Abstract
Metal ion-DNA interactions play a crucial role in modulating the structure and function of genetic material in the natural environment. In this study, we report on the favorable electrochemical activity of holmium(III) (Ho3+) on a glassy carbon electrode (GCE) and its interaction with double-stranded DNA. The interaction between DNA and Ho3+ was investigated for the first time using cyclic voltammetry and differential pulse voltammetry. The electrochemical behavior of Ho3+ ions on a GCE exhibited a reversible electron transfer process, indicative of its redox activity. A linear correlation between the peak current and the square root of the scan rate was observed, suggesting a diffusion-controlled kinetic regime for the electrochemical process. Additionally, fluorescence and absorption spectroscopy were employed to confirm the binding of Ho3+ to DNA. Our findings demonstrate that, at pH 7.2, specific DNA bases and phosphate groups can interact with Ho3+ ions. Moreover, electrochemical measurements suggest that Ho3+ ions bind to DNA via a groove binding mode, with a calculated binding ratio of 1:1 between Ho3+ and DNA. Notably, under optimal conditions, an increase in the amount of DNA leads to a significant reduction in the current intensity of Ho3+ ions.
Similar content being viewed by others
Introduction
Interactions between metal ions and DNA have been implicated in the development of various diseases1,2,3,4. Metal ions, possessing positive charges, exhibit reactivity towards sites with high electron density. The presence of such sites on DNA molecules enables the identification of potential metal binding sites without the need for experimental investigations. These high-electron density sites encompass N1, N3, and NH2 in adenine; N3, N7, and NH2 in guanine; and N1 and O7 in thymine and cytosine. Within double-stranded DNA, only N7 of guanine and adenine, as well as O7 of thymine, are accessible for binding, with N7 being the most reactive center5.
Studies on metal ion-DNA interactions have been conducted at two levels: the addition of metal ions to natural DNA preparations and the investigation of resulting metal-DNA complexes, as well as the identification of metal ions already present in DNA molecules extracted from various biological materials5,6. Although the biochemical aspects of the toxicity of heavy metal ions and their cellular uptake mechanisms remain largely unknown, specific metal ion-DNA complexes' existence and properties may contribute to the toxic effects associated with these metal ions5. Recent investigations have unveiled three distinct binding modes to DNA: (1) interstrand penetration, (2) inter-groove binding, and (3) external binding through aggregate formation. External binding can occur throughout the double-stranded DNA structure, while interstrand binding primarily occurs in regions rich in guanine-cytosine bases and at the connections between grooves in regions rich in adenine–thymine bases (Fig. 1)7,8.
The binding of metal ions to DNA poses a complex scenario, giving rise to numerous challenges, particularly when metal ions are associated with bases. Based on their properties, metal ions can be roughly categorized into three classes: (1) those that exclusively bind to phosphate groups, such as alkali metals (Na+, K+, etc.) and alkaline earth metals (Ca2+, Mg2+); (2) those that exclusively bind to bases, including Ag+ and Hg2+; and (3) those that bind to both phosphates and bases. Metal ions, in the sequence Co2+ < Ni2+ < Zn2+ < Mn2+ < Cd2+ < Cu2+, have been reported to exhibit an increasing affinity for bases over phosphate sites9,10.
The lanthanide series demonstrates significant bioinorganic similarity to Ca2+, with ionic radii that are nearly equivalent but higher charge density. This similarity raises concerns regarding the displacement of calcium in physiological environments, especially in various therapeutic and diagnostic applications11. Lanthanides are inner transition elements characterized by the filling of 4f. orbitals, including La itself with the electron shell configuration La:[Xe]4f05d16s2. A distinctive feature of the lanthanide series is the contraction of ionic radii across the series12. Extensive investigations have been conducted to understand the interaction of these metal ions with DNA, employing various methods such as luminescent techniques13,14,15, UV–Vis spectroscopy16, NMR17,18, and fluorescence spectroscopy19. However, the evidence for metal ion-DNA binding remains somewhat indirect and subject to ongoing discussion. In recent times, modified electrodes have been employed to monitor DNA interactions or other molecules using electrochemical techniques20,21,22,23. The binding of a metal to DNA is frequently observed through alterations in the redox process (electrochemical behavior) of the metal in the presence and absence of DNA. These changes typically include shifts in the formal potential of the redox couple and a decrease in peak current, resulting from a sudden decrease in the diffusion coefficient after binding to DNA24,25. For instance, cyclic voltammetry, differential pulse voltammetry, chronocoulometry, and electro-generated chemiluminescence have been employed to study the chelation of Eu(III)2, Yb(III)26, Sm(III)27, Co(III)28, Fe(II)29, Ru(II)30, and Os(II)31 complexes with 1,10-phenanthroline and 2,2'-bipyridyl bound to DNA. Maeda et al.32 explored the binding of K+, Na+, Ca2+, Mg2+, and Ba2+ to native DNA on DNA-modified gold electrodes, prepared through self-assembly of DNA with a mercapto group, using indirect electrochemical methods based on their interactions with DNA. Their findings revealed the binding order of these metal ions to DNA-modified electrodes as Mg2+, Ca2+, Ba2+, K+, and Na+. Barton and colleagues34 reported on a gold electrode derivative with 15-base-pair double-stranded DNA oligonucleotides containing a pendant 5'-hexanethiol linker.
