Abstract
Staphylococcus aureus (S. aureus), particularly Methicillin-resistant S. aureus (MRSA), poses a significant global public health threat, necessitating advanced methodologies to enhance our understanding of this organism at the omics levels. This study introduces a refined protocol for constructing and curing high-density transposon mutant (tn-mutant) libraries in S. aureus, addressing the challenges associated with low transductant yields, and the complex genetic manipulation mechanism in Gram-positive bacteria. Our methodology employs a Himar1 transposon based on a two-plasmid system, leveraging Himar1’s high insertional efficiency in AT-rich organisms. Enhanced transduction efficiency was achieved through chloramphenicol pre-treatment and the use of modified enriched media. Complementing this, an optimized plasmid curing procedure ensured a representative and stable tn-mutant library. The protocol was successfully applied to multiple S. aureus strains, demonstrating an increase in mutant recovery and reduced post-curing impact. The method offers a robust approach for Transposon Insertion Sequencing (TIS) applications in S. aureus, enabling deeper insights into survival, resistance, and pathogenicity mechanisms. This protocol holds a significant potential for accelerating the construction of tn-mutant libraries in various S. aureus strains.
Similar content being viewed by others
Introduction
Staphylococcus aureus (S. aureus), in particular, Methicillin-resistant S. aureus (MRSA), represents a significant global public health challenge. This bacterium is responsible for a broad spectrum of concerning diseases, including bacteremia, endocarditis, skin and soft tissue infections, bone and joint infections1. Given the significant health threat posed by this bacterium, there is a pressing need for advanced, high-throughput methodologies to augment our understanding and knowledge at the omics levels.
Since the discovery of the first well-characterized transposable element, Tn551, by Novick et al., transposon mutagenesis has revolutionized the field of S. aureus biology2,3. This began with the use of Tn917 in the early 90s and the construction of the first transposon mutant (tn-mutant) library containing 1248 mutants and led to the development of high-density tn-mutant libraries4,5.
Over the last decade, there has been a growing focus on exploring the relationship between bacterial genotype and phenotype on a large scale. This exploration has been facilitated by Transposon Insertion Sequencing (TIS), a revolutionary technology that has significantly broadened our understanding of gene-behavior connections on a genome-wide scale6,7.
A thorough understanding of these genes is crucial for the comprehension of the genetic mechanisms affecting survival, infection, resistance, and adaptation8. A central prerequisite for TIS is to establish robust and saturated tn-mutant libraries9. A successful TIS experiment requires a comprehensive library of transposon insertions at all possible genomic sites10. However, constructing such libraries for S. aureus and other Gram-positive bacteria presents a significant challenge, in contrast to Gram-negative bacteria where kits such as EZ-tn5 can be easily employed11,12. The application of these methods to S. aureus is considerably less effective13. A two-plasmid system for transposition has proven to be effective in generating S. aureus tn-mutant libraries13. However, constructing such a library using this system involves several challenges14,15. The first critical step is the efficient insertion of the transposition system into the bacterial genome. The phage-based delivery system has proven to be a successful approach for recovering mutants, which is essential for generating a high-density tn-mutant library12. The second step is curing the transposition system plasmids without affecting the representation of the mutants, which can be particularly challenging for temperature-sensitive mutants13. This step is necessary to ensure plasmid-free mutants, thereby maintaining the stability of the transposon insertions within the tn-mutant library. These challenges highlight the need to refine this approach to obtain a stable, high-saturated tn-mutant library.
Our work aims to optimize the above key steps involved in constructing a tn-mutant library in S. aureus, by boosting transduction efficiency and minimizing the heat-induced effect during plasmid curing. This protocol, successfully applied to various S. aureus strains with different phylogenetic backgrounds; is expected to accelerate the generation of highly saturated tn-mutant libraries for TIS screening.
Results
We optimized a protocol for constructing and curing a tn-mutant library as shown in Fig. 1. We focused on enhancing transduction efficiency by fine-tuning the multiplicity of infection (MOI), pre-treating with chloramphenicol, and adjusting the transduction medium composition. Additionally, we validated the efficacy of the Himar1 transposition system in increasing the complexity and diversity of tn-mutant libraries using Transposon directed insertion sequencing (TraDIS). Furthermore, we compared our plasmid curing protocol to the conventional method, demonstrating its effectiveness in reducing heat-induced bias and ensuring stable mutant recovery. The following subsections provide detailed insights into our findings, highlighting the improvements achieved through these optimizations.
Schematic workflow of proposed optimizations for constructing tn-mutant libraries in S. aureus. This figure illustrates the optimized workflow for constructing tn-mutant libraries in S. aureus. The key steps are sequentially numbered, beginning with the introduction of pFA545 carrying the mariner transposase into the recipient strain of interest (1). This was followed by the introduction of the Himar1 transposon by transduction using ɸ11/pMARGK2b-Himar1 lysate (2) after pretreatment of recipient strain with chloramphenicol (3) and the use of a transduction enhancer SN medium (4). Once the transposition system is introduced, insertions are generated, and each mutant contains a single transposon insertion within the bacterial genome (5). The process continues with a stepwise thermal increase and passage reduction during plasmids curing process (6), concluding with TraDIS (7). Details of the optimization are included in the annotation boxes.
