Abstract
Smart weed-crop discrimination is crucial for modern precision weed management. In this study, we aimed to develop a robust system for site-specific weed control in saffron fields by utilizing color images and a deep learning approach to distinguish saffron from four common weeds: flixweed, hoary cress, mouse barley, and wild garlic. A total of 504 images were taken in natural and unstructured field settings. Eight state-of-the-art deep learning networks – VGG19, ResNet152, Xception, InceptionResNetV2, EfficientNetB0, EfficientNetB1, EfficientNetV2B0, and EfficientNetV2B1 were evaluated as potential base networks. These networks underwent pre-training on ImageNet using transfer learning, followed by fine-tuning and improvement with additional layers to optimize performance on our dataset. The improved EfficientNetB0 model stood out as the top performer among the eight models, achieving an accuracy rate of 94.06% and a loss value of 0.513 on the test dataset. This proposed model excelled in accurately classifying plant categories, obtaining f1-scores ranging from 82 to 100%. We scrutinized fifteen scenarios of weed presence in saffron fields, focusing on various weed types, to propose efficient management tactics using the model. These discoveries lay the groundwork for precise saffron weed management strategies that reduce herbicide use, environmental impact, and boost yield and quality.
Similar content being viewed by others
Introduction
Saffron (Crocus sativus L.), a member of the Iridaceae family, is one of the most valuable medicinal and spice plants in Asia, particularly in Iran1. Saffron is a weak competitor against weeds, so weed control in saffron fields using methods with the least dependence on herbicide usage is crucial for producing a healthy crop and is an important aspect of managing the production of this crop2. Manual removal of weeds in fields is not a viable option due to the risk of crop damage and the high labor costs involved. An alternative approach to reducing herbicide use is to capitalize on the spatial patterns of weed distribution within fields. Understanding the spatial distribution of weeds is a key factor in implementing site-specific weed management (SSWM), a strategy that enables targeted herbicide application, minimizes environmental impact, and helps mitigate the development of herbicide-resistant weed populations3. According to a study by Nordmeyer4, that analyzed five cereal fields, targeted herbicide application was necessary for 39% of the area to control grass weeds, 49% to manage Galium aparine L., and 44% to control other broadleaf weed species. Precise identification and differentiation of weeds from crop plants is a crucial component of effective site-specific weed management. By precisely distinguishing weeds from main crop, farmers can target herbicide applications to specific areas where weeds are present, thereby optimizing weed control.
By harnessing the power of advanced technologies like computer vision and image processing, researchers can achieve high precision in weed detection, enabling the development of targeted management strategies that accurately distinguish between crops and weeds, and ultimately optimizing the effectiveness of site-specific weed control techniques3,5. Notably, RGB or color images, which capture 2D information, have been extensively utilized in this field6, laying the groundwork for further advancements. Research has demonstrated that distinct morphological characteristics, such as leaf shape, color, and size, can be leveraged as effective features for object identification and classification models7. Specifically, features like leaf area, length, perimeter, width, and color can be employed to differentiate between plants and inform plant management strategies8. The complexity of weed detection is heightened by the diversity of weed species and their morphological differences, as well as the visual similarities between weeds and coexisting crop plants9. This task is further complicated by environmental factors such as complex backgrounds, variable lighting conditions10, and occlusions with other plant components11, which can compromise detection accuracy. Moreover, algorithms must be able to generalize across different features and operate in real-time, enabling applications such as autonomous weed-fighting robots to rapidly identify various types of weeds. Additionally, timing is critical in robotic weed control, and researchers are working to develop solutions that determine the optimal moment to implement control measures. Despite these challenges, significant progress has been made in developing object recognition algorithms for precision agriculture, with varying degrees of accuracy achieved in different applications. The application of deep learning techniques has demonstrated significant potential in achieving these objectives. Over the past decade, deep learning has transformed the field of image processing, achieving unprecedented success and finding numerous practical applications in precision agriculture12. While deep learning algorithms require large datasets for training and validation13, advances in computational power and data availability have made them increasingly popular. As a subset of machine learning, deep learning leverages hierarchical representations of data through sequential layers, typically employing convolutional neural networks (CNNs) to extract complex, non-linear image features. This approach has been shown to exhibit exceptional generalization capabilities, making it well-suited for classification tasks14,15,16.
