Abstract
Salsola plants are halophytic crops that are distributed worldwide, with more than 100 species figured out in Asia, the Mediterranean region and North Africa. Different Salsola species were reported to exert marked anti-inflammatory activities, whereas the potential anti-inflammatory activities of the three species, S. tetrandra, S. tetragona and S. vermiculata, have not been evaluated. This study provides a comprehensive metabolic study of the shoots and roots of those three species to identify potential anti-inflammatory candidates. An ultra-high performance liquid chromatography mass-mass spectrometry (UHPLC MS/MS) method in conjunction with multivariate analysis principles was utilized in an attempt to decipher their bio-active metabolites and their relevant anti-inflammatory activities. Eighty metabolites were identified in the tested extracts, where nitrogenous compounds and phenolics were highly detected in S. tetragona samples, meanwhile, saponins and phenolic acids were highly dominant in S. tetrendra sample and S. vermiculata samples have a similar chemical profile as S. tetrandra. Concerning the anti-inflammatory activity of the tested extracts, the safety margin of all the tested extracts was higher than that of the standard drug piroxicam. The shoots of the three species demonstrated more potent anti-inflammatory activities compared to the roots. The shoot extract of S. tetrandra was the most biologically active fraction. The obtained results revealed the shoots of the three Salosla species to be promising anti-inflammatory drug candidates of high safety and efficacy that could be used in the pharmaceutical industry.
Similar content being viewed by others
Introduction
Halophytes (salt-tolerant plants) are a group of plants that can survive in saline soils in coastal regions. They represent considerable biomass with a high degree of diversity. They represent renewable economic platforms that sustainably offer pastoral, feed, agricultural and medical supplies1. Yet, few research work was carried out in order to investigate halophytes2. Genus Salsola,which belongs to Chenopodiaceae or Amaranthaceae family, is a typical representative of halophytes. Being contributing to more than 100 species distributed over the world, it represents the major genus in the family. Namely; “Salsola” is derived from the Latin name “Salsus” meaning salty due to the high tolerability of these plants to saline soils. Concerning the plant ecology, Salsola species naturally inhabit temperate, arid and semi-arid regions, which are commonly, characterized by extremely harsh conditions. Geographically, these plants are distributed in central and southwestern Asia, the Mediterranean region and North Africa. That genus is cosmopolitan, where diversity was observed in its species ranging from perennial herbs to annuals, sub-shrubs, shrubs and small trees. Unfortunately, discrimination between different species of Salsola is challenging because of their close morphological similarities2,3. Moreover, fourteen species of the investigated genus are endemic to Egypt, located on the Mediterranean Coast, Sinai and the Red Sea region4.
Previous phytochemical investigations of Salsola revealed the tremendous diversity in their matrices. Previous phytochemical investigations of different Salsola species exhibited that flavonoids, nitrogenous compounds, phenolic compounds and triterpenoids were abundant chemical classes in these extracts2,3,5. Furthermore, the biological screening of some Salsola plants has demonstrated profound activities as hypoglycemic, antioxidant, anti-microbial, cytotoxic and anti-inflammatory putative5,6,7,8,9. In the same context, the protective activities of these extracts on the cardiovascular, neurological and digestive systems were recorded3,8.
A wide range of research work effectively clarifies the anti-inflammatory activity of crude extracts and phytochemicals, exhibiting their ability to relieve different conditions of inflammation ranging from being localized to a generalized condition and from being an acute problem to a chronic matter. They also represent drugs and lead compounds for designing safe and effective anti-inflammatory agents, devoid of the serious side effects of existing synthetic molecules with anti-inflammatory potential10.
Salsola is extensively consumed in folk medicine to treat various stages of inflammatory conditions, targeting the skin, the digestive system and the respiratory system3. Most Salsola species exhibited auspicious results when testing their anti-inflammatory activity; considering the in-vitro studies, S. soda markedly inhibited aldose reductase enzyme, an enzyme responsible for the synthesis of pro-inflammatory mediators6. Moreover, the butanol extract of S. imbricta inhibited the release of nitric oxide, known to largely contribute to tissue injury. As well as S. komarovii extract, it could reduce mucosal damage in an ulcerative colitis simulating model. Whereas in-vivo studies revealed that S. cycophylla and S. imbricta extracts reduced edema, pain and inflammatory conditions in experimental animals3,9,11,12.
Despite the marked antioxidant activity of S. tetrandra and S. vermiculata, up to our knowledge, no studies for their potential anti-inflammatory activities, as well as the biological activity of S. tetragona, have been screened yet. Moreover, no previous study was able to provide a comprehensive chemical profile of these three plants. Hence, our work aimed to provide a comprehensive metabolomic study of these three Egyptian species for the first time. To fulfill this objective, an ultra-high performance liquid chromatography-mass-mass (UHPLC MS/MS) analysis was implemented in conjunction with multivariate analysis. The potential anti-inflammatory activity of the tested samples was screened to correlate the chemical metabolomic profiles of the tested extracts to their biological matrices, revealing the most biologically active fraction that could be a promising safe and effective anti-inflammatory candidate.
Methods
Plant material
Three species of Salsola were investigated in this work, namely, Salsola tetrandra Forssek, Salsola vermiculata L and Salsola tetragona Delile.
The samples were collected from Al-Amaya Zawyt Abdelkader and Burg Al-Arab in August 2021, during the vegetative periods of the three species. All samples were authenticated by Prof. Dr. Sania Ahmed, Professor of Plant Ecology, Faculty of Science, Alexandria University, through comparable identification with authentic samples allocated at the Alexandria University Herbarium. Samples of the collected species were deposited in the same herbarium, voucher numbers: Sal. tetrandra 123, Sal. tetragona 124 and Sal. vermiculata 125.
Sample preparation
Shoots were separated from the roots for each species and dried in the shed at 20 °C. Fifty grams of each shoot and root part from the three Salsola species were independently ground using an electric grinder and macerated in 70% ethanol for 14 days. We performed a pilot study to find out the optimum conditions for extracting the three species of Salsola, including extracting them for 24 h, 3 days, 7 days and 14 days as mentioned in the previous work2,13,14. The maximum number of metabolites was obtained after 14 days of maceration, as described by13.
The hydro-alcoholic extracts were dried under vacuum using Buchi rotavap RT 210 rotary evaporator13.
Sample and authentic compound preparation for UHPLC-MS/MS analysis
Dilution of each extract of the three species in HPLC-grade methanol was performed followed by filtration through a membrane disc filter of 0.2 μm pore size, followed by degassing by ultrasonic sonication and finally, 10 µL of the sample volume was loaded at a concentration of 1 mg/ml on the reversed-phase column in the full loop mode injection. The injection process of each sample occurred in five replicates on 5 consecutive days to ensure the reproducibility of the method15. Standardization, repeatability and stability of the analytical work were guaranteed by pooling a 5µL aliquot of each sample as a quality control sample that could be used as authentic data to be extracted, as it had all the sample information16.
For the authentic compounds, The procedure was performed as follows: Dilution of each extract in HPLC-grade was in methanol followed by filtration through a membrane disc filter of 0.2 m pore size, then degassing by sonication, and finally, 10 µL of the sample volume was loaded at a concentration of 1 mg/ml on the reversed-phase column in the full loop mode injection. The injection process for each sample occurred in five replicates.
The conditions of ultra-high performance liquid chromatography utilized to chemically profile Salsola species
To identify the Salsola metabolites in different extracts, a Shimadzu 8045 UPLC triple quadrupole (TQD) instrument composed of a pump (LC-2040), an autosampler (LC-2040), a detector (LC-2030/2040 PDA detector), a degasser and a Shimpak UPLC C18 column of 1.7 μm particle size, 2.1 mm internal diameter and 150 mm length was used. The pump worked in a low-pressure gradient mode with a flow rate of 0.2 ml/min and a maximum pressure of 66 MPa in auto-LPGE mode. The autosampler’s rinsing volume, rinsing speed, sampling speed, purge time and cooler temperature were 500 µL, 35 µL/s, 5 µL/s, 2 min and 15 °C, respectively. In addition, the rinse mode was repeated before and after aspiration. The photodiode array detector (PDA) was operated by a D2 lamp covering a wavelength range from 200 to 800 nm. The cell temperature was 40 °C and the slit width was 8 nm, which was used in both positive and negative modes. Channels 1 and 2 wavelengths, bandwidths and output ranges were 254 nm, 4 nm and 1.0 AU/V, respectively. The mobile phase system was ultra-pure water (Phase A) and HPLC-grade acetonitrile (Phase B) and the total chromatographic run lasted for 35 min. The elution gradient was set as the following: 10% of B at the retention time of 0.01–5.00 min, 30% of B at 5.00–15.00 min, 70% of B at 15.00–22.00, 80% of B at 22.00–26.00 and 10% of B at 29.00–30.00.
