Abstract
Lung disease analysis in chest X-rays (CXR) using deep learning presents significant challenges due to the wide variation in lung appearance caused by disease progression and differing X-ray settings. While deep learning models have shown remarkable success in segmenting lungs from CXR images with normal or mildly abnormal findings, their performance declines when faced with complex structures, such as pulmonary opacifications. In this study, we propose AMRU++, an attention-based multi-residual UNet++ network designed for robust and accurate lung segmentation in CXR images with both normal and severe abnormalities. The model incorporates attention modules to capture relevant spatial information and multi-residual blocks to extract rich contextual and discriminative features of lung regions. To further enhance segmentation performance, we introduce a data augmentation technique that simulates the features and characteristics of CXR pathologies, addressing the issue of limited annotated data. Extensive experiments on public and private datasets comprising 350 cases of pneumoconiosis, COVID-19, and tuberculosis validate the effectiveness of our proposed framework and data augmentation technique.
Similar content being viewed by others
Introduction
Lung diseases are one of the leading causes of death and disability worldwide1,2. Recently, the COVID-19 pandemic killed many people and burdened healthcare systems3,4. Chest X-ray (CXR) images are widely used for analysis of many pulmonary diseases due to low radiation dosage, availability, and low cost5,6. Due to the subtle features of lung diseases, and proximity of other anatomical regions, accurate lung segmentation is an important step of chest X-ray image analysis for lung disease diagnosis5,7. For segmentation tasks, manual image annotation, and particularly at pixel-level, is labor- intensive, tedious, time consuming, and thus expensive. High inter-observer and intra-observer variations have been reported due to blurred lung boundaries8,9. An automatic robust lung segmentation tool would be valuable to enable computer-aided diagnosis in detecting and analyzing pulmonary disorders, monitoring their progression, and recovery for improved patient outcomes.
In this study, we focus on lung segmentation in challenging CXR images that include pneumoconiosis, COVID-19, and tuberculosis. This task is especially challenging due to the complexity of the images and scarcity of annotated data. For example, inhalation of respirable particles, like coal dust, can lead to inflammation in the lungs, known as pulmonary opacification2. CXR images with such opacifications have ambiguous lung boundaries and therefore are difficult to segment7,8. Patient age, co-morbidities, poor contrast, artefacts, and overlapping of the lungs with other anatomic structures, such as heart and rib cage, also contribute to the challenge of lung segmentation7. Finally, the lack of standardised acquisition and limited availability of public dataset with annotation, often hamper the performances of machine learning based system2. In Fig. 1, some examples of CXR images with pneumoconiosis, COVID-19, and tuberculosis are shown. The main contributions of this work are summarised as follows:
-
1.
Addressing the challenging problem of segmentation of lungs with severe abnormalities, especially pulmonary opacification. Most studies have worked on segmenting images with normal or mild conditions8,10, while this work addresses the segmentation of severely diseased lungs.
-
2.
Development of a novel deep learning-based model, suitable for challenging medical image segmentation tasks, including cases with ambiguous lung boundaries.
-
3.
Introducing a novel data augmentation technique to simulate the features and characteristics of CXR images with complex structures, particularly clusters of lesions. The effectiveness of the proposed data augmentation technique is demonstrated by training the model on a small number of CXR images with normal or mild abnormalities and testing on independent data sets that contain images with various conditions including extreme cases of opacification and low-quality images.
Related works
This section presents related work for segmentation models and data augmentation techniques.
Segmentation models
Lung segmentation in CXR images has received substantial attention7,8,9, however it remains a challenging problem, especially for pathological lungs with blurred lung boundaries8,10. Using convolutional networks, Ronneberger et al.11 proposed a U-Net architecture that has achieved remarkable success for lung segmentation and that we considered state-of-the-art (SOTA). In our experiment, that model (U-Net) does not always work consistently on images with complex abnormalities, which might be due to identified limitations, including the semantic gap between the encoder and decoder caused by skip connections, and the loss of spatial information due to repetitive down-sampling operations12,13.