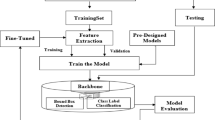
Convolutional neural networks are a specialized type of deep learning technique, specifically designed for supervised classification tasks17. The process begins with the input of an image, which can vary in size, and undergoes preprocessing before entering the feature extraction stage. This phase involves a series of layers that progressively extract features from the image, with the complexity of the features increasing with each additional layer. Key components of this phase include Convolutional and Max Pooling layers, although other layers such as Global Average Pooling, Batch Normalization, and more may also be utilized. Following feature extraction, the classification process begins, employing traditional fully connected layers commonly found in neural networks. The arrangement and configuration of these layers, as well as their parameters and properties, give rise to a diverse range of networks with distinct characteristics and capabilities. Nevertheless, standard networks developed by companies and research institutions have demonstrated their effectiveness through proven performance17. CNNs have been extensively explored for identification and classification tasks in agriculture. Research has demonstrated the effectiveness of these networks in various applications, including fruit detection and recognition. For instance, Saedi & Khosravi18, developed an image-based CNN model to recognize six different types of on-branch fruits in orchards, achieving an impressive accuracy of 99.8%. Also, Khosravi et al.19 employed a deep neural network approach to classify two olive fruit cultivars at various growth stages, with their designed model achieving an accuracy of over 91% in accurately identifying each stage. Other studies have utilized deep learning algorithms, such as ResNet10115, Fast R-CNN20, and Faster R-CNN6, to detect and classify various fruits, including apples and olives. Transfer learning has also been applied to develop fruit detection algorithms using RGB and NIR images21. Additionally, researchers have used deep neural networks, such as Xception22 and lightweight CNNs23, to recognize and classify fruits with high accuracy, demonstrating the potential of these models in image classification tasks.
Deep learning algorithms have been successfully applied to weed detection and classification, demonstrating superior performance in distinguishing between crop plants and weeds9,24. For instance, Quan et al.25, (2019) employed the Faster R-CNN model to detect corn seedlings and weeds in complex environments, while Fan et al.26 improved this model to achieve an accuracy of over 98.43% in identifying weeds in cotton fields. G C et al.27 developed a cloud-based automatic data acquisition system to capture images of weeds and crops at regular intervals, allowing for the consideration of plant growth stages in weed identification. Other studies have utilized deep learning-based models to identify common weeds in sugar beet cultivation28, distinguish between weeds and pepper plants29, and detect weeds in sugar beet fields30. Additionally, researchers have evaluated the performance of deep learning models, such as Yolov8, Yolov9, and customized Yolov9, in real-field conditions for eight crop species27. Similarly, a modified Xception deep learning approach was successfully utilized to distinguish saffron from its broadleaf weeds, yielding promising results with f1-scores ranging from 91 to 96%. In a practical study, Rai & Sun,31 designed a single-stage deep learning (DL) architecture that accomplishes two tasks simultaneously: detecting weed presence through bounding boxes and achieving pixel-wise instance segmentation on unmanned aerial system (UAS) acquired remote sensing images. The model achieved the best detection and segmentation scores of 85.4 and 82.1%, respectively.
The mentioned works demonstrate the potential of deep learning algorithms in plant identification and classification, and highlight the need for further research in this area. Therefore, the current study introduces advanced deep learning models designed to differentiate between saffron plants and four common weed species: Flixweed, Hoary cress, Mouse barley, Saffron, and Wild garlic. Utilizing color images, these models can accurately classify each plant category across various scenarios, paving the way for the creation of precise weed management systems for automated control in saffron fields.
Materials and methods
Data preparation and pre-processing
To develop a deep neural network model, images were collected from the five classes understudy: Flixweed, Hoary cress, Mouse barley, Saffron, and Wild garlic. A selection of these images and their total quantity are illustrated in Fig. 1. The photos were taken using a Samsung Galaxy A12 (model number SM-A125F/DS) mobile phone camera with a 48-megapixel lens, between March and April 2024, at 10 am every day. To obtain images of weeds and saffron, saffron fields were selected from three regions of Iran. The first region was South Khorasan, Boshrooyeh County, including saffron fields in two areas of Eresk City (Longitude: 57°21 × 19˝E. Latitude: 33°42 × 8˝N.). The second region was Razavi Khorasan Province, Torbat Heydariyeh County, covering saffron fields in the Roshtkhar and Malekabad areae (Longitude: 59°13′10″E. Latitude: 35°16′26″N). The third region was Semnan Province, Shahroud County, focusing on saffron fields in the Bastam area (Longitude: 54°59′54″E. Latitude: 36°29′5″N). In each region, the authors captured photos of weeds from at least five different fields. The images analyzed in this study were obtained with the supervision and permission of the relevant university, adhering to national guidelines. Additionally, all authors ensured full compliance with local and national regulations.
