Abstract
Antibodies are widely used as therapeutic agents to tackle various diseases. In the present study, to enhance their clinical values, we rationally designed pH-responsivity by exploiting the idiosyncratic protonation/deprotonation profiles of non-natural amino acids. 3-Nitro-l-tyrosine, 3-cyano-l-tyrosine, and 3, 5-halogenated-l-tyrosine, each with near neutral pKa, were thus incorporated into Fab fragments in place of tyrosines and other residues in the variable regions. Cell-based assays showed that these modifications achieved up to 140-fold tighter binding to antigens and several-fold tighter cytotoxicity to antigen-expressing cell at pH 6.0 than pH 7.4. The pH-dependent binding effect was retained in full-length antibodies. In silico structural analyses revealed electrostatic repulsion at neutral pH between antigens and antibodies or inside the antibody as the underlying mechanisms of the acid preference, and this finding increases the designability of pH-dependent antigen binding. The development of antibodies responsive to the microenvironments of diseased tissues will allow more disease-related antigens to be targeted in treatments, because of the reduced cross-reactivity toward healthy tissues.
Similar content being viewed by others
Introduction
To date, more than 100 monoclonal antibodies have been approved as therapeutics for the treatment of cancers, chronic diseases, and autoimmune disorders, while > 500 more antibodies are now under clinical trials1. The functions of antibodies, such as antibody-dependent cellular cytotoxicity and complement-dependent cytotoxicity, facilitate treatments, and the antigen-dependent targeting of disease-related cells is pivotal for the efficacy of antibodies. Although the importance of therapeutic antibodies to medical are rising, the number of therapeutic targets is limited because of their side effect. Actually, some approved therapeutics targeted same antigen such as B lymphocyte antigen CD20, epidermal growth factor receptor (EGFR), human epidermal growth factor 2 (HER2), etc.2. Most antigens are found across different cell types, and there are very few of them expressed in only disease-related cells. The cross-reactivity of antibodies with healthy tissues seriously limits the availability of antigens as therapeutic targets3,4. To overcome this problem, more precise targeting of cells is necessary. Bispecific antibodies have been developed to recognize the combinations of two different antigens5, while taking advantage of the idiosyncratic microenvironments surrounding diseased cells is also a promising direction6,7. For example, “ProBody” is one of the prodrug technologies focusing the cancer-specific protease. While “ProBody” is masked against antigen binding in normal tissue, it is locally activated by protease in tumor6. The extracellular environments of tumor tissues are often acidic, approaching pH 6.0, due to the high glycolytic activities of cancer cells8. Acidic microenvironments are also prevalent among inflammation and ischemia9. Low-pH targeting peptide, pH-sensitive liposomes were reported for diagnostic, surgical imaging and therapeutic agents10,11. Acidic condition in several disease states is getting more attention as an intriguing environmental factor and technology utilizing acidic condition would be needed. Antibodies showing pH-responsivity would facilitate precise targeting of cells in the treatments of these major illnesses.
The genetic encoding of non-natural amino acids allows the introduction of novel functionalities and chemical structures into proteins in designable manners. These amino acids are incorporated into proteins site-specifically, and the incorporation sites are defined by specific codons in their genes12,13. The synthesis of modified proteins in this manner guarantees site-specificity and homogeneity in modification among the products, which makes this technology suitable for biologics development. The substitution on position 3 of the tyrosine ring affects pKa of the 4-hydroxyl group in tyrosine derivatives. When the substituent is an electron attractor, the pKa is lower than that of tyrosine (10.1). 3-Nitro-l-tyrosine, 3-cyano-l-tyrosine, and 3, 5-halogenated l-tyrosine have the pKa values of around 7, and would work as a pH sensor over the desirable pH range from pH 6.0 to pH 7.4 (pKa is an ACD/Percepta calculation.). Environmentally responsive antibodies might thus be created by choosing appropriate sites for incorporating these tyrosines derivatives.
In the present study, we systematically searched for the sites where the incorporated tyrosine derivatives exerted pH-dependent binding activities. The variable regions of Fab fragments were examined first, and the reproducibility of the pH- dependent binding effects was confirmed in full-length antibodies. These analyses provide new knowledge of some criteria to exert pH dependency such as pKa of tyrosines derivatives and the incorporation site. The results showed that the 3-substituted tyrosine derivatives can work as a pH sensor in antibodies, and thus promise the development of therapeutic agents with reduced cross-reactivity toward healthy tissues.