Several variations of U-Net have been proposed14,15. UNet++13 solved the semantic gap problem by redesigning the U-Net architecture with nested and dense skip connections. Oktay et al.16 proposed Attention U-Net to improve the performance of medical image segmentation. They modified U-Net using an attention gate to focus on target structures of various shapes and sizes. MultiResUNet17 is another extended version of U-Net that replaced the convolutional layers of the standard U-Net architecture with MultiRes blocks, each of which consists of convolutional layers with different kernel sizes and a residual connection to extract multi-scale features. Dual Channel U-Net (DC-UNet)18 is another potential successor to the U-Net model, which is based on the concept of different-scale features and residual connections. The authors reported improved segmentation performance compared to the SOTA model, particularly on challenging images. Based on the Deep Residual U-Net (ResUNet), Jha et al.12 designed ResUNet++ to segment medical images. They claimed significant improvement of segmentation performance using this architecture compared to U-Net and ResUNet. In 2023, Xu et al. introduced DCSAU-Net19, a deeper and more compact split-attention U-shaped network, for medical image segmentation. Recently, Dai et al.20 developed a dual-path U-Net with rich information interaction for medical image segmentation. In this paper, we refer to this network as I2U-Net for convenience. Different from these studies, in this work we aim for accurate lung segmentation in challenging CXR images by capturing rich context information through attention gate and our proposed multi-scale residual block that can capture multi-scale information at a granular level.
Data augmentation techniques
Deep learning networks trained on a small number of CXR images perform poorly on unseen test images that contain variations not observed during training21. Specifically, when trained on CXR images with normal or mild opacities performance often degrades to segment other images with dense opacities due to large feature variations, as shown in Fig. 1. Data augmentation techniques attempt to address this issue21,22,23.
In 2017, DeVries24 introduced cutout, which is a regularisation technique that involves random masking of some regions of the input images to generate partially occluded versions of existing samples. Using this method, the authors showed improved robustness and overall classification performance of CNN-based networks. Varkarakis et al.23 explored the effectiveness of different data augmentation techniques for iris segmentation. Due to the lack of training samples with ground truth, they simulated the effects of real-world conditions in iris images with different techniques including varying image contrast, spatial stretching, and tilting. Bae et al.22 used Perlin noise-based data augmentation to mimic different patterns of diffuse interstitial lung disease (DILD) in high-resolution computed tomography (HRCT) scans. They reported improved classification performance of deep neural networks with Perlin noise compared to several conventional augmentation methods. Unlike other studies, we explored the effects of different data augmentation techniques comprehensively on lung segmentation performance of challenging CXR images and proposed a new data augmentation technique.
Methodology
In this section, details of the proposed attention-based multi-residual UNet++ (AMRU++) network are provided, followed by the description of the proposed data augmentation technique for lung segmentation on CXR images.
Proposed network architecture
For segmenting challenging images, there is a gap between the results provided by radiologists and those produced using CNN. This could be due to the fixed geometric structures of convolution blocks, which may not capture optimal spatial features, and the loss of spatial information due to consecutive pooling and convolution striding operations8,12,13. This can in turn affect the segmentation of lungs with blurred boundaries caused by poor image quality or pathological conditions. A solution to the information recession problem is to extract multi-scale contextual features and concatenate them into a dense feature map25. Extraction of discriminative features and rich semantic context information is critical to segmenting images with complex structure26.
Several feature extraction blocks have been proposed17,25,27,28. He et al.29 introduced the residual block that alleviates the vanishing gradient problem, propagates low-level fine details, and converges faster. Ibtehaz et al.17 and Li et al.25 proposed a multi-scale residual block to extract rich contextual features to improve model performance. Different from these studies, our proposed block can capture multi-scale information at a granular level and focus on relevant important information through an attention module that is found to be effective for lung segmentation. Both Squeeze-and-Excitation (SE)27 and attention gate (AG) modules16 can focus more on essential features of varying shapes and sizes, which are necessary in biomedical image segmentation30.
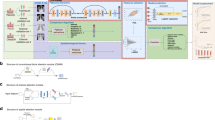
Motivated by the success of UNet++13 and different feature extraction blocks, a framework for robust lung segmentation called AMRU++ is proposed, and depicted in Fig. 2a, where the UNet++13 architecture is used as the baseline. A soft attention gate (AG) module30 is inserted between convolution blocks to focus relevant spatial information from the encoder path and propagate information to the decoding path. To extract discriminative features, the basic building blocks in the UNet++ architecture13 are replaced with the proposed multi-scale residual block (Fig. 2b), now discussed.