The images captured underwent a review process where inappropriate samples were excluded. An agronomist with weed and crop properties expertise contributed to this selection process. Ultimately, 504 images were chosen for model development and network training. From the total data (504), 20% i.e. 101 data were selected as test data. The remaining 80% were divided into two groups, 85% of which (342) were selected as training data and 15% (61) as validation data. Validation data, derived from the training set, is crucial for assessing the model’s performance during training. It aids in fine-tuning the model’s hyper parameters and configurations based on feedback obtained during validation. It’s worth emphasizing that the test set is reserved exclusively for evaluating the final model performance and testing its accuracy on unseen data. This separation ensures a reliable assessment of the model’s generalization capabilities.
Preprocessing input images is essential for improving the model’s accuracy, mitigating overfitting, and enhancing its generalization ability. Initially, we standardized the image sizes to 299 × 299 and 224 × 224 pixels to ensure uniform resolution across all images. Subsequently, we normalized the resolutions to fall within the range [0,1] by dividing their values by the maximum value present in the captured images, as outlined in Eq. 1.
where \(\:{x}_{i}^{{\prime\:}}\) symbolizes the normalized value within the dataset, \(\:{x}_{i}\) represents the i-th value in the dataset, and\(\:\:{x}_{min}\) and \(\:{x}_{max}\:\)denote the minimum and maximum values in the dataset. This formula guarantees that after normalization the normalized value for the dataset’s minimum value will consistently be 0, and the normalized value for the dataset’s maximum value will invariably be 1.
One of the primary challenges faced by deep neural networks is their reliance on a substantial amount of labeled data to achieve satisfactory accuracy. This large dataset facilitates the training process by enabling the network to learn all the necessary parameters and minimize the risk of overfitting. However, acquiring such a large number of labeled images can be a laborious, expensive, and time-consuming endeavor. Data augmentation, a well-established technique in computer vision, offers an effective solution to this problem. It helps reduce overfitting by generating additional training data from existing samples through the application of random transformations, ensuring that the generated images remain plausible. This process improves the model’s generalization because the same image is not encountered more than once17. In this study, an automated data augmentation method was employed, which included random height and width translation, random flip, random contrast, and random rotation. The resulting images were then used to train the model. The specific data augmentation parameters are presented in Table 1. A sample result of this step in outlined in Fig. 2.
Base networks
The initial step towards accomplishing our goal involves selecting a base network from a range of standard options through transfer learning, where their weights are loaded from the ImageNet dataset. To this goal, we considered testing eight innovative deep learning candidate networks including VGG19, ResNet152, Xception, InceptionResNetV2, EfficientNetB0, EfficientNetB1, EfficientNetV2B0, and EfficientNetV2B1.
Fine-tuning
The objective of fine-tuning process was to customize the candidate base networks for the particular dataset and task at hand, leveraging the previously learned features while adapting to the new data. This prevented overfitting of the pre-trained layers on the new dataset while enabling the newly added layers to adapt to the new data. During the fine-tuning process, our goal was to selectively freeze certain layers (the early ones) while enabling the training of others (the later ones) when incorporating a pre-trained model into transfer learning within a deep learning framework. Initially, we disabled the `trainable` attribute for the first 20 layers, locking them during training to prevent their weights from being altered, and preserved the valuable features they have already acquired from the original training data. Subsequently, we advanced to fine-tune the latter layers of the base model. Then, we iterated through the layers starting from the 20th position and activate the `trainable` attribute for each of these layers. This adjustment allows these layers to undergo training and updates throughout the training phase.