Results
Effect of amino acid substitution from Tyr to tyrosine derivatives in Fabs on pH dependency
We first replaced tyrosine (Tyr) residues with 3-nitro-l-tyrosine in the CDR or framework regions of Trastuzumab-Fab (anti-Her2 Fab, Tra-Fab). These mutants showed the expected band sizes in sodium dodecyl sulfate polyacrylamide gel electrophoresis and the incorporation of synthetic amino acids into Fabs was confirmed by mass spectrometry (see Supplementary Fig. 1). The binding activities of the 3-nitro-l-tyrosine incorporated Tra-Fabs to A431 cells were analyzed by flow cytometry. The binding affinity ratio pH 7.4/pH 6.0 vs wild-type below 0.8 were set to have acidic pH dependency. Some mutants with amino acid substitutions in their CDR regions (Tra-Fab-H-Y33NY and Tra-Fab-H-Y56NY) showed the pH-dependent binding capabilities, while all mutants with amino acid substitutions in their framework regions (Tra-Fab-H-Y79NY, Tra-Fab-H-Y90NY, Tra-Fab-H-Y91NY, Tra-Fab-L-Y49NY, Tra-Fab-L-Y86NY and Tra-Fab-L-Y87NY) did not show any pH dependency (Fig. 1). These results suggested that the amino acid substitution from Tyr to tyrosine derivatives in CDR region could impart the pH-dependent binding activities, then we primarily focused on CDR region for further study.
Binding affinity ratio of pH 7.4 to pH 6.0 in nitro-tyrosine incorporated Trastuzumab Fab variants. Binding affinities of several nonnatural tyrosine-incorporated trastuzumab variants to SKBR3 cells at pH 6.0 and pH 7.4 were determined using flowcytometer (n = 2). The binding affinity ratio pH 7.4 to pH 6.0 represents the relative binding affinity with respect to wild-type. 1.0 was indicated by dashed line. The value below 0.8 are shown with * as acidic pH dependent value.
Importance of pKa values of tyrosine derivatives
To examine whether the pKa values of tyrosine derivatives affect the pH-dependent binding activity, several tyrosine derivatives such as 3-cyano-l-tyrosine (pKa value of around 7.0), and 3, 5-halogenated l-tyrosine (pKa value of around 7.0), 3-iode-l-tyrosine (pKa value of around 8.4), were incorporated into Tra-Fab CDR regions. As shown Fig. 1, Tra-Fab-H-Y33NY and Tra-Fab-H-Y56NY showed the pH-dependent binding capabilities, while Tra-Fab-H-Y79NY did not show any pH dependency. The position of Tyr33, Tyr56 were selected as positive incorporation site, and the position of Tyr79 was selected as negative incorporation site. 3-nitro-l-tyrosine, 3-cyano-l-tyrosine, and 3, 5-halogenated l-tyrosine showed almost the same pH-dependent binding phenomena when incorporated in Tra-Fab-H-Y33 and Tra-Fab-H-Y56, while 3-iode-l-tyrosine have less pH-dependency (Fig. 2). The results suggest that the pKa of Tyr derivative plays an important role in conferring the pH- dependency for Tra-Fabs.
Binding affinity ratio of pH 7.4 to pH 6.0 in several non-natural tyrosine-incorporated trastuzumab Fab variants. Binding affinities of several nonnatural tyrosine-incorporated trastuzumab variants to SKBR3 cells at pH 6.0 and pH 7.4 were determined using flowcytometer (n = 2). The binding affinity ratio pH 6.0 and pH 7.4 represents the relative binding affinity with respect to wild-type. 1.0 was indicated by dashed line. The value below 0.8 are shown with * as acidic pH dependent value.
Hypothetical mechanism of pH dependency elucidated by in silico analysis and its application to mutant design
To elucidate the hypothetical mechanism of pH dependency observed in Tra-Fab-H-Y33NY and Tra-Fab-H-Y56NY, in silico analysis was performed based on X-ray crystal structure of Tra -Fab – HER2 complex (PDB ID: 1N8Z). First, the residues in the Fab-antigen interface as well as the pairs of interaction were identified using inter-atomic distance smaller than 8.0 Å (see Supplementary Table 1). Fab-Tyr-interacting residues in antigen were, then, compared in the presence and absence of pH dependency upon introducing 3-nitro-l-tyrosine in the Fab. Interestingly, both of antibody tyrosines (Tyr33 and 56 in heavy chain) showing pH dependency have Glu558 in their proximity. Electrostatic potential of epitope was calculated using the same X-ray structure. Intriguingly, it shows that negatively charged patch composed of Glu558 and Asp 560 indeed exist in the interface with Tyr33 and Tyr56 (Fig. 3), implying that electrostatic repulsion at neutral pH could be a new hypothetical mechanism of pH dependency that was not described previously. No more promising Tra-Fab Tyr shows significant pH dependency being replaced by 3-nitro-l-tyrosine. Trp95 in heavy chain, however, exists in the interface with Glu558 and Asp560 in antigen. In addition, His91 in light chain is located near Asp 570 in antigen (Fig. 3). Although these are non-Tyr residues, there is a possibility that replacing them with 3-nitro-l-tyrosine may reveal the similar pH dependency observed in Tyr. To test the hypothesis, two non-Tyr residues (His91 in light chain and Trp95 in heavy chain) were selected in Tra-Fab. We replaced His91 in light chain and Trp95 in heavy chain of Tra-Fab with 3-nitro-l-tyrosine. The pH-dependent binding to SKBR3 cells was experimentally validated in 3-nitro-l-tyrosine incorporated Tra-Fab mutants (Tra-Fab-L-H91NY and Tra-Fab-H-W95NY) (Fig. 4 a). Supplementary Table 1 summarized the list of antibody residues showing pH-dependency through 3-nitro-l-tyrosine substitution, and interaction pairs in antibody-antigen interface. The list of mutants showing pH-dependency was shown in Supplementary Table 2.