The conditions of electro-spray ionization-mass spectrometry (ESI-MS)
A mass spectrometer with a (TQD) coupled to an (ESI) source was used to analyze Salsola metabolites in both positive and negative modes. A TQD mass analyzer was used for further fragmentation of the ions, producing daughter ion fragments that facilitate the identification process. The first and third quadrupoles act as mass filters, meanwhile, the second one acts as a collision-induced dissociation cell15,17.
ESI conditions were 2.51 L/min nebulizing gas flow, 10.00 L/min drying gas flow, 300 °C interface temperature and 526 °C de-solvation temperature. The conversion dynode was 2.00 kV, the acquisition mass range was from m/z 100.00 to m/z 1500.00, which is the auto-range, the scan speed was 1428 u/s, the interface volt was 4.00Kv, the DUIS corona needle volt was 4.50 kV, the acquisition mode was Q3 scan in both segment 1 Event 1 and segment 1 Event 2, the capillary’s voltage was 3 kv, the cone’s voltage was 35 v, the ion source’s temperature was 150 °C and the nitrogen gas nebulizer’s pressure was 35 psi.
It is worth mentioning that six external standards, namely, caffeic acid, hesperidin, scopoletin glucoside, tyramine, azalaic acid and oleanolic acid (Sigma-Aldrich Co., St. Louis, MO, USA)], were used to construct calibration curves expressed in (mg standard equivalent/g) dry extract using an exact weight of the reference (10 mg) was dissolved in 10 mL of HPLC- grade methanol to create each reference solution. For generation of calibration curves from reference compounds, five µLs from each concentration level were injected onto the HPLC column in duplicate after the solution had been serially diluted to working concentrations. Those curves obtained from the reference compounds provided a tool for peak area calculations of all detected metabolites according to their chemical class. Plotting the peak area of each standard concentration on the Y-axis against the known concentration on the X-axis was performed to construct the calibration curve of each standard. The limit of quantitation (LOQ) and the limit of detection (LOD) parameters were detected to ensure method validation. Furthermore, all parameters were assessed based on the FDA guidelines on bioanalytical method validation15. Standard preparation and calibration curve validation are discussed in Table 1. The regression equation is y = ax + b, where y is the peak area, x is the standard concentration in mg/ml, a is the slope, and b is the intercept.
Determination of cytotoxicity and anti-inflammatory activities of Salsola species
3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl-2 H-tetrazolium bromide (MTT) assay was carried out to establish the cytotoxicity of all Salsola extracts and their results were compared to those of standard piroxicam. The effective anti-inflammatory concentrations (EAICs) were estimated for each fraction extract in lipopolysaccharide (LPS)-stimulated human white blood cell culture. The expression levels of genes coding pro-inflammatory mediators IL-1β, IL6, TNF-α and TNF-γ were determined by real-time polymerase chain reaction (PCR). Results were expressed as means ± standard deviation (SD) of three individual replicates.
Isolation and cultivation techniques of human white blood cells WBCs
To prevent coagulation, the obtained human blood specimen was kept in a heparin tube. In a 15 ml centrifuge tube, a 1 ml blood specimen was treated with a fresh lysing solution to get rid of red blood cells. The lysing solution filled the tube to capacity. Then, the centrifuge tube was inverted for about 10 min at room temperature until the liquid became clear red. This was followed by centrifugation of lysed blood specimen at 4 °C, 2000 rounds per minute (rpm) for 10 min, decantation of the supernatant, and then drainage of the tube. In 10 ml cold phosphate buffered solution (PBS), the WBCs were suspended, then centrifugated. The isolated WBCs were re-suspended in Roswell Park Memorial Institute Medium (PRMI).
Viability and WBCs count were assessed by Dye Exclusion Method by mixing 50 µL of cell suspension with an equal volume of 0.5% trypan blue stain solution. After that, the mix was loaded onto a hemocytometer. Only the non-viable cells stained and counting process occurred in the four corner quadrants (A, B, C, and D).
N is the number of viable or non-viable cells, and D is the sample dilution (1:1 trypan blue solution).
Only mediums containing at least 90% viable cells were used for further investigations. The incubation conditions of the RPMI culture medium were 37 °C temperature, 5% carbon dioxide, and 90% relative humidity for six days in a carbon dioxide incubator. The cell titer was 100.000 cells/well in a 96-well plate.
Cytotoxicity assessment of the crude extracts compared to standard piroxicam
200 µL of cultured medium containing 100.000 WBCs/well were treated with different concentrations (0, 3.125, 6.25, 12.5, 25, and 50 µg/ml) of the crude extracts in an RPMI medium without bovine fetal serum or piroxicam (standard anti-inflammatory drug), then the plates were incubated at 37 °C temperature, 5% CO2, and 90% relative humidity for 72 h in a CO2 incubator.
After three days of incubation, 20 µL of MMT solution was added to each well followed by plate incubation to allow the progress of the MTT reaction. This was followed by centrifuging the incubated medium at 1650 rpm for 10 min and discarding the medium. MMT byproducts (formazan needles) were re-suspended in 100µL dimethyl formamide (DMSO). The absorbance of re-suspended crystals was measured at 570 nm using a spectrophotometer to detect a safe dose, which causes 100% viability.
The mathematical equation for calculation of % cell viability:
AT = mean absorbance of cells treated with different concentrations of each crude extract. Ac = mean absorbance of untreated cells with culture medium only. Ab = mean absorbance of cells treated with the vehicle of plant extract (PRMI without fetal bovine serum).
The cytotoxicity assay of the compound expressed as EC100 was calculated by the Graphed Instat software using % viable calculated from the serial dilutions of each plant extract.
Determination of the effective anti-inflammatory concentration (EAICs) of the crude extracts in lipopolysaccharides (LPS) stimulated human WBCs culture
In a 96-well plate, a volume of 50 µL of the culture medium containing 100,000 human WBCs was dispensed per well. Induction of inflammation occurred by adding 50 µL of LPS to the plated cells and incubation them in a CO2 incubator for 24 h. This was followed by centrifugation at 1650 rpm for 5 min, then discarding of the supernatant, and adding 200 µL of different concentrations (0,3.125, 6.25, 12.5, 25, and 50 µg/ml) of crude extracts and piroxicam.
The control cells (which were the negative control) containing the culture medium only and the tested plates were incubi ted for 72 h in the CO2 incubator.
The proliferation of the stimulated cells was determined using an MTT assay (please refer to the previous section dealing with the steps of the MTT assay).
Cell proliferation was assessed using the stimulation index (SI), which is the mean absorbance of LPS-stimulated WBCs treated with different concentrations of crude extracts/absorbance of control untreated cells.
EAIC of each crude extract is the ability to retrieve the abnormal proliferation of LPS-stimulated cells to the normal proliferation of un-stimulated cells. SI = 1 was calculated using an Instate graph pad.
Extraction of RNA of untreated and treated LPS stimulated human white blood cells
Extraction of RNA started with re-suspension of cell pellets in 50 µL solution R1, mixing for 30 s, then incubation at room temperature for 1 min. This was followed by adding 300 µL of solution R2 and incubation at 4 °C for 3–5 min. The obtained supernatant was loaded on a spin column and centrifuged for 30 s at 4 °C, 14.000 rpm. After that, 300 µL of working wash buffer was added to the spin column and the flow-through was discarded. Centrifugation was done for 30 s, and the described process was repeated twice.
Another centrifugation for 1 min at 10.000 rpm was done and transferring the centrifuge into a sterile 1.50 ml microcentrifuge tube was followed. To the center of the membrane, 30µL of elution buffer was added and incubated at room temperature for 1 min, then centrifugated for 30 s at 14.000 rpm, 4 °C.
The optical density of the extracted RNA was determined by mixing the absorbance and purity at A260, and A260/A280 nm respectively using a spectrophotometer and kept at −80 °C until real-time polymerase chain reaction (PCR).
cDNA synthesis from RNA extracted from untreated and treated LPS stimulated white blood cells
In a PCR tube, 2 µg of total RNA or nuclease-free water and 1 µg of oligo dT primer were added to the nuclease-free water to obtain a total of 12 µL. This was followed by gentle mixing, centrifugation, incubation at 65 °C for 5 min in a PCR machine, and placing back on ice immediately.
The gentle mixing of 4 µl of 5× reaction buffer, 1 µl of RNase inhibitor, 2 µl of the dNTPs mix, and 1 µl of reverse transcriptase or 1 µl of nuclease-free water instead of reverse-transcriptase for reverse transcriptase negative control with the previous mixture was performed. After that, spin down and incubation for 60 min at 42 °C followed by heat inactivation at 70 °C for 5 min in the PCR machine was carried out.