Multi-scale residual (MR) block consists of two-bypass networks that use different dilation rates and a convolution kernel of the same size \([ 3\times 3]\). To take advantage of multi-scale feature extraction at the granular level, the input feature map of \(x_i\) is split into two equal parts \(x_{i1}\) and \(x_{i2}\) empirically- the latter are two subsets of \(x_i\) with the same spatial size and half the number of channels. For the first group of feature maps \(x_{i1}\), the standard convolution operation (dilation rate = 1) is used; for the second group of feature maps \(x_{i2}\), a convolution operation with a dilation rate = 2, chosen empirically, is used to extract features using large size receptive fields. To extract rich contextual information, features from the two groups are shared with each other. The operations of the bypass networks can be defined by the following transformations:
where \(\beta (\cdot )\) denotes the batch normalisation function and \(\sigma (\cdot )\) represents the rectified linear units (ReLU) activation function. Batch normalisation is used before activation to speed up network convergence12. Similarly, w and b are the weights and biases respectively. The subscripts of w represent the convolution filter size \([3\times 3]\) and the DR used in the layer, the superscripts represent the number of the layers at which they are located and [,] represents the concatenation operation. To focus on more informative features and discard redundant ones, an SE unit is inserted as follows:
where \(f(\cdot )\) represents the residual learning function that performs a nonlinear transformation with a series of operations, and \(\epsilon (\cdot )\) denotes the Squeeze-and-Excitation function. Finally, to increase the gradient flow, a residual connection is adopted for each block. Each MR block can then be expressed as follows:
where \(x_i\) and \(x_{i+1}\) represent the input and output of the \(i\)-th MR block. The operation \(f(x_i)+x_i\) is performed using elementwise addition. Both the input and output have the same resolution.
Proposed data augmentation method
The effectiveness of conventional data augmentation techniques (i.e., rotation, flip) on CXR images for segmentation was examined using the U-Net architecture11, (see Fig. 3 for illustrative examples). The CXR images were randomly rotated by selecting a rotation angle from the uniform distribution over (\(-15\), \(+15\)) degrees, and the images were flipped along the X-axes. For shifting and scaling, factors of 0.05 and 0.05 respectively were used.
The cutout method, which randomly masks square regions of images, can simulate extreme levels of opacification that obscure lung areas. However, this technique was initially designed for natural images, such as those in the CIFAR10 and SVHN datasets. To better mimic features caused by lesions in the medical imaging domain, the standard cutout algorithm is modified and is referred to as Selective Cutout (Scutout). Scutout creates an equal number of masked regions (referred to as holes) in both the left and right lungs, with 20% of the holes placed inside the lungs and 80% along the lung borders (see Fig. 3). In this study, Scutout is also referred to as the proposed data augmentation technique.
Materials and system implementation
This section describes the datasets used in this study and the experimental setup.
Datasets
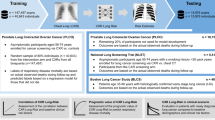
In our study, five CXR datasets were used for comparison and their details are summarized in Table 1. For lung segmentation, all models were developed on a combined dataset (denoted as MJ) from Montgomery32 and Japanese Society of Radiological Technology (JSRT)31, and consists of 385 CXR images (138+247). We observed that most of the lung regions in the MJ datasets are normal or have mild disease conditions. However, in real life scenarios, lung diseases like pneumoconiosis can cause severe lung damage34; therefore, we used three independent test sets that contain images with more challenging conditions with severe lung damage. The Shenzhen set32 dataset contains X-rays images showing manifestations of tuberculosis. We used 100 abnormal images from the Shenzhen set dataset to evaluate the performance of lung segmentation. We used an additional publicly available COVID-19 dataset33, containing 50 CXR images, referred to as COVID.