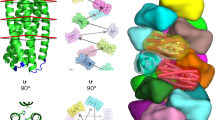
Electrostatic potential of HER2 (antigen) within 6 Å from trastuzumab-Fab. Electrostatic potential was calculated and displayed to characterize the surface property of the epitope. The scale of the potential is set from − 40 kcal/mol/e (red) to + 40 kcal/mol/e (blue). Tyr33, Tyr56, and Trp95 in heavy chain of Trastuzumab-Fab are located near negatively charged patch composed of Glu558 and Asp560 in HER2 (indicated by yellow circle). His91 in light chain of Trastuzumab-Fab is located near Asp570 in HER2 (indicated by green circle). Introducing 3-nitro-l-tyrone to each of the four residues in Trastuzumab-Fab reveals pH-dependent binding characteristics to HER2 presumably due to the electrostatic repulsion between antibody and antigen.
Binding affinity ratio of pH 7.4 to pH 6.0 in nitro-tyrosine-incorporated Trastuzumab Fab variants (a) and anti-Her2 antibody (Clone, 3wlw) Fab variant (b). Binding affinities of nitro tyrosine-incorporated trastuzumab variants to SKBR3 cells at pH 6.0 and pH 7.4 were determined using flowcytometer (n = 2). Binding affinity ratio of pH 7.4 to pH 6.0 represents relative binding affinity with respect to wild-type. 1.0 was indicated by dashed line. The value below 0.8 are shown with * as acidic pH dependent value.
So far pH-dependency was identified only when incorporating 3-nitro-l-tyrosine into Tra-Fab. We explored the possibility of introducing 3-nitro-l-tyrosine into other antibody-Fab besides Tra-Fab. Based on the X-ray crystal structures of anti-HER2 antibody-Fab-HER2 complex (PDB ID: 3WLW), 3-nitro-l-tyrosine was incorporated into anti-Her2 antibody-Fab in the paratope adjacent to acidic amino acid in the epitope. To test the hypothesis, one Tyr residue was selected in anti-Her2 antibody-Fab. The pH-dependent binding to SKBR3 cells was detected in 3-nitro-l-tyrosine incorporated anti-Her2 antibody-Fab mutant (3wlw-Fab-L-Y91NY) (Fig. 4b). Therefore, the range of application of this technology such as the incorporation site and kind of antibody could be expanded.
Versatility of pH dependency by incorporating tyrosine derivatives into several Fabs
Rituximab-Fab (anti-CD20 Fab, 2OSL-Fab), obinutuzumab-Fab (anti-CD20 Fab, 3PP4-Fab) and avelumab-Fab (Anti-PD-L1 Fab, 5GRJ-Fab) were used to confirm the versatility of pH dependency by incorporating tyrosine derivatives into Fabs. We incorporated 3-nitro-l-tyrosine into these three Fabs at the position from acidic amino acid of epitope to 8.0 Å or less or in place of Tyr (see Supplementary Table 1). The binding profiles of these modified Fabs to each antigen expressing cell were analyzed by flow cytometry. For quantitative evaluation, the mean EC50 values of binding affinity were calculated from the flow cytometry date using 4-parameter logistic model. EC50 ratio of binding affinity determined at pH 6.0 and pH 7.4 (or pH 8.0) of some modified Fabs were several times higher than that of wild-type Fabs (Tables 1, 2, 3). Especially, 2OSL-Fab-H-Y97NY, 3PP4-Fab-L-Y96NY and 5GRJ-Fab-L-Y91NY showed the significant pH-dependent binding. Relative EC50 ratio of the binding affinity at pH 6.0 and pH 7.4 compared to wild-type were 5.6, 48 and 140 for 2OSL-Fab-H-Y97NY, 3PP4-Fab-L-Y96NY and 5GRJ-Fab-L-Y91NY, respectively. These results showed that the strategy of conferring the pH-dependency to Fabs via rational incorporation of tyrosine derivative was reproducible.