Determination of IL-1β, IL 6, TNF, and INF-γ expression levels by real-time polymerase chain reaction (PCR)
30 µl of 2× SYBR green master mix was mixed with 5 µl of cDNA, 0.5 µl of 10 pmoles/ml forward primer, and 0.5 µl of pmoles/ml reverse primer for each primer in PCR tubes. As for the reference tube, 0.5 µl of 10 pmoles/ml forward primer of β-actin and 0.5 µl of 10 pmoles/ml for reverse primer of β- actin were added. To assess for reagent contamination or primer dimers, another tube was used as a non-template control (NTC) by adding 1 µl of nuclease-free water instead of a template used. After that, the gentle mixing of the tubes with 6.5 µl nuclease-free water without creating bubbles was done and then spun for a few seconds. In the cycler, samples were placed, and the program was started as the following: initial denaturation (1 cycle of 95 °C for 10 min), followed by denaturation (40 cycles of 95 °C for 15 s), annealing (at 60 °C for 30s), and extension (at 72 °C for 30 s). the effect of LPS and extracts on gene expression was expressed as a Fold change in gene expression which was calculated according to the following equations:
Calculation
-
Expressions fold levels of gene calculated by
-
ΔCt normal = Ct normal untreated cells – Ct reference
-
ΔCt tested plant extract = Ct tested plant extract−treated cells – Ct reference
-
ΔCt induced = − Ct LPS−exposed cells – Ct reference
-
-
In case of genes:
-
ΔΔCTtested plant extract = ΔCt tested plant extract – ΔCt normal
-
ΔΔCTinduced = ΔCt induced – ΔCt normal
-
-
In case of GAPDH:
-
ΔΔCTtested plant extract = ΔCt normal – ΔCt tested plant extract
-
-
ΔΔCTinduced = ΔCt normal – ΔCt induced
Where: Ct tested plant extract: threshold cycle value of genes of extracted mRNA of plant extract treated-LPS-stimulated WBCs which is defined as the cycle number at which the fluorescence generated within a reaction crosses the fluorescence threshold. Ct reference: threshold cycle value of GAPDH which is used for normalization. Ct normal: threshold cycle value of genes of extracted mRNA of untreated control WBCs. Ct induced: threshold cycle value of gene of extracted mRNA of LPS-stimulated WBCs.
The primers used:
TNF-α | F-CTCTTC TGCCTGCTGCACTTTG |
R- ATGGG CTACAGGCTTGTCACTC | |
IL-6 | F, 5′-TGAA CTCCTTCTCCACAAGCG-3′ |
R, 5′-TCTG AAGAGGTGAGTGGCTGTC-3′ | |
IL-1β | F, CCACA GACCTTCCAGGAGAATG |
R, GTGCA GTTCAGTGATCGTACAGG | |
INF-γ | F, GAGTG TGGAGACCATCAAGGAAG |
R, TGCTT TGCGTTGGACATTCAAGTC | |
R, GGAAGATGGTGATGGGATT | |
GAPDH | F, GGATTTGGTCGTATTGGG |
R, GGAAGATGGTGATGGGATT |
Statistical analysis
One-way analysis of variance (ANOVA) is the statistical tool that was used for semi-quantitative analysis and anti-inflammatory activity investigation. The peak intensities of the identified metabolites were used to calculate the relative concentration of each metabolite as mg equivalent (Eq.)/g dry weight dry extract and the data of the retrieving power of each extract to bring the LPS-stimulated WBCs to the normal state of each extract was used to calculate the effective anti-inflammatory concentration of each one. Multivariate analysis of the metabolomics data was performed using SIMCA 14 software (Umetrics, Malmo, Sweden). The heat map was constructed using Metaboanalyst 5.0 (http://www.metaboanalyst.cal/), an online free tool for data analysis.
The relative concentrations obtained from semi-quantitative analysis were used to construct the heat map.
Results and discussion
Metabolite annotation of different Salsola extracts
Six Salsola samples, namely, shoot and root 70% ethanolic extracts of S. tetrandra, S. vermiculata and S. tetragona, were subjected to UHPLC-MS/MS analysis to identify their principle active constituents. Eighty compounds were tentatively identified in all extracts of the aforementioned three Salsola species through investigating both positive and negative ESI-MS modes, where they included phenolic compounds, flavonoids, triterpenes, saponins, coumarins, fatty acids and nitrogenous compounds in the negative and positive ion modes (Table 2, Fig. S1). The compounds were assigned in accordance with their retention times, recorded MS/MS fragment ions and fragmentation patterns as compared to an in-house database with about 300 metabolites of varying chemical classes from the entire Salsola genus, as well as Mass bank database reference and the Dictionary of Natural Products. The in-house database composed only metabolites identified in Salsola species, ignoring any other species. The total number of compounds in the library was 312 compounds of all identified chemical classes in genus Salsola. During constructing the in-house library, we selected only compounds that were identified in different Salsola species using LC-MS technique, where ESI conditions were applied, the reversed-phase column C18 was used and the same composition of the mobile phase or even the same polarity and the same gradient elution mode The elution order of the metabolites was also a main factor in the compound elucidation.
The identified compounds are numbered according to the experimental elution order. Fragmentation patterns of the principle components are described in the supplementary materials form of schematic diagrams. The peak intensity of each identified metabolite in each Salsola extract was mentioned in Table S1, calculated as (expressed as mg Equivalents/g dry extract). Furthermore, of the eighty identified metabolites, sixteen and twenty-nine compounds were identified in S. tetrandra shoots and roots, respectively. Also, twenty-seven and twenty- four phytochemicals of the total eighty metabolites were investigated in the shoots and roots of S. vermiculata, accordingly, whereas seventeen and thirty-nine secondary metabolites were known in the shoots and roots of S. tetragona.
Flavonoids
Flavonoids, being a commonly found secondary metabolite class in the plant kingdom, have the highest occurrence in our tested Salsola samples. Fourteen flavonoids were identified in our samples. They are a product of condensation between hydroxycinnamic acids and malonyl residues, comprising the structure of the C6-C3-C6 scaffold. Based on the oxidation state of ring C, they are classified, and the retro-Diels Alder (RDA) reaction is the main fragmentation pathway18. Compound 24 exhibited an intense peak at m/z 609 [M−H]− and daughter ions 447 [M−H-162]−, 301 [M−H-162-146]−, 151 due to C-C bond cleavage at ring C19. Therefore, it was elucidated as hesperidin. Similarly, compound 28 was tentatively annotated as diosmin. It displayed a base peak at m/z 609 [M+H]+, in addition to 447 [M+H-162]+, 301 [M+H-162-146]+ for diosmetin and 191 for 7- etoxycoumarin20. Compounds 10 and 17 were annotated as dirhamnosyl quercetin and glucosyl rhamnosyl quercetin, respectively. They showed deprotonated quasi-molecular ions of m/z 593 and 609, respectively. In addition to the daughter ions [M−H-146]−, [M−H- 146–146]− corresponding to the loss of one and two rhamnose units, respectively, [M−H-162]− corresponding to the loss of a glucose unit and [M−H-162-146]− relevant to the loss of glucose and rhamnose units, quercetin aglycone ions were spotted in the spectra of the aforementioned two compounds, 10 and 1721. Compound 38 was elucidated as catechin due to the presence of daughter ions at 291 [M+H]+, 273 [M+H-H2O]+ and 139 [M+H-152]+ due to 1,2 cleavage of ring C22. Compounds 45 and 77 were elucidated as kaempferol and kaempferol methyl ether. They exhibited significant peaks of deprotonated ions [M−H]− at m/z 285 and 299, respectively, alongside the RDA ions 257,151 and [M−H-CH3]−23. Compound 76 was tentatively annotated as isorhamnetin as it displayed a base peak at m/z 315 [M−H]− and [M−H-CH3]− to produce rhamnetin ion23. Compound 50 was elucidated as naringenin, which displayed a base peak at m/z 273 [M+H]+ and RDA ion 15324. Similarly, Compound 72 was identified as apigenin, as it exhibited a significant ion at m/z 269 [M−H]−, then it was subjected to 1,2 cleavage of ring C to produce RDA ion at m/z 11725. Compound 68 was annotated as cyanidin as it exhibited daughter ion peaks [M-CO]+, [M−H2O-CO]+, [M-CO-CO]+, [M−H2O-CO-CO]+ and an RDA fragments at m/z 259, 241, 231, 213, 150, and 137, respectively26. Furthermore, the mass spectrum of compound 79 was noticed to ensure the presence of daughter ions m/z 176 and 144, corresponding to the formation of RDA ions of chalcones, respectively. Hence, it was known as pinocembrin chalcone13,27.
Compound 64 displayed daughter peaks at m/z 316, 301, 286, 121, and 120, corresponding to [M−H]−, [M−H-CH3]−, [M−H-CH3-CH3]−, [M−H-C10H11O4]− and [M−H-C10H12O4]−, respectively. In the same manner, as described by28. Compound 42, which was identified as tri methoxy-dimethoxy isoflavone, exhibited characteristic peaks at m/z 329, 314, 299, 165, and 164 relevant to fragment ions [M−H]−, [M−H-CH3]−, [M−H-CH3-CH3]− and RDA ions, respectively29.
