Abstract
Cyanobacteria possess special defense mechanisms to protect themselves against ultraviolet (UV) radiation. This study combines experimental and computational methods to identify the role of protective strategies in Nostoc species against UV-C radiation. To achieve this goal, various species of the genus Nostoc from diverse natural habitats in Iran were exposed to artificial UV-C radiation. The results indicated that UV-C treatment significantly reduced the photosynthetic pigments while simultaneously increasing the activity of antioxidant enzymes. Notably, N. sphaericum ISB97 and Nostoc sp. ISB99, the brown Nostoc species isolated from habitats with high solar radiations, exhibited greater resistance compared to the green-colored species. Additionally, an increase in scytonemin content occurred with a high expression of key genes associated with its synthesis (scyF and scyD) during the later stages of UV-C exposure in these species. The molecular docking of scytonemin with lipopolysaccharides of the cyanobacteria that mainly cover the extracellular matrix revealed the top/side positioning of scytonemin on the glycans of these lipopolysaccharides to form a UV-protective shield. These findings pave the way for exploring the molecular effects of scytonemin in forming the UV protection shield in cyanobacteria, an aspect that has been ambiguous until now.
Similar content being viewed by others
Introduction
Cyanobacteria, the most primitive group of gram-negative photosynthetic prokaryotes, presumably appeared on Earth during the Precambrian Era1. Nostoc species (Nostocales), filamentous nitrogen-fixing cyanobacteria, colonize diverse habitats and can be found almost in every ecosystem, even under extreme conditions2,3. These microorganisms harvest solar light for photosynthesis and are exposed to high doses of solar ultraviolet radiation (UVR). It seems that during the Precambrian era, UV-C radiation was an environmental stress factor due to the lack of the ozone layer. Therefore, Nostoc ancestors may have evolved various strategies to survive in environments with high levels of solar radiation4.
In cyanobacteria, higher doses of all forms of solar ultraviolet radiation (UV-A, -B, and -C) cause an increase in production of reactive oxygen species (ROS) and damage biomolecules such as proteins, lipids, DNA, and RNA. Furthermore, these radiations disturb the activity of the photosynthetic reaction center (PSII) and photo-bleaching of pigments (chlorophylls, carotenoids, and phycobiliproteins)5.
Synthesizing UV-absorbing pigments such as mycosporine-like amino acids (MAAs) and scytonemin, as well as producing antioxidant enzymes, are some of Nostoc species protective strategies to reduce damages induced by UVR6,7,8.
The cyanobacterial cell wall shows a large overall resemblance to gram-negative bacteria, with Lipopolysaccharides (LPS) covering the 3/4th of the total cell surface area of the outer membrane layer. Nostoc species have LPS in their membranes and peptidoglycans in their cell walls, which are common Microbe-Associated Molecular Patterns (MAMPs). LPS plays a crucial role in shaping and maintaining the functionality of the membrane, serving as the first line of defense system necessary for their survival9,10,11.
LPS is a glycolipid composed of three parts: lipid A, the core oligosaccharide, and the O antigen. Lipid A, the hydrophobic portion of the molecule, is linked to a glucosamine disaccharide that forms the leaflet of the outer membrane (OM). The core oligosaccharide is a structure that binds with the glucosamines in lipid A. Typically, this core composition consists of 3-deoxy-D-manno-oct-2-ulosonic acid (KDO) units, heptoses, and various hexoses. These elements can be altered through processes that include phosphates and additional substances such as phosphoethanolamine. Although variations may occur at the species and strain levels, the general structure of LPS remains consistent12,13. Furthermore, cyanobacterial extracellular polysaccharides (EPS) play a significant role in protecting them against abnormal conditions and have biotechnological applications in industries14,15. Cyanobacterial EPS can be classified into two morphological forms: (1) Capsular Polysaccharides (CPS), which form a thin or thick sheath attached to the external cell surface, and (2) Released Polysaccharides (RPS), the slimy and soluble polysaccharides released into the surrounding environment by cyanobacterial cells10,16.
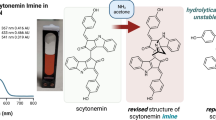
Scytonemin is an important lipid-soluble, yellow–brown-colored, UV-absorbing pigment found in the protective layer of certain cyanobacteria, including Nostoc species. This compound consists of two connected units made up of carbon atoms forming a bond17. Scytonemin exhibited absorption in both the UV-A and violet-blue regions of light, playing a crucial role as a shield against ultraviolet radiation to safeguard the cellular components of cyanobacteria18.
Additionally, Scytonemin can be found in the layer of exopolysaccharides surrounding Nostoc19. However, due to the lack of specific information about the function of scytonemin related to other molecular components in a cyanobacterial model system, the overall understanding remains unclear.
Among Nostoc species, the genome of N. punctiforme has been completely sequenced and is a good model for studying the genes related to cyanobacterial UV-absorbing pigments. According to past researches, in the cyanobacterium N. punctiforme, the genomic region of 18-gene cluster (from Npun_R1276 to Npun_R1259) is responsible for scytonemin biosynthesis20. The key genes (ScyD, ScyE, and ScyF) have been reported to be involved in the late biosynthetic steps4. A gene cluster with four genes has also been found to be associated with the biosynthesis of MAA in cyanobacteria, including Nostoc species21.
Additionally, cyanobacteria have enzymatic antioxidant defense mechanisms against UV stress and reactive oxygen species (ROS) 22,23. In this defense, enzymes such as superoxide-dismutase (SOD) and catalase (CAT) reduce oxidative damage and protect these organisms24,25.
In recent decades, ozone layer depletion has occurred due to atmospheric pollutants and climate change. As a result, solar ultraviolet radiation, such as UV-C (180–280 nm) and UV-B (280–315 nm), penetrates beyond the ozone layer. Therefore, the increasing penetration of UV radiation has raised concerns about its negative effects on ecosystems and organisms26,27 . Many studies have reported the effects of UV-A and UV-B radiation on some Nostoc species28,29,30,31, but little information is available about the response of this group of microorganisms to UV-C radiation. The harmful effect of UV-C radiation on growth, photosynthesis apparatus (pigments and proteins), and nitrogen metabolism in Nostoc muscorum Meg1, and Nostoc flagelliforme have been investigated in recent years5,32. The changes that occur in the morphology of Nostoc sp. LBALBR-2 under UV-C radiation also have been reported by Sousa et al.33. However, limited information is available on the effect of UV-C exposure on Nostoc species and their resistance to this harmful radiation.
The genus Nostoc is recognized as a producer of several active compounds, which enable it to thrive in varied and highly competitive ecological niches8. Considering the varying gradients of UV radiation observed in different geographical regions of Iran, the presence of microalgae with different adaptation potential in these regions can be predicted. Despite the importance of investigating useful cyanobacteria in the study area, there has been no comprehensive study in this field so far. Therefore, investigating the resistance and ecological adaptation of Nostoc species in Iran's natural habitats is one of the main goals of this study. Additionally, the hypothesis is that different Nostoc species collected from Iran's natural habitats, with varying levels of solar radiation, exhibit differing resistance to UV-C radiation due to their diverse protection strategies, such as the production of extracellular polymeric substances (EPS), UV-absorbing pigments (scytonemin and mycosporine-like amino acids), and antioxidant enzymes. Therefore, this study integrates experimental and computational methods to assess the role of habitat characteristics and protective strategies in enhancing the resistance of Nostoc species to UV-C radiation. The study evaluates the effects of UV-C exposure on: (1) tolerance and viability, (2) EPS contents, (3) chlorophyll-a and carotenoid contents, (4) antioxidant enzymes activity, and (5) UV-absorbing pigment levels, as well as the expression of key genes associated with the biosynthesis of scytonemin (scyF, scyE, scyD), mycosporine-like amino acids (mysD), and capsular polysaccharides (Nos7107-0132 and Nos7524-0337) in the isolated Nostoc species.
Another important issue that has not been fully explained in previous studies is how scytonemin, a crucial photoprotective compound, interacts with LPS in the cyanobacterial cell wall. Therefore, these aspects were investigated in the present research. We employed molecular modeling and docking techniques to understand how scytonemin interacts with LPS, an essential component of the cyanobacterial cell wall. Thus, computational modeling of the molecular mechanism underlying scytonemin binding to LPS was conducted for the first time.
The findings of the present study contribute to understanding the impact of natural habitat solar radiation conditions on the development and adjustment cyanobacterial genomes towards ecological adaptations, particularly in forming a scytonemin photoprotective shield in Nostoc species. Additionally, the molecular modeling and docking results from this research provide novel insights into the potential molecular mechanisms underlying scytonemin's protective role against UV radiation in cyanobacteria.