Binding affinities of NO2-tyrosine-incorporated rituximab variants to Raji cells at pH 6.0, pH 7.4 and pH 8.0 were determined using flowcytometer. The mean EC50 values (n = 2) were calculated from the data set of concentrations and corresponding measured values using 4-parametoer logistic model. EC50 of rituximab variants and their pH 7.4 (pH 8.0) to pH 6.0 EC50 ratio to wild-type are shown.
Binding affinities of NO2-tyrosine-incorporated obinutuzumab variants to Raji cells at pH 6.0, pH 7.4 and pH 8.0 were determined using flowcytometer. The mean EC50 values (n = 2) were calculated from the data set of concentrations and corresponding measured values using 4-parametoer logistic model. EC50 of rituximab variants and their pH 7.4 (pH 8.0) to pH 6.0 EC50 ratio to wild-type are shown.
Binding affinities of NO2-tyrosine-incorporated avelumab variants to A431 cells at pH 6.0, pH 7.4 and pH 8.0 were determined using flowcytometer. The mean EC50 values (n = 2) were calculated from the data set of concentrations and corresponding measured values using 4-parametoer logistic model. EC50 of rituximab variants and their pH 7.4 (pH 8.0) to pH 6.0 EC50 ratio to wild-type are shown.
Additional potential mechanisms of pH-dependency observed in anti-PD-L1 and anti-CD20 Fabs via in silico analyses
To explore the underlying mechanisms of the remarkable pH dependency of 2OSL-Fab-H-Y97NY, 3PP4-Fab-L-Y96NY, and 5GRJ-Fab-L-Y91NY, further in silico analyses were conducted. The electrostatic potential of paratope in 5GRJ-Fab-L-Y91NY assumes more negative charge at pH 7.4 than pH 6.0 (Fig. 5a,b). This seems to be the same mechanism as one observed in anti-HER2 Fabs. It, however, showed tremendous pH dependency like 140-fold possibly due to the formation of acidic amino acid cluster composed of Glu58, Glu60, and Asp61 in PD-L1 at the interface with tyrosine derivative in Fab (Fig. 5c). In contrast, as for 2OSL-Fab-H-Y97NY and 3PP4-Fab-L-Y96NY, no acidic amino acids exist in antigen within 12 Å from each of Fabs. Thus, different types of mechanism seem to be involved to induce pH dependency. In silico analysis implies that within the 2OSL-Fab-H-Y97NY Asp100A in heavy chain forms hydrogen bonds with His34 in light chain that may be disrupted by introducing 3-nitro-l-tyrosine into Tyr97 in heavy chain especially at pH 7.4 (Fig. 6a). In silico analysis also identified that chloride ion near Tyr96 in light chain seems to stabilize the interaction between wild type of 3PP4-Fab and CD-20 (Fig. 6b). Substituting Tyr96 with 3-nitro-l-tyrosine may exclude the crucial chloride ion in Fab-antigen interface at pH 7.4, thereby leading to the striking pH-dependent decrease in antigen binding at pH 7.4.
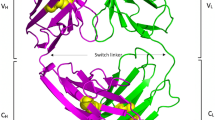
Characteristic of antibody (variant 6 of 5GRJ-Fab)-antigen (PD-L1) interface revealing the remarkable pH dependency of 140-fold. (a) Electrostatic potential of paratope at pH 6.0. (b) Electrostatic potential of paratope at pH 7.4. 3-nitro-l-tyrosine is indicated by yellow circle. The scale of the potential is set from − 20 kcal/mol/e (red) to + 20 kcal/mol/e (blue). Antibody heavy/light chains and antigen are shown in magenta, cyan, and green respectively. Antigen is just shown for structural guidance at pH 7.4. (c) Cluster of acidic amino acids (Glu58, Glu60, and Asp61) in PD-L1 putatively close to nitro Tyr91 in light chain of variant 6 of 5GRJ-Fab) at pH 6.0. Strong electrostatic repulsion may occur between antibody and antigen at pH 7.4, potentially leading to the tremendous pH dependency like 140-fold.