The successful use of lanthanides and lanthanide complexes as electron shuttles for oxidoreductase enzymes has encouraged some researchers to investigate the biological activity of these compounds. These complexes have shown slight IC50 concentrations against various cancer cells such as A-172, HCT-116, and RDM-4. These results indicate that lanthanides and lanthanide complexes could be suitable alternatives to cisplatin as anticancer drugs, at least for those cancer cells that have shown resistance to cisplatin. Lanthanides interact with DNA in laboratory conditions and cause DNA damage. Therefore, they could provide a new chemotherapeutic method or complementary anticancer agent. Cerium, lanthanum, and neodymium coumarin complexes have been investigated for this purpose, and preliminary preclinical studies have shown their inhibitory properties against brain cancer cells34,35.
Understanding the mechanism of DNA-metal interactions holds crucial importance in biological studies encompassing drug design and pharmaceutical development processes. Voltammetry techniques prove highly valuable for this purpose, as they enable the investigation of DNA-metal interactions and the assessment of changes in the electronic properties induced by metal association with DNA. Such investigations are of fundamental significance in the fields of biochemistry and analytical biochemistry36.
Holmium ([Xe]4f116s2) is naturally found in association with other rare-earth metals, such as thulium, and is classified as a relatively scarce lanthanide element, comprising only 1.4 parts per million of the Earth's crust. Its abundance is comparable to that of tungsten37. In recent years, there has been an increasing interest in holmium due to its emerging applications in various technologies, including catalyst production, glass polishing, and biological systems, prompting extensive investigations into its properties38.
To the best of our knowledge, no previous studies have explored the electrochemical behavior of holmium for investigating its interactions with DNA using electrochemical methods. In this study, we aimed to investigate the electrochemical behavior of Ho3+ ions and their interaction with human DNA using cyclic voltammetry and differential pulse voltammetry. The obtained results were further confirmed through UV–Vis and fluorescence spectroscopy.
Experimental section
Instrumentation
Cyclic voltammetry and differential pulse voltammetry measurements were performed using the PalmSens Electroanalyzer system. All voltammetric experiments were conducted at room temperature (25 ± 2 °C) employing a three-electrode setup comprising a glassy carbon electrode (GCE) as the working electrode, a platinum wire as the counter electrode, and Ag/AgCl/KCl 3 mol L–1 as the reference electrode. Prior to usage, the GCE underwent polishing with 0.3 mm alumina powder on a polishing cloth, followed by thorough washing with water.
Fluorescence emission and excitation measurements were carried out using a Perkin Elmer luminescence spectrometer, utilizing a high-pressure xenon lamp as the light source. The instrument exhibited no detectable drift over a 4 h measuring period and demonstrated low levels of noise at the sensitivities employed. Spectrophotometric determinations were conducted using a Shimadzu UV–Vis spectrophotometer equipped with a 1 cm path cell. pH measurements were performed using a Metrohm AG digital pH-meter (Model 913) that underwent appropriate standardization with 0.05 mol L–1 potassium hydrogen phthalate (pH 4.0 at 25 °C) and 0.01 mol L–1 borax (pH 9.2 at 25 °C).
Reagents
High-purity HoCl3·6H2O (powder, 99.9%) was obtained from Sigma-Aldrich. A stock solution of Ho3+ (1.0 × 10−2 mol L–1) was prepared by dissolving 0.2713 g of HoCl3·6H2O in a minimum volume of Tris–HCl buffer (0.01 mol L–1, obtained from Sigma-Aldrich) and subsequently diluting it with double-distilled water in a 100 mL volumetric flask. Tris–HCl buffer (pH 7.2) containing 5 × 10–3 mol L–1 Tris–HCl and 5 × 10–2 mol L–1 NaCl was utilized in all experiments investigating the interaction of holmium with DNA.