Enhancing mutant recovery by optimizing multiplicity of infection in S. aureus strains
The MOI, defined as the ratio of infecting phages to host bacterial cells, can significantly affect the infection process. We conducted experiments to determine the optimal MOI to enhance mutant recovery. A range of MOI from 0.1 to 0.6 was explored for various S. aureus strains, including JE2, ST398, Newman, COL, and MW2. It was observed that a high MOI was associated with an increased cell lysis, while a very low MOI was associated with reduced transduction efficiency. As depicted in Fig. 2, most of the tested S. aureus strains showed an optimal MOI within the investigated range, with minor variations. These variations in optimal MOI among different S. aureus strains could be linked to several factors, such as the presence and abundance of specific phage receptors on the bacterial cell surface and the effectiveness of bacterial immune mechanisms against phages.
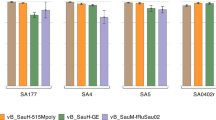
Optimal MOI for maximal transduction efficiency in S. aureus strains. A significant variation in transduction efficiency across S. aureus strains was noted at different MOI (0.1, 0.2, 0.3, 0.6). The most substantial number of transductants is observed at an MOI of 0.2 for COL strain, MOI of 0.3 for JE2 and Newman strains, and MOI of 0.6 for ST398 strain. Although the MW2 strain generally exhibits a lower number of transductants, its highest efficiency is achieved at an MOI of 0.2.
Chloramphenicol pre-treatment and transduction medium composition improve ϕ11 transduction efficiency in S. aureus
Phage ϕ11 binds to its receptor on the wall teichoic acid (WTA) of bacterial cells. To increase the bacterial WTA content, a sub-inhibitory concentration of chloramphenicol was employed in parallel with tunicamycin and fosfomycin, known to inhibit WTA and decrease cell wall content, respectively. An increase in transduction efficiency of JE2 was observed in the cultures pre-treated with chloramphenicol while a significant reduction was observed with fosfomycin or tunicamycin (Fig. 3). Pre-treatment with chloramphenicol showed a consistent enhancement in transduction efficiency across all tested strains (Figs. 3 and 4).
Differential impact of antibiotics pre-treatment on bacterial transduction efficiency. Comparison of the number of S. aureus JE2 pMARGk2b-Himar1/pFA545 transductants obtained using SN transduction medium, following cultures pre-treatment with chloramphenicol, fosfomycin, or tunicamycin.The highest yield of transductants was obtained after chloramphenicol pre-treatment. The statistical significance of these differences was determined using one-way ANOVA, followed by Tukey’s multiple comparisons test. The three asterisks ‘***’ corresponds to an adjusted p-value < 0.001.
Variations in transduction efficiency in TSB and SN media among S. aureus strains with or without chloramphenicol pre-treatment. The use of SN medium with sub-inhibitory concentrations of chloramphenicol (SN + CAM) showed a significant increase in the number of transductants in strains JE2, MW2, ST398, and COL compared to TSB alone, TSB supplemented with chloramphenicol (TSB + CAM), and SN medium used independently. The Newman strain did not exhibit a significant difference in transductant numbers between SN + CAM and TSB + CAM conditions. However, all S. aureus strains tested demonstrate higher transduction efficiency in presence of SN + CAM compared to SN alone. Statistical analysis was performed using one-way ANOVA, adjusted by Tukey’s multiple comparison test. The significance levels are indicated as follows: * corresponds to a p-value < 0.01, ** corresponds to a p-value < 0.001, *** corresponds to a p-value < 0.0001, and **** corresponds to a p-value < 0.00001.
The transduction efficiency of phage ϕ11 in S. aureus was also significantly influenced by the composition of the transduction SN medium with 5 mM CaCl2. A significant increase in transduction efficiency was observed across all tested strains, except Newman strain (Fig. 3).
Himar1 transposition system enhances complexity and diversity in S. aureus tn-mutant libraries
To measure the library complexity and transposition efficiency of the Himar1 transposition system. TraDIS was employed for both the optimized and conventional plasmid curing approaches of JE2 tn-mutant libraries (Tn-Library 1 and Tn-Library 2, respectively (Fig. 5). No significant differences were observed in the quantitative aspects such as plasmids cure percentage, library size, and number of unique insertion sites (UISs) (Table 1). Linker PCR confirmed the presence of unique transposon insertions in ten randomly selected mutants prior to sequencing (data not shown). The number of unique transposon mutants in both libraries was significantly high, with more than 400,000 UISs and a coverage of more than 75% of all possible insertion sites. This was significantly higher than the two previous studies16,17, with at least a more than 1.5-fold increase. In Tn-Library 1 and Tn-Library 2, 317 and 401 essential genes were respectively identified using the tradis_essentiality.R script, with a shared set of 310 genes (Fig. 5). A total of 281 genes were common between both libraries and the essential genes reported by Grosser et al.16 (Fig. 6, Supplementary Table S2).
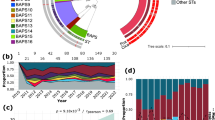
Mapping of mariner transposon insertions in S. aureus JE2 genome. The plot was generated using Artemis 18.2.0. The two tracks represent the frequency and location of transposon insertions in Tn-Library 1 (a) and Tn-Library 2 (b), in both sense and antisense orientations, which are colored in green and red, respectively. The raw transposon junction sequences were mapped and processed onto the USA300_FPR3757 reference genome.