Illustrations of environment around Tyr to reveal pH dependency upon introducing 3-nitro-l-tyrosine possibly through different mechanism from inter-molecular electrostatic repulsion between antibody and antigen. (a) Environment around Tyr97 in heavy chain of 2OSL-Fab. Asp100A forms hydrogen bonds with His34 that may be disrupted upon introducing 3-nitro-l-tyrosine. (b) Environment around Tyr96 in light chain of 3PP4-Fab. Chloride ion near Tyr96 seems to stabilize the interaction between 3PP4-Fab and CD20. Introducing 3-nitro-l-tyrosine to Tyr96 may exclude chloride ion at pH 7.4, resulting in the great decrease in antigen binding. Antibody heavy/light chains and antigen are shown in magenta, cyan, and green respectively. CDR loops in heavy and light chains were shown in red and blue respectively. Atoms involved in the antibody-antigen interaction are shown in purple bubbles.
pH-dependent binding observed in the tyrosine derivatives-incorporated full-length antibodies
We made the 3-cyano-l-tyrosine incorporated 3wlw-Mab (3wlw-Mab-L-Y91CNY) with the same amino acid substitution as 3wlw-Fab-L-Y91CNY and measured the binding activities to SKBR3 cells. EC50 ratio of binding affinity at pH 6.0 and pH 7.4 of 3wlw-Mab-L-Y91CNY was almost the same as 3wlw-Fab-L-Y91CNY (Table 4). This suggests that the pH-dependent binding effect is also confirmed in the full-length antibodies.
Binding affinities of CN-tyrosine-incorporated anti HER2 antibody (Clone, 3wlw) variants to SKBR3 cells at pH 6.0, pH 7.4 and pH 8.0 were determined using flowcytometer. The mean EC50 values (n = 2) were calculated from the data set of concentrations and corresponding measured values using 4-parametoer logistic model. N.D.: not determined.
Cytotoxicity of tyrosine derivatives incorporated anti-CD20 Fab
Obinutuzumab is known to induce direct cell death in tumor cells14. To address the pH-dependent cytotoxicity of 3PP4-Fab-L-Y96NY, we measured the apoptosis-inducing activity in Raji cells at several pH. Intriguingly, 3PP4-Fab-L-Y96NY showed the pH-dependent apoptosis-inducing capability (Fig. 7). Relative EC50 ratio of the cytotoxicity at pH 6.0 to pH 7.4 (or pH 8.0) in 3PP4-Fab-L-Y96NY was several times higher than that of wild-type Fab (Table 5). These results suggest that the incorporation of non-natural tyrosine into Fab could show the pH-dependency not only in binding profile but also in functional activity.
Effect of nitro-tyrosine-incorporation into obinutuzumab variants on initiation of apoptosis in vitro. Representative flow cytometry histogram plot showed the effect of wild-type (closed circle) and nitro-tyrosine-incorporated obinutuzumab Fab variant (open circle) on apoptosis induction in pH 6.0 (a), pH 7.4 (b) or pH 8.0 (c).
Apoptosis-inducing activities of NO2-tyrosine-incorporated obinutuzumab variants to Raji cells at pH 6.0, pH 7.4 and pH 8.0 were determined using flowcytometer. The mean EC50 values (n = 2) were calculated from the data set of concentrations and corresponding measured values using 4-parametoer logistic model.
Discussion
Tyr is one of the most frequent amino acids in the third complementarity-determining region of the heavy chain (CDR-H3) that often determine the antigen binding and specificity15. It has multiple modes of interaction such as hydrogen bond, hydrophobic, van der Waals, aromatic hydrogen bond and stacking. In this study, we focused on the tyrosine derivatives with pH sensitivity added to the multiple functions of natural Tyr. As expected, the rational incorporation of these tyrosine derivatives added pH-sensitive activity to the conventional antibodies. In silico structural analyses revealed the following three hypothetical mechanisms of the pH-dependency: (A) controlling hydrogen bond of Tyr; (B) “intermolecular” electrostatic repulsion; (C) “intramolecular” electrostatic repulsion) (Fig. 8). In addition, this study shows that using 3D structural information of antibody-antigen complex was a key to increase the success rate when selecting the sites of incorporation of 3-nitro-l-tyrosine into three antibodies to induce pH-dependency.
Dan et al. reported that the stochastically nitrated antibodies by tetranitromethane (TNM) showed pH-dependent binding (decreased at pH > 8, restored at pH < 6) with the same mechanism as (A)7. From therapeutic standpoint, our rational and specific incorporation is superior to their probabilistic approach. Stochastic modification is not desirable for the therapeutic application because it may lead to the heterogeneous molecules, each of which having different efficacies, pharmacokinetics, and therapeutic windows. Indeed, the randomly nitrated antibodies were heterogeneous and had significantly decreased binding affinity (see Supplementary Fig. 2). Furthermore, whereas only the amino acid of Tyr could be nitrated probabilistically, the amino acid near the acidic epitope besides Tyr could be selected as the incorporation site in site-specific incorporation strategy, leading to the expanded feasibility. The approach based on “intermolecular” electrostatic repulsion of positive charge between antibody and antigen was reported previously by Igawa et al.16. They replaced amino acids in variable region with histidine (His) and produced the neutral preferable antibodies that was the opposite to our acid preferable antibodies based on mechanism B. As the neutral preferable antibodies change their intracellular dynamics, it would be the clinically beneficial for efficient depletion of soluble factor, increased lysosomal delivery of payload of ADC, and reduction of side effect by inhibition of degradation in endosome16,17,18. Their approach, however, cannot target disease-related acidic condition due to their decreased binding affinity at acidic condition.