Human genome extraction method
The extraction of DNA is a fundamental and indispensable technique in DNA studies, with whole blood samples being a primary source for DNA isolation. Biological samples of blood were obtained from the research team working on this project. The laboratory protocols were approved by the Ethics Committee of the University of Guilan prior to the commencement of the study, in accordance with the regulations published regarding clinical studies involving humans and laboratory animals on 17, Feb, 2021. This research project is under the oversight of the ethics committee with the number 2407-1192 in the Biomedical Research Committee of the University of Guilan. Blood samples treated with ethylenediaminetetraacetic acid (EDTA) were stored at − 70 °C for subsequent use. All necessary reagents were procured from Merck (Germany). The DNA isolation procedure from blood entails four key steps as follows: (1) Cell lysis using a detergent such as sodium dodecyl sulfate (SDS), (2) Protein digestion with proteinase K, targeting the proteins released during cell lysis, (3) Protein removal using phenol, (4) DNA precipitation with alcohol.
In this protocol, DNA extraction was performed according to the method described by Ghaheri et al.39. A mixture of 1 mL of distilled water and 1 mL of blood was prepared in a 2 mL micro-tube. The tube was then subjected to centrifugation at 4000 rpm for 10 min, and the supernatant was discarded. This step was repeated 2 or 3 times. Subsequently, 1 mL of lysis buffer (1 × 10–2 mol L–1 Tris–HCl, 1 × 10–3 mol L–1 disodium EDTA) was added to the tube, followed by centrifugation at 4000 rpm for 5 min. The supernatant was then removed. Next, 0.8 mL of lysis buffer, 0.1 mL of 10% SDS, and 2.5 × 10–2 mL of proteinase K were added to the tube. The tube was shaken, and 0.4 mL of phenol was introduced. After shaking, the tube underwent centrifugation at 4000 rpm for 5 min. The upper phases were carefully transferred to a new tube, and 0.4 mL of chloroform-isoamyl alcohol (24:1) and 0.05 mL of sodium acetate were added. The tube was shaken and centrifuged at 4000 rpm for 5 min. Cold absolute ethanol was added in three successive increments to the resulting solution from the previous step. The tube was gently shaken and centrifuged at 3000 rpm for 3 min. The supernatant was discarded, and 1 mL of 70% ethanol was added. Following centrifugation at 3000 rpm for 3 min, the supernatant was discarded. The DNA was air-dried, re-suspended in 0.1 mL of H2O, and stored at − 20 °C. UV–Vis spectrometry was employed to assess the purity of DNA (A260:A280 > 1.80) as well as its concentration (ɛ = 6600 L mol–1 cm–1 at 260 nm) (Fig. 2).
Results and discussion
Electrochemical study
Holmium ions exhibit distinct cathodic and anodic peaks on the surface of a GCE, attributed to the facile electron donating characteristics between Ho(III) and Ho(IV), as depicted in Fig. 3. Despite the potential significance of holmium metal in nucleic acid research, its electrochemical properties have not been extensively investigated, leaving a significant knowledge gap in this area. Therefore, this study presents a comprehensive analysis of the electrochemical behavior of Ho3+ on the GCE at pH 7.2, providing novel insights into its redox characteristics. Figure 3 showcases the cyclic voltammograms of Ho3+ at various concentrations, employing a potential scan range of − 0.1 to 0.7 V. Notably, the inset of Fig. 3 reveals a linear correlation between the peak current (Ip) and the concentration of Ho3+, further validating the observed electrochemical behavior.
To delve deeper into the electrochemical properties, Fig. 4 portrays the cyclic voltammograms of 2.5 × 10–3 mol L–1 Ho3+ at pH 7.2, in the presence of 0.1 mol L–1 NaCl on the GCE. The potential range spans from − 0.1 to 0.5 V, while the scan rates vary from 0.005 to 0.15 V s–1. Remarkably, an evident augmentation in the redox current peaks is observed with increasing scan rate. Furthermore, the inset of Fig. 4 demonstrates that the peak current (Ip), as well as the current at any other point on the wave, is proportional to the square root of the scan rate (υ), indicating a diffusion-controlled process associated with a reversible redox peak. This reversible redox behavior of Ho3+ ions aligns with previous research findings40. However, owing to the charge density and hydration effects of Ho3+ ions, wherein they are surrounded by 6–12 water molecules, their solvated radii become considerably large. Consequently, their mobility near the electrode surface becomes sluggish, particularly at elevated concentrations, necessitating an extended time for their departure from the electrode surface. As a result, quasi-reversible cyclic voltammograms are observed.