Comparative analysis of the number of UISs and library composition in JE2. (a) This part of the Figure presents a comparative analysis of the number of UISs in Tn-Library 1 and 2, along with other studies16,17,18. (b) This part of the Figure shows a Venn diagram that illustrates the distinct and shared fitness genes following the plasmid curing step among Tn-Library 1 (represented in green), Tn-Library 2 (represented in orange), and the study conducted by Grosser et al.13 (represented in grey).
Gradual thermal increase and passage limitation reduce heat-induced bias in S. aureus tn-mutant library
After normalizing and comparing the read numbers from Tn-Library 1 and Tn-Library 2, a significant mutant depletion in Tn-Library 2 was observed compared to the slight depletion in Tn-Library 1 (Fig. 7).
Differential gene abundance and fitness between Tn-Library 1 and Tn-Library 2. (a) This chart displays the differential abundance of normalized reads for genes identified by Santiago et al.13, as being depleted at high temperatures, comparing Tn-Library 1 (green) and Tn-Library 2 (orange). Each gene is represented by a pair of bars showing its relative abundance in each library. (b) The volcano plot represents the comparative analysis of Tn-Library 1 and Tn-Library 2. Mutants that were depleted in Tn-Library 2, with a log2 fold change (log2FC) less than − 2, are highlighted in jungle green. Conversely, those in Tn-Library 1 with a log2FC greater than 2 are highlighted in sage green. (c) Numerous transposon insertions were present in pbp4 in Tn-Library 1, whereas they were almost absent in Tn-Library 2.
By using the tradis_essentiality.R script, 91 genes were identified as conditionally essential in Tn-Library 2; but not in Tn-Library 1, under their respective curing conditions. This substantial depletion of mutants in Tn-Library 2 particularly affected mutants mainly involved in cell wall biosynthesis and homeostasis (pbp3, pbp4, fmtC, and auxA) and heme biosynthesis pathway (hemB, hemC, hemD, hemE, hemL) (Fig. 7, Supplementary Table S3).
Conversely, other genes were more enriched in Tn-Library 2 such as serine protease htrA, lyrA, sgtB and regulators such as the membrane two-component system member srrB, rot, SarS (Supplementary Table S3).
Discussion
TIS was pioneered by emerging tn-mutant libraries using high-throughput sequencing, providing a comprehensive perspective on a range of bacterial species6. Numerous studies on both human and livestock strains of S. aureus have used various TIS methodologies, including Transposon Sequencing (Tn-seq), and TraDIS18,19,20. Despite its promising potential, TIS has continually encountered obstacles due to the strict restrictions imposed on the genetic manipulation of multiple S. aureus strains10,21. This demonstrates the necessity for continued improvements, particularly in the generation of highly saturated and unbiased libraries to ensure the success of TIS in vitro and in vivo screens.
In this study, we present an enhanced methodology for constructing tn-mutant libraries and validate this approach in commonly studied S. aureus strains. The efficiency of mutagenesis protocols is influenced by several factors, including the type of transposon, the transposition system employed, transduction efficiency, and curing method to generate highly cured, comprehensive tn-mutant libraries. Although the Tn5 transposon was used to create tn-mutant libraries in S. aureus, enabling insertion at diverse genomic sites, it demonstrated a bias for regions with high GC content22. In contrast, the Himar1 transposon is constrained to insertions specifically at TA dinucleotides23. In S. aureus, the AT content constitutes approximately 67% of the entire genome, underscoring the significance of employing the Himar1 transposon to study AT-rich organisms24.
We tested five S. aureus strains representing three clonal complexes (CC): CC8 (JE2, Newman, and COL), CC1 (MW2), and CC398 (ST398). These strains included both methicillin-resistant and methicillin-sensitive variants. Considering that S. aureus possesses a thick cell wall, a robust restriction system, and lacks an efficient delivery system via conjugation13,25, we used a phage-based system to achieve higher transposition efficiency. The phage ϕ11-based delivery method has been successfully used to construct S. aureus transposon libraries13,26,27. Consequently, we optimized the transduction protocol for this phage to enhance the transduction efficiency across different S. aureus strains.
Phage ϕ11, one of the most thoroughly characterized staphylococcal phages, is frequently used as a laboratory tool for genetic manipulations. The interaction between S. aureus and phage ϕ11 occurs via its receptor-binding protein, which binds to α- or β-N-acetylglucosamine moieties on the cell wall WTA28,29. To enhance transduction efficiency, we conducted transduction experiments following pre-treatment with three different antibiotics: chloramphenicol, tunicamycin, and fosfomycin, in comparison with a no-treatment control. Our results showed a significant improvement in transduction efficiency with a sub-inhibitory concentration of chloramphenicol compared to other pre-treatments. Tunicamycin and fosfomycin were used as negative controls, known for reducing cellular WTA levels and polysaccharide cell-wall content, respectively, at sub-inhibitory concentrations30. Our findings align with Nygaard et al., who demonstrated using whole-cell nuclear magnetic resonance spectroscopy, that exposure to chloramphenicol results in a relative increase in the content of polysaccharides, which constitute the primary component of the staphylococcal WTA31,32. This increase in cell wall WTA content augments the surface area available for phage ϕ11 attachment to its receptor, thereby enhancing the transduction process.