Materials and methods
Experimental methods
Site description and sample collection
Based on the De Martonne climate classification of Iran, a large area of this country (approximately 70% of the 164 million hectares) is located in arid and semi-arid regions with high received solar radiation34,35,36. Iran, with 280 sunny days/year, is situated within the global solar belt, where solar radiation is at its highest level compared to other countries37. In the present research, soil samples were collected from natural habitats in Iran, such as Mazandaran, Alborz, and Kerman provinces .These sampling sites were located along a gradient of solar radiation with different levels of radiation and climatic conditions38,39.
Geographically, Galugah (Mazandaran, Iran) is located at 36° 43′ 11″ N 53° 48′ 57″ W, with an average altitude of 37 m above sea level (a.s.l.) and low solar radiation. Dasht-e Qazvin (Alborz, Iran) is situated at 36° 19′ 46″ N 50° 01′ 35″ W, with an average altitude of 1.41 m a.s.l and medium solar radiation. Additionally, Nosratabad (Kerman, Iran) is located at 29° 30′ 33″ N 55° 36′ 01″ W, with an average altitude of 1.73 m a.s.l. and high solar radiation. Soil samples from each site were collected based on the methodology provided by Rangaswamy40. Soil sampling was done in the summer at a depth of 5 cm, and samples sieved for culturing cyanobacteria.
Cyanobacteria isolation and culture condition
Cyanobacteria were isolated from soil samples following the method described by Stanier et al.41. Initially, ten grams of sieved soil that collected from each natural site were transferred to sterile petri dishes containing 50 mL sterile liquid nitrate-free BG-11 medium. The soil samples were then incubated at 25 ± 2 °C under artificial illumination at 74 μmol photons m-2 s-1 with a 12/12 h light/dark cycle. After three weeks, cyanobacterial colonies appeared on the surface of the soil and liquid culture medium. Then, colonies were inoculated and subsequently streaked on fresh solid BG-11 medium 42.
For morphometric determinations, semi-permanent slides of all isolated colonies were inspected, and the morphometric study was conducted using light microscopy (Olympus, Model BH-2), following previous studies2,43,44,45. UV-tolerant cyanobacteria were accurately detected using 16S rRNA gene sequencing and molecular methods. Genomic DNA was extracted from fresh cyanobacterial mass using a genomic DNA extraction kit (GeneAll, Korea). Polymerase chain reaction (PCR) amplification was carried out with primers 27F (5′-AGAGTTTGATCCTGGCTCAG-3′) and 781R (5′-GACTACTGG GGTATCTAATCCCATT-3′)46,47. The PCR mixture contained 5 μL 10 × Taq buffer, 2 μL purified DNA (50 ng/μL), 4 μL of 2.5 mM dNTP, 2.8 μL of 50mM MgCl2, 500 ng of each primer, and 0.5 μL Taq polymerase (2.5 U/μL), adjusted to a final volume of 50 μL with ultra-pure water. The PCR reactions were conducted with the following program: initial denaturation at 95°C for 4 min, followed by 30 cycles of denaturation at 95°C for 40 s, annealing at 58°C for 40 s, and extension at 72°C for 35 s. The final cycle included an extension at 72°C for 10 min. The PCR products were analyzed by electrophoresis on 1% (w/v) agarose gel in TBE buffer. Selected PCR products were sequenced using the Sanger sequencing method48 by Vista Gene Enzyme Company (Tehran, Iran). The obtained sequences were edited using BioEdit ver. 7.0.9.049 and compared with available sequence information in the National Center for Biotechnology Information (NCBI) using BLAST analysis (http://www.ncbi.nlm.nih.gov/BLAST). Each cyanobacterial isolate corresponded to an individual Operational Taxonomic Unit (OTU) and was assigned a distinct accession number in GenBank. Further details can be found in the supplemental information S1.
Sample preparation and UV-C irradiation
To select UV-C resistant species, we investigated the effect of UV-C radiation on the survival of nine Nostoc species isolated from soil. The species were cultivated in 500-mL Erlenmeyer flasks containing 200 mL of sterile liquid nitrate-free BG-11 medium at 25 ± 2 °C and under artificial illumination with an intensity of 74 μmol photons m−2 s−1 with a 12/12 h light/dark cycle for 15 days. The cultures were shaken daily to prevent self-shading and clumping.
The tolerance of cyanobacteria to UV-C radiation was assessed following the protocol of Monsalves et al.50 Cyanobacterial samples were exposed to artificial UV-C radiation in a designed dark chamber equipped with a UV-C lamp with an irradiance intensity of 400 μW/cm2 and a peak emission at 254 nm (General Electric, Cleveland, Ohio). Each sample (cyanobacterial biomass in 100 mL sterile nitrate-free BG-11 medium) was placed in separate uncovered petri dishes positioned 30 cm away from the UV-C lamp and exposed to radiation for 0 (control), 10, 20, 30, 40, 50, 60, 90 and 120 min, respectively. The experiments were conducted at a temperature of 25 ± 2°C, and the cultures were shaken during exposure to prevent self-shading of the filaments.
Determination of viability
The survival and colony-forming ability of cyanobacteria following UV-C radiation were studied using the protocol previously described by Richa and Sinha51. From each cyanobacterial sample, 100 μL aliquots were withdrawn at predetermined time intervals after UV-C exposure and transferred to sterile plates with solid medium. The plates were then incubated in the culture room, and after 10 days, colonies appeared. Colony counts were used to calculate the percent survival according to the formula:
where Nt is number of colonies in the treated samples with UV-C and No is number of colonies in control.
Measurement of pigments
Chlorophyll-a and total carotenoid contents were determined according to Han et al.52 with some modifications. In our modified method, 10 mL of each sample after each UV-C treatment interval was taken and centrifuged at 10,000 × g for 5 min instead of 5mL sample and 4000 × g for 8 min, respectively. The supernatant was discarded, and 10 mL of cold methanol (4 °C) was added. Due to the rapid degradation of pigments, we reduced the incubation time in our modification, and instead of 12 h as described in the previous method, samples were stored in dark at 4 °C for 45 min. After incubation, samples were centrifuged at 10,000 × g for 10 min. The supernatant was transferred into a cuvette and measured at 470 and 665 nm in a UV–visible spectrophotometer (Hitachi 2900, Japan), using methanol as a blank. Contents were calculated using the following Eqs. 53:
The content of UV-absorbing pigments, such as MAAs and Scytonemin, was extracted and calculated according to the procedure elaborated by Mushir and Fatma54,55. For determination of MAAs content, cells were harvested after each UV-C treatment interval from each cyanobacterial sample and washed twice with distilled water. Then, the cells suspended in 20% (vol/vol) aqueous methanol and incubated in a water bath at 45ºC for 2.5 h. After incubation, samples were centrifuged at 10,000 × g for 10 min. The absorbance of the supernatant was measured spectrophotometrically at 362 nm (maximum absorbance), and the MAAs content was calculated according to the following Eq. 54,56:
For determination of scytonemin content, cells were harvested after each UV-C treatment interval from each cyanobacterial sample and washed twice with distilled water. The cells were then suspended in 100% acetone and stored overnight in dark at 4 °C. After incubation, samples were centrifuged at 10,000 × g for 10 min, and the absorbance of the supernatant was measured at 384 nm (scytonemin maximum), 470 nm (carotenoids) and 665 nm (chlorophyll-a) on a UV–visible spectrophotometer. The scytonemin content of samples was calculated according to the present Eq. 55:
Extraction and quantification of capsular polysaccharides and released polysaccharides
Cyanobacterial samples were exposed to artificial UV-C radiation as described above and then harvested by centrifugation at 15,000 × g for 10 min. Due to the sticky form of the N. carneum ISB88 medium culture, the supernatant was retained for the extraction of the released polysaccharides (RPS) secreted into the medium. The pellet of N. commune ISB98 was suspended in distilled water and then incubated at 70°C for 12 h to extract the capsular polysaccharides (CPS) embedded in the outer layer of filaments. The CPS and the RPS were precipitated at 4 °C overnight with four volumes of absolute ethanol (95%)28 and then dialyzed with distilled water for 12 h to remove salts and other small molecules. The isolated CPS and RPS were quantified using the phenol–sulfuric acid assay57.