It was also reported that His-mediated “intramolecular” electrostatic repulsion improved the sensitivity of protein G, affinity ligand for Immunoglobulin G (IgG)19. At acidic conditions, His introduced into Protein G causes “intramolecular” electrostatic repulsion between positive charge, thereby disrupting the interaction within Protein G and decreasing the binding affinity to IgG by the similar mechanism to C. Although this His-mediated “intramolecular” electrostatic repulsion strategy is worth in some cases such as mild purification of IgG, it is difficult to apply to targeting acidic site.
Recent investigations have demonstrated that the improvement of binding at acidic pH relative to physiological pH was achieved by His mutagenesis20,21. Introduced positively charged His was favorable interaction with negatively charged amino acids at acidic condition while it could not form salt bridge to negatively charged amino acids at neutral condition. But this His-mediated strategy might be applied to the antibody having somewhat weak or no binding activity at physiological condition, because introduced His could strengthen the binding at acidic pH but could not weaken the binding at neutral pH.
In summary, we rationally designed acidic preferable antibodies with pH-sensing tyrosine derivatives by in silico analyses. Although it remains to be how effectively our strategy can work in vivo, we believe that acidic preferable binding with reduced cross-reactivity toward healthy tissues should possibly decrease the side effect and/or improve the pharmacokinetics of currently approved antibody. Furthermore, it redevelops the antibody failed in past clinical trials, and develop the antibody which is thought to be difficult to develop because of systemic expression of recognized antigen. To achieve ideal therapeutic antibodies described above, further studies such as in vivo test and optimization of binding affinity should be needed.
Materials and methods
Reagents and cells
3-Nitro-l-tyrosine and 3-iode-l-tyrosine were purchased from SIGMA (MO, USA). 3-cyano-l-tyrosine and 3-fluoro-5-fluoro l-tyrosine were obtained from NetChem (Edenvale, South Africa) and PepTech (MA, USA), respectively. Human adenocarcinoma (SK-BR-3) cell lines, human epidermoid carcinoma (A431) cell lines and human Burkitt’s lymphoma (Raji) cell lines were obtained from ATCC (VA, USA).
Cell culture
SK-BR-3 cells and Raji cells were cultured at 37 °C with 5% CO2 in RPMI1640 (Nacalai tesqu, Kyoto, JAPAN) supplemented with 10% heat-inactivated fetal bovine serum and 50 μg/mL gentamicin (Nacalai tesqu). A431 cells were cultured at 37 °C with 5% CO2 in DMEM, high glucose (Thermo Fisher Scientific, MA, USA) supplemented with 10% heat-inactivated fetal bovine serum and 50 μg/mL gentamicin (Nacalai tesqu).
Construction, expression, and purification of antibodies
The genes encoding the heavy- and light-chain variable regions of the anti-Her2 antibodies (trastuzumab and 3wlw), anti-CD20 antibodies (rituximab and obinutuzumab), and anti-PD-L1 antibody (avelumab) were cloned into E. coli or mammalian expression vector. Each gene of antibodies had an amber codon for the site-specific incorporation of tyrosine derivatives into antibodies. These expression vectors were introduced into RF-1-knockout E. coli cells or CHO-s cells with the vector having UAG-reading tRNA and tyrosyl-tRNA synthetase (TyrRS) variant specific for each tyrosine derivative, and the tyrosine derivative incorporated antibodies were produced and purified as previously reported12,13. The purity of modified antibodies and the incorporation of tyrosine derivatives were examined by sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) and mass spectrometry analysis, respectively.
Measurement of each modified antibody binding affinity
The antigen-binding capability of each modified antibody was measured using flow cytometry. SK-BR-3 cells, Raji cells and A431 cells were stained with anti-Her2 antibodies, anti-CD20 antibodies and anti-PD-L1 antibodies, respectively, at 4 °C for 30 min in pH 6.0, pH 7.4 or pH 8.0 phosphate buffered saline (PBS) containing 1% bovine serum albumin (BSA). Fluorescein isothiocyanate (FITC)-conjugated anti-human IgG (H + L) (R&D systems, MN, USA) was used as the secondary antibody to detect the cell-bound modified antibodies. The stained cells were then analyzed with a BD FACSVerse (BD Biosciences, CA, USA). The mean EC50 values were calculated from the data set of concentrations and corresponding measured values using 4-parametoer logistic model.