Another aspect that we opted for optimizing the transduction step, is the use of the appropriate MOI for each strain. This helps maintain a balance between maximizing the recovery of tn-mutants and minimizing negative impacts on the bacterial cells.
Moreover, to facilitate the accessibility of the phage to bacterial cells and increase the transduction efficiency, we prepared an in-house transduction SN medium that contains extra carbon and nitrogen sources, along with divalent cations. We observed a significant increase in the number of transductants in JE2, ST398, COL, and MW2 when SN medium with 5 mM CaCl2 was employed in the transduction step compared to TSB. This enhancement can probably be attributed to the addition of carbon and phosphorus sources, such as β-glycerophosphate disodium salt, which induces physical changes to the surface area of Staphylococcus, positively influencing phage adsorption33. Additionally, it acts as a chelating agent for calcium residuals34, preventing reinfection and interrupting the infectious cycle of the phages. Furthermore, we included divalent cations (Ca2+ and Mg2+) in the transduction medium to facilitate phage attachment by neutralizing the charge between the host cell surface polymer and the phage surface33,35 Evaluating transduction enhancers, that suit the specific phage and strain in use, is an essential process that can greatly influence the process of generating a tn-mutant library.
Beyond transduction, the cure process of temperature-sensitive plasmids is considered the tipping point in the construction of a successful tn-mutant library. Specifically, this event can introduce significant bias into the library representation, potentially leading to the underrepresentation of certain mutants that are sensitive to high-temperatures13. To evaluate this, we chose JE2 as a model strain to evaluate this process. We initially implemented a gradual increase in temperature and skipped passage after a brief time at 43 °C to mitigate the bias introduced during the curing step. Parallelly, we imitated the conventional plasmid curing process employed in previous studies18,36. We found that numerous genes became defective due to a sudden temperature change and a short exposure at 43 °C before the first overnight incubation. This intensifies the defective effects of slow mutants at high temperatures during the curing process, resulting in a skewed library. Transposon reads of 91 genes were depleted in Tn-Library 2 compared to Tn-Library 1. Defining essential or fitness genes can be challenging, as it depends on the medium, conditions, and the competition-based approach used in tn-library construction. However, within the context of our tn-library construction, these genes may play a role in responding to high-temperature treatment under the conventional curing protocol. This observation aligns with Santiago et al. findings, who showed that a sudden temperature rise to 43 °C resulted in a significant depletion of mutants, especially those involved in heat-shock response13. These heat survival genes were reported to significantly impact cell wall biosynthesis and signaling systems, protein folding, and c-di-AMP metabolism13,37. For instance, transposon insertions in cell wall biosynthesis genes, such as pbp3, pbp4, mreC, ssaA, lcpA, lytH, fmtC, auxA and auxB were found in Tn-Library 1 but not in Tn-Library 2. This indicates that, despite the reduced fitness of these genes at high temperatures, the adjustments in the curing process led to better recovery of these genes. Nonetheless, no or few transposon insertions were detected in genes involved in protein folding such as dnaJ, grpE, and dnaK13,37,38. Previous reports indicate that these genes are non-essential at 37 °C and essential at 43 °C. Our findings show that transposon insertions in these genes were detrimental in both libraries. Furthermore, it has been observed that certain mutants in the menaquinone biosynthesis pathway (including menC, menF, aroB, aroF, and ubiE) and in the pyrimidine pathway (such as pyrB, pyrC, and carB) were significantly enriched in both libraries. These genes have been reported to be essential at 30 °C13. Previous studies have established a connection between mutations in these pathways and the small colony variant phenotype39. Interestingly, Tn-Library 2 showed an enrichment of certain genes, including serine protease htrA, lyrA, and sgtB, and regulators, such as the membrane two-component system members srrB, rot, and sarS13,40. This implies that the curing process of the plasmids can qualitatively influence the representation of mutants in the tn-mutant library.
It’s important to note that this process did not significantly impact the quantitative aspects of the tn-mutant library. For instance, the unique number of transposon insertions in both libraries represents a coverage of nearly 76% of all TA sites (with ca. 6 bp distance between transposon insertions). This coverage was significantly higher in both libraries compared to previous studies conducted on the USA300 background, where the coverage of insertions was around 55% and 28%16,17.
To summarize, our optimized protocol offers a reliable basis for constructing tn-mutant libraries in S. aureus, which is considered the rate-limiting step of a successful TIS application. This will accelerate functional genomics studies enabling more efficient exploration of bacterial survival, resistance, and pathogenicity mechanisms in S. aureus and other staphylococcal pathogens.