Structure of the proposed models
To create a robust and improved deep learning model, a modifying block was added to the last layer of the base networks. We tested different layer configurations for the modifying block, exploring various types, placements, and parameters to find the most efficient setup. Various types and combinations of layers, such as 2D Convolution, Global Average Pooling, Dropout, Batch Normalization, among others, were examined. The parameters for this stage were selected through trial-and-error testing, which is not elaborated here to maintain brevity. Ultimately, the optimized modifying block involved three sets of layers including Convolution, Batch Normalization, Max Pooling, and Dropout layers, followed by Fully Connected and Global Average Pooling layers. These modifying layers were appended to the final layer of all the eight fine-tuned base networks to create the proposed architecture for the outlined study (Fig. 3).
In this study, a total of eight improved models were examined using two different optimizers, SGD and Adam, along with two image sizes (224 × 224 and 299 × 299), resulting in the evaluation of 32 unique models. From these models, the top eight performing ones (models one to eight) were identified and compared to meet the objectives outlined of the study (Table 2).
we evaluated these models using a batch size of 32, and 200 epochs to identify the optimal model. During this process, we employed selection criteria that included assessing the model’s stability during training, as well as analyzing accuracy on the test dataset. Considering the accuracy of the above 8 models (Results and Discussion section), finally, Model 5 with EfficientNetB0 base network was selected as the best model for the purposes of this research.
The architecture of the EfficientNetB0 network
EfficientNetB0 is a cutting-edge convolutional neural network (CNN) architecture that excels in efficient and accurate image classification. It employs a novel compound scaling method to enhance performance while keeping computational costs low. The core components of this network are the MBConv blocks, which include: Depthwise Convolution: This layer reduces parameters by applying filters to each input channel separately. Swish Activation Function: Enhances gradient flow during training, improving model performance. Squeeze-and-Excitation Layer: Dynamically recalibrates channel-wise feature responses to focus on the most informative features.
The EfficientNetB0 architecture begins with multiple MBConv blocks, followed by downsampling layers that reduce spatial dimensions. The final layers consist of global average pooling and a fully connected layer for outputting class probabilities. This design enables EfficientNetB0 to achieve high accuracy with fewer parameters, making it ideal for resource-constrained environments and transfer learning applications. Figure 4 illustrates the architecture of EfficientNetB0 network.
The final structure of the proposed deep learning model for the classification of five plant species under study is depicted in Fig. 5.
Table 3 offers a comprehensive overview of the layers comprising the model 5 (EfficieentNetB0 – based network), including the output shapes and parameter count. The model includes approximately 46,000 non-trainable parameters (0.54%) from a total exceeding 8.4 million. Specific layers, such as Dropout, Max Pooling, and Global Average Pooling, do not add to the number of trainable parameters. This is because these layers do not possess any trainable parameters themselves.
The number of parameters is a key factor in model performance, but it must be balanced with considerations like dataset size, task complexity, and computational resources. While more parameters can enable a model to learn more complex patterns, they also increase the risk of overfitting, computational cost, and optimization challenges. Therefore, the number of parameters is the primary factor influencing the response speed of a deep learning model. Trainable parameters refer to the weights and biases that the model can adjust and optimize during the training process. These parameters directly influence the ability of the model to learn and make predictions. A higher number of trainable parameters will increase the model’s capacity to capture complex patterns and relationships in the data, but it can also slow down the training and inference processes. On the other hand, non-trainable parameters are typically fixed values that are not updated during training. These parameters do not directly impact the model’s learning process or response speed. Therefore, the number of trainable parameters is the primary factor influencing the response speed of a deep learning model. EfficientNetB0 stands out with its few parameters, resulting in reduced memory usage and quicker inference. Despite its compact size, EfficientNetB0 delivers strong accuracy in tasks such as image classification, thanks to its efficient design.
Network training
The CNN training procedure adheres to the conventional neural network structure, encompassing two main phases: Feed-Forward and Back-Propagation. During the Feed-Forward phase, the network evaluates its error by contrasting the output produced from the input image with the expected output. Following this, in the Back-Propagation phase, the gradients of the parameters are determined based on the network’s error, and each weight matrix is adjusted accordingly. This cyclic process involves feeding the input into the network, passing it through multiple layers, comparing the resulting output with the target output to ascertain the error. This error is then leveraged to fine-tune the network parameters, with the iteration repeated numerous times, termed as an epoch. The error computation entails employing various formulas and functions, while the optimization technique entails making gradual modifications to the weights to minimize errors. To achieve this objective, we meticulously selected the most appropriate optimizer among established options such as RMSprop, SGD, Adam, and Nadam. The model’s performance was assessed using accuracy on the test dataset, with the categorical cross-entropy function acting as the designated loss function. As previously noted, SGD and Adam optimizers outperformed other options, making them the preferred choices for model development in this study. The specific arguments and values used for these optimizers are detailed in Table 4.