The diffusion coefficient (Do) of the electroactive species can be determined by applying the dependencies defined by the Randles-Ševčík Eq. 40:
In Eq. (1), the symbol 'n' represents the number of electrons involved in the half reaction of the redox couple, 'υ' denotes the rate at which the potential is swept (V s–1), 'A' signifies the electrode area (cm2), and Do corresponds to the diffusion coefficient of the analyte (cm2 s–1). By analyzing the plot of Ip versus υ1/2 as depicted in the inset of Fig. 4, the diffusion coefficient is calculated to be 5.1 × 10–2 cm2 s–1.
Furthermore, Fig. 5 illustrates the cyclic voltammograms of Ho3+ in the absence and presence of DNA in Tris buffer at pH 7.2. Intriguingly, the addition of DNA to Ho3+ results in a decrease in peak current, accompanied by the absence of new reduction peaks. This decrease in peak current implies the formation of a novel complex, indicating an interaction between DNA and Ho3+. Specifically, it is proposed that Ho3+ and DNA form an electrochemically inactive complex that cannot be reduced at the GCE.
Measurement of the stoichiometry of the DNA-nHo3+
To determine the binding number and the equilibrium constant of the binding reaction, the following equations were utilized. Among the three types of binding modes between small molecules and DNA, Bard reported40 that if the E° shifted to more negative values following interaction of small molecules with DNA, the interaction mode was electrostatic binding. Conversely, if the E° shifted to more positive values, the interaction mode was intercalative binding. The decrease in peak current may be due to the competitive adsorption between Ho3+ and DNA40. The binding number and the equilibrium constant of the binding reaction can be determined as follows:
The equilibrium constant was
According to the Ilkovĭc equation, Id = 708nDo1/2m2/3t1/6Co* = kCo*, the peak current (Id) is proportional to the concentration (Co*). From this equation, the following equations can be deduced:
Introducing Eqs. (3), (5) and (7) gave
The difference in peak current (ΔI) between the presence and absence of DNA, and the maximum obtained value of peak current difference (ΔImax) at an extremely high concentration of Ho3+ compared to DNA were considered in this study. The equilibrium constant (βs) was calculated as the intercept of the plot of log(ΔI/(ΔImax-I)) versus log[DNA]. A higher βs value indicates intercalation-based interactions, whereas a lower value suggests weaker groove or electrostatic interactions41. The inset in Fig. 5 demonstrates a linear relationship, indicating the formation of a single complex between Ho3+ and DNA at various DNA concentrations. The slope of the equation was determined to be − 1.119, the intercept as 1.129. The slope value suggests the binding of only one Ho3+ ion to each DNA molecule. Furthermore, the equilibrium constant was obtained from the intercept of this equation (βs = 1.34 × 101 L mol–1). The lower value of the equilibrium constant obtained for the interaction between the studied metal ion and the DNA chain suggests a groove binding interaction mode42.
In voltammetric techniques, the measured electrochemical signal comprises both the undesired capacitive current and the desired current associated with the proper electrode reaction, known as the Faraday current. To further investigate this phenomenon, the differential pulse voltammetry method was employed (Fig. 6). The potential range used was 0.1 to 0.5 V. Curve (a) represents the differential pulse voltammogram of a 2.5 × 10−3 mol L–1 Ho3+ solution, while curves (b-f) represent the differential pulse voltammograms of Ho3+ solution with varying concentrations of DNA added (1, 2, 4, 7, and 9 µmol L–1, respectively). The figure illustrates a decrease in peak currents as the DNA concentration increases.
Differential pulse voltammograms of (a) 2.5 × 10−3 mol L–1 Ho3+, (b) a + 1 × 10–6 mol L–1 DNA, (c) a + 2 × 10–6 mol L–1 DNA and (d) a + 4 × 10–6 mol L–1 DNA (e) a + 7 × 10–6 mol L–1 DNA, (f) a + 9 × 10–6 mol L–1 DNA at pH 7.2. The initial potential, 0.1 V, the end potential, 0.4 V, the step potential, 0.05 V, the modulation time, 0.02 s.
pH effect
This study aimed to investigate the influence of pH variations on the electrochemical behavior of Ho3+ ions. Two fixed concentrations, specifically 1.0 × 10−2 and 1.0 × 10−3 mol L–1, were chosen for analysis. Solutions with consistent analyte ion concentrations were prepared at different pH values ranging from 2.0 to 12.0. Cyclic voltammetry was employed to examine the electrochemical characteristics of Ho3+ at lower pH values. However, obtaining cyclic voltammograms of Ho3+ proved challenging due to the protonation effect on the anion phosphate groups. This effect arises because the majority of phosphate groups in DNA and RNA molecules are involved in phosphodiester bonds, leaving only one hydroxyl group available for dissociation. At higher pH values, an increase in hydroxide ion concentration ([OH−]) can result in the formation of insoluble holmium hydroxide complexes (Ho(OH)3, Ho(OH)2+, Ho(OH)2+). These compounds can cover the electrode surface, impeding the access of Ho3+ to the electrode. Consequently, all experimental investigations in this study were conducted at pH 7.2. This pH value was selected to be compatible with intracellular conditions and deemed suitable for in vivo studies (Fig. 7).