In brief, 1 mg of each isolated CPS and RPS was suspended in 1mL of deionized water to prepare a 1000 μg mL-1 concentration, and then other concentrations (50- 800 μg mL-1) were made from them. For quantification, 50 μL of each sample was transferred to a 96-well plate, and 150 μL of 96% sulfuric acid and 30 μL of 5% phenol were added to each well. The plate was incubated in a water bath at 90 °C for 5 min. After incubation, the absorbance of the samples was measured at 490 nm on a UV–visible spectrophotometer.
The standard curve was drawn using 10–800 μg mL-1 of glucose, and results were expressed as grams of glucose equivalents per 100 g of extract.
Scanning electron microscopy of Nostoc species and the extracted polysaccharides
Scanning electron microscopy (SEM) was performed to examine the morphology of the non-irradiated cyanobacterium, as well as the extracted CPS and RPS, to understanding their surface structure. Samples were fixed with 0.1 M phosphate buffer (pH 7.2) containing 1% glutaraldehyde for 4 h and then washed with distilled water. The samples were dehydrated through a graded ethanol series (10%, 30%, 50%, 70%, 90%, and 100%) for 15 min. Subsequently, the dried samples were fixed to SEM stubs using double-sided tape, coated with a 15 nm thick layer of gold, and subjected to SEM analysis. A Hitachi S-4160 microscope (Japan) fitted with an LFD detector was used.
Effect of UV-C radiation on antioxidant enzymes activity.
The effect of UV-C radiation on the activity of antioxidant enzymes (SOD and CAT) was studied in the species. Samples were taken at each time interval after UV-C exposure and assayed for SOD and CAT activities. Enzyme activities were compared to the enzyme activities of control (time 0), which were unexposed. Measurements were performed in triplicate. For sample preparation, 0.3 g of cells was homogenized with 1.5 mL of cold 100 mM sodium phosphate buffer (pH 7.0). The homogenate was centrifuged at 13,000 × g for 10 min at 4 °C, and the supernatant was collected for enzyme assays58.
CAT activity was assessed using the method outlined by Aebi59.The reaction mixture consisted of 1mL of 50 mM sodium phosphate buffer (pH 7.0) and 0.1 mL of enzyme extract. The mixture was activated by adding 0.1 mL of 100 mM H2O2. Changes in absorbance were measured at 240 nm on a UV–visible spectrophotometer at time intervals of 15 s and 1 min. The results were expressed as unit mg protein−1; one unit of enzyme activity was equivalent to a decrease of 0.1 of absorbance at 240 nm per minute at 25 °C58.
SOD activity was measured following the procedure of Giannopolitis and Ries60 with slight modification. The reaction mixture consisted of 50 mM potassium phosphate buffer (pH 7.0), 0.1 mM EDTA, 0.75 mM nitroblue tetrazolium (NBT), 4 µM riboflavin, 13 mM methionine with 0.1 mL of enzyme extract incubated at 25°C under a fluorescent lamp for 8 min (instead of 5 min as described in the previous method). Enzyme activity was expressed as unit mg protein -1; one unit of SOD enzyme activity was defined as the amount of the enzyme required to cause 50% inhibition of the rate of NBT reduction at 560 nm58.
The protein concentration in the enzyme extract was determined using the Bradford method 61, with bovine serum albumin as the standard. Samples were prepared at concentrations ranging 50–1000 μg mL-1 in phosphate buffer. To prepare the Bradford reagent, 100 mg of Coomassie Brilliant Blue was dissolved in 50 mL of 95% ethanol, followed by addition of 100 mL of 85% phosphoric acid. The solution was then diluted to one liter with distilled water. For quantification, 50 µl of each sample was transferred to a 96-well plate, and 200 μL of the Bradford reagent was added to each well. The plate was shaken for 60 s, and the absorbance of the samples was measured at 595 nm on a UV–visible spectrophotometer. A standard curve was drawn using bovine serum albumin concentrations ranging from 50 to 1000 μg mL-1, and the protein concentration of the samples was determined using the equation derived from the standard curve.
Gene expression analysis
Total RNA was extracted from the cyanobacterial cells treated under different UV-C radiation intervals (0, 20 and 120 min) using trizol reagent (Azma Elixir, Iran) following the manufacturer’s instructions. The cDNA synthesis was performed using the PrimeScript RT reagent kit (Takara, Japan), following the manufacturer’s instructions. The sequences and accession numbers of scyF, scyE , scyD, and mysD genes involved in scytonemin and MAAs biosynthesis, reported from Nostoc punctiforme PCC73102, Nos7107-0132, and Nos7524-0337 genes related to biosynthesis of capsular polysaccharides from Nostoc sp. PCC7107 and Nostoc sp. PCC7524, respectively, and rnpB gene as reference were derived from the cyanobase (www.cyanobase.com) and the National Center for Biotechnology Information (NCBI) (www.ncbi.nlm.nihv.gov).
The primers for the genes were designed and used for real-time quantitative RT-PCR (Table 1). Quantitative RT-PCR was performed on a Step One Plus™ Real-Time PCR System (Applied Biosystems, USA) using SYBR Green PCR Master Mix (Ampliqon, Denmark). The amplification of the genes was done according to PCR program (95◦C for 5 min, followed by 35 cycles at 95 °C for 20 s, 60 °C for 30 s and 72 °C for 15s). The relative expression of each gene was analyzed by the 2−ΔΔCt method62. The gene expression analyses were performed in triplicate for each sample.
Computational methods
Preparation of the LPS and scytonemin 3D structures
The 3D structure of scytonemin was obtained from the PubChem database (www.pubchem.ncbi.nlm.nihv.gov) in PDB format. Furthermore, the 3D structure of the ligand of interest was converted to PDBQT format using the OPEN BABEL GUI software. Due to the lack of information regarding LPS sequences in Nostoc species in this study, the well-established structure of LPS from gram-negative bacteria was selected. The LPS setup provided by the CHARMM LPS modeling framework was used to obtain the complex structure of LPS. The LPS complex studied in our research is a product of lipid A type 1, and the core oligosaccharide is also constructed from D-glucose, D-galactosamine, D-glucosamine, N-acetyl-D-glucosamine, and 3-Deoxy- d-manno-oct-2-ulosonic acid63.
Molecular docking
The interaction between the LPS complex and scytonemin was studied using AutoDock 4.2. A grid box was created around a point in the LPS complex with dimensions of 70 × 70 × 70 points, in the X, Y, and Z directions using AutoGrid 4.2. The docking process utilized an algorithm with the results saved using a genetic algorithm. Default docking parameters were used, with a population size of 300 individuals and 50 GA runs performed. Scytonemin remained flexible while LPS was held rigid during the docking process. The results were assessed based on their docking score and binding energy values.
Atomistic interactions of the LPS-scytonemin complex
The selected binding modes were chosen based on the binding free energy. The Python-Based Molecular Visualization System (PyMol) was used to visualize the interactions between the receptor LPS and the ligand scytonemin. A thorough analysis of each docking output cluster was conducted to identify the interacting residues in each cluster.
Statistical analysis
The experiments were performed with three replications, and the data were subjected to One-way analysis of variance (ANOVA) using SPSS 20.0 (SPSS Inc., Chicago, IL, USA). To assess the significance of the data at P < 0.05, the Duncan test was applied. Normality and homoscedasticity were checked, and no data transformation was needed. The Statistical results were presented as the mean ± standard deviation. Histograms were created using GraphPad Prism 8.4.2 (GraphPad Software Inc., San Diego, CA, USA).
Results
Experimental results
Nostoc species morphology and their survival
Based on morphological and molecular studies, nine species of Nostoc from natural habitats with different solar radiation levels were purified and selected for UV-C viability test. These selected species were various in their filaments and colony colors (Table 2).
Among the selected Nostoc species, N. carneum ISB88, characterized by blue-green filaments with a thin gelatinous sheath, released a significant amount of extracellular polysaccharides (RPS) into the medium culture. In contrast, N. commune ISB98, with olive-green filaments, had filaments embedded one by one in capsular polysaccharides (CPS), forming a thick sheath. Additionally, N. sphaericum ISB97 and Nostoc sp. ISB99 exhibited brown-colored filaments without an obvious gelatinous sheath. N. linckia ISB96, N. paludosum ISB95, Nostoc sp. ISB89, Nostoc sp. ISB87, and Nostoc sp. ISB86, characterized by blue-green filaments, displayed thin gelatinous sheaths around their filaments.