The model training encompassed a methodical exploration of hyperparameters and architectural components to pinpoint the optimal configuration. Throughout the training phase, we diligently tracked the trends in loss and accuracy for both the training and validation datasets at each epoch. This meticulous approach enabled us to conduct a comprehensive assessment of the models’ performance and make well-informed decisions regarding their suitability for our specific task.
The detailed outcomes of the model selection process have been extensively discussed in the results and discussion section. Within our thorough evaluation of the potential models, we scrutinized various performance metrics, encompassing accuracy, loss, and model stability. Accuracy functions as a gauge of the model’s correct prediction percentage, whereas loss denotes the mean error per instance. Lower loss values typically indicate superior model performance. However, it is essential to recognize that a model with high accuracy but elevated loss may imply potential overfitting issues, which can compromise its ability to generalize on unseen data. Hence, we meticulously assessed the risk of overfitting during the comparative analysis of the candidate models.
To ensure the credibility and reproducibility of our findings, we conducted the training, validation, and testing procedures utilizing Python version 3.10.12 within the cloud-based computing platform, Google Collaboratory. The computational setup featured an NVIDIA K80 GPU and 12 GB of memory. We leveraged Keras with TensorFlow’s backend (version 2.18.1) in conjunction with supplementary libraries like OpenCV to streamline the execution of our experiments.
Results and discussion
In this section, the results of developing deep learning models for distinguishing between saffron and four prevalent weeds, namely Flixweed, Hoary cress, Mouse barley, Saffron, and Wild garlic, using color images are presented. The proposed model would be suitable for precision saffron weed management in various scenarios where any of these weeds are present.
Evaluation of models
The initial step in developing an effective deep learning model involves choosing a base network from various standard options through transfer learning, where their weights are imported from the ImageNet dataset. We evaluated 32 state-of-the-art standard deep learning base networks derived from VGG19, ResNet152, Xception, InceptionResNetV2, EfficientNetB0, EfficientNetB1, EfficientNetV2B0, and EfficientNetV2B1 networks using two input image sizes (299 × 299 pixels and 224 × 224 pixels) and two optimizers (SGD and Adam). Initially, eight top-performing networks were assessed to decide on the best model for the purpose of our study.
The results of the accuracy study of the above 8 models on the test data are shown in Fig. 6. Accordingly, model 5 with an accuracy of 94% on the test dataset was selected as the proposed model to meet the objectives of this research.
Train performance of the proposed model
The train performance of the best model (Model 5) is illustrated in Table 5; Fig. 7. As observed in Table 5, the model achieved 100% accuracy on the training and validation datasets on epoch 90 and epoch 186, respectively. The minimum loss on train and validation dataset were 0.1282 on epoch 167 and 0.2899 on epoch 80, respectively. It is worth mentioning that the proposed model represented the accuracy and loss of 0.9406 and 0.5132 on test dataset.
Figure 7 illustrates the training performance of model 5 over 200 epochs, displaying both accuracy and loss metrics for the training and validation sets. As shown in this figure, the accuracy of both the training dataset and validation dataset increases over time, indicating that the model is learning effectively. The accuracy on the training dataset consistently surpasses that of the validation dataset, which is typical since models generally perform better on data they were trained on. The difference between training and validation accuracies suggests some degree of overfitting, where performance is better on training data compared to validation data. However, this gap does not appear excessively large, suggesting that the model generalizes reasonably well. Regarding the loss plot, both losses for the training and validation datasets decrease over epochs—a positive sign indicating that error minimization is occurring. Similar to accuracy trends, losses are lower in training than in validation due to optimization being performed on training data. Importantly, there is no significant increase in validation loss over time; this suggests that severe overfitting does not occur.