Coumarins
Coumarins are a group of polyphenolic compounds characterized by the presence of a benzo-γ-pyrone ring (benzo chromone)30. Seven coumarins were identified in the three Salsola species. Coumarinolignanoids undergo fragmentation31. Hence, compound 6 was assigned as cleomiscon B due to the presence of daughter ions at m/z 367, 207, 179, 136, and 12313. Compound 33 was annotated as daphnoretin as it displayed a base peak at m/z 353 corresponding to [M+H]+ and daughter peaks of m/z 336 and 191 due to loss of the CH3 group and de-dimerization, respectively32.
Compound 37 was annotated as furanocoumarin; known as bergaptol, as it formed a protonated ion peak at m/z 203 and then the elimination of small molecules such as CO and 2 CO was observed13,33. Similarly, Compound 41, which was elucidated as fraxetin, exhibited CH3 and CO molecule loss, meanwhile, Compound 47, which was known as isofraxidin, exhibited only CH3 group loss34,35. On the other side, compound 49 base peak at m/z 353 exhibited a loss of 162 Da, corresponding to a hexose unit loss, followed by demethylation and loss of CO molecule; hence, it was annotated as scopoletin glycoside36. Compound 75 exhibited an intense peak at m/z 163. This peak was fragmented by eliminating CO2 and CO and C2O2 (−56 Da) molecules; therefore, it was annotated as umbelliferone37.
Triterpenoids
Seven triterpenoids were annotated in the sample extracts. Compound 58 displayed a peak of m/z 425 corresponding to [M−H]− and other peaks of 407, 389, and 363 corresponding to [M−H-H2O]−, [M−H-2H2O]− and [M−H-CH2CHOH]−, respectively. So, it was tentatively annotated as lupeol38.
Compounds 27, 48, 70, and 52 were elucidated as glucopyranosyl oleanolic acid, olean-en-diol, oleanolic acid and salolic acid, respectively, as they displayed intense peaks at m/z 619, 443, 457, and 489, respectively corresponding to the protonated ions of each one. MS/MS fragmentation revealed daughter ions due to the loss of one glucose unit (−162 Da), followed by the loss of a water molecule and formic acid (−46 Da)39,40. Compound 36 was tentatively identified as salsolin A. It exhibited a base peak of m/z 504 corresponding to [M+H]+, followed by the loss of one carboxyl group (−46 Da) with the characteristic retro-Diels-Alder fragments at m/z 224 and 280, corresponding to the two hydroxyl groups in ring A and B and two hydroxyl groups and a carboxyl group in ring C and D or E41. Compound 65 exhibited an intense peak at m/z 503, which conforms to a molecular formula of C30H47O2. Therefore, it was tentatively identified as mediagenic acid12.
Phenolics and phenolic acids
Phenolic acids represent a major class of secondary metabolites. They are distributed in our Salsola samples as seven phenolic compounds and five phenolic acids. Compound 40 exhibited an intense peak at m/z 359; its MS/MS fragmentation revealed the formation of 3-(2,3-dihydroxy-phenyl) lactic acid (m/z 197), caffeic acid (m/z179) and their dehydrated form (m/z 161). Therefore, it was tentatively identified as rosmarinic acid42. Compounds 25, 46, 54, and 57 were tentatively identified as tachioside, dihydroxy phenyl-ethyl-β-D-glucopyranoside, cunataside C and canthoside D as they exhibited precursor ions at m/z 301,315, 551,433, respectively, followed by loss of glucose unit [M−H-162]−13,43,44. Compound 34 showed an intense peak at m/z 164 [M+H]+ and a daughter peak at m/z 146; hence, it was elucidated as p-coumaramide45. Compound 59 was identified as gingerol as it displayed a precursor ion peak at m/z 293 [M−H]− and daughter ions at m/z 193 and 139 due to the cleavage of C4-C5 bond46.
Each phenolic acid molecule is characterized by the presence of hydroxylated aromatic rings linked to a carboxyl group. They occur in the form of C6-C1 and C6-C3. Compound 8 displayed a deprotonated ion (m/z 179) that was fragmented to form the thermodynamically most favorable pathway of CO2 loss (−44 Da) and a consecutive loss of CO molecules (−28 Da). This peak ion, in another pathway, showed H2O molecule loss (m/z 161) through triple bond formation (C2 or C6) by remote hydrogen re-arrangement, so it was tentatively annotated as caffeic acid47. Similarly, compound 7 was identified as orsellic acid, as its deprotonated ion exhibited CO2 loss, followed by an H2O molecule in one pathway and an H2O molecule in the other one48. In addition to that, compound 4, which was annotated as hypogallic acid, showed an intense deprotonated peak at m/z 153, followed by a loss of CO2 molecules49 and compound 13 showed a deprotonated ion at m/z 183, followed by the loss of CH3, CH3 and CO and CH3 and CO2 molecules, so it was known as O-methyl gallic acid50. Compound 9 was annotated as anisic acid as its protonated ion exhibited a loss of CO2 molecules besides the formation of protonated carbon dioxide (HO + C ≡ O, m/z 45)51.
Nitrogenous compounds
Eleven nitrogenous compounds were identified in Salsola extracts. Compounds 16, 19, 21 and 22 were characterized as tyramine derivatives, as their MS/MS fragmentation displayed fragments corresponding to tyramine moiety (m/z 136) and loss of caffeoyl moiety [M−H-166]− for N-caffeoyl tyramine and feruloyl moiety [M−H-177]− for N-feruloyl tyramine and loss of feruloyl and methoxyl groups [M−H-177]− and [tyramine moiety-31]− for N-feruloyl-3′′′-methoxy tyramine and loss of hydroxy feruloyl moiety [M−H-193]−for hydroxy moupinamide2. Compound 14 was characterized as feruloyl octopamine as it exhibited a protonated base peak and daughter ions relevant to the loss of the feruloyl moiety and the octopamine moiety, which in turn showed the loss of a water molecule to produce the tyramine moiety. Similarly, compound 12, which was identified as N-(4’-methoxcinnamoyl)-norepinephrine, showed a precursor ion at m/z 328 [M−H]− and daughter ions at m/z 153 due to the loss of methoxy cinnamoyl and m/z 107 corresponding to norepinephrine13. Compounds 5 and 29 were elucidated as pericamylinone A and salasmine, respectively. Their MS/MS fragmentation spectra exhibited the characteristic RDA fragments at m/z 192, 178, and 14913. Compound 39 exhibited a loss of 132 Da, corresponding to a ribose sugar unit. Therefore, it was elucidated as uridine, as described by52. Compound 3 protonated precursor ion exhibited a characteristic acetyl group loss to produce a daughter ion fragment at m/z 58. Therefore, it was elucidated as betaine53. Compound 18 was tentatively annotated as hernandine because of the characteristic fragmentation pattern of protonated aporphine alkaloids [M+H]+, [M+H-CH3]+, [M+H-CH3-CH3OH] and [M+H-CH3OH-CO]54.
Fatty acids
Salsola species are rich in fatty acids and eleven fatty acids were identified in our samples. Compound 20 was elucidated as a dicarboxylic acid metabolite (Azelaic acid), as it exhibited a deprotonated ion fragment at m/z 187, with a sequential loss of H2O, CO2 and H2O + CO2 molecules55. Compound 62, which was known as hydroxy octadecanoic acid, showed an intense peak at m/z 295 corresponding to [M−H]−, accompanied by a loss of water molecule (−18 Da), which was an indicator of the presence of one hydroxyl group, yielding a daughter peak at m/z 277 corresponding to octadecadienoic acid. In the same way, compound 43 MS/MS fragmentation spectrum revealed three consecutive water molecules loss. Hence, it was elucidated as trihydroxydecosan-ienoic acid2. Compound 69 displayed a precursor ion peak at m/z 353 [M+H]+. Further fragmentation gave rise to daughter peaks at m/z 261 as a result of a glycerol moiety loss and 243 due to a water molecule loss. Therefore, this compound was annotated as 9,12,15-octadecatrienoic acid, 2,3-hydroxypropyl ester (1-monolinolenin)56. The base peak is the most sensitive peak for free fatty acid detection. Hence, compounds 71, 73, 74, 78, 80, 81 and 82 were annotated as ecosenoic acid, margaric, linoleic, behenic, lignoceric, hexacosanoic and octacosanoic acids, respectively57,58.
Saponins
Four saponins were annotated in Salsola extracts. Compounds 30, 31 and 35 were tentatively elucidated as salsoloside D, salsoloside E and salsoloside C, respectively. They exhibited base peaks at m/z 941, 1087 and 925, respectively. MS/MS fragmentation spectrum of compound 30 revealed a hedragenin aglycone ion moiety (m/z 471) alongside other ions of m/z [M−H-162]−, corresponding to the loss of a hexose unit, [M−H-162-132]−, corresponding to the loss of a pentose unit and [M−H-162-132-176]−, corresponding to the loss of one glucuronic acid molecule6,59. Compounds 31 and 35 MS/MS fragmentation revealed an oleanolic acid ion (m/z 455), [M−H-162]−, [M−H-162-162]−, [M−H-162-162-132]− and [M−H-162-162-132-176]− as compound 306. Compound 32 showed a base peak at m/z 681 [M−H]− and daughter ions at m/z 519 [M−H-162]− and 471[M−H-162-HCHO-H2O]−; therefore, it was elucidated as pentahydroxy-oleanen-oic acid glucopyranoside13.