The results indicated that the viability of N. carneum ISB88 was significantly affected, with 50% inhibition recorded after 10 min of exposure. Complete killing of N. carneum ISB88 cells occurred after 40 min of exposure, while N. commune ISB98 showed 100% killing after 60 min of radiation exposure. On the other hand, N. sphaericum ISB97 and Nostoc sp. ISB99 remained alive until the end of the experiment, forming colonies after 120 min of UV-C radiation exposure. For other species, including N. linckia ISB96, N. paludosum ISB95, Nostoc sp. ISB89, Nostoc sp. ISB87, and Nostoc sp. ISB86, complete killing occurred after 30 min of exposure, and they were no longer able to form colonies.
Based on the viability test for the 9 species and their varying resistance under UV-C exposure, we decided to select four species with brown, olive-green, and blue-green filaments that exhibited higher resistance than other species from habitats with high, medium, and low levels of solar radiations to compare with each other for further investigations in this study.
Thus, N. sphaericum ISB97, and Nostoc sp. ISB99 were selected as resistant brown species that were isolated from natural habitats with high solar radiations. Additionally, N. carneum ISB88 and N. commune ISB98 were selected as resistant blue-olive green species that were isolated from habitats with low-medium solar radiations.
Extracted polysaccharides contents
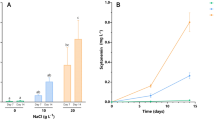
In this study, filaments of N. sphaericum ISB97 and Nostoc sp. ISB99 were examined for EPS extraction, but no EPS was isolated from these species. The RPS isolated from N. carneum ISB88 exhibited a looser structure compared to the dense CPS of N. commune ISB98. The extracted CPS amounted to approximately 681.8 mg L−1, which was higher than the RPS content of approximately 470.2 mg L−1. Both CPS and RPS contents remained constant with no significant difference observed during the UV-C radiation intervals (P > 0.05) (Fig. 1). These results suggested that UV-C exposure did not have negative or positive effects on the quantity of these polysaccharides.
Morphology of Nostoc species cells and the extracted EPS: scanning electron microscopy
Scanning electron microscopy (SEM) images of the cyanobacterial samples and the extracted EPS for investigation of variations are presented in Fig. 2. A difference in the surface morphology of the RPS and the CPS was revealed with SEM (Fig. 2a, b). The RPS surface is decorated with rigid, bar-shaped aggregations, while the CPS surface appears meshwork and smooth. The morphology of the studied species filaments in SEM analysis are shown in Fig. 2c, d, e and f. A halo around the N. commune ISB98 filament can be attributed to the presence of a thick CPS (Fig. 2c).
Effects of UV-C irradiation on the photosynthetic pigments content
The chlorophyll-a and total carotenoids contents of N. commune ISB98, N. carneum ISB88, N. sphaericum ISB97and, Nostoc sp. ISB99 are shown in Fig. 3a, b.
Effect of UV-C on the contents of chlorophyll-a (a), carotenoids (b), Scytonemin (c), and MAAs (d) in Nostoc commune ISB98, Nostoc carneum ISB88, Nostoc sphaericum ISB97, and Nostoc sp. ISB99. Results are expressed as the means of three replicates. Vertical bars indicate standard deviation of the means.
It was observed that the initial level of chlorophyll-a was significantly higher in N. carneum ISB88 than in the other species, approximately 1.6 folds (P < 0.05) (Fig. 3a). The initial level of carotenoids content was significantly higher in N. sphaericum ISB97 and Nostoc sp. ISB99 than in the other species, 2.27 and 1.90 folds, respectively (Fig. 3b). Results showed that the contents of chlorophyll-a and carotenoids decreased with an increase in UV-C radiation time. The total carotenoids contents in species were found to be less affected by UV-C radiation than chlorophyll-a, especially in N. sphaericum ISB97 and Nostoc sp. ISB99, where the predominant color of their colonies was brown. Chlorophyll-a depletion in N. carneum ISB88 occurred more rapidly than in the others, so that after ten minutes of radiation, the amount of chlorophyll-a in N. carneum ISB88 decreased from 10.6 mg L−1 to 7.36 mg L−1.The photosynthetic pigments were significantly reduced after 120 min of UV-C irradiation in N. commune ISB98 and N. carneum ISB88, so the discoloration in theses colonies was observed.
UV-C irradiation and UV- absorbing pigments
The effect of UV-C on the scytonemin and MAAs contents in the Nostoc species cells are shown in Fig. 3c, d. The results suggest that UV-C can induce the synthesis of scytonemin. In the first 50 min, no significant change was observed in the scytonemin contents of the species (Fig. 3c). After 60 min of exposure, the scytonemin content increased in N. sphaericum ISB97and Nostoc sp., reaching maximum values after 120 min (0.27 and 0.26 µg mL-1, respectively, P < 0.05). However, a decrease in the scytonemin content of N. commune ISB98 and N. carneum ISB88 could be attributed to their cell death. Changes in the MAAs contents were not evident during UV-C exposure, except in N. commune ISB98 and N. carneum ISB88, where the MAAs content decreased after cell death (Fig. 3d).
Antioxidant enzymes activity
After ten minutes of UV-C radiation, SOD and CAT activities in all treated species showed a multifold increase (P < 0.05) compared to controls (Fig. 4a, b). Although in N. sphaericum ISB97 and Nostoc sp. ISB99, this increase in values retained an upward trend until 120 min. A sharp peak in CAT and SOD activities was found at 30 min of UV-C radiation in N. carneum ISB88 and N. commune ISB98, and then persistently declined to low levels until 120 min (Fig. 4b).
Effect of UV-C radiation on (a) Catalase (CAT) and (b) Superoxide Dismutase (SOD) activities. SOD and CAT activities were measured at different time intervals and compared to the enzyme activitiy at time 0 (control). Measurements were performed in triplicate, and vertical bars indicate the standard deviation of the means.
Quantitative PCR analysis
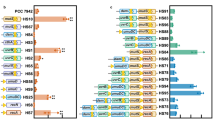
Expression analysis of genes associated with scytonemin, MAAs and capsular polysaccharides biosynthesis under different UV-C intervals (0, 20 and 120 min) is shown in Fig. 5. In N. sphaericum ISB97 and Nostoc sp. ISB99, after 20 min of exposure, the expression of scytonemin biosynthesis-related genes (scyF and scyD) continuously increased up to 120 min (P < 0.05) (Fig. 5a, c). Results indicate that long UV-C exposure time decreased the expression of the scytonemin biosynthesis-related gene (scyE) in all species, whereas the expression of scyF and scyD was not significantly changed in N. commune ISB98 and N. carneum ISB88 under UV-C exposure and control conditions (Fig. 5a, b, c). High expression of the mysD gene, involved in MAAs synthesis, was observed in N. sphaericum ISB97 and Nostoc sp. ISB99 under control condition (Fig. 5d). However, mysD expression decreased during UV-C radiation. Regarding CPS biosynthesis-related genes, high expression of Nos7107 and Nos7524 was shown in N. commune ISB88 and N. carneum ISB98 under control condition, respectively. UV-C exposure decreased the expression of these two genes in all species (Fig. 5e, f).
qRT-PCR expression analysis of genes involved in scytonemin biosynthesis (scyF, scyE , scyD) (a, b, c), MAAs biosynthesis (mysD) (d), and capsular polysaccharides biosynthesis (Nos7107 and Nos7524) (e, f) in N. commune ISB98, N. carneum ISB88, N. sphaericum ISB97, and Nostoc sp. ISB99. Results are expressed as the means of three replicates.
Computational results
Molecular Docking of LPS-scytonemin
The LPS-scytonemin binding poses were assessed by binding free energy to determine the best binding mode. All 50 structural poses were sorted into separate clusters based on a root mean square deviation (RMSD) limit of 2.0 Å compared to the ligand conformation. The summary of clustering results has been represented in Tables 3 and Fig. 7. The binding free energies of the leading docking poses across all clusters are listed in Table 3. The topmost binding energies within these 11 clusters range from − 6.38 kcal/mol to − 5.21 kcal/mol (Fig. 7). The best binding pose within each cluster is determined based on the lowest binding energy and is regarded as the representative choice (Fig. 6) (Table 4).
Among these eleven identified binding poses, the first cluster displays the most favorable binding energy, with scytonemin is located at the side of glycans chains (Fig. 6A). However, due to the possible variations in the LPS structure, variations in scytonemin placement could occur if the o chain length increases. When scytonemin interacts with a pre-assembled LPS, the side placement is less likely due to spatial clashes. Therefore, it might be more likely for the scytonemin to be positioned at the top of the LPS complex (Fig. 6). The binding pose where scytonemin is located at the top of LPS complex is also among the selected binding poses, presenting a binding energy of -5.21 in cluster 11 (Table 4 and Fig. 6K). Additionally, Fig. 6 B, C, D, E, F, J, H, I, and J represent the side positioning of scytonemin onto the LPS (Fig. 7).