Spectroscopic study
Electronic absorption spectroscopy
The UV–Vis absorption spectrum of DNA exhibits a broad band in the UV region (200–350 nm) with a maximum peak at 265 nm. This peak arises from the chromophoric groups present in purine and pyrimidine moieties, responsible for electronic transitions. Slight variations in pH or ionic strength can cause changes in the absorption maximum and molar absorptivity of DNA. Comparing the UV–Vis absorption spectra of free metal ions and DNA-metal complexes provides valuable insights into the interaction between DNA and metal ions, as they often exhibit distinct spectra. Intercalation binding of metal ions to DNA typically leads to hypochromism and hypsochromism (blue shift) or bathochromism (red shift). Additionally, denaturation of the DNA double helix, achieved through the use of denaturing agents, reduces base–base interactions, resulting in an increase in DNA absorbance. Single-stranded DNA (ss-DNA) exhibits 40% higher absorbance than double-stranded DNA (ds-DNA) at equivalent concentrations. The hyperchromic effect is primarily attributed to the presence of charged cations that bind to the major groove or the minor groove of DNA through coordination with specific functional groups on the bases or backbone, causing structural damage and contraction43.
To investigate the behavior of Ho3+ ions in Tris–HCl buffer solution, UV–Vis spectroscopy was performed at room temperature for 24 h prior to the reaction with DNA. An absorbance spectrum of 6.0 × 10−5 mol L–1 Ho3+ was obtained at 230 nm. Subsequently, titration experiments were conducted by adding different concentrations of DNA (ranging from 5.2 to 17.5 µmol L–1) at pH 7.2. Each sample was allowed to equilibrate for 5 min, and the absorption spectra were recorded in the range of 220–275 nm. Figure 8 presents the absorption spectra of Ho3+ in the absence and presence of DNA, illustrating spectroscopic changes indicative of the interaction between Ho3+ and DNA.
Fluorescence spectroscopy
Fluorescence spectroscopy, known for its sensitivity, wide linear concentration range, and selectivity, was employed to further investigate the interaction between Ho3+ and DNA. Fluorescence quenching experiments were conducted to provide additional insights into the localization of the metal ions and their mode of interaction with DNA44. Ho3+ fluorescence intensity was measured in aqueous solution after optimizing the concentration and pH. The interaction between DNA and Ho3+ was examined by titrating a constant concentration of Ho3+ (6.0 × 10−5 mol L–1) with varying amounts of DNA (ranging from 0.4 to 16.9 µmol L–1). The effects on the emission intensity were recorded after 5 min at 299 K, monitoring the fluorescence emission at 364 nm. Fluorescence quenching can occur through various molecular interactions, including excited-state reactions, molecular rearrangements, energy transfer, and complex formation. The Stern–Volmer equation (Eq. 9) was utilized to describe the fluorescence quenching process45,46:
where Fo represents the fluorescence intensity in the absence of the quencher, F represents the fluorescence intensity in the presence of the quencher, Kq is the bimolecular quenching constant, τ0 is the fluorophore’s lifetime, KSV is the Stern–Volmer constant, and [Q] is the concentration of the quencher.
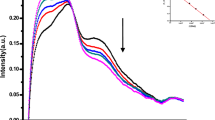
A solution of 6.0 × 10−5 mol L–1 HoCl3 was selected as the optimal concentration for fluorescence studies. The effect of pH on Ho3+ fluorescence was investigated, and a pH of 7.2 was determined to be suitable for the investigations. The emission titration curves (Fig. 9) present the spectral results of the fluorescence experiments, showing a decrease in emission intensity at 364 nm with increasing concentrations of DNA. However, the shape of the emission bands remained unchanged. This quenching phenomenon can be attributed to photoelectron transfer from the guanine base of DNA to the excited levels of Ho3+ ions47. Various interactions, such as groove binding48,49, electrostatic interactions50, hydrogen bonding, or hydrophobic interactions, can lead to a decrease in fluorescence intensity in the presence of DNA close to the sugar-phosphate backbone51. The binding constant (Kb) and the number of binding sites (n) for the DNA- Ho3+ system were determined using Eq. (10): 46
The fluorescent spectral characteristics of Ho(III)-DNA. (a) Ho(III) concentration: 6.0 × 10–5 mol L–1 in pH 7.2 Tris-buffer. DNA concentrations: (b) = 0.4, (c) = 3.1, (d) = 5.7, (e) = 8.3, (f) = 11.7, (g) = 12.9, (h) = 14.2, (i) = 15.8, and (j) = 16.9 µmol L–1. Inset: plot of log(Fo–F/F) vs. log[DNA].