Sugars
Compound 1 showed an intense peak at m/z 365, which corresponds to the formation of the [M+Na]+ adduct of trehalose. Further fragmentation of these adducts yielded another peak at m/z 203, which corresponds to the formation of the [M+Na]+ adduct of glucose60. On the other hand, Compound 2 displayed a precursor ion peak at 193 corresponding to [M−H]− and daughter ions corresponding to a non-reducing end and a reducing end, respectively. So, it was tentatively annotated as galacturonic acid61.
Cardenolides
Caredenolides are a group of cardiac active metabolites. They are characterized by the presence of a steroidal nucleus, a five-membered lactone ring and an oligosaccharide moiety attached to position 362. Cardenolides displayed very similar MS/MS spectra. Compounds 26 and 67 were elucidated as salsoteragonin and calactin, respectively. That was observed through the loss of sugar units, H2O and CO molecules13,63,64.
Miscellaneous compounds
Compound 56 was elucidated as hydroxy pseudoguaien olide didehydro ketone as it showed an intense peak at m/z 263 corresponding to [M+H]+ and daughter peaks [M+H-H2O]+, [M+H-H2O-CH3]+ and [M+H-H2O-CH3-CO]+65. Compound 61 was annotated as sitostanol. Phytosterol displayed similar mass spectra, including the loss of water molecules and the cleavage of C17 and C20. Therefore, the obtained ions 417 [M+H]+ and 399 [M+H-H2O]+ confirmed the presence of sitostanol66,67. Compounds 53, 55 and 60 were tentatively annotated as blumenyl glucopyranoside, staphylinoside D and oxo-ionol glucopyranoside, respectively, with the reference to literature, as they exhibited precursor ion peaks at m/z 387, 385 and 371, respectively68. Compound 63 was tentatively annotated as phytol, displaying a prominent peak at m/z 318 [M+Na]+. It was.
confirmed from data on Mass Bank and NIST database alongside the literature21.
Comprehensive metabolomic study of the shoot and root samples of the three Salsola species
A metabolomics-based approach linked to multivariate analysis was implemented to explore the clustering pattern of the three Salsola species. Unsupervised recognition analysis was performed by constructing the principal component analysis (PCA) score scatter plot model. PC1 and PC2 represented 69.75% of the total variation. Model validation was ensured as R2 (goodness of fit) and Q2 (goodness of prediction) were valued at 1 and 0.999, respectively, proving the model’s reliability for fitting and predicting future samples. As depicted in Fig. 1A, Salsola samples were segregated into three clusters, revealing the variation in their chemical matrices. PC2 separates S. tetragona from S. tetrandra and S. vermiculata. S. tetragona is the only species showing separation between its organs in the PCA, although the other two species were clustered together in the form of a single holistic cluster combining both shoot and root extracts of both species. The samples of S. tetragona roots were clustered along the positive side of PC1 and the negative side of PC2, meanwhile, the shoots of the same species were clustered along the positive sides of both PC1 and PC2. S. tetrandra and S. vermiculata shoot and root samples were clustered along the negative sides of both PC1 and PC2. Principal components 1 and 2 were used to classify the extracts according to the quartile where they are clustered, hence the clustering quartiles were referred to as the positive and negative side in relation to each PC. The positive side of both was the upper right quartile and the negative side of both was the lower left quartile.
Semi-quantitative data on the identified metabolites (Table S2) was used to construct a hierarchical cluster analysis heat map (HCA). As depicted in Fig. 1B, the highest amount of nitrogenous compounds was detected in the root sample of S. tetrandra and the shoot sample of S. tetragona, meanwhile, phenolics and phenolic acids were mostly detected in the shoot sample of S. tetragona and the samples of both organs of S. tetrandra. The shoot sample of S. tetrandra overwhelmed the vast quantity of flavonoids. It is clear that the proportion of most classes was markedly different between the shoot and root samples of S. tetragona, particularly fatty acids, nitrogenous compounds and phenolics. The root sample of S. tetragona possessed double the amounts of fatty acids compared to the shoots. On the other hand, S. tetragona shoots contained roughly double the amounts of phenolics and nitrogenous compounds in comparison to the same species’ root. It is worth mentioning that megastigmane glycosides were exclusively detected in the shoot and root samples of S. tetragona. Back to the literature, no previous studies could present a comprehensive profiling of that species, giving us the credit of providing the first attempt for investigating S. tetragona. S. tetrandra and S. vermiculata shared a common chemical profile, recording comparable amounts of triterpenoids and phenolics. A previous study utilized the LC-MS/MS technique to unravel the chemical complexity of S. tetrandra and S. vermiculata and recommended further investigation of their chemical profile2. The study in hand, was able to deeply compare between the two species and ensured the chemical similarities between them. Based on the aforementioned discussion, the marked variation in the chemical profile between the species of S. tetragona and the noticeable similarities among S. tetrandra and S. vermiculata samples were the reasons behind the clustering pattern. The heat map (Fig. 2) represented the relative concentration of the identified metabolites in each extract. Each standard of certain chemical class was used to calculate the relative concentration of the compounds of the same chemical class by dividing the peak area of the compound over the peak area of the standard, then utilizing the slope and intercept of each standard, the concentration of each metabolite of the same chemical class was calculated as mg/equivalent, then the value was divided by the amount of the total extract, which is 0.02 g (the amount used in the UHPLC study from each extract), and the value would be mg equivalent (Eq.)/g dry weight dry extract.
As depicted in Fig. 2, the species effect had the upper hand in the clustering pattern of the investigated Salsola species, holding a great variation in the chemical profile of S. tetragona and some similarities in the profiles of S. tetrandra and S. tetragona. The organ effect was observed in the sub-clustering pattern of each species.
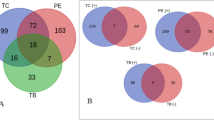
Eight compounds of the eighty identified metabolites were common in both organs of the three tested species; two nitrogenous compounds, betaine and feruloyl octopamine, had the highest occurrence in S. tetrandra roots and S. tetragona shoots, respectively. The saponin salsolside D, dicarboxylic azelaic acid and the phenolic compound gingerol possessed the highest prevalence in S. tetrandra roots. The acyclic diterpene phytol was mostly detected in S. vermiculata roots. The phenolic compound dihydroxyphenyl-ethyl glucopyranoside was highly concentrated in S. tetrandra shoots. Moreover, D-galacturonic acid was common in all samples but was accumulated in comparable concentrations in both S. tetragona and S. tetrandra shoots.
Also, studying Fig. 2 revealed that N-caffeoyl tyramine, hydroxy octadecadienoic acid and dirhamnosyl quercetin were the major compounds in S. tetrandra shoot samples. On the other hand, salsoloside E, glucopyranosyl oleanolic acid and hypogallic acid were the main metabolites in S. tetrandra root samples. Whereas salsoloside E, hypogallic acid and O-methyl gallic acid were the main chemical components in S. vermiculata shoot samples and hydroxy octadecadienoic acid, catechin and salsoloside E were the major constituents in S. vermiculata root samples. On the contrary, hexacosanoic acid, canthoside D, N-caffeoyl tyramine and behenic acid were the major secondary metabolites in S. tetragona shoot samples, meanwhile, margaric acid, eicosenoic acid, taxiphyllin and hydroxy octadecadienoic were the major chemical compounds in S. tetragona root sample.
Investigating the chemical matrices of each species’ extract, orthogonal projection to latent structure discriminatory analysis (OPLS DA) scores scatter plot models of total extracts to assign any in-between or within-class discrimination. The species was used as the main class identifier for constructing a validated discriminatory model. The validation of the model was assured with R2X, R2Y and Q2 values of 1,1,0.999. As observed in Fig. 3, the shoot and root extracts of S. tetrandra were clustered together and both organs of S. vermiculata were clustered together. Both species exhibited two separated clusters on the positive side of LV1 and the negative side of LV2, controversy, the roots extract of S. tetragona was completely separated from the shoots, as the shoot extract of S.tetragona was clustered along on the positive side of LV2 and the negative side of LV1 and the root extract of the same species was clustered along the negative sides of both LV1 and LV2. To sum it up, OPLS DA confirmed that the species effect has been profoundly able to discriminate between the three species, recording a great variation in the chemistry between the shoots and roots of S. tetragona and some similarities between S. tetrandra and S. vermiculata.
The coefficient plots find a proportional correlation of the amount of any compound in the extract; therefore, as the length of the bar corresponding to a certain metabolite was the highest, that meant the high abundance of that compound in the extract.