Methods
Bacterial strains, plasmids, phage and culture conditions
In this study, we used MRSA strains (JE2, MW2, COL, ST398, Mu50) and methicillin-susceptible S. aureus (MSSA) strains (Newman, SH1000, SH1000 pMARGK2b-Himar1, SH1000 pFA545, SH1000 pFA545gen), and Escherichia coli (E. coli) IM08B strain (Supplementary Table S1). All wild-type (WT) strains of S. aureus were cultured in Brain Heart Infusion (BHI) broth or BHI agar (BHI-A), and Tryptic Soy broth (TSB) or Tryptic Soy agar (TSA), at 37 °C. E. coli IM08B was cultured in Luria broth (LB) or Luria agar (LA) agar at 37 °C. Additionally, BHI, TSB, BHI-A, and TSA were used as well for growing SH1000 strains harboring temperature-sensitive plasmids at 30 °C. The appropriate antibiotics used included tetracycline, chloramphenicol, erythromycin, tunicamycin, gentamicin and fosfomycin (Sigma Aldrich). Two temperature-sensitive plasmids transposition system were used: pMARGK2b-Himar1, carrying Himar1 transposon and 33 base pairs (bp) mariner mosaic ends (GAGACCGGGGACTTATCAGCCAACCTGTTAATC), and pFA545, which contains a mariner transposase. Transduction experiments were conducted using phage ϕ11. For propagation and transduction, an in-house medium, designed SN [TSB supplemented with filtered components: 1 mg/mL gelatin (Sigma Aldrich), 13.48 mg/mL β-glycerophosphate disodium salt (Sigma Aldrich), and 72 mg/mL MgSO4 was used] and CaCl2 (Sigma Aldrich). Transductants were selectively grown on BHI-A plates supplemented with 5 mg/L erythromycin, and either 5 mg/L tetracycline or 5 mg/L gentamicin.
Plasmid-based transposase introduction into S. aureus strains
The temperature-sensitive pFA545 plasmid containing mariner transposase was introduced into the S. aureus strain of interest, following two rounds of transformation via the E. coli IM08B shuttle strain. First, pFA545 plasmid was extracted from an overnight culture of S. aureus SH1000 at 30 °C using the Qiagen mini-prep plasmid DNA extraction Kit (Qiagen) according to the manufacturer’s instructions. The extracted pFA545 plasmid was used for heat shock transformation of chemically competent cells of E. coli IM08B according to the method described by Monk et al.41. E. coli IM08B pFA545 transformants were then selected on LA plates containing 5 mg/L of tetracycline following 48 h incubation at 30 °C. Once again, the pFA545 plasmid was isolated from E. coli IM08B transformant using the Qiagen Midi-Prep Plasmid DNA Extraction Kit (Qiagen). This plasmid was then used for the electroporation of electrocompetent S. aureus strains (JE2, MW2, COL, ST398, and Newman), following the method described by Monk et al.42.
High-concentration pFA545 DNA (up to 5 µg) was then added to 50 µl of freshly competent cells, followed by electroporation under specific conditions (18 kV/cm, 200 Ω, 25 µF). Subsequently, the cells were incubated in preheated BHI at 30 °C for two hours. Ultimately, the transformed cells were plated on BHI-A supplemented with 5 mg/L of tetracycline and incubated at 30 °C for 48 h.
Due to the resistance of ST398 to tetracycline, the selection marker cassette was replaced with aac6′-aph2′ gene, which encodes gentamicin resistance and was obtained from the Mu50 strain. This was done to construct pFA545gen, following the method of Christiansen et al.18. For colony screening and the subsequent experiments, tetracycline was substituted with gentamicin.
Introduction of Himar1 transposon into S. aureus strains containing transposase using phage ϕ11
The propagation and harvest of phage ϕ11/pMARGK2b-Himar1 were conducted using freshly prepared SN medium with 5 mM CaCl2. SH1000 pMARGK2b-Himar1 strain containing the Himar1 transposon, was cultured overnight at 30 °C on TSA agar and a single colony was resuspended in 1 mL of SN medium with 5 mM CaCl2. The phage ϕ11 stock (approximately 1010 pfu/mL) was then serially diluted in SN medium with 5 mM CaCl2. A mixture of 10 µL of SH1000 pMARGK2b-Himar1 resuspended cells and 100 µL of phage ϕ11 serial dilutions (up to 10− 9 ) was prepared in 4 mL of pre-maintained TSA soft agar at 50 °C and poured onto TSA plates supplemented with 5 mM CaCl2 followed by incubation at 30 °C for 24 h.
For lysate ϕ11/pMARGK2b-Himar1 harvest and titer determination, plates exhibiting near confluent lysis were selected. The lysate was harvested by adding 3 mL of SN medium to the plates, centrifuged for 10 min at 10,000 x g. The supernatant was filtered through a 0.22 μm filter and titrated using the aforementioned propagation method and then stored at 4 °C.
The introduction of ϕ11/pMARGK2b-Himar1 into S. aureus strains (JE2, MW2, COL, ST398 and Newman) was performed with high efficiency using transduction, following the Krausz and Bose protocol with modifications43. An overnight culture of the recipient strains was diluted 1:100 in 400 mL TSB with 1.5 mg/L chloramphenicol and incubated at 30 °C with shaking (200 rpm) until reaching the mid-exponential phase (OD600 0.2 :0.3). The culture was then transferred to eight 50-mL conical tubes and centrifuged at 4,500 x g for 10 min. The supernatant was discarded, the pellets of each tube were resuspended in 0.5 mL of freshly prepared SN medium with 5 mM CaCl2, and an appropriate aliquot of phage ϕ11/pMARGK2b-Himar1 was added to achieve the desired MOI for each strain. The tubes were incubated at room temperature for 10 min and then at 30 °C for 35 min without shaking. An additional 2.5 mL of TSB was added to each tube, followed by centrifugation at 4,500 x g for 10 min to pellet the cells. After discarding the supernatant, the pellets of each tube were resuspended in 5 mL of TSB, incubated with shaking (200 rpm) for 2 h at 30 °C, and then centrifuged at 4,500 x g for 10 min. Finally, the pellets from each tube were resuspended in 250 uL of TSB and plated each on 15 cm Ø BHI-A supplemented with 5 mg/L erythromycin and 5 mg/L tetracycline and incubated at 30 °C for 48 h. This process was repeated 20 times, resulting in 160 plates. Colonies were harvested from each plate by adding 3 mL of BHI supplemented with 5 mg/L erythromycin. All aliquots were then combined into a single pool, distributed into 1 mL glycerol stocks, and stored at -80 °C for future use.