Furthermore, we incorporated a private pneumoconiosis dataset named GMH, comprising 200 CXR images obtained from Good Morning Hospital, South Korea. In contrast to the COVID dataset, the images from the GMH dataset presented greater challenges due to the presence of opacities of different sizes and shapes, existence of other diseases, and the fact that many of the X-ray images were acquired from aged patients. The publicly available datasets, MJ and Shenzhen set, came with ground truth annotations. Two radiologists from St Vincent’s Hospital (Sydney) assisted us to obtain ground truth masks for the remaining 250 CXR images (50 images from the COVID dataset and 200 images from GMH).
Experimental setup
All experiments were run on a Dell C4140 server in a High-Performance Computing (HPC) cluster with 4 \(\times\) Nvidia V100 GPUs, 2 \(\times\) Intel Xeon 6130 CPUs, and 192 GB RAM (12 \(\times\) 16 GB). TensorFlow35 was used to implement the proposed model. We investigated the performance of different models using both default and customised hyperparameters. For example, learning rates ranging from \(4 \times 10^{-2}\) to \(4\times 10^{-6}\) and batch sizes of 4, 8, and 16 were explored. While the performance was comparable across different hyperparameter settings, we observed slightly better results with a learning rate of \(4 \times 10^{-4}\) and a batch size of 8. For all networks, Adam optimiser36 was used to optimise the network with batch size 8 and trained for 50 epochs. The initial learning rate was set to \(4 \times 10^{-4}\). An early stop technique was used to avoid overfitting and save training time. To reduce the memory and computation overhead, all images were down sampled to size 256 x 256. We evaluated the models using Dice similarity coefficient (DSC)26 and Jaccard Index (JI)26. To unify the contrast level of CXR images from different datasets, we used histogram matching. All training datasets were split using five-fold cross validation.
We examined the impact of both the number of augmented image ratios and the types of augmentation methods. Different augmented training sets were generated based on varying augmentation percentages. For instance, with a training set of 245 original images, 100% augmentation means that 245 augmented images were created and added to the original set, resulting in 490 (245+245) total images for model training. For lung segmentation, post-processing techniques were applied to fill gaps in the lung regions using a flood-fill algorithm37 and to eliminate small, irrelevant objects outside the lung field.
Ethics approval and consent to participate
CSIRO Health and Medical Human Research Ethics Committee, Australia, granted approval for this research (approval number: LR 22/2016), and waived a requirement of informed consent since data were evaluated retrospectively, pseudonymously, and was solely obtained for treatment purposes. All methods were performed in accordance with the relevant institutional guidelines and regulations, and all data used for the research are de-identified.
Results
This section presents the results of various models without data augmentation and with the application of data augmentation techniques.
Results without Augmentation
To evaluate the performance of different architectures, we first trained all models without applying any data augmentation techniques. Table 2 presents a comparison of the proposed AMRU++ model against other SOTA models across four datasets (with the Montgomery and JSRT datasets combined as one). The results show that, for images with normal or mild conditions, such as those from the MJ dataset, ResUNet++ performs the best, while the performance of our proposed AMRU++ is comparable. However, AMRU++ significantly outperforms all other models on the remaining test datasets. For images with mild to moderate conditions, such as those from the COVID and Shenzhen set datasets, performance is comparable among different networks though AMRU++ outperforms all other networks. In more challenging cases, such as the GMH dataset, the performance of other models declines notably, with the exception of I2U-Net. However, remarkable improvements are observed on the GMH dataset using the proposed AMRU++ architecture, where DC-UNet and DCSAU-Net achieved the worst results. The images from the Shenzhen set and GMH datasets are challenging for segmentation due to blurred object boundaries, which negatively affects the performance of most models. Experimental results suggest that the AMRU++ architecture is especially effective in segmenting lungs from challenging images with blurred boundaries. This is likely due to the contextual information and strong discriminative features extracted through the multi-scale context and attention mechanisms, which are critical for medical image segmentation. Additionally, AMRU++ consistently excels across all datasets, while other models tend to perform well on some datasets but struggle on others, highlighting the robustness of the proposed architecture. However, the proposed architecture utilizes more parameters than other networks, making it computationally more expensive in terms of Floating Point Operations Per Second (FLOPs) (see Table 2).