Classification performance
To evaluate the efficacy of the proposed model in discriminating among the five classes, various parameters were considered. These parameters include True Positive (TP), representing the count of instances correctly identified as positive; True Negative (TN), denoting instances correctly identified as negative; False Positive (FP), indicating instances incorrectly identified as positive; and False Negative (FN), signifying instances incorrectly identified as negative. These parameters are utilized in the development of two classification metrics: the classification report and the confusion matrix. The classification report furnishes insights into the model’s performance based on precision, recall, and f1-score, as outlined in Eqs. 2–4. Precision quantifies the model’s accuracy in predicting positive cases. A high precision value suggests a lower ratio of incorrect to correct predictions, and conversely. Recall measures the proportion of correctly predicted positive instances relative to the total actual positive instances, emphasizing the model’s ability to accurately identify all positive instances. Furthermore, recall provides information on the model’s capacity to minimize false negatives. The f1-score, calculated as the geometric mean of precision and recall, serves as a metric that balances these two parameters to achieve an optimal trade-off between precision and recall.
The classification report detailing the categorization of Flixweed, Hoary cress, Mouse barley, Saffron, and Wild garlic classes based on color images by the proposed model is outlined in Table 6. There are four averaging methods in this table: Micro Avg., which aggregates metrics globally, giving equal weight to each instance. Macro Avg., which averages metrics per class, giving equal weight to each class. Weighted Avg., which averages metrics per class, weighted by the number of instances in each class, and finally, Samples Avg., which averages metrics per instance. Each averaging method provides a different perspective on the model’s performance.
Based on the classification report in Table 6, the model demonstrates strong performance for the Flixweed, Hoary Cress, Mouse Barley, and Wild Garlic classes, achieving precision, recall, and F1-scores above 0.90. However, the Saffron class presents a challenge, with a lower precision of 0.70 but a perfect recall of 1.00. This indicates that while the model successfully identifies all true Saffron cases, it tends to over-predict them, resulting in more false positives. The high Micro Average (0.94) reflects strong overall performance when treating all samples equally, while the Macro Average (0.94 for precision and 0.93 for F1-score) shows balanced performance across categories despite the precision issue with Saffron. Additionally, the Weighted Average (0.96 for precision and 0.94 for F1-score) highlights the influence of classes with more samples, such as Mouse Barley, on the final scores. The Samples Average (0.94) aligns with the other metrics, further confirming the model’s robustness. In conclusion, the model generally performs well but requires improvement for the Saffron class, particularly in reducing false positives to enhance precision.
Figure 8 presents the confusion matrix for the classification of five classes using the proposed model. In this matrix, the rows represent true classes, while columns show predicted classes. Examining this matrix reveals that all 23 Flixweed instances were correctly predicted without any misclassifications, indicating perfect performance for this class. For Hoary Cress, there were 18 correct predictions with three instances misclassified as Saffron. This highlights a potential weakness in distinguishing between Hoary Cress and Saffron despite strong overall performance. Mouse Barley showed excellent performance with 26 correct predictions and only one instance misclassified as Saffron. This suggests minimal errors in its classification. Regarding Saffron, it is evident that there were 14 correct predictions but two instances misclassified as Wild Garlic. Although all actual Saffron cases were identified (perfect recall), confusion with Wild Garlic indicates over-prediction and aligns with lower precision observed in the classification report. For Wild Garlic, there were 14 correct predictions with two misclassifications as Saffron. This indicates a slight difficulty in distinguishing Wild Garlic from Saffron. Overall, the matrix demonstrates strong performance for Flixweed, Mouse Barley, and Wild Garlic with minimal classification errors. Misclassifications primarily involve confusion between Saffron and both Hoary Cress and Wild Garlic, suggesting a need for further refinement to better distinguish Saffron from similar classes. The model shows high precision and recall for most categories but requires improvement in handling Saffron to reduce false positives and enhance differentiation from similar classes.
Figure 9 represents the class saliency map of the model 5. The saliency map is a heatmap that shows which parts of the image the model focused on to make its prediction. In the saliency map, the brighter (hotter) regions indicate areas that had the most influence on the model’s decision. This is achieved by computing the gradient of the output class score with respect to the input image pixels. The magnitude of these gradients indicates the importance of each pixel in the image for the classification decision. The saliency map in Fig. 9 highlighted the regions correspond to distinctive features of the plants (e.g., leaves, flowers, or stems). It shows bright regions around the leaves or flowers, suggesting that these parts of the plant were crucial for the model’s classification. This makes sense because leaves and flowers often contain distinctive patterns and shapes that help in identifying plant species. As the saliency map does not highlight irrelevant or noisy regions (e.g., background elements), it indicates that the model is not overfitting to non-discriminative features. Furthermore, the map suggests that the model is capable of distinguishing between different plant species by focusing on class-specific features. This is a positive sign of the model’s ability to generalize and differentiate between classes.