Atomistic interactions of LPS-scytonemin
Hydrogen bonding is the main interaction between LPS and scytonemin (Fig. 8). The hydrogen bond interactions are illustrated in PyMOL to present the details of interactions between the LPS and Scytonemin. Hydrogen bonding between scytonemin and LPS complex and van der Waals interactions, are represented in (Table 4). The van der Waals energy falls within the range of -7.6 to -6.4, as indicated in Table 4. The top conformation (Fig. 8A) forms 13 hydrogen bonds within the complex, which is the highest number among all binding poses. Additionally, Fig. 8B–K represent the side positions of scytonemin onto the LPS.
3D structure of scytonemin binding to LPS complex with hydrogen bond interactions. The blue atom name sticks indicates the lipid A section of the LPS complex. The green atom name sticks indicate the core oligosaccharide section of the LPS complex and the yellow atom name sticks show the scytonemin molecule. The purple dashed line one shows the H-bonds interactions in the LPS complex and the white color indicates the hydrogen atom.
Discussion
Considering the ozone depletion in the atmosphere and the increase in solar UV radiation on the Earth’s surface, the investigation of UV-resistant microorganisms and their defense strategies has received much attention in the last decade. Cyanobacteria are well-known microorganisms with a wide distribution and high resistance in extreme habitats, even with high solar radiation64,65. According to Mloszewska et al.66, UV radiation stress in natural habitats is an important factor shaping ecosystems and the rates of primary productivity. Photosynthesizing microorganisms, such as cyanobacteria, have required the evolution of defense mechanisms to decrease the damage caused by solar UV radiation and protect them against the harmful effects of radiation in natural habitats. Since limited information is available on the effect of UV-C radiation on Nostoc species, we used experimental and computational methods to identify the role of protective strategies in these microorganisms against UV-C radiation.
At first, we compared the sensitivity of nine species of the genus Nostoc isolated from natural habitats in Iran against artificial UV-C radiation. The results showed varying sensitivity among taxa, with some species demonstrating higher resistance to UV-C radiation (Table 2). Under a short duration of UV-C exposure (less than 60 min), we compared N. commune ISB98 with N. carneum ISB88. Considering that N. commune ISB98, with its thick layers of CPS isolated from a habitat with medium solar radiation, showed better survival compared to N. carneum ISB88. The presence of extracellular polysaccharides in the form of a gelatinous sheath appeared to protect cells against intense conditions such as UV-C radiation. Previous studies have demonstrated that the massive CPS around N. commune filaments plays an essential role in extreme desiccation tolerance, as well as in low or high temperatures. Removal of the gelatinous sheath from N. commune filaments increased sensitivity to environmental conditions67,68,69,70. Although the chlorophyll-a and carotenoids contents were affected by UV-C radiation in all tested species, the degradation rates of photosynthetic pigments and the photo-bleaching cells in N. carneum ISB88 were faster than in the other species. These observations indicated that the production of extracellular polysaccharides in the form of RPS does not play a significant role in protecting cells from intense UV radiation. Therefore, CPS is different in shape and function from RPS in Nostoc species71. Susa et al. (2022) demonstrated that the formation of thick and dense mucilaginous sheath (CPS) around filaments in Nostoc culture under UV-C exposure is one of the defense strategies in this microorganism to protect cells from lethal effects of UV-C 33.
Based on our qRT-PCR results for the expression of genes related to capsular polysaccharide biosynthesis, Nos7524 showed high expression in N. carneum ISB88 and may be related to the production of RPS. It is possible that EPS, initially produced as CPS, is subsequently secreted into the culture medium in the form of RPS. Although the exact mechanism of RPS assembly in Nostoc species remains unclear, the expression of genes associated with CPS biosynthesis in the studied brown Nostoc species suggests the presence of a capsular sheath around their filaments, although it may be potentially too thin to be visibly apparent.
The results indicated that the gelatinous sheath alone is not a sufficient strategy to protect Nostoc cells against high solar radiation, while the brown species belonging to habitats with high solar radiation survived for up to 120 min of UV-C exposure. This resistance to radiation may be attributed to the presence of high amounts of UV radiation-blocking sunscreen pigments such as scytonemin, which play a crucial role in mitigating the lethal effects of UV radiation4. The results revealed a rapid increase in scytonemin content in Nostoc sp. ISB99 and N. sphaericum ISB97 after 50 min of UV-C exposure, continuing until the end of the experiment, suggesting that prolonged UV-C exposure induces scytonemin synthesis in these species. Similarly, previous studies have reported that UV-C radiation stimulates scytonemin production in N. flagelliforme, although high levels of UV-C radiation can inhibit its production32. Our qRT-PCR results showed the crucial role of scytonemin as a potent protective pigment against UV-C in brown Nostoc species. However, the levels of mycosporine-like amino acids (MAAs) remained constant across the four species during UV-C radiation. Previous research has suggested that UV-B exposure is more effective than other forms of UV radiation in inducing MAAs synthesis 72,73. Hence, it appears that the resistance of Nostoc species to UV radiation is associated with the expression of scytonemin biosynthesis genes, as evidenced by the increased expression of scyF and scyD genes under UV-C radiation.
The presented results are in line with the findings of Soule et al.74, who compared wild-type and scytonemin-deficient mutant of the cyanobacterium Nostoc punctiforme ATCC 29,133 and demonstrated the significant involvement of ScyD, ScyE, and ScyF genes in the scytonemin biosynthesis pathway. However, the results of Orellana et al. (2020) revealed a stimulatory effect of UV radiation on the increased transcription of a gene involved in scytonemin biosynthesis in cyanobacteria75. Herein, molecular modeling and molecular docking techniques were used for the first time to elucidate the interactions between scytonemin and the LPS complex in Nostoc species. 11 binding poses were identified for the LPS-scytonemin binding mechanism. Where all of the binding poses except one represent the side positioning of scytonemin on the carbohydrate region of the LPS complex. While one binding pose represents the top positioning of scytonemin on the glycans of LPS. These findings pave the way for exploring the molecular effects of scytonemin in forming the UV protection shield in cyanobacteria, which has been ambiguous until now.
Due to the lack of available three-dimensional (3D) structural data on LipoPolySaccharides (LPS) from the cyanobacterial genus Nostoc, researchers often rely on the well-established 3D structures of LPS from gram-negative bacteria to draw comparisons and inferences. This approach is reasonable considering that the cell walls of cyanobacteria and gram-negative bacteria possess LPS in their outer membranes, contributing to their structural aspects. However, while LPS have been well-characterized in a significant number of proteobacteria, they remain poorly understood in cyanobacteria76,77,78. Several methods have been applied to elucidate the structure of LPS, including Nuclear Magnetic Resonance (NMR) spectroscopy, Mass Spectrometry (MS), which includes gas chromatography and Matrix-Assisted Laser Desorption/Ionization, (MALDI)-MS79. However, the majority of work has been done on LPS from Gram-negative bacteria, while information on cyanobacterial LPSs has been limited and sometimes contradictory9. Computational modeling methods play a significant role in unveiling the 3D structures of glycolipids such as LPS80. Glycolipids are compounds comprising a glycan headgroup attached to a lipid segment. Due to their diverse and often isomeric nature resulting from biosynthesis, these molecules possess various biological functions81. While the complexity of glycolipids poses challenges in analysis and isolation, computational tools excel at handling the structures of glycans that may be challenging to study using traditional methods82. In addition to the studies mentioned earlier, the simulation of glycolipid structures utilizing bioinformatics tools, such as CHARMM-GUI, has been a common approach83.
Typically, the inner part of the LPS molecule in proteobacteria contains conserved elements such as 3-deoxy-D-manno-octulosonic acid (KDO), heptose and phosphate. The inner core area connects the polysaccharide, which can differ among strains, to the lipid A part that remains stable and attaches the LPS to the membrane77. While some cyanobacterial LPS have been found, through chemical analysis, to lack the phosphate group, KDO, and heptose, which are common in LPS of gram negative bacteria, specific cyanobacteria, such as Phormidium spp84, Agmenellum quadruplicatum85, and Anacystis nidulans86, exhibit similar chemical compositions as gram-negative bacterial LPS9,87,88,89. This distinction may manifest as variability in the structure of LPS molecules produced by species or even strains of the species coexisting in a shared environment9. The detailed discussion earlier highlighted the information on the structures of LPS from Gram-negative bacteria, which serves as a valuable reference point for examining cyanobacterial LPS. It is crucial to consider the structures of these two types of LPS and to further investigate the unique characteristics of cyanobacterial LPS through upcoming research and analysis77,90. Apart from the mentioned studies, another report also addresses the utilization of established Gram-negative LPS structures as benchmarks for structural and biophysical inquiries into the bacterial outer membrane and its interactions with antimicrobial substances91.