Statistical analysis
To evaluate the effectiveness of our proposed model without data augmentation, we measured the statistical significance of the difference between model performances on three datasets: namely GMH, COVID, and Shenzhen set. Mann-Whitney U tests38, the nonparametric equivalent of the independent two-sample t-test, were used to measure the statistical differences in the segmentation performance of each pair of networks. The bubble plots in Fig. 4 present the statistical significance in terms of DSC. Statistical differences in terms of JI were similar to DSC and therefore omitted. In the figure, ‘no significance’ indicates that the difference is not statistically significant (p-value greater than 0.05), while bubbles of different colours and sizes represent four levels of significance (0.05, 0.01, 0.001, and 0.0001) measured by p-values. For the GMH dataset, the proposed architecture outperforms all other networks significantly with p-value \(<0.0001\) except I2U-Net. There is no statistically significant difference between AMRU++ and I2U-Net. For the COVID dataset, a significantly different score between AMRU++ and U-Net is observed (\(p < 0.05\)). There is no statistically significant difference between AMRU++ and (UNet++, AttentionUNet, ResUNet++, DCUNet, DCSAU-Net and I2U-Net) for this dataset. The proposed AMRU++ architecture outperforms all other architectures, except ResUNet++, DCSAU-Net and I2U-Net, at different significance levels, for the Shenzhen set dataset.
Bubble plots showing statistical significance based on Mann-Whitney U test on each pair of networks in terms of DSC on three datasets. ‘no significance’ indicates not statistically significant, bubbles with different colours and sizes indicate different levels of significance calculated by p-values.
Ablation study
This subsection briefly describes the ablation study (Table 3) conducted to validate the individual contribution of different strategies of the proposed framework.
-
1.
Impact of AG: First, the effectiveness of AG was explored, compared to the baseline, see row 2 in Table 3. Results suggest that AG is effective in improving the performance of segmenting images from GMH and COVID datasets. However, for the Shenzhen set dataset there is no effect of the AG module.
-
2.
Impact of DC: To investigate the effectiveness of DC, two dilation rates of DR = 1 and DR = 2 were used empirically for two different branches. From Table 3, it is observed that Base + DC (row 3) can boost the segmentation performance for all datasets. This suggests that extracting multi-scale context features with varying sizes of receptive fields using different DR is effective for the segmentation of images with a complex structure that could be due to various reasons such as opacities. This result is consistent with the previous experiments.
-
3.
Impact of multi-scale feature extraction at a granular level: The importance of multi-scale feature extraction at a granular level was evaluated by splitting feature maps (SFM) into two groups and applying convolution operations with different DR on the two groups. The results in row 4 (Base + DC + SFM) in Table 3 indicate that the multiscale representation ability at a more granular level is necessary for improving the segmentation of all images.
-
4.
Impact of SE unit: The performance of the SE unit was also investigated by integrating it with DC and SFM components (row 5 in Table 3) and its effectiveness validated in segmenting images from all datasets. The SE module slightly improves the segmentation performance on all datasets. Finally, all components were combined (row 6 in Table 3) to develop the proposed model and the results suggest that this combination does segment images more accurately.
Results with augmentation
The segmentation performance of U-Net trained on different augmented datasets, with varying percentages of data augmentation ranging from 50 to 200%, is presented in Fig. 5. As the aim of this study is to segment lungs in complex images, the GMH, COVID and Shenzhen set datasets were selected for evaluation, with DSC used as the primary evaluation metric. For all datasets, as expected, the worst performance was observed when no data augmentation (denoted No_aug) was used.
For images with mild or moderate conditions from the COVID dataset, the proposed Scutout data augmentation method performs comparably to conventional and standard cutout techniques. However, for more challenging images from the GMH dataset and Shenzhen set, the Scutout method outperforms all other augmentation techniques. This superior performance may be attributed to Scutout’s ability to effectively mimic opacities in CXR images from the GMH and Shenzhen datasets. In contrast, the performance of standard cutout methods is inconsistent across different augmentation percentages. This inconsistency may arise from the fact that the standard cutout was originally designed for natural images, where random regions are removed. If the masked regions do not cover key lung areas, particularly in diseased lungs, the segmentation may be less effective, leading to variable performance. On the other hand, conventional data augmentation techniques achieved the worst performance on these images. This means that the contextual information of the images generated using the traditional augmentation techniques does not change much and therefore the models trained using the generated images are not effective in segmenting unseen images with complex structures due to opacities.