To optimize this process, transduction experiments were conducted in SN medium with 5 mM CaCl2, with and without a pre-treatment of TSB containing sub-inhibitory concentration of chloramphenicol, tunicamycin (4 mg/L), or fosfomycin (4 mg/L) as above. Additionally, a range of MOI ratios (from 0.1 to 0.6) with approximately 109 CFU/mL of S. aureus JE2, ST398, Newman, COL, and MW2 strains, to identify the optimal MOI that yields the highest number of transductants.
Curing of temperature-sensitive plasmids from transposon mutants
JE2 was chosen as a model to investigate the thermal effects of plasmid curing on the tn-mutant library. An aliquot of glycerol stock of an uncured tn-mutant library of JE2 was thawed on ice and subsequently inoculated into 600 mL of BHI medium containing three antibiotics (chloramphenicol, tetracycline, and erythromycin). The culture was then subjected to a stepwise temperature increase starting at 35 °C for 20 min, then increased to 38 °C for 20 min, further to 41 °C for 20 min, and finally a step at 43 °C for 20 min. The OD600 was monitored for the subsequent 100 min until it reached 0.4. A 30 mL aliquot of the culture was then recovered by centrifugation (4000 rpm for 10 min), resuspended in 600 mL of pre-warmed BHI (43 °C) containing 5 mg/L erythromycin, and incubated with aeration overnight at 43 °C. This procedure was repeated for two days to yield a third-generation tn-mutant library, designed as Tn-Library (1) In parallel, an aliquot of the uncured library of JE2 was subjected to a conventional curing process without the gradual increase to 43 °C and included an extra passage before the first overnight incubation at 43 °C. This library was assigned as Tn-Library (2) For both libraries, cells were plated, each day, on mono-antibiotic BHI-A plates containing 5 mg/L of erythromycin, chloramphenicol, or tetracycline and incubated at 37 °C overnight. The curing percentage was calculated for each step. Aliquots of the cured transposon libraries containing 20% glycerol were stored at -80 °C.
Verification of tn-mutant library complexity by Linker PCR
The complexity of the tn-mutant library was confirmed using a method known as Linker PCR, as described by Christiansen et al.18. Random selection of ten colonies from BHI-A plates containing 5 mg/L erythromycin, were cultured in BHI broth at 37 °C overnight. DNA was extracted from these overnight cultures using the DNeasy Blood & Tissue kit (Qiagen), with the addition of 0.5 mg/mL (250 units/mL) of lysostaphin (Sigma Aldrich). The DNA was then digested with the RsaI restriction enzyme (Thermo Fisher Scientific) at 37 °C for 16 h, followed by purification using the GenJet PCR Purification Kit (Thermo Fisher Scientific).
Oligo 254 and 256 adaptors (Supplementary Table S4) were prepared by resuspending them in an annealing buffer composed of 100 mM Tris-HCl (pH 8), 500 mM NaCl, and 20 mM EDTA, and then diluting each adaptor to a concentration of 100 µM. The adaptor solutions were combined in a 1:1 ratio and denatured at 95 °C for 3 min, followed by an hour-long incubation at room temperature. The annealed adaptors were then ligated to the digested DNA using T4 ligase (Thermo Fisher Scientific) at 22 °C for an hour. Post-ligation, DNA was purified using the GenJet PCR Purification Kit (Thermo Fisher Scientific) and subjected to PCR using Phusion high-fidelity DNA Polymerase (Thermo Fisher Scientific) with specific primers (forward TnL and reverse primer 258) (Supplementary Table S4). The PCR conditions were as follows: an initial denaturation for 3 min, followed by 30 cycles of 45-sec denaturation at 94 °C, 1 min annealing at 53 °C, and 2 min elongation at 72 °C, with a final 10 min elongation at 72 °C. PCR products were then visualized on a 1% agarose gel run at 90 volts for 30 min.
TraDIS sequencing and data processing
DNA was extracted from two aliquots of the previously mentioned two cured libraries, separately. Two µg of each extracted DNA sample in 100 µL H2O were sheared to an average fragment size of 300–400 bp using a sonication device (Bioruptor® Pico, Diagenode) with the following profile: 10 cycles of 30 s ON, 90 s OFF at low frequency. Following the methodology outlined by Barquist, et al.44, these fragmented DNA samples were prepared for TraDIS with adapters and primers previously designed18. The libraries were sequenced on an Illumina MiSeq platform using Miseq V2 50 cycles kit resulting in reads with the initial 10 bp (CAACCTGTTA) being transposon insert-specific followed by the junction region.
The resulting data were subsequently processed for bioinformatic analysis using the Bio-Tradis pipeline (https://github.com/sanger-pathogens/Bio-Tradis). This analysis generated files that facilitated the determination of the number of reads and the accurate enumeration of UISs for each gene within the JE2 reference genome (Genbank: CP000255.1). To compare output reads of the optimized curing method with those of the conventional method, the tradis_comparison.R script was used. The UISs were visualized at both the individual gene level and within a circular representation of the genome using Artemis (version 18.2.0) and Artemis DNAPlotter (version 18.2.0), respectively.