The plot of log((F0-F)/F) against log[Q] yielded a slope and intercept, from which the binding constant and number of binding sites were calculated as Kb = 2.3 × 105 and n = 1.143, respectively (inset of Fig. 9).
Conclusions
The investigation of the interaction between Ho3+ ions and DNA utilizing a variety of experimental techniques has yielded significant insights. The findings underscore the efficacy of electrochemical methods in exploring the intricate interplay between DNA and metal ions. The electrochemical behavior of Ho3+ ions on a GCE exhibited a reversible electron transfer process, indicative of its redox activity. Furthermore, an intriguing linear correlation between the peak current and the square root of the scan rate was consistently observed across different scan rates, suggesting a diffusion-controlled kinetic regime for the electrochemical process. Complementary UV–Vis and fluorescence spectroscopy experiments provided valuable information regarding the nature of the interaction between Ho3+ ions and DNA. Specifically, the results indicate that Ho3+ ions can engage in a groove binding interaction mode with DNA. The fluorescence emission studies further revealed that the presence of DNA leads to the quenching of fluorescence exhibited by Ho3+ ions. These findings contribute to the broader understanding of the intermolecular interactions involving metal ions and DNA, offering valuable insights for future studies in the field of nucleic acid research.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Landegren, U., Kaiser, R., Sanders, J. & Hood, L. A ligase-mediated gene detection technique. Science 241, 1077–1080 (1988).
Ilkhani, H., Ganjali, M. R., Arvand, M., Faridbod, F. & Norouzi, P. The effect of pH on the interaction between Eu3+ ions and short single-stranded DNA sequence, studied with electrochemical, spectroscopic and computational methods. Mater. Sci. Eng. C 32, 653–658 (2012).
Millan, K. M. & Mikkelsen, S. R. Sequence-selective biosensor for DNA based on electroactive hybridization indicators. Anal. Chem. 65, 2317–2323 (1993).
Zhang, S. S. et al. Studies on the interaction mechanism between hexakis(imidazole)manganese(II) terephthalate and DNA and preparation of DNA electrochemical sensor. J. Inorg. Biochem. 99, 2340–2347 (2005).
Sissoeff, I., Grisvard, J. & Guille, E. Studies on metal ions-DNA interactions: specific behaviour of reiterative DNA sequences. Prog. Biophys. Mol. Biolog. 31, 165–199 (1978).
Manning, G. S. The molecular theory of polyelectrolyte solutions with applications to the electrostatic properties of polynucleotides. Q. Rev. Biophys. 11, 179–246 (1978).
Montalban, A. G. et al. Bis(dimethylamino)porphyrazines: Synthetic, structural, and spectroscopic investigations. J. Org. Chem. 65, 2472–2478 (2000).
Mohammadi, S., Perrée-Fauvet, M., Gresh, N., Hillairet, K. & Taillandier, E. Joint molecular modeling and spectroscopic studies of DNA complexes of a bis(arginyl) conjugate of a tricationic porphyrin designed to target the major groove. Biochemistry 37, 6165–6178 (1998).
Eichhorn, G. L. & Shin, Y. A. Interaction of metal ions with polynucleotides and related compounds. XII. The relative effect of various metal ions on DNA helicity. J. Am. Chem. Soc. 90, 7323–7328 (1968).
Izatt, R. M., Christensen, J. J. & Rytting, J. H. Sites and thermodynamic quantities associated with proton and metal ion interaction with ribonucleic acid, deoxyribonucleic acid, and their constituent bases, nucleosides, and nucleotides. Chem. Rev. 71, 439–481 (1971).
Thompson, K. H., Barta, C. A. & Orvig, C. Metal complexes of maltol and close analogues in medicinal inorganic chemistry. Chem. Soc. Rev. 35, 545–556 (2006).
Bünzli, J. G. & Eliseeva, S. V. Basics of lanthanide photophysics. In Lanthanide luminescence: Photophysical, photophysical, analytical and biological aspects (Springer, Berlin, 2011).
Zhang, L. Z. & Cheng, P. Study of Ni(II) ion-DNA interactions with methylene blue as fluorescent probe. J. Inorg. Biochem. 98, 569–574 (2004).
Bloomfield, V. A. DNA condensation by multivalent cations. Biopolymers 44, 269–282 (1997).