Model construction and computational methods for in silico analysis
Original X-ray crystal structures were obtained from the Protein Data Bank (PDB): anti-HER2 antibody Fab-HER2 complexes (PDB ID: 1N8Z, 3WLW), anti-CD20 antibody Fab-CD20 complexes (PDB ID: 2OSL, 3BKY, 3PP4), and anti-PD-L1 antibody Fab-PD-L1 complexes (PDB ID: 5X8L, 5X8M, 5GRJ). The following computational procedures were performed in MOE 2018.010122. In each structure, missing residues and atoms were restored, and hydrogen atoms were added by using Protonate3D module at pH 7.4, 298 K, and 150 mM ionic strength. Atomic partial charge was assigned based on AMBER10:EHT force field. Dielectric constant for protein was set to 2. The aqueous environment was treated implicitly by utilizing the generalized Born/volume integration (GB/VI) method23 with dielectric constant of 80. Electrostatic potential was calculated by Poisson Boltzmann solver in MOE, and visualized on the molecular surface. 3-Nitro-tyrosine incorporated mutants were prepared by either introducing nitro-group into Tyr or substituting non-Tyr with Tyr then adding nitro-group. Inter- or Intra-atomic distances were measured between phenolic oxygen in 3-Nitro-tyrosine and carboxyl oxygen in Asp/Glu, whereas the equivalent distances were measured upon substituting non-Tyr, such as His and Phe, with Tyr. As for the sites of antibody revealing pH-dependency when incorporating 3-Nitro-tyrosine, the lists of interaction pairs between antibody and antigen within 8 Å were created with respect to the original residues in antibodies. In this study, inter-atomic distance below 12 Å was considered as it is a reasonable cutoff distance for non-bonded interactions for biomacromolecules.
Evaluation of modified anti- CD20 antibody cytotoxicity
The cytotoxic activity of anti-CD20 antibody Fabs was evaluated using flow cytometer. Raji cells were incubated with the Fabs cross-linked by anti-Human Kappa Light Chain Antibody (Bethyl Laboratories, TX, USA) at 37 °C for 24 h in pH 6.0, pH 7,4 or pH 8.0 medium (45% RPMI-1640 (Thermo Fisher Scientific, MA, USA), 45% CO2-independent Medium (Thermo Fisher Scientific, MA, USA), 10% FBS, 10 mM HEPES). After the incubation, the cells were washed with cold phosphate-buffered saline (PBS) and cold binding buffer (10 mM HEPES, 140 mM NaCl, and 2.5 mM CaCl2, pH 7.4) and then stained with Annexin V-Alexa Fluor 488 conjugate (Molecular Probes, OR, USA) for 15 min in dark. The stained cells were analyzed with a BD FACSVerse (BD Biosciences, CA, USA). The mean EC50 values were calculated as described above.
Data availability
All of the data and information from this study are available from the corresponding authors on reasonable request.
References
Kaplon, H., Chenoweth, A., Crescioli, S. & Reichert, J. M. Antibodies to watch in 2022. MAbs 14(1), 2014296. https://doi.org/10.1080/19420862.2021.2014296 (2022).
Carter, P. J. & Lazar, G. A. Next generation antibody drugs: Pursuit of the “high-hanging fruit”. Nat. Rev. Drug Discov. 17, 197–223. https://doi.org/10.1038/nrd.2017.227 (2018).
Münz, M. et al. Side-by-side analysis of five clinically tested anti-EpCAM monoclonal antibodies. Cancer Cell Int. 10, 44. https://doi.org/10.1186/1475-2867-10-44 (2010).
Segaert, S. & Van Cutsem, E. Clinical signs, pathophysiology and management of skin toxicity during therapy with epidermal growth factor receptor inhibitors. Ann. Oncol. 16, 1425–1433. https://doi.org/10.1093/annonc/mdi279 (2005).
Labrijn, A. F., Janmaat, M. L., Reichert, J. M. & Parren, P. Bispecific antibodies: A mechanistic review of the pipeline. Nat. Rev. Drug Discov. 18, 585–608. https://doi.org/10.1038/s41573-019-0028-1 (2019).
Desnoyers, L. R. et al. Tumor-specific activation of an EGFR-targeting probody enhances therapeutic index. Sci. Transl. Med. 5, 207ra144. https://doi.org/10.1126/scitranslmed.3006682 (2013).