Data availability
Raw sequencing data and metadata are available in the NCBI Sequence Read Archive (SRA), under BioProject accession no. PRJNA1137847, https://www.ncbi.nlm.nih.gov/bioproject/PRJNA1137847.
References
Turner, N. A. et al. Methicillin-resistant Staphylococcus aureus: an overview of basic and clinical research. Nat. Rev. Microbiol.17, 203–218 (2019).
Novick, R. P., Edelman, I., Latta, P. D., Swanson, E. C. & Pattee, P. A. Translocatable elements in Staphylococcus aureus. Contrib. Microbiol. Immunol.6, 41–55 (1979).
Berger-Bächi, B. Insertional inactivation of staphylococcal methicillin resistance by Tn551. J. Bacteriol.154, 479–487 (1983).
Mani, N., Tobin, P. & Jayaswal, R. K. Isolation and characterization of autolysis-defective mutants of Staphylococcus aureus created by Tn917-lacZ mutagenesis. J. Bacteriol.175, 1493–1499 (1993).
Mei, J., Nourbakhsh, F., Ford, C. W. & Holden, D. W. Identification of Staphylococcus aureus virulence genes in a murine model of bacteraemia using signature-tagged mutagenesis. Mol. Microbiol.26, 399–407 (1997).
Cain, A. K. et al. A decade of advances in transposon-insertion sequencing. Nat. Rev. Genet.21, 526–540 (2020).
Champie, A. et al. Enabling low-cost and robust essentiality studies with high-throughput transposon mutagenesis (HTTM). PLoS ONE. 18, e0283990 (2023).
Shields, R. C. & Jensen, P. A. The bare necessities: uncovering essential and condition-critical genes with transposon sequencing. Mol. Oral Microbiol.34, 39–50 (2019).
Cho, H. Transposon insertion site sequencing (TIS) of Pseudomonas aeruginosa. J. Microbiol.59, 1067–1074 (2021).
Yajjala, V. K., Widhelm, T. J., Endres, J. L., Fey, P. D. & Bayles, K. W. Generation of a Transposon Mutant Library in Staphylococcus aureus and Staphylococcus epidermidis using bursa aurealis. In The Genetic Manipulation of Staphylococci (ed Bose, J. L.) Vol. 1373, 103–110 (Springer New York, 2014).
Khatiwara, A. et al. Genome scanning for conditionally essential genes in Salmonella enterica Serotype Typhimurium. Appl. Environ. Microbiol.78, 3098–3107 (2012).
Maliszewski, K. L. Batch transduction of transposon mutant libraries for rapid phenotype screening in Staphylococcus. In The Genetic Manipulation of Staphylococci (ed Bose, J. L.) Vol. 1373, 75–81 (Springer New York, 2015).
Santiago, M. et al. A new platform for ultra-high density Staphylococcus aureus transposon libraries. BMC Genom.16, 252 (2015).
Bowring, J. Z. et al. Screening for highly transduced genes in Staphylococcus aureus revealed both lateral and specialized transduction. Microbiol. Spectr.10, e02423–e02421 (2022).
Mašlaňová, I., Stříbná, S., Doškař, J. & Pantůček, R. Efficient plasmid transduction to Staphylococcus aureus strains insensitive to the lytic action of transducing phage. FEMS Microbiol. Lett.363, fnw211 (2016).
Grosser, M. R. et al. Genetic requirements for Staphylococcus aureus nitric oxide resistance and virulence. PLoS Pathog. 14, e1006907 (2018).
Freiberg, J. A. et al. Restriction of arginine induces antibiotic tolerance in Staphylococcus aureus. Nat. Commun.15, 6734 (2024).
Christiansen, M. T. et al. Genome-wide high-throughput screening to investigate essential genes involved in Methicillin-Resistant Staphylococcus aureus sequence type 398 survival. PLoS ONE. 9, e89018 (2014).
Lo, H. Y. et al. Transposon sequencing identifies genes impacting Staphylococcus aureus invasion in a human macrophage model. Infect. Immun.91, e00228–e00223 (2023).
Monk, I. R. Genetic manipulation of Staphylococci—breaking through the barrier. Front. Cell. Inf. Microbiol.2, (2012).
Chaudhuri, R. R. et al. Comprehensive identification of Salmonella enterica Serovar typhimurium genes required for infection of BALB/c mice. PLoS Pathog. 5, e1000529 (2009).
Herron, P. R. Transposon express, a software application to report the identity of insertions obtained by comprehensive transposon mutagenesis of sequenced genomes: analysis of the preference for in vitro Tn5 transposition into GC-rich DNA. Nucleic Acids Res.32, e113–e113 (2004).
Choudhery, S. et al. Modeling site-specific nucleotide biases affecting Himar1 transposon insertion frequencies in TnSeq data sets. mSystems. 6, e00876–e00821 (2021).
Bossé, J. T. et al. Rationally designed mariner vectors for functional genomic analysis of Actinobacillus pleuropneumoniae and other Pasteurellaceae species by transposon-directed insertion-site sequencing (TraDIS). Anim. Dis.1, 29 (2021).