As depicted in Fig. S2, OPLS DA coefficient plots revealed that salsoloside D, salsoloside E, salsolic acid and gingerol were positively correlated for the discrimination of S. vermiculata shoots; meanwile, betaine, catechin, salisoflvan, margaric acid, linoleic acid and N-feruloyl-3′′′-methoxy tyramine were positively attributed to the species roots. Moreover, D-galacturonic acid, cleomiscosin B, N-caffeoyl tyramine, feruloyl octopamine and monolinolenin were discriminatory metabolites of S. tetragona shoots, while trehalose, azelaic acid, dihydroxy phenyl-ethyl glucopyranoside, eicosenoic acid, cuneataside D, salsoloside D, Oxo-ionol glucopyranoside, hydroxy octadecadienoic acid and margaric acid were strongly correlated to that species roots. In addition to that, taxiphyllin, hydroxy octadecadienoic acid, salsoloside D, salsoloside E and pinocembrin chalcone were positively correlated to S. tetrandra shoots; betaine, hypogallic acid, anisic acid, pericampylinone A, hernandine, tachioside, glucopyranosyl oleanolic acid, pentahydroxy oleanen-oic acid glucopyranoside, salsoloside C and azelaic acid were strongly correlated to S. tetrandra roots.
Anti-inflammatory activity of Salsola extracts
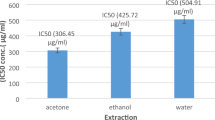
Determining a safe dose of each extract is a crucial step in the biological scanning process to exclude any toxicological interference. Hence, the MTT assay was carried out to assign the dose producing 100% viability of the tested WBCs (IC50), where high values determine high safety. It was observed in Fig. 4A, that IC50s of all extracts were relatively higher than the standard anti-inflammatory drug piroxicam, which indicates a high safety margin. The extract of Salsola tetragona roots showed an 8-fold higher IC50 than the piroxicam. Moreover, all Salosla extracts exhibited higher safety margins than the standard chemical drug. The results were in agreement with previous publications on the safety profile of Salsola69,70. To date, the study in hand has provided safety evidence for the three investigated species for the first time. 1/10 of the IC50 dose of each fraction was used to evaluate its anti-inflammatory activity by calculating the EAIC. EAIC is defined as the extract concentration required to bring the LPS-stimulated WBCs to the normal proliferation condition, considering that potent activity is reflected by low EAIC values. As depicted in Fig. 4B, the shoots of the three species demonstrated higher activities than the roots and piroxicam, recording 1.02, 2.67 and 14.6 μg/ml for S. tetrandra, S. vermiculata and S. tetragona, respectively, while the standard recorded 15.43 μg/ml and the root extracts recorded results over 100 μg/ml, revealing the potent anti-inflammatory activity of S. tetrandra, S. vermiculata and S. tetragona shoot extracts as they were able to bring the LPS-stimulated WBCs to normal proliferation in very low concentrations compared to that recored by the standard drug, piroxicam and the weak activity of the roots, which were used in concentrations more than six folds than the reference to normalize the proliferation of stimulated cells. Tracking the chemical profiles of these active extracts revealed their overwhelming richness in polyhydroxylated metabolites, especially flavonoids and saponin glycosides. Four pro-inflammatory biomarkers TNF-α, IL1β, IL6 and INF-γ, were measured in the presence of each extract and piroxicam through a mechanistic study searching for the best anti-inflammatory drug candidate. This procedure included the utilization of LPS to stimulate the cultivated WBCs and then adding the extract and the reference to screen the activity. Biochemically, LPS is a part of the cell wall of gram-negative bacteria, comprising three parts lipid: A, core protein and O-antigen. Lipid A stimulated WBCs and the aforementioned four pro-inflammatory mediators were released15. Regarding Fig. 5: the fold gene expression level of the four pro-inflammatory cytokines was significantly downregulated in the presence of the shoot extracts of the three species compared to roots and piroxicam, spotting out the potential anti-inflammatory activity of the shoot extracts and their ability to decrease the expression level of the four pro-inflammatory biomarkers. Among all the extracts, the shoot extract of S. tetrandra was the most biologically active extract as the fold expression of TNF-α, IL 1β, INF-γ and IL 6 was 0.01256, 0.04, 0.03 and 0.002728 fold, respectively. Furthermore, in the presence of all other extracts the expression level was higher and even in the presence of piroxicam, which recorded 1.53, 2.5, 1.3 and 1.8 fold for TNF-α, IL 1β, INF-γ and IL 6. Therefore, this ex-vivo study revealed the potential anti-inflammatory activity of S. tetrandra and its potent activity that exceeded the activity of piroxicam.
Predicting the anti-inflammatory activity of Salsola extracts
An OPLS biplot was constructed to correlate the tested Salsola samples to the investigated pro-inflammatory cytokines. Figure 6 and S3 reveal the model validation for prediction. R2X, R2Y and Q2 were 0.958, 0.988 and 0.986, respectively. Furthermore, four permutation plots were constructed for each investigated cytokine, where R2 and Q2 of the permuted data were less than 0.6 and 0, respectively, in each model, excluding data overfitting and confirming the validity. Studying Fig. 6 revealed that the shoot extract of Salsola tetrandra was the most biologically active extract due to its close proximity to the investigated pro-inflammatory mediators. Also, 4-methoxy cinnamoyl norepinephrine was positively correlated to the inhibition of the gene expression levels of TNF-α, IL6 and INF-γ. Moreover, salsolosides D and E were positively correlated to TNF-α, IL1β and IL 6 gene expression downregulation. Biphenyl salsinol was positively correlated with inhibiting TNF-α and INF-γ gene expression levels, meanwhile, calactin and medicagenic acid were positively correlated to downregulate IL6 expression levels. In addition, 4-O-methyl gallic acid, hypogallic acid and hernandine were exclusively correlated to inhibit INF-γ gene expression level. The anti-inflammatory activities of these biomarkers were reported in the literature. These anti-inflammatory markers have previously been reported to exert an in-vitro/in-vivo anti-inflammatory effect. Regarding the nitrogenous compounds, N-4-methoxy cinnamoyl norepinephrine was proven to inhibit the release of TNFα, IL1β, INFγ and IL613 and that chemical compound’s antioxidant activity was clearly reported by 71. Hernandine anti-inflammatory effect was reported by12. Moreover, medicagenic acid was reported to have a significant anti-inflammatory effect in S. cyclophylla holistic extract12. In addition to that, the anti-inflammatory activity of triterpene glycosides was reported to be comparable to that produced by glucocorticoids72. The anti-inflammatory activity of the cardenolides calactin and the phenolic compound biphenyl salsinol was statistically reported for the first time and further biological investigations regarding their possible mechanism of action are required.
Conclusion
In our study, three Salsola species, namely, S. tetrandra, S. tetragona and S. vermiculata shoots and roots were comprehensively chemically profiled for the first time using an ultra-high performance liquid chromatography-mass-mass spectrometry (UHPLC-MS/MS) method. Eighty metabolites were tentatively identified in our investigated extracts, where flavonoids, saponins, nitrogenous compounds, phenolics and phenolic acids were detected in all Salsola samples, except for megasigmane glycosides, which were detected only in S. tetragona samples. HCA revealed that the species effect was profoundly observed in the clustering patterns of Salsola extracts, while the organ effect was noticed in the sub-clustering pattern of each species. OPLS DA models revealed in-between and within-class discrimination between the two organs of each species. The ex-vivo anti-inflammatory activity of the tested samples was screened for the first time, demonstrating the mechanism of action of each extract on the gene expression level. It is worth mentioning that Salsola extracts represented candidates for a remarkably safer anti-inflammatory drug than the chemical standard piroxicam, whereas all extracts. The shoots of the three species exerted more potent anti-inflammatory activity than the roots. Using a chemometric-based statistical analysis, it was found that the shoot extract of S. tetrandra was the most biologically active extract.
Data availability
The datasets used to support this study are available from the corresponding author upon request and after satisfying ethical requirements for their release.
Abbreviations
- STA:
-
Salsola tetrandra shoots
- STR:
-
Salsola tetrandra roots
- SVA:
-
Salsola vermiculata shoots
- SVR:
-
Salsola vermiculata roots
- SGA:
-
Salsola tetragona shoots
- SGR:
-
Salsola tetragona roots
References
Vineeth, T. V., Kumar, S., Shukla, M., Chinchmalatpure, A. & Sharma, P. C. Ecological and economic potential of major halophytes and salt tolerant vegetation in India. in Abiotic Stress in Plants (eds. Fahad, S., Saud, S., Chen, Y., Wu, C. & Wang, D.) Ch. 8 (IntechOpen, 2020). https://doi.org/10.5772/intechopen.93841
Rasheed, D. M., Zalabani, E., Koheil, S. M., El-Hefnawy, M. A., Farag, M. A. & H. M. & Metabolite profiling driven analysis of Salsola species and their anti-acetylcholinesterase potential. Nat. Prod. Res. 27, 2320–2327 (2013).
Murshid, S. et al. Genus Salsola: chemistry, biological activities and future prospective—a review. Plants 11, 714 (2022).