Performance improved gradually with the increase in the number of augmented images up to a certain percentage, and further increases did not improve the performance any more. When the number of images synthesised from the augmentation method increases above a certain percentage, the model may be overfitted and therefore does not perform well on new data. Therefore, we considered upto 200% augmented data to train all the models.
Table 4 presents the performance of our proposed Scutout data augmentation method across various network architectures for lung segmentation. All models were trained using 200% augmented data generated with the Scutout technique. Our experimental results demonstrate that the proposed Scutout data augmentation technique improves segmentation performance across all models. For instance, using the U-Net model on the GMH dataset, a DSC of 0.9065 is achieved with Scutout augmentation, compared to 0.8475 without augmentation (see Table 2 and Table 4). Similarly, the DSC score of the proposed model improved from 0.9097 to 0.9363 with Scutout augmentation. The results indicate that the Scutout technique is particularly effective for the GMH and Shenzhen set datasets (see Table 2 and Table 4).
For images with moderate to severe disease conditions (Shenzhen sets and GMH), the proposed model consistently outperformed all others in terms of DSC. In contrast, for the COVID dataset, where conditions were milder, performance across different models was comparable, though the proposed model achieved the best results. On the other hand, AttentionUNet and DC-UNet achieved the worst performance across all datasets. I2U-Net ranked second on the GMH and COVID datasets, while ResUNet++ achieved the second-best results for the Shenzhen set dataset.
Qualitative results
Visual comparison of segmentation performance of the proposed AMRU++ and the SOTA methods on different medical applications is depicted in Fig. 6. Three images were selected from three different datasets. For the images, two predicted masks are shown - one was produced without any data augmentation denoted as ‘w/o DA’ and another was generated with our proposed data augmentation denoted as ‘with PDA’ (200% augmented data generated with the Scutout technique). The results suggest that without data augmentation, the proposed architecture shows the best performance for images from all datasets used in the experiments. However, for the GMH and COVID datasets, the performance of the proposed model is comparable with I2U-Net, and UNet++ respectively. And for Shenzhen set dataset, the performance of the proposed model is comparable with AttentionUNet. On the other hand, the proposed data augmentation technique can improve the segmentation performance of all evaluated models on all image datasets. The proposed data augmentation technique is effective for images without clear lung boundaries due to opacities or artifacts. The experimental results reveal that the proposed model together with the proposed data augmentation technique can be used to segment lungs accurately.
Visual comparison of segmentation performance for models w/o DA and with PDA. One image was selected from each of the three datasets. For CXR images, two predicted masks are shown, one without any data augmentation denoted as ‘w/o DA’ and another with the proposed data augmentation denoted as ‘with PDA’.
Conclusion
Over the past few decades, numerous methods have been developed for lung segmentation. While SOTA models achieve good performance on normal lung images, their effectiveness diminishes when dealing with images that feature complex structures. Additionally, the performance of deep learning-based models is often constrained by the limited availability and diversity of training data. This study focuses on lung segmentation in diseased lung images, addressing these challenges.
The main contributions of this study are twofold: first, we have proposed a novel deep learning architecture, namely AMRU++, which effectively captures rich contextual information and strong discriminative features, outperforming existing segmentation models. Second, we have introduced an innovative data augmentation technique that generates synthetic images capable of mimicking complex structures such as pulmonary opacities. Experimental results demonstrate the superior performance of the proposed network and augmentation technique.
The proposed network architecture can be trained to segment other types of 2D medical images, such as skin lesions in dermoscopy images. However, a limitation of the AMRU++ architecture is its increased number of parameters compared to other models, which results in longer training time. In future, we shall develop a simpler architecture with fewer parameters while maintaining performance. The current evaluation was limited to small datasets from the X-ray domain. Future work will extend to other types of 2D medical images, such as retinal images with vascular structures. Additionally, we plan to assess the proposed data augmentation method for disease classification using CXR images.