Zhang, L. Z. & Cheng, P. Spectroscopic studies of the interactions of Zn(II) ion with herring sperm DNA toward a better understanding of the M-DNA complex. Inorg. Chem. Commun. 7, 392–394 (2004).
Hong, F. et al. Direct evidence for interaction between lead ions and kidney DNA from silver crucian carp. Chemosphere 68, 1442–1446 (2007).
Takasaki, B. K. & Chin, J. Cleavage of the phosphate diester backbone of DNA with cerium(III) and molecular oxygen. J. Am. Chem. Soc. 116, 1121–1122 (1994).
Geraldes, C. F. & Williams, R. J. Nucleotide torsional flexibility in solution and the use of the lanthanides as nuclear-magnetic-resonance conformational probes: The case of adenosine 5′-monophosphate. Eur. J. Biochem. 85, 463–470 (1978).
Niroomand, S., Khorasani-Motlagh, M., Noroozifar, M. & Moodi, A. Spectroscopic studies on the binding of holmium-1,10-phenanthroline complex with DNA. J. Photochem. Photobiol. B: Biology 117, 132–139 (2012).
Goyal, R. N., Gupta, V. K. & Chatterjee, S. Simultaneous determination of adenosine and inosine using single-wall carbon nanotubes modified pyrolytic graphite electrode. Talanta 76, 662–668 (2008).
Jain, A. K., Gupta, V. K., Singh, L. P., Srivastava, P. & Raisoni, J. R. Anion recognition through novel C-thiophenecalix[4]resorcinarene: PVC based sensor for chromate ions. Talanta 65, 716–721 (2005).
Prasad, R., Gupta, V. K. & Kumar, A. Metallo-tetraazaporphyrin based anion sensors: Regulation of sensor characteristics through central metal ion coordination. Anal. Chim. Acta 508, 61–70 (2004).
Arvand, M., Ilkhani, H., Ganjali, M. R. & Pourhabib, A. Electrochemical investigation of DNA-metal complex interactions and development of a highly sensitive electrochemical biosensor. Anal. Biochem. 652, 114738 (2022).
Hashemzadeh, N. et al. Graphene quantum dot modified glassy carbon electrode for the determination of doxorubicin hydrochloride in human plasma. J. Pharm. Anal. 6, 235–241 (2016).
Gupta, V. K., Jain, R., Radhapyari, K., Jadon, N. & Agarwal, S. Voltammetric techniques for the assay of pharmaceuticals-a review. Anal. Biochem. 408, 179–196 (2011).
Ilkhani, H. et al. Electrochemical spectroscopic investigations on the interaction of an ytterbium complex with DNA and their analytical applications such as biosensor. Int. J. Biolog. Macromol. 49, 1117–1123 (2011).
Ilkhani, H., Ganjali, M. R., Arvand, M. & Norouzi, P. Electrochemical and spectroscopic study of samarium ion interaction with DNA in different pHs. Int. J. Electrochem. Sci. 5, 168–176 (2010).
Carter, M. T. & Bard, A. J. Voltammetric studies of the interaction of tris(1,10-phenanthroline)cobalt(III) with DNA. J. Am. Chem. Soc. 109, 7528–7530 (1987).
Carter, M. T., Rodriguez, M. & Bard, A. J. Voltammetric studies of the interaction of metal chelates with DNA. 2. Tris-chelated complexes of cobalt(III) and iron(II) with 1,10-phenanthroline and 2,2’-bipyridine. J. Am. Chem. Soc. 111, 8901–8911 (1989).
Carter, M. T. & Bard, A. J. Electrochemical investigations of the interaction of metal chelates with DNA. 3. Electrogenerated chemiluminescent investigation of the interaction of tris(1,10-phenanthroline)ruthenium(II) with DNA. Bioconjug. Chem. 1, 257–263 (1990).
Rodriguez, M. & Bard, A. J. Electrochemical studies of the interaction of metal chelates with DNA 4 Voltammetric and electrogenerated chemiluminescent studies of the interaction of tris(2,2’-bipyridine)osmium(II) with DNA. Anal. Chem. 62, 2658–2662 (1990).
Maeda, M., Nakano, K., Uchida, S. & Takagi, M. Mg2+-selective electrode comprising double-helical DNA as receptive entity. Chem. Lett. 23, 1805–1808 (1994).
Kelley, S. O., Barton, J. K., Jackson, N. M. & Hill, M. G. Electrochemistry of methylene blue bound to a DNA-modified electrode. Bioconjug. Chem. 8, 31–37 (1997).