Tawfik, D. S., Chap, R., Eshhar, Z. & Green, B. S. pH on-off switching of antibody-hapten binding by site-specific chemical modification of tyrosine. Protein Eng. 7, 431–434. https://doi.org/10.1093/protein/7.3.431 (1994).
Gatenby, R. A. & Gillies, R. J. Why do cancers have high aerobic glycolysis?. Nat. Rev. Cancer 4, 891–899. https://doi.org/10.1038/nrc1478 (2004).
Riemann, A. et al. Acidic environment activates inflammatory programs in fibroblasts via a cAMP-MAPK pathway. Biochim. Biophys. Acta 1853, 299–307. https://doi.org/10.1016/j.bbamcr.2014.11.022 (2015).
Deacon, J. C., Engelman, D. M. & Barrera, F. N. Targeting acidity in diseased tissues: Mechanism and applications of the membrane-inserting peptide, pHLIP. Arch. Biochem. Biophys. 565, 40–48. https://doi.org/10.1016/j.abb.2014.11.002 (2015).
Paliwal, S. R., Paliwal, R. & Vyas, S. P. A review of mechanistic insight and application of pH-sensitive liposomes in drug delivery. Drug Deliv. 22, 231–242. https://doi.org/10.3109/10717544.2014.882469 (2015).
Hino, N., Hayashi, A., Sakamoto, K. & Yokoyama, S. Site-specific incorporation of non-natural amino acids into proteins in mammalian cells with an expanded genetic code. Nat. Protoc. 1, 2957–2962. https://doi.org/10.1038/nprot.2006.424 (2006).
Mukai, T. et al. Highly reproductive Escherichia coli cells with no specific assignment to the UAG codon. Sci. Rep. 5, 9699. https://doi.org/10.1038/srep09699 (2015).
Herter, S. et al. Preclinical activity of the type II CD20 antibody GA101 (obinutuzumab) compared with rituximab and ofatumumab in vitro and in xenograft models. Mol. Cancer Ther. 12, 2031–2042. https://doi.org/10.1158/1535-7163.mct-12-1182 (2013).
Zemlin, M. et al. Expressed murine and human CDR-H3 intervals of equal length exhibit distinct repertoires that differ in their amino acid composition and predicted range of structures. J. Mol. Biol. 334, 733–749. https://doi.org/10.1016/j.jmb.2003.10.007 (2003).
Igawa, T. et al. Antibody recycling by engineered pH-dependent antigen binding improves the duration of antigen neutralization. Nat. Biotechnol. 28, 1203–1207. https://doi.org/10.1038/nbt.1691 (2010).
Kang, J. C. et al. Engineering a HER2-specific antibody-drug conjugate to increase lysosomal delivery and therapeutic efficacy. Nat. Biotechnol. 37, 523–526. https://doi.org/10.1038/s41587-019-0073-7 (2019).
Zhang, Y. et al. Hijacking antibody-induced CTLA-4 lysosomal degradation for safer and more effective cancer immunotherapy. Cell Res. 29, 609–627. https://doi.org/10.1038/s41422-019-0184-1 (2019).
Watanabe, H. et al. Histidine-mediated intramolecular electrostatic repulsion for controlling pH-dependent protein–protein interaction. ACS Chem. Biol. 14, 2729–2736. https://doi.org/10.1021/acschembio.9b00652 (2019).
Sulea, T. et al. Structure-based engineering of pH-dependent antibody binding for selective targeting of solid-tumor microenvironment. MAbs 12, 1682866. https://doi.org/10.1080/19420862.2019.1682866 (2020).
Johnston, R. J. et al. VISTA is an acidic pH-selective ligand for PSGL-1. Nature 574, 565–570. https://doi.org/10.1038/s41586-019-1674-5 (2019).
Molecular Operating Environment (MOE). Chemical Computing Group ULC (2018).
Labute, P. The generalized Born/volume integral implicit solvent model: Estimation of the free energy of hydration using London dispersion instead of atomic surface area. J. Comput. Chem. 29, 1693–1698. https://doi.org/10.1002/jcc.20933 (2008).
Author information
Authors and Affiliations
Contributions
Y.I and K.O. contributed equally to this work. M. I, K. M., K. S and Y. S. conceived the study. Y.I, K.O., W.P., F. K. and A. U. conducted experiments, collected and analyzed data. T. O. contributed to in silico data analysis and interpretation. Y.I, K.O., W.P., F.K., K. S. and Y.S. drafted the manuscript. All authors critically revised the manuscript and approved the final version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Isoda, Y., Ohtake, K., Piao, W. et al. Rational design of environmentally responsive antibodies with pH-sensing synthetic amino acids. Sci Rep 14, 19428 (2024). https://doi.org/10.1038/s41598-024-70271-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-70271-3