Another important mechanism observed in these studied Nostoc species is the increase in the activity of antioxidant enzymes such as SOD and CAT. Wang et al. (2015) demonstrated significant enhancement in ROS production in the cyanobacterium Microcystis aeruginosa under UV-C exposure92. Phukan & Syiem (2019) evaluated the antioxidant responses of the cyanobacteria Nostoc muscorum against UV radiation and demonstrated that UV-C significantly induced enzymatic and non-enzymatic antioxidant production in this cyanobacterium.
According to Phukan et al. (2018), the D1 protein of PSII in the photosynthetic apparatus is a highly sensitive target for UV-C induced intracellular ROS such as singlet oxygen, superoxide, hydroxyl radical, and hydrogen peroxide93,94,95.
In the present study, SOD and CAT activities in all species were multiplied under UV-C treatment until they were alive. The activity of these enzymes in N. carneum ISB88 and N. commune ISB98 decreased after their death, along with the contents of photosynthetic pigments (chlorophyll-a and carotenoids), until the end of the UV-C exposure (120 min), and cell photo- bleaching occurred in these species.
The studies by Ye et al. (2021) have found a high survival rate for Nostoc sp. after exposure to a Mars-like stratosphere environment, inducing the expression of some genes relevant to the key products of the antioxidant system96.
These results suggest that effective ROS scavenging by antioxidant enzymes in Nostoc cells protects these microorganisms in stressful situations, and the adequate amount of these antioxidants is important to decrease damage from ROS produce during UV irradiation70,97,98,99,100.
Conclusion
In conclusion, as predicted, the brown Nostoc species originating from habitats with high solar radiation are well-adapted to UV radiation, including UV-C exposure. These adaptations enable cyanobacteria to defend themselves against the damaging effects of intense solar UV radiation. Therefore, habitat conditions have the potential to to influence the expression of cyanobacterial genomes, promoting ecological adaptations that allow these microorganisms to colonize and thrive in extreme environments on Earth. Based on our investigations, scytonemin appears to be sufficient in Nostoc species as a valuable UV-C protection strategy, suggesting that the high production of scytonemin forms the UV shield.
Since lipopolysaccharide (LPS) covers the outer membrane in cyanobacteria, it may be the primary target of the highly expressed scytonemin under UV stress. Several studies, including this one, suggest that scytonemin is crucial for protecting cyanobacteria from UV radiation. However, previous researches has not fully explained how exactly scytonemin carries out this role. To bridge this knowledge gap, we carried out modeling techniques to understand how scytonemin interacts with LPS, an essential part of the cyanobacterial cell wall. In this study, we computationally modeled the molecular mechanism of scytonemin binding to LPS for the first time, revealing novel findings. Our molecular docking studies showed a strong interaction between scytonemin and LPS, with the best binding pose exhibiting a favorable binding energy of -6.38 kcal/mol. Additionally, we observed that scytonemin tends to be located on the side of the glycan chains within the LPS structure, providing new information about the spatial arrangement of this interaction. The complex formed by LPS and scytonemin displayed hydrogen bond interactions, with the binding position showing two hydrogen bonds which enhance the binding stability. Our investigation delved into atomic level interactions between scytonemin and specific parts of the LPS molecule, particularly focusing on core regions, offering a deeper insight into their binding mechanism. Furthermore, the strong binding affinity and specific interactions observed suggest that scytonemin could influence the structure or function of LPS, potentially aiding in UV protection mechanisms in cyanobacteria. These results provide novel insights into the scytonemin's protective role against UV radiation in cyanobacteria, enhancing our experimental findings at a molecular level.
Data availability
Data regarding the main Nostoc species of this research, including Nostoc sp.4 ISB211 (accession number OQ225494), Nostoc sphaericum ISB210 (accession number OQ225510), Nostoc carneum ISB92 (accession number MK771137), and Nostoc commune ISB212 (accession number OQ225490), have been deposited in GenBank. More detail and information can be found in the Supplementary information S1.
References
Rastogi, R. P. et al. Ultraviolet radiation and cyanobacteria. J. Photochem. Photobiol. B. 141, 154–169 (2014).
Komárek, J. Süßwasserfora von Mitteleuropa. Cyanoprokaryota: 3 Teil/Part 3: Heterocystous genera (Springer, 2013).
Dabravolski, S. A. & Isayenkov, S. V. Metabolites facilitating adaptation of desert cyanobacteria to extremely arid environments. Plants. 11, 3225 (2022).
Pathak, J. et al. Genetic regulation of scytonemin and mycosporine-like amino acids (MAAs) biosynthesis in cyanobacteria. Plant Gene. 17, 100172 (2019).
Phukan, T., Rai, A. N. & Syiem, M. B. Dose dependent variance in UV-C radiation induced effects on carbon and nitrogen metabolism in the cyanobacterium Nostoc muscorum Meg1. Ecotoxicol. Environ. Saf. 155, 171–179 (2018).
Singh, S. P., Häder, D. P. & Sinha, R. P. Cyanobacteria and ultraviolet radiation (UVR) stress: mitigation strategies. Ageing Res. Rev. 9, 79–90 (2010).
Jain, S. et al. Cyanobacteria as efficient producers of mycosporine-like amino acids. J. Basic Microbiol. 57, 715–727 (2017).
Liu, Y. et al. Non-random genetic alterations in the cyanobacterium Nostoc sp. exposed to space conditions. Sci Rep. 12, 12580. https://doi.org/10.1038/s41598-022-16789-w (2022).
Durai, P., Batool, M. & Choi, S. Structure and effects of cyanobacterial lipopolysaccharides. Mar. Drugs. 13, 4217–4230 (2015).
Kehr, J. C. & Dittmann, E. Biosynthesis and function of extracellular glycans in cyanobacteria. Life 5, 164–180 (2015).
Belton, S., McCabe, P. F. & Ng, C. K. Y. The cyanobacterium, Nostoc punctiforme can protect against programmed cell death and induce defence genes in Arabidopsis thaliana. J. Plant Interact. 16, 64–74 (2021).
Raetz, C. R. & Whitfield, C. Lipopolysaccharide endotoxins. Annu. Rev. Biochem. 71, 635–700 (2002).
Bertani, B. & Ruiz, N. Function and Biogenesis of Lipopolysaccharides. EcoSal Plus https://doi.org/10.1128/ecosalplus.esp-0001-2018 (2018).
Singh, S., Verma, E., Niveshika Tiwari, B. & Mishra, A. K. Exopolysaccharide production in Anabaena sp. PCC 7120 under different CaCl2 regimes. Physiol Mol Biol Plants 22, 557–566 (2016).
Xiao, R. et al. Investigation of composition, structure and bioactivity of extracellular polymeric substances from original and stress-induced strains of Thraustochytrium striatum. Carbohydr. Polym. 195, 515–524 (2018).
Cruz, D., Vasconcelos, V., Pierre, G., Michaud, P. & Delattre, C. Exopolysaccharides from cyanobacteria: Strategies for bioprocess development. Appl. Sci. 10, 3763 (2020).
Sinha, R. & Häder, D. Natural bioactive compounds: Technological advancements (Elsevier, Amsterdam, 2021).
Abdulla, M. H. & Sumayya, N. S. Antioxidants from marine cyanobacteria (Elsevier, Amsterdam, 2023).
Gao, X., Jing, X., Liu, X. & Lindblad, P. Biotechnological production of the sunscreen pigment scytonemin in cyanobacteria: Progress and strategy. Nat. Rev. Microbiol. 19, 791–802 (2021).
Soule, T., Garcia-Pichel, F. & Stout, V. Gene expression patterns associated with the biosynthesis of the sunscreen scytonemin in Nostoc punctiforme ATCC 29133 in response to UVA radiation. J. Bacteriol. 191, 4639–4646 (2009).
Rosic, N. N. Mycosporine-like amino acids: Making the foundation for organic personalised sunscreens. Mar. Drugs. 17, 638 (2019).
Janknegt, P. J., Van De Poll, W. H., Visser, R. J., Rijstenbil, J. W. & Buma, A. G. Oxidative stress responses in the marine Antarctic diatom Chaetocerosbrevis (Bacillariophyceae) during photoacclimation. J. Phycol. 44, 957–966 (2008).