Saffron weed management scenarios
It is crucial to highlight that the findings of this study can be broadly applied to diverse weed appearance scenarios in saffron fields. By examining four distinct weed species, we demonstrate that the proposed model serves as a reliable tool for effective weed management across 15 potential scenarios. These scenarios, outlined in Table 7, consider the coexistence of one to four weeds alongside saffron. Clearly, scenario 15 reflects the general condition explored in this study, providing a comprehensive framework for understanding and addressing weed management challenges in saffron cultivation.
For this analysis, we utilized the confusion matrix (Fig. 8) and extracted a separate subset for each scenario, ensuring that saffron was included in each model (Table 7). This involved creating 15 subsets of the confusion matrix. The results are presented Fig. 10. Accordingly, when only one weed coexisted with saffron (Scenarios 1 to 4), the best performance was observed in the saffron-flixweed scenario, achieving a perfect accuracy of 100%. In contrast, the saffron-hoary cress coexistence scenario had the lowest accuracy at 91%. In scenarios where two weeds coexisted alongside saffron (Scenarios 5 to 10), the highest accuracy of 98% was achieved when Mouse Barley and Flixweed were present together with saffron. Conversely, Wild Garlic and Hoary Cress coexisting with saffron resulted in the lowest accuracy of 90%. When three weeds were present alongside saffron (Scenarios 11 to14), the highest accuracy again reached up to 98%, specifically in the case of Wild Garlic, Mouse Barley, and Flixweed. The least accurate scenario in this sub category involved Wild Garlic, Mouse Barley, and Hoary Cress (0.92 of accuracy). Calculating across all scenarios revealed an average accuracy of approximately 0.95 with a standard deviation of 0.027, indicating that the accuracy of overall condition (Scenario 15) (0.94) is comparable with the average.
Identifying different weed and crop species from RGB images poses a significant challenge, particularly when dealing with a large number of species. This difficulty stems from shared phenotypic traits, which complicate accurate differentiation. Additionally, weed management systems need to identify weeds in natural field conditions rather than controlled settings, adding another layer of complexity. Relying on images captured under laboratory conditions with fixed parameters like lighting is impractical due to these varying environmental factors. Recent research has shown promise by focusing on a broader range of species and achieving higher accuracy while minimizing the impact of specific environmental conditions. Various studies have addressed these challenges using advanced deep learning techniques, as highlighted in the comparative analysis presented in Table 8. This table compares different deep learning approaches for identifying weeds and crops in natural environments. It’s important to note that a successful weed discrimination model for one crop may not perform adequately for others. However, developing models that consider various weed appearance scenarios for distinct crops could be advantageous because it caters to diverse conditions that may arise for those crops. This aspect could be a significant strength of our study compared to similar research. For example, Shao et al.32 attempted to distinguish six weed species in paddy fields using deep learning methods under similar natural field conditions but achieved lower accuracy compared to our study. Fan et al.26 studied eight classes under cotton field conditions and reported accuracy higher than our findings. G C et al.33 applied five different deep learning frameworks to differentiate between crops and weed species in a greenhouse environment but reported relatively low accuracy despite considering 14 crop and weed species compared to our work. In another study by G C et al.34 images of four weed and six crop species were pre-processed using a feature selection algorithm before applying the VGG16 deep learning network in a greenhouse setting. While their classification results were impressive with f1-scores ranging from 93 to 97.5%, the pre-processing steps likely contributed significantly to their success. In contrast, our study achieved promising accuracy without such pre-processing steps. Finally, Makarian & Saedi35 attained perfect accuracy under natural conditions but limited their model’s generalizability by studying only a small number of classes compared to other studies like ours that handle more diverse scenarios effectively without extensive preprocessing or controlled environments.
