rev. Neurosci. 45, 23–39 (2022).
Hofer, M. & Lutolf, M. P. Engineering organoids. Nat. Rev. Mater. 6(5), 402–420 (2021).
Buxbaum, J. N. & Reixach, N. Transthyretin: The servant of many masters. Cell. Mol. Life Sci. 66(19), 3095–3101 (2009).
Loyo-Celis, V. et al. Biophysical characterization of chloride intracellular channel 6 (CLIC6). J. Biol. Chem. 299(11), 105349 (2023).
Lun, M. P., Monuki, E. S. & Lehtinen, M. K. Development and functions of the choroid plexus–cerebrospinal fluid system. Nat. Rev. Neurosci. 16(8), 445–457 (2015).
Bak, R. O., Dever, D. P. & Porteus, M. H. CRISPR/Cas9 genome editing in human hematopoietic stem cells. Nat. Protoc. 13(2), 358 (2018).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339(6121), 819–823 (2013).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34(5), 525–527 (2016).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15(12), 1–21 (2014).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS : J. Integrat. Biol. 16(5), 284 (2012).
Wu, T. et al. clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innovation 2(3), 100141 (2021).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25(14), 1754–1760 (2009).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31(2), 166 (2015).
Velmeshev, D. et al. Single-cell genomics identifies cell type-specific molecular changes in autism. Sci. (New York, N.Y.) 364(6441), 685–689 (2019).
Chung, N. C. & Storey, J. D. Statistical significance of variables driving systematic variation in high-dimensional data. Bioinformatics 31(4), 545 (2015).
van der Maaten, L. & Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 9(86), 2579–2605 (2008).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32(4), 381–386 (2014).
Acknowledgements
We would like to extend our appreciation to Dr. Sirisha Pochareddy, Dr. Yuka Imamura, and the Genome Sciences Facility (RRID:SCR_021123) for their valuable assistance in RNA-seq and scRNA-seq analysis. We are also grateful to the Flow Cytometry Core (RRID:SCR_021134) at the College of Medicine, Penn State Health, for their support. Our thanks also go to Dr. Greg Yochum and his lab for their help with the TOP-Flash/FOP-Flash reporter assay. This work was supported by the following grants to Z. Gao: R35GM133496 from NIGMS and the Brad Hollinger Autism Research Endowment. This project is funded, in part, under a grant from the Pennsylvania Department of Health using Tobacco CURE Funds. The Department specifically disclaims responsibility for any analyses, interpretations, or conclusions.
Author information
Authors and Affiliations
Contributions
Z.Geng, Y.T., and Q.W. designed and performed the experiments, analyzed the data, and wrote the manuscript. Z.Geng performed the bioinformatics analysis. Z.Gao supervised the project and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Geng, Z., Tai, Y.T., Wang, Q. et al. AUTS2 disruption causes neuronal differentiation defects in human cerebral organoids through hyperactivation of the WNT/β-catenin pathway. Sci Rep 14, 19522 (2024). https://doi.org/10.1038/s41598-024-69912-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-69912-4