Wang, G. et al. The response of antioxidant systems in Nostoc sphaeroides against UV-B radiation and the protective effects of exogenous antioxidants. Adv. Space Res. 39, 1034–1042 (2007).
Monsalves, M. T., Amenábar, M. J., Ollivet-Besson, G. P. & Blamey, J. M. Effect of UV radiation on a thermostable superoxide dismutase purified from a thermophilic bacterium isolated from a sterilization drying oven. Protein pept. lett. 20, 749–754 (2013).
Correa-Llantén, D. N., Amenábar, M. J. & Blamey, J. M. Antioxidant capacity of novel pigments from an Antarctic bacterium. J. Microbiol. 50, 374–379 (2012).
Pattanaik, B., Schumann, R. & Karsten, U. Effects of ultraviolet radiation on cyanobacteria and their protective mechanisms 29–45 (Springer, Berlin, 2007).
Singh, V. K. et al. Resilience and mitigation strategies of cyanobacteria under ultraviolet radiation stress. Int. J. Mol. Sci. 24, 12381. https://doi.org/10.3390/ijms241512381 (2023).
Feng, Y. N., Zhang, Z. C., Feng, J. L. & Qiu, B. S. Effects of UV-B radiation and periodic desiccation on the morphogenesis of the edible terrestrial cyanobacterium Nostoc flagelliforme. Appl. Environ. Microbiol. 78, 7075–7081 (2012).
Mansouri, H. & Talebizadeh, R. The effects of UVB on growth and anti-UV compounds contents in cyanobacteria Nostoc linckia. J. oceanogr. 12, 1–12 (2022).
Yu, H. & Liu, R. Effect of UV-B radiation on the synthesis of UV-absorbing compounds in a terrestrial cyanobacterium. Nostoc flagelliforme. J. Appl. Phycol. 25, 1441–1446 (2013).
Sheeba, Ruhil, K. & Prasad, S. M. Nostoc muscorum and Phormidium foveolarum differentially respond to butachlor and UV-B stress. Physiol. Mol. Biol. Plants. 26, 841–856 (2020).
Yu, H., Pang, S. & Liu, R. Effects of UV-B and UV-C radiation on the accumulation of scytonemin in a terrestrial cyanobacterium, Nostoc flagelliforme. Preprint at https://ieeexplore.ieee.org/document/6098596 (2011).
de Sousa, E. B. et al. Effect of ultraviolet-C radiation on the morphology of cyanobacteria nostoc sp. LBALBR-2 isolated from supply reservoir (Belém, Pará, Brazil). Res. Soc. Dev. 11, e447111234391. https://doi.org/10.33448/rsd-v11i12.34391 (2022).
Khalili, A., Bazrafshan, J. & Chraghalizadeh, M. A Comparative study on climate maps of Iran in extended de Martonne classification and application of the method for world climate zoning. J Agricul. Meteorol. 10, 3–16 (2022).
Feizi, V., Mollashahi, M., Farajzadeh, M. & Azizi, G. Spatial and temporal trend analysis of temperature and precipitation in Iran. Ecopersia 2, 727–742 (2014).
Etemadi-Khah, A., Pourbabaee, A., Noroozi, M., Alikhani, H. & Bruno, L. Biodiversity of Isolated Cyanobacteria from Desert Soils in Iran. Geomicrobiol. J. 34, 90546809. https://doi.org/10.1080/01490451.2016.1271064 (2017).
Ahmad, F. A. Valuation of solar power generating potential in Iran desert areas. J. Appl. Sci. Environ. Manag. 22, 6. https://doi.org/10.4314/jasem.v22i6.21 (2018).
Irankhahi, P., Riahi, H., Shariatmadari, Z. & Shariatmadari, Z. A. Diversity and distribution of heterocystous cyanobacteria across solar radiation gradient in terrestrial habitats of Iran. Rostaniha 23, 264–281 (2022).
Aghashariatmadari, Z. Evaluation of model for estimating total solar radiation at horizontal surfaces based on meteorological data, with emphasis on the performance of the angstrom model over Iran (Tehran University, 2011).
Rangaswami, G. Agricultural microbiology (Asia Publishing House, Mumbai, 2011).
Stanier, R. Y., Kunisawa, R., Mandel, M. & Cohen-Bazire, G. Purification and properties of unicellular blue-green algae (order Chroococcales). Bacteriolo. Rev. 35, 171–205 (1971).
Andersen, R. A. Algal culturing techniques (Elsevier Science, Amsterdam, 2005).
Wehr, J., Sheath, R. & Kociolek, P. Freshwater Algae of North America: Ecology and Classification (Elsevier, Amsterdam, 2015).
John, D. M., Whitton, B. A. & Brook, A. The freshwater algal flora of the British Isles: an identification guide to freshwater and terrestrial algae (Cambridge University Press, Cambridge, 2002).
Hauer, T. & Komárek, J. CyanoDB 2.0 - On-line database of cyanobacterial genera. http://www.cyanodb.cz (2022).
Dos Santos, H. R. M., Argolo, C. S., Argôlo-Filho, R. C. & Loguercio, L. L. A 16S rDNA PCR-based theoretical to actual delta approach on culturable mock communities revealed severe losses of diversity information. BMC Microbiol. 19, 74 (2019).
Nübel, U., Garcia-Pichel, F. & Muyzer, G. PCR primers to amplify 16S rRNA genes from cyanobacteria. Appl. Environ. Microbiol. 63, 3327–3332 (1997).
Sanger, F. & Coulson, A. R. A rapid method for determining sequences in DNA by primed synthesis with DNA polymerase. J. Mol. Biol. 94, 441–448 (1975).
Hall, T. A. BioEdit: A user-friendly biological sequence alignment editor and analysis program for windows 95/98/NT. Nucl. Acids Symp. Ser. 41, 95–98 (1999).
Monsalves, M. T., Ollivet-Besson, G. P., Amenabar, M. J. & Blamey, J. M. Isolation of a psychrotolerant and UV-C-resistant bacterium from Elephant Island, Antarctica with a highly thermoactive and thermostable catalase. Microorganisms 8, 95 (2020).
Richa Sinha, R. P. Sensitivity of two Nostoc species harbouring diverse habitats to ultraviolet-B radiation. Microbiology 84, 398–4075 (2015).
Han, P. et al. Applying the strategy of light environment control to improve the biomass and polysaccharide production of Nostoc flagelliforme. J. Appl. Phycol. 29, 55–65 (2017).
Wellburn, A. R. The spectral determination of chlorophylls a and b, as well as total carotenoids, using various solvents with spectrophotometers of different resolution. J. Plant Physiol. 144, 307–313 (1994).
Mushir, S. & Fatma, T. Ultraviolet Radiation-absorbing Mycosporine-like Amino Acids in cyanobacterium Aulosira fertilissima: environmental perspective and characterization. Curr. Res. J. Biol. Sci. 3, 165–171 (2011).
Mushir, S. & Fatma, T. Monitoring stress responses in cyanobacterial scytonemin – screening and characterization. Environ. Technol. 33, 153–157 (2012).
Garcia-Pichel, F. & Castenholz, R. W. Occurrence of UV-absorbing, mycosporine-like compounds among cyanobacterial isolates and an estimate of their screening capacity. Appl. Environ. Microbiol. 59, 163–169 (1993).
DuBois, M., Gilles, K. A., Hamilton, J. K., Rebers, P. A. & Smith, F. Colorimetric method for determination of sugars and related substances. Anal. Chem. 28, 350–356 (1956).
Chairat, B., Pongprasert, N. & Srilaong, V. Effect of UV-C treatment on chlorophyll degradation, antioxidant enzyme activities and senescence in Chinese kale (Brassica oleracea var. alboglabra). Int. Food Res. J. 20, 623–628 (2013).
Aebi, H. Catalase in vitro. Meth. Enzymol. 105, 121–126 (1984).
Giannopolitis, C. N. & Ries, S. K. Superoxide dismutases: I. occurrence in higher plants. Plant Physiol. 59, 309–314 (1977).
Bradford, M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72, 248–254 (1976).
Pfaffl, M. W. A new mathematical model for relative quantification in real-time RT-PCR. Nucl. Acids Res. 29, e45. https://doi.org/10.1093/nar/29.9.e45 (2001).
Jo, S. et al. Lipopolysaccharide membrane building and simulation. Methods Mol. Biol. 1273, 391–406 (2015).
Garcia-Pichel, F. & Belnap, J. Microenvironments and microscale productivity of cyanobacterial desert crusts. J. Phycol. 32, 774–782 (2008).