Chundawat, N. S., Jadoun, S., Zarrintaj, P. & Chauhan, N. P. S. Lanthanide complexes as anticancer agents: A review. Polyhedron 207, 115387 (2021).
Fricker, S. P. The therapeutic application of lanthanides. Chem. Soc. Rev. 35, 524–533 (2006).
Harper, M., Boyce, J. D. & Adler, B. Pasteurella multocida pathogenesis: 125 years after Pasteur. FEMS Microbiol. Lett. 265, 1–10 (1997).
Gilling, P. J., Kennett, K., Das, A. K., Thompson, D. & Fraundorfer, M. R. Holmium laser enucleation of the prostate (HoLEP) combined with transurethral tissue morcellation: An update on the early clinical experience. J. Endourol. 12, 457–459 (1998).
Fraundorfer, M. R. & Gilling, P. J. Holmium: YAG laser nucleation of the prostate combined with mechanical morcellation: Preliminary results. Eur. Urol. 33, 69–72 (1998).
Ghaheri, M. et al. A comparative evaluation of four DNA extraction protocols from whole blood sample. Cell. Mol. Biol. 62, 120–124 (2016).
Bard, A. J., Faulkner, L. R. & White, H. S. Electrochemical methods: fundamentals and applications (Wiley, Hoboken, 2022).
Ramotowska, S., Ciesielska, A. & Makowski, M. What can electrochemical methods offer in determining DNA-drug interactions?. Molecules 26, 3478 (2021).
Qin, D. A., Cai, X. Q., Miao, Q., Wang, Z. H. & Hu, M. L. Cyclic voltammetry and molecular docking study of the interactions of two derivatives of 5-fluorouracil with DNA. Int. J. Electrochem. Sci. 9, 1608–1620 (2014).
Aleksić, M. M. & Kapetanović, V. An overview of the optical and electrochemical methods for detection of DNA-drug interactions. Acta Chim. Slov. 61, 555–573 (2014).
Yunus, G., Srivastava, S. & Gupta, V. D. Stability of DNA binding with dipyrandium: A theoretical study. Int. J. Phys. Sci. 6, 8151–8156 (2011).
Lakowicz, J. R. Principles of fluorescence spectroscopy (Springer, Boston, MA, 2006).
Zhang, G., Hu, X., Zhao, N., Li, W. & He, L. Studies on the interaction of aminocarb with calf thymus DNA by spectroscopic methods. Pestic. Biochem. Phys. 98, 206–212 (2010).
Xi, P. X. et al. Synthesis, characterization, antioxidant activity and DNA-binding studies of three rare earth (III) complexes with 1-(4-aminoantipyrine)-3-tosylurea ligand. J. Fluoresc. 19, 63–72 (2009).
Joshi, H., Sengupta, A., Gavvala, K. & Hazra, P. Unraveling the mode of binding of the anticancer drug topotecan with dsDNA. RSC Adv. 4, 1015–1024 (2014).
Mati, S. S., Roy, S. S., Chall, S., Bhattacharya, S. & Bhattacharya, S. C. Unveiling the groove binding mechanism of a biocompatible naphthalimide-based organoselenocyanate with calf thymus DNA: An “ex vivo” fluorescence imaging application appended by biophysical experiments and molecular docking simulations. J. Phys. Chem. B 117, 14655–14665 (2013).
Paul, B. K. & Guchhait, N. Exploring the strength, mode, dynamics, and kinetics of binding interaction of a cationic biological photosensitizer with DNA: Implication on dissociation of the drug-DNA complex via detergent sequestration. J. Phys. Chem. B 115, 11938–11949 (2011).
Cui, F. et al. Study on the interaction between aglycon of daunorubicin and calf thymus DNA by spectroscopy. J. Mol. Struct. 1001, 104–110 (2011).
Acknowledgements
The authors are thankful to the post-graduate office of Guilan University for the support of this work.
Author information
Authors and Affiliations
Contributions
Ali Tilehkan: Data curation, Methodology, Investigation, Writing-original draft. Majid Arvand: Conceptualization, Visualization, Supervision, Writing-review & editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Tilehkan, A., Arvand, M. Study on the electrochemical and spectroscopic characteristics of holmium ion and its interaction with DNA. Sci Rep 14, 20383 (2024). https://doi.org/10.1038/s41598-024-71412-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-71412-4
Keywords
This article is cited by
-
Probing the interaction of Iron Oxide Nanoparticles (IONPs) with dsDNA in presence of UV–Vis light
Surface Science and Technology (2025)
-
DNA binding, and apoptosis-inducing activities of a β-ionone-derived ester in human myeloid leukemia cells: multispectral and molecular dynamic simulation analyses
Scientific Reports (2024)