Data availability
Private datasets are not publicly available due to restrictions in data sharing agreements with third-party X-ray providers. Access to the data can be requested by contacting D.W., subject to approval under the terms of the data-sharing agreement. Public dataset NIOSH is available at https://www.cdc.gov/niosh/learning/b-reader/start/1.html
References
Aboyans, V. & Causes, C. Global, regional, and national age-sex specific all-cause and cause-specific mortality for 240 causes of death, 1990–2013: A systematic analysis for the global burden of disease study 2013. The Lancet 385, 117–171 (2013).
Zosky, G. R. et al. Coal workers’ pneumoconiosis: An Australian perspective. Med. J. Aust. 204, 414–418 (2016).
Fan, D.-P. et al. Inf-Net: Automatic COVID-19 lung infection segmentation from CT images. IEEE Trans. Med. Imaging 39, 2626–2637 (2020).
Wang, G. et al. A noise-robust framework for automatic segmentation of COVID-19 pneumonia lesions from CT images. IEEE Trans. Med. Imaging 39, 2653–2663 (2020).
Neal, R. D. et al. Immediate chest X-ray for patients at risk of lung cancer presenting in primary care: Randomised controlled feasibility trial. Br. J. Cancer 116, 293–302 (2017).
Eslami, M. et al. Image-to-images translation for multi-task organ segmentation and bone suppression in chest X-ray radiography. IEEE Trans. Med. Imaging 39, 2553–2565 (2020).
Novikov, A. A. et al. Fully convolutional architectures for multiclass segmentation in chest radiographs. IEEE Trans. Med. Imaging 37, 1865–1876 (2018).
Mittal, A., Hooda, R. & Sofat, S. Lung field segmentation in chest radiographs: A historical review, current status, and expectations from deep learning. IET Image Process. 11, 937–952 (2017).
Anis, S. et al. An overview of deep learning approaches in chest radiograph. IEEE Access 8, 182347–182354 (2020).
Tang, Y.-B., Tang, Y.-X., Xiao, J. & Summers, R. M. Xlsor: A robust and accurate lung segmentor on chest X-rays using criss-cross attention and customized radiorealistic abnormalities generation. In International Conference on Medical Imaging with Deep Learning 457–467 (PMLR, 2019).
Ronneberger, O., Fischer, P. & Brox, T. U-net: Convolutional networks for biomedical image segmentation. In International Conference on Medical Image Computing and Computer-Assisted Intervention 234–241 (Springer, 2015).
Jha, D. Resunet++: An advanced architecture for medical image segmentation. In 2019 IEEE International Symposium on Multimedia (ISM) 225–2255 (IEEE, 2019).
Zhou, Z., Siddiquee, M. M. R., Tajbakhsh, N. & Liang, J. UNet++: Redesigning skip connections to exploit multiscale features in image segmentation. IEEE Trans. Med. Imaging 39, 1856–1867 (2020).
Ullah, Z., Usman, M., Latif, S., Khan, A. & Gwak, J. SSMD-UNet: Semi-supervised multi-task decoders network for diabetic retinopathy segmentation. Sci. Rep. 13, 9087 (2023).
Ullah, Z., Usman, M., Jeon, M. & Gwak, J. Cascade multiscale residual attention CNNs with adaptive ROI for automatic brain tumor segmentation. Inf. Sci. 608, 1541–1556 (2022).
Oktay, O. Attention u-net: Learning where to look for the pancreas. arXiv:1804.03999 (2018).
Ibtehaz, N. & Rahman, M. S. MultiResUNet: Rethinking the U-Net architecture for multimodal biomedical image segmentation. Neural Netw. 121, 74–87 (2020).
Lou, A., Guan, S. & Loew, M. H. DC-UNet: Rethinking the U-Net architecture with dual channel efficient CNN for medical image segmentation. In Medical Imaging 2021: Image Processing (eds Landman, B. A. & Išgum, I.) (SPIE, 2021).
Xu, Q., Ma, Z., Na, H. & Duan, W. DCSAU-Net: A deeper and more compact split-attention U-Net for medical image segmentation. Comput. Biol. Med. 154, 106626 (2023).
Dai, D. et al. I2u-net: A dual-path u-net with rich information interaction for medical image segmentation. Med. Image Anal. 103241 (2024).