Wu, H., Gao, K., Villafañe, V. E., Watanabe, T. & Helbling, E. W. Effects of solar UV radiation on morphology and photosynthesis of filamentous cyanobacterium Arthrospira platensis. Appl. Environ. Microbiol. 71, 5004–5013 (2005).
Mloszewska, A. M. et al. UV radiation limited the expansion of cyanobacteria in early marine photic environments. Nat. Commun. 9, 3088 (2018).
Tamaru, Y., Takani, Y., Yoshida, T. & Sakamoto, T. Crucial role of extracellular polysaccharides in desiccation and freezing tolerance in the terrestrial cyanobacterium Nostoc commune. Appl. Environ. Microbiol. 71, 7327–7333 (2005).
Helm, R. F. et al. Structural characterization of the released polysaccharide of desiccation-tolerant Nostoc commune DRH-1. J. Bacteriol. 182, 974–982 (2000).
Sakamoto, T. et al. The extracellular-matrix-retaining cyanobacterium Nostoc verrucosum accumulates trehalose, but is sensitive to desiccation. FEMS Microbiol. Ecol. 77, 385–394 (2011).
Li, H. et al. Antioxidant and moisture-retention activities of the polysaccharide from Nostoc commune. Carbohydr. Polym. 83, 1821–1827 (2011).
Han, P. P., Sun, Y., Jia, S. R., Zhong, C. & Tan, Z. L. Effects of light wavelengths on extracellular and capsular polysaccharide production by Nostoc flagelliforme. Carbohydr. Polym. 105, 145–151 (2014).
Chandra, R., Fernanda, P. F., Parra-Saldívar, R. & Rittmann, B. E. Effect of ultra-violet exposure on production of mycosporine-like amino acids and lipids by Lyngbya purpurem. Biomass Bioenergy 134, 105475 (2020).
Sinha, R. P., Klisch, M., Helbling, E. W. & Häder, D. Induction of mycosporine-like amino acids (MAAs) in cyanobacteria by solar ultraviolet-B radiation. J. Photochem. Photobiol. B. 60, 129–135 (2001).
Soule, T., Stout, V., Swingley, W. D., Meeks, J. C. & Garcia-Pichel, F. Molecular genetics and genomic analysis of scytonemin biosynthesis in Nostoc punctiforme ATCC 29133. J. Bacteriol. 189, 4465–4472 (2007).
Orellana, G., Gómez-Silva, B., Urrutia, M. & Galetović, A. UV-A irradiation increases scytonemin biosynthesis in cyanobacteria inhabiting halites at Salar Grande Atacama desert. Microorganisms 8, 1690 (2020).
Hoiczyk, E. & Hansel, A. Cyanobacterial cell walls: news from an unusual prokaryotic envelope. J. Bacteriol. 182, 1191–1199 (2000).
Snyder, D. S., Brahamsha, B., Azadi, P. & Palenik, B. Structure of compositionally simple lipopolysaccharide from marine synechococcus. J. Bacteriol. 191, 5499–5509 (2009).
McCarren, J. et al. Inactivation of swmA results in the loss of an outer cell layer in a swimming synechococcus strain. J. Bacteriol. 187, 224–230 (2005).
Anwar, M. A. & Choi, S. Gram-negative marine bacteria: Structural features of lipopolysaccharides and their relevance for economically important diseases. Mar. Drugs. 12, 2485–2514 (2014).
DeMarco, M. L. Three-dimensional structure of glycolipids in biological membranes. Biochemistry 51, 5725–5732 (2012).
Kirschbaum, C. et al. Unravelling the structural complexity of glycolipids with cryogenic infrared spectroscopy. Nat. Commun. 12, 1201 (2021).
Azimzadeh Irani, M. Correlation between experimentally indicated and atomistically simulated roles of EGFR N-glycosylation. Mol. Simul. 44, 743–748 (2018).
Lee, J. et al. CHARMM-GUI membrane builder for complex biological membrane simulations with glycolipids and lipoglycans. J. Chem. Theory Comput. 15, 775–786 (2019).
Mikheyskaya, L. V., Ovodova, R. G. & Ovodov, Y. S. Isolation and characterization of lipopolysaccharides from cell walls of blue-green algae of the genus. Phormidium J. Bacteriol. 130, 1–3 (1977).
Buttke, T. M. & Ingram, L. O. Comparison of lipopolysaccharides from Agmenellum quadruplicatum to Escherichia coli and Salmonella typhimurium by using thin-layer chromatography. J. Bacteriol. 124, 1566–1573 (1975).
Weise, G., Drews, G., Jann, B. & Jann, K. Identification and analysis of a lipopolysaccharide in cell walls of the blue-green alga Anacystis nidulans. Arch. Microbiol. 71, 89–98 (1970).
Keleti, G. & Sykora, J. L. Production and properties of cyanobacterial endotoxins. Appl. Environ. Microbiol. 43, 104–109 (1982).
Keleti, G., Sykora, J. L., Lippy, E. C. & Shapiro, M. A. Composition and biological properties of lipopolysaccharides isolated from Schizothrix calcicola (Ag) Gomont (Cyanobacteria). Appl. Environ. Microbiol. 38, 471–477 (1979).
Weckesser, J., Katz, A., Drews, G., Mayer, H. & Fromme, I. Lipopolysaccharide containing L-acofriose in the filamentous blue-green alga Anabaena variabilis. J. Bacteriol. 120, 672–678 (1974).
Gemma, S., Molteni, M. & Rossetti, C. Lipopolysaccharides in cyanobacteria: A brief overview. Adv. Microbiol. 6, 391–397 (2016).
Paracini, N., Schneck, E., Imberty, A. & Micciulla, S. Lipopolysaccharides at solid and liquid interfaces: Models for biophysical studies of the gram-negative bacterial outer membrane. Adv. Colloid Interface Sci. 301, 102603 (2022).
Wang, B. et al. The combined effects of UV-C radiation and H2O2 on Microcystis aeruginosa, a bloom-forming cyanobacterium. Chemosphere 141, 34–43 (2015).
Phukan, T., Rai, A. N. & Syiem, M. B. Dose dependent variance in UV-C radiation induced effects on carbon and nitrogen metabolism in the cyanobacterium Nostoc muscorum Meg1. Ecotoxicol. Environ. Saf. 155, 171–179 (2018).
Hakkila, K. et al. Oxidative stress and photoinhibition can be separated in the cyanobacterium Synechocystis sp. PCC 6803. Biochimica et Biophysica Acta BBA Bioenerget. 137, 217–225 (2014).
Lupínková, L. & Komenda, J. Oxidative modifications of the Photosystem II D1 protein by reactive oxygen species: From isolated protein to cyanobacterial cells. Photochem. Photobiol. 79, 152–162 (2004).
Ye, T. et al. Exposure of cyanobacterium Nostoc sp. to the Mars-like stratosphere environment. J. Photochem. Photobiol. B. 224, 112307 (2021).
Apel, K. & Hirt, H. Reactive oxygen species: Metabolism, oxidative stress, and signal transduction. Annu. Rev. Plant Biol. 55, 373–399 (2004).
Rath, J. & Adhikary, S. Response of the estuarine cyanobacterium Lyngbya aestuarii to UV-B radiation. J. Appl. Phycol. 19, 529–536 (2007).
Shen, S. G. et al. The physiological responses of terrestrial cyanobacterium Nostoc flagelliforme to different intensities of ultraviolet-B radiation. RSC Adv. 8, 21065–21074 (2018).
Wang, G. et al. The involvement of the antioxidant system in protection of desert cyanobacterium Nostoc sp. against UV-B radiation and the effects of exogenous antioxidants. Ecotoxicol. Environ. Saf. 69, 150–157 (2008).
Author information
Authors and Affiliations
Contributions
P.I.: conceptualization, methodology, investigation, formal analysis, writing original draf, and editing, visualization. H.R.: conceptualization, methodology, resources, writing original draf, writing review and editing, supervision. S.H.: Conceptualization, methodology, investigation, formal analysis. M.E.: Investigation, molecular analysis, resources, writing original draf. M.A.: Investigation, methodology, molecular analysis, resources, writing original draf, writing review and editing. Z.S.: Conceptualization, methodology, resources, writing original draf, writing-review and editing, supervision.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Irankhahi, P., Riahi, H., Hassani, S.B. et al. The role of the protective shield against UV-C radiation and its molecular interactions in Nostoc species (Cyanobacteria). Sci Rep 14, 19258 (2024). https://doi.org/10.1038/s41598-024-70002-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-70002-8