Turki, Z. Chemotaxonomical studies of the genus Salsola (Chenopodiaceae) in Egypt. Feddes Repert. 110, 81–87 (2008).
Boulaaba, M. et al. Biological activities and phytochemical analysis of phenolic extracts from Salsola kali L. Role of endogenous factors in the selection of the best plant extracts. South Afr. J. Bot. 123, 193–199 (2019).
Iannuzzi, A. M. et al. Chemical profile and nutraceutical features of Salsola soda (agretti): anti-inflammatory and antidiabetic potential of its flavonoids. Food Biosci. 37, 100713 (2020).
Jin, Y. S. et al. Chemical and biologically active constituents of Salsola collina. Chem. Nat. Compd. 47, 257–260 (2011).
Tundis, R., Loizzo, M., Statti, G. & Menichini, F. Inhibitory effects on the digestive enzyme α-amylase of three Salsola species (Chenopodiaceae) in vitro. Pharmazie 62, 473–475 (2007).
Seo, J. H., Jin, M. H. & Chang, Y. H. Anti-inflammatory effect of Salsola Komarovii extract with dissociated glucocorticoid activity. BMC Complement. Med. Ther. 20, 176 (2020).
Gonfa, Y. H. et al. Anti-inflammatory activity of phytochemicals from medicinal plants and their nanoparticles: a review. Curr. Res. Biotechnol. 6, 100152 (2023).
Osman, S. M., el Kashak, W. A., Wink, M. & el Raey, M. A. New isorhamnetin derivatives from Salsola Imbricata Forssk. Leaves with distinct anti-inflammatory activity. Pharmacogn. Mag. 12, S47–S51 (2016).
Mohammed, H. et al. Phytochemical analysis, pharmacological and safety evaluations of Halophytic Plant, Salsola Cyclophylla. Molecules 26, 2384 (2021).
Selim, D. A., Shawky, E., Ghareeb, D. A., Abdulmalek, S. A. & Abu El-Khair, R. M. Comparative metabolomics of the different fractions of two saltwort (Salsola L.) species in relation to their anti-inflammatory activity. Food Biosci. 51, 102306 (2023).
Shehab, N. & Abu-Gharbieh, E. Phenolic profiling and evaluation of contraceptive effect of the ethanolic extract of Salsola imbricata Forssk. in male albino rats. Evid.-based Complement. Altern. Med. (2014).
Darwish, R. et al. Comparative metabolomics reveals the cytotoxic and anti-inflammatory discriminatory chemical markers of raw and roasted colocynth fruit (Citrullus colocynthis L). RSC Adv. 11, 37049–37062 (2021).
Shawky, E., Sobhy, A., Ghareeb, D., Shams, S. & Selim, D. Comparative metabolomics analysis of bioactive constituents of the leaves of different Trigonella species: correlation study to α-amylase and α-glycosidase inhibitory effects. Ind. Crops Prod. 182, 114947 (2022).
Selim, D. A., Shawky, E. & Abu El-Khair, R. M. Identification of the discriminatory chemical markers of different grades of Sri Lankan white, green and black tea (Camellia sinenesis L.) via metabolomics combined to chemometrics. J. Food Compos. Anal. 109, 104473 (2022).
Cuyckens, F. & Claeys, M. Mass spectrometry in the structural analysis of flavonoids. J. Mass Spectrom. 39, 1–15 (2004).
Xu, F., Liu, Y., Zhang, Z., Yang, C. & Tian, Y. Quasi-MS identification of flavanone 7-glycoside isomers in Da Chengqi Tang by high performance liquid chromatography-tandem mass spectrometry. Chin. Med. 4, 15 (2009).
Campanero, M. et al. Simultaneous determination of diosmin and diosmetin in human plasma by ion trap liquid chromatography-atmospheric pressure chemical ionization tandem mass spectrometry: application to a clinical pharmacokinetic study. J. Pharm. Biomed. Anal. 51, 875–881 (2009).
Ibrahim, R. M. et al. HPLC-DAD-MS/MS profiling of phenolics from Securigera securidaca flowers and its anti-hyperglycemic and anti-hyperlipidemic activities. Revista Brasileira De Farmacognosia 25, 134–141 (2015).
Chang, C. L. & Wu, R. T. Quantification of (+)-catechin and (–)-epicatechin in coconut water by LC–MS. Food Chem. 126, 710–717 (2011).
Falcão, S. et al. Phenolic profiling of Portuguese Propolis by LC-MS spectrometry: uncommon Propolis rich in flavonoid glycosides. Phytochem. Anal. 24 (2013).
Ma, Y. et al. LC/MS/MS quantitation assay for pharmacokinetics of naringenin and double peaks phenomenon in rat plasma. Int. J. Pharm. 307, 292–299 (2006).
Liu, W. et al. Determination and quantification of active phenolic compounds in pigeon pea leaves and its medicinal product using liquid chromatography-tandem mass spectrometry. J. Chromatogr. A 1217, 4723–4732 (2010).
Barnes, J. S. & Schug, K. A. Structural characterization of cyanidin-3,5-diglucoside and pelargonidin-3,5-diglucoside anthocyanins: multi-dimensional fragmentation pathways using high performance liquid chromatography-electrospray ionization-ion trap-time of flight mass spectrometry. Int. J. Mass Spectrom. 308, 71–80 (2011).
Zhang, J. Y. et al. Simultaneous screening and identifying four categories of Particular flavonoids in the leaves of Murraya exotica L. by HPLC-DAD-ESI-MS-MS. J. Chromatogr. Sci. 52 (2013).
Xia, M. et al. Characterization of chemical constituents of Oxytropis microphylla (pall.) DC. by ultra-high-performance liquid chromatography coupled with quadrupole-time-of flight tandem mass spectrometry. Separations 9, 297 (2022).
Campbell, R., Harper, S. & Kemp, A. Isoflavonoid constituents of the heartwood of Cordyla Africana. J. Chem. Soc. C, 1787–1795. https://doi.org/10.1039/J39690001787 (1969).
Matos, M. J. et al. Coumarins—an important class of phytochemicals, Ch. 5. In Phytochemicals (eds. Rao, A. V. & Rao, L. G.) (IntechOpen, 2015). https://doi.org/10.5772/59982
Ahmad, I. Repenins A-D, four new antioxidative coumarinolignoids from Duranta repens Linn. Bioorg. Med. Chem. Lett. 19, 3521–3524 (2009). ISSN 0960-894X.
Lin, L. C., Yang, K. Y., Chen, Y. F., Wang, S. C. & Tsai, T. H. Measurement of daphnoretin in plasma of freely moving rat by liquid chromatography. J. Chromatogr. A. 1073, 285–289 (2005).
Tine, Y., Renucci, F., Costa, J., Wele, A. & Paolini, J. A method for LC-MS/MS profiling of coumarins in Zanthoxylum zanthoxyloides (Lam.) B. Zepernich and Timler extracts and essential oils. Molecules 22, 174 (2017).
Wang, H., Xiao, B., Hao, Z. & Sun, Z. Simultaneous determination of fraxin and its metabolite, fraxetin, in rat plasma by liquid chromatography-tandem mass spectrometry and its application in a pharmacokinetic study. J. Chromatogr. B 1017–1018, 70–74 (2016).
Kang, J. et al. Characterization of compounds from the roots of Saposhnikovia divaricata by high-performance liquid chromatography coupled with electrospray ionization tandem mass spectrometry. Rapid Commun. Mass Spectrom. 22, 1899–1911 (2008).
Cao, S. et al. Chemical constituent analysis of Ranunculus sceleratus L. using ultra-high-performance liquid chromatography coupled with quadrupole-orbitrap high-resolution mass spectrometry. Molecules 27, 3299 (2022).
Yi, T. et al. An integrated strategy based on UPLC–DAD–QTOF-MS for metabolism and pharmacokinetic studies of herbal medicines: Tibetan Snow Lotus herb (Saussurea laniceps), a case study. J. Ethnopharmacol. 153, 701–713 (2014).
Huang, H. M. et al. HPLC/ESI-MS and NMR analysis of chemical constitutes in bioactive extract from the root nodule of Vaccinium emarginatum. Pharmaceuticals 14 (2021).
Ahmad, Z. et al. ChemInform abstract: butyrylcholinesterase inhibitory triterpenes from Salsola baryosma. ChemInform 39 (2008).
Pham, H. et al. UHPLC-Q‐TOF‐MS/MS dereplication to identify chemical constituents of Hedera helix leaves in Vietnam. J. Anal. Methods Chem. 1–18 (2022).
Ahmad, Z. et al. Structural determination of salsolins A and B, new antioxidant polyoxygenated triterpenes from Salsola baryosma, by 1D and 2D NMR spectroscopy. Magn. Reson. Chem. 46, 94–98. https://doi.org/10.1002/mrc.2119 (2008).
Lecomte, J., López-Giraldo, L., Laguerre, M., Baréa, B. & Villeneuve, P. Synthesis, characterization and free radical scavenging properties of rosmarinic acid fatty esters. J. Am. Oil Chem. Soc. 87, 615–620 (2010).