Chlap, P. et al. A review of medical image data augmentation techniques for deep learning applications. J. Med. Imaging Radiat. Oncol. 65, 545–563 (2021).
Bae, H.-J. et al. A Perlin noise-based augmentation strategy for deep learning with small data samples of HRCT images. Sci. Rep. 8, 17687 (2018).
Varkarakis, V., Bazrafkan, S. & Corcoran, P. Deep neural network and data augmentation methodology for off-axis iris segmentation in wearable headsets. Neural Netw. 121, 101–121 (2020).
DeVries, T. Improved regularization of convolutional neural networks with cutout. arXiv:1708.04552 (2017).
Li, J., Fang, F., Mei, K. & Zhang, G. Multi-scale residual network for image super-resolution. In Proceedings of the European Conference on Computer Vision (ECCV) 517–532 (2018).
Alam, M. S., Wang, D., Liao, Q. & Sowmya, A. A multi-scale context aware attention model for medical image segmentation. IEEE J. Biomed. Health Inf. (2022).
Hu, J., Shen, L. & Sun, G. Squeeze-and-excitation networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition 7132–7141 (2018).
Gao, S.-H. et al. Res2Net: A new multi-scale backbone architecture. IEEE Trans. Pattern Anal. Mach. Intell. 43, 652–662 (2021).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR) (IEEE, 2016).
Abraham, N. & Khan, N. M. A novel focal tversky loss function with improved attention u-net for lesion segmentation. In 2019 IEEE 16th International Symposium on Biomedical Imaging (ISBI 2019) (IEEE, 2019).
Shiraishi, J. et al. Development of a digital image database for chest radiographs with and without a lung nodule: Receiver operating characteristic analysis of radiologists’ detection of pulmonary nodules: Receiver operating characteristic analysis of radiologists’ detection of pulmonary nodules. AJR Am. J. Roentgenol. 174, 71–74 (2000).
Jaeger, S. et al. Two public chest x-ray datasets for computer-aided screening of pulmonary diseases. Quant. Imaging Med. Surg. 4, 475–477 (2014).
Cohen, J. P. et al. COVID-19 image data collection: Prospective predictions are the future. arXiv:2006.11988 (2020).
Alam, M. S., Wang, D. & Sowmya, A. Bidirectional convolutional-LSTM based network for lung segmentation of chest X-ray images. In 2021 IEEE 33rd International Conference on Tools with Artificial Intelligence (ICTAI) (IEEE, 2021).
Singh, P., Manure, A., Singh, P. & Manure, A. Introduction to tensorflow 2.0. In Learn TensorFlow 2.0: Implement Machine Learning and Deep Learning Models with Python 1–24 (2020).
Kingma, D. P. & Ba, J. Adam: A method for stochastic optimization. arXiv:1412.6980 (2014).
Smith, A. R. Tint fill. In Proceedings of the 6th Annual Conference on Computer Graphics and Interactive Techniques (ACM, New York, 1979).
Dodge, Y. The Concise Encyclopedia of Statistics (Springer Science & Business Media, Berlin, 2008).
Acknowledgements
This work was funded in part by Coal Services Health and Safety Trust Project No. 20656, and approved by CSIRO Health and Medical Human Research Ethics Committee, approval number: LR 22/2016. We would like to thank Good Morning Hospital, South Korea, for providing chest X-rays of coal mine workers and St Vincent’s Hospital, Sydney, for annotating the X-ray images.
Author information
Authors and Affiliations
Contributions
Conceptualization, and formal analysis, M.S.A., D.W., and A.S.; methodology, M.S.A.; resources and data curation, M.S.A., D.W., Y.A., J.A.E., J.K., L.S. and D.Y.; writing: original draft and design of figures, M.S.A.; writing-review and editing, M.S.A., D.W., Y.A., O.S. and A.S.; supervision, M.S.A., D.W., O.S. and A.S.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Alam, M.S., Wang, D., Arzhaeva, Y. et al. Attention-based multi-residual network for lung segmentation in diseased lungs with custom data augmentation. Sci Rep 14, 28983 (2024). https://doi.org/10.1038/s41598-024-79494-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-79494-w