Pajunen, M. I. Generation of transposon insertion mutant libraries for Gram-positive bacteria by electroporation of phage Mu DNA transposition complexes. Microbiology. 151, 1209–1218 (2005).
Coe, K. A. et al. Multi-strain Tn-Seq reveals common daptomycin resistance determinants in Staphylococcus aureus. PLoS Pathog. 15, e1007862 (2019).
Lee, W. Construction of high-density transposon mutant library of Staphylococcus aureus using bacteriophage ϕ11. J. Microbiol.60, 1123–1129 (2022).
Ingmer, H., Gerlach, D. & Wolz, C. Temperate phages of Staphylococcus aureus. Microbiol. Spectr.7, 7.5.1 (2019).
Li, X. et al. An essential role for the baseplate protein Gp45 in phage adsorption to Staphylococcus aureus. Sci. Rep.6, 26455 (2016).
Zhu, X. et al. Tunicamycin mediated inhibition of Wall Teichoic Acid affects Staphylococcus aureus and Listeria monocytogenes cell morphology, biofilm formation and virulence. Front. Microbiol.9, 1352 (2018).
Nygaard, R., Romaniuk, J. A. H., Rice, D. M. & Cegelski, L. Spectral snapshots of bacterial cell-wall composition and the influence of antibiotics by whole-cell NMR. Biophys. J.108, 1380–1389 (2015).
Van Dalen, R., Peschel, A. & Van Sorge, N. M. Wall Teichoic Acid in Staphylococcus aureus host Interaction. Trends Microbiol.28, 985–998 (2020).
Kim, S. G. et al. Strategy for mass production of lytic Staphylococcus aureus bacteriophage pSa-3: contribution of multiplicity of infection and response surface methodology. Microb. Cell. Fact.20, 56 (2021).
Ding, G. J. et al. Amorphous calcium phosphate nanowires prepared using beta-glycerophosphate disodium salt as an organic phosphate source by a microwave-assisted hydrothermal method and adsorption of heavy metals in water treatment. RSC Adv.5, 40154–40162 (2015).
Li, X. et al. Exploring the benefits of metal ions in phage cocktail for the treatment of methicillin-resistant Staphylococcus aureus (MRSA) infection. Infect. Drug Resist.15, 2689–2702 (2022).
Ba, X. et al. High-throughput mutagenesis reveals a role for antimicrobial eesistance- and virulence-associated mobile genetic elements in Staphylococcus aureus host adaptation. Microbiol. Spectr.11, e04213–e04222 (2023).
Singh, V. K. et al. An insight into the significance of the DnaK heat shock system in Staphylococcus aureus. Int. J. Med. Microbiol.302, 242–252 (2012).
Matthews, T., Baumann, R. G. & Domanski, T. L. Analysis and characterization of Staphylococcus aureus ClpB and associated Co-chaperone proteins dnaK, dnaJ and grpE. FASEB J.21, (2007).
Proctor, R. A. et al. Small colony variants: a pathogenic form of bacteria that facilitates persistent and recurrent infections. Nat. Rev. Microbiol.4, 295–305 (2006).
Kim, D. Y., Kwon, E., Shin, Y. K., Kweon, D. H. & Kim, K. K. The mechanism of temperature-induced bacterial HtrA activation. J. Mol. Biol.377, 410–420 (2008).
Monk, I. R., Tree, J. J., Howden, B. P., Stinear, T. P. & Foster, T. J. Complete Bypass of Restriction Systems for Major Staphylococcus aureus Lineages. mBio6, e00308-15 (2015).
Monk, I. R. & Stinear, T. P. From cloning to mutant in 5 days: rapid allelic exchange in Staphylococcus aureus. Access. Microbiol.3, (2020).
Krausz, K. L. & Bose, J. L. Bacteriophage transduction in Staphylococcus aureus: broth-based method. In The Genetic Manipulation of Staphylococci (ed Bose, J. L.) Vol. 1373, 63–68 (Springer New York, 2014).
Barquist, L. et al. The TraDIS toolkit: sequencing and analysis for dense transposon mutant libraries. Bioinformatics. 32, 1109–1111 (2016).
Acknowledgements
This study has received funding from the European Union’s Horizon 2020 research and innovation program under the Marie Sklodowska-Curie grant agreement No. 956154. Special thanks are extended to Prof. Simon Foster (University of Sheffield, UK) for kindly providing the transposon and transposase plasmids and phage. We would also like to thank Prof. John Elmerdahl Olsen (University of Copenhagen, Denmark) for providing the wild-type strains and for reviewing the manuscript. We extend our profound gratitude to the OHAR group led by Prof. Luca Guardabassi (University of Copenhagen, Denmark), for hosting and providing access to the sequencing platform.
Author information
Authors and Affiliations
Contributions
SY and NA were involved in the conception and design of the study, acquisition of data, and data analysis and interpretation, drafted the manuscript and revised it critically for important intellectual content. BP provided overall supervision, contributed to the study’s design and coordination, and was responsible for the final approval of the version to be published. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yousief, S.W., Abdelmalek, N. & Paglietti, B. Optimizing phage-based mutant recovery and minimizing heat effect in the construction of transposon libraries in Staphylococcus aureus. Sci Rep 14, 22831 (2024). https://doi.org/10.1038/s41598-024-73731-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-73731-y