Anuar, T. A. F. T. et al. LCMS dataset on compounds in Syzygium polyanthum (Wight) Walp. leaves variant from the East coast of Peninsular Malaysia. Data Brief. 39, 107485 (2021).
Rekik, O., Ben Mansour, A. & Bouaziz, M. Evaluation of phenolic composition and antioxidant activity changes in olive flowers during development using HPLC/DAD and LC-MS/MS. Electrophoresis 39, 1663–1672 (2018).
Park, S., Kim, B., So, H.-Y., Kim, Y.-J. & Kim, J. Development of isotope dilution-liquid chromatography/tandem mass spectrometry as a candidate reference method for the determination of acrylamide in …. Bull Korean Chem. Soc. 28 (2007).
Tao, Y., Li, W., Liang, W. & van Breemen, R. Identification and quantification of gingerols and related compounds in ginger dietary supplements using high-performance liquid chromatography-tandem mass spectrometry. J. Agric. Food Chem. 57, 10014–10021 (2009).
Sinosaki, N. B. M. et al. Structural study of phenolic acids by triple quadrupole mass spectrometry with electrospray ionization in negative mode and H/D isotopic exchange. J. Braz. Chem. Soc. 31 (2020).
Jin, Y. et al. UHPLC-Q-TOF-MS/MS-oriented characteristic components dataset and multivariate statistical techniques for the holistic quality control of Usnea. RSC Adv. 8, 15487–15500 (2018).
Ruan, J. et al. Comprehensive chemical profiling in the ethanol extract of Pluchea indica aerial parts by liquid chromatography/mass spectrometry analysis of its silica gel column chromatography fractions. Molecules 24, 2784 (2019).
Hong, Y., Liao, X. & Chen, Z. Determination of bioactive components in the fruits of Cercis Chinensis Bunge by HPLC-MS/MS and quality evaluation by principal components and hierarchical cluster analyses. J. Pharm. Anal. 11, 465–471 (2021).
Xu, S., Pavlov, J. & Attygalle, A. B. Collision-induced dissociation processes of protonated benzoic acid and related compounds: competitive generation of protonated carbon dioxide or protonated benzene. J. Mass Spectrom. 52, 230–238 (2017).
Yin, F. et al. Quantitation of uridine and L-dihydroorotic acid in human plasma by LC–MS/MS using a surrogate matrix approach. J. Pharm. Biomed. Anal. 192, 113669 (2021).
Ocque, A. J., Stubbs, J. R. & Nolin, T. D. Development and validation of a simple UHPLC–MS/MS method for the simultaneous determination of trimethylamine N-oxide, choline, and betaine in human plasma and urine. J. Pharm. Biomed. Anal. 109, 128–135 (2015).
Qing, Z. et al. Investigation of fragmentation behaviours of isoquinoline alkaloids by mass spectrometry combined with computational chemistry. Sci. Rep. 10 (2020).
Grossert, S., Fancy, P. & White, R. Fragmentation pathways of negative ions produced by electrospray ionization of acyclic dicarboxylic acids and derivatives. Can. J. Chem. 83, 1878–1890 (2011).
Norazhar, A. I., Lee, S. Y., Faudzi, S. M. M. & Shaari, K. Metabolite profiling of christia Vespertilionis leaf metabolome via molecular network approach. Appl. Sci. (Switzerland) 11 (2021).
Bi, X. et al. Rapid and sensitive determination of fatty acids in edible oil by liquid chromatography-electrospray ionization tandem mass spectrometry. Sci. China Chem. 57, 447–452 (2014).
Takahashi, H. et al. Long-chain free fatty acid profiling analysis by liquid chromatography–mass spectrometry in mouse treated with peroxisome proliferator-activated receptor α agonist. Biosci. Biotechnol. Biochem. 77, 2288–2293 (2013).
Li, H. L. et al. Determination of α-hederin in rat plasma using liquid chromatography electrospray ionization tandem mass spectrometry (LC-ESI-MS/MS) and its application to a pharmacokinetic study. Anal. Methods 7 (2015).
Kretschmer, P. M., Bannister, A. M., Brien, O., MacManus-Spencer, M. K., Paulick, M. G. & L. A. & A liquid chromatography-tandem mass spectrometry assay for the detection and quantification of trehalose in biological samples. J. Chromatogr. B Analyt Technol. Biomed. Life Sci. 1033–1034, 9–16 (2016).
Ralet, M. C., Lerouge, P. & Quéméner, B. Mass spectrometry for pectin structure analysis. Carbohydr. Res. 344, 1798–1807 (2008).
Brown, P., Brüschweiler, F., Pettit, G. & Reichstein, T. Field ionization mass spectrometry—III: Cardenolides. Org. Mass Spectrom. 5, 573–597 (1971).
Bader, A. et al. Systematic phytochemical screening of different organs of Calotropis procera and the ovicidal effect of their extracts to the foodstuff pest cadra cautella. Molecules 26, 905 (2021).
Hosseini, S. H. et al. LC-MS-based metabolite profiling of aqueous extract of Pergularia tomentosa L. and its anti-hyperglycemic effect. Iran. J. Basic Med. Sci. 25, 1433–1441 (2022).
Toribio, F. P. & Geissman, T. A. Sesquiterpene lactones. New lactones from Hymenoclea salsola T. and G. Phytochemistry 7, 1623–1630 (1968).
Palmgrén, J. J., Töyräs, A., Mauriala, T., Mönkkönen, J. & Auriola, S. Quantitative determination of cholesterol, sitosterol, and sitostanol in cultured Caco-2 cells by liquid chromatography–atmospheric pressure chemical ionization mass spectrometry. J. Chromatogr. B 821, 144–152 (2005).
Lukitaningsih, E. Phytosterol content in Bengkoang (Pachyrhizus erosus). Pharmacon: Jurnal Farmasi Indonesia UMS. 13, 47–54 (2012).
Cho, H. K., Suh, W. S., Kim, K. H., Kim, S. Y. & Lee, K. R. Phytochemical constituents of Salsola komarovii and their effects on NGF induction. Nat. Prod. Sci. 20, 95–101 (2014).
Soliman, M. et al. The protective impact of Salsola imbricata leaf extract from taif against acrylamide-induced hepatic inflammation and oxidative damage: the role of antioxidants, cytokines, and apoptosis-associated genes. Front. Vet. Sci. 8 (2022).
Hanif, Z., Ali, H. H., Rasool, G., Tanveer, A. & Chauhan, B. Genus Salsola: its benefits, uses, environmental perspectives and future aspects—a review. J. Rangel. Sci. 8, 315–328 (2018).
Kalinowska, M. et al. Plant-derived and dietary hydroxybenzoic acids—a comprehensive study of structural, anti-/pro-oxidant, lipophilic, antimicrobial, and cytotoxic activity in MDA-MB-231 and MCF-7 cell lines. Nutrients 13, 3107 (2021).
Lacaille-Dubois, M. A. & Wagner, H. New perspectives for natural triterpene glycosides as potential adjuvants. Phytomedicine 37 (2017).
Oueslati, M., Bouajila, J. & Ben Jannet, H. Two new bioactive biphenylpropanoids from the roots of Salsola imbricata (Chenopodiaceae) growing in Saudi Arabia. Oriental J. Chem. 33, 1871–1878. https://doi.org/10.13005/ojc/330432 (2017).
Wang, W.-Z. et al. Three new compounds isolated from the whole plants of Salsola collina pall. Nat. Prod. Res. 1–8. https://doi.org/10.1080/14786419.2022.2055556 (2022).
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB).
This work was supported by Science, Technology & Innovation Funding Authority [Call 6, 2022], where that authority didn’t contribute to any step of the work.
Author information
Authors and Affiliations
Contributions
Abdelrhman Zakaria: Conceptualization, Methodology, Data curation, Writing-original draft preparation, Supervision, Validation, Writing- review and editing. Fahima F. Kassem: Methodology, Data curation, Writing-original draft preparation. Doaa A. Ghareeb: Methodology, Data curation, Writing-original draft preparation. Safa M. Shams Eldin: Conceptualization, Methodology, Data curation, Writing-original draft preparation, Supervision, Validation, Writing- review and editing. Dina A. Selim: Conceptualization, Methodology, Data curation, Writing-original draft preparation, Supervision, Validation, Writing- review and editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics declaration
Human leukocyte cells were isolated from peripheral blood samples obtained from human participants. Informed consent was obtained from all subjects. All methods were carried out following relevant guidelines and regulations. The study was approved by the bioethics committee of the Faculty of Pharmacy, Alexandria University (approval number AU06202311937).
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zakaria, A., Kassem, F.F., Ghareeb, D.A. et al. A comprehensive metabolomic study of three Egyptian Salsola species revealed their potential anti-inflammatory activity. Sci Rep 15, 5056 (2025). https://doi.org/10.1038/s41598-024-80807-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-80807-2