Conclusion
In this research, we developed a deep neural network model based on the EfficientNetB0 architecture to classify saffron and four common weeds: Flixweed, Hoary cress, Mouse barley, and Wild garlic. Using color images as input data, we trained and evaluated the model. The results from testing on unseen data (test dataset) revealed that the model effectively identified the five plant species in images, achieving 94.06% accuracy with a loss of 0.513. Additionally, we explored fifteen different weed occurrence scenarios in saffron fields, considering the varying numbers of weeds alongside saffron, to identify optimal weed management strategies. The model consistently performed well across these scenarios, treating saffron as a distinct class. This method also allows for the identification of non-saffron plants as weeds, supporting a targeted weed control approach. Given the significance of producing herbicide-free saffron and minimizing environmental pollution, this study advances precision weed management and the potential for automated weed removal systems. This technology promises to reduce herbicide use, lessen environmental harm, and enhance saffron yield and quality.
Data availability
The datasets used and analysed during the current study available from the corresponding author on reasonable request.
References
Marrone, G. et al. Saffron (Crocus sativus L.) and its By-Products: healthy effects in internal medicine. Nutrients 16 (14), 2319. https://doi.org/10.3390/nu16142319 (2024).
Makarian, H., Rashed Mohassel, M. H., Bannayan, M. & Nassiri, M. Soil seed bank and seedling populations of Hordeum murinum and Cardaria draba in saffron fields. Agric. Ecosyst. Environ. 120 (2–4), 307–312. https://doi.org/10.1016/j.agee.2006.10.020 (2007).
Somerville, G. J., Sønderskov, M., Mathiassen, S. K. & Metcalfe, H. Spatial Modelling of Within-Field Weed Populations; a Review. In Agronomy 10,7. (2020). https://doi.org/10.3390/agronomy10071044
Nordmeyer, H. Patchy weed distribution and site-specific weed control in winter cereals. Precision Agric. 7 (3), 219–231. https://doi.org/10.1007/s11119-006-9015-8 (2006).
Juwono, F. H. et al. Machine learning for weed–plant discrimination in agriculture 5.0: an in-depth review. Artif. Intell. Agric. 10, 13–25. https://doi.org/10.1016/j.aiia.2023.09.002 (2023).
Gené-Mola, J. et al. Multi-modal deep learning for Fuji Apple detection using RGB-D cameras and their radiometric capabilities. Comput. Electron. Agric. 162, 689–698. https://doi.org/10.1016/j.compag.2019.05.016 (2019).
Susetyarini, E., Wahyono, P., Latifa, R. & Nurrohman, E. The identification of morphological and anatomical structures of Pluchea indica. J. Phys: Conf. Ser. 1539 (1), 12001. https://doi.org/10.1088/1742-6596/1539/1/012001 (2020).
Speck, O. & Speck, T. Functional morphology of plants - a key to biomimetic applications. New Phytol. 231 (3), 950–956. https://doi.org/10.1111/nph.17396 (2021).
Li, Z. et al. Winter wheat weed detection based on deep learning models. Comput. Electron. Agric. 227, 109448. https://doi.org/10.1016/j.compag.2024.109448 (2024).
Feng, J., Zeng, L. & He, L. Apple fruit recognition algorithm based on Multi-Spectral dynamic image analysis. Sens. (Basel Switzerland). 19 (4), 949. https://doi.org/10.3390/s19040949 (2019).
Wang, C., Tang, Y., Zou, X., Luo, L. & Chen, X. Recognition and matching of clustered mature litchi fruits using binocular charge-coupled device (CCD) color cameras. In Sensors 17, 11. (2017). https://doi.org/10.3390/s17112564
Kamilaris, A. & Prenafeta-Boldú, F. X. Deep learning in agriculture: A survey. Comput. Electron. Agric. 147, 70–90. https://doi.org/10.1016/j.compag.2018.02.016 (2018).
Dyrmann, M., Karstoft, H. & Midtiby, H. S. Plant species classification using deep convolutional neural network. Biosyst. Eng. 151, 72–80. https://doi.org/10.1016/j.biosystemseng.2016.08.024 (2016).
Nasiri, A., Taheri-Garavand, A. & Zhang, Y. D. Image-based deep learning automated sorting of date fruit. Postharvest Biol. Technol. 153, 133–141. https://doi.org/10.1016/j.postharvbio.2019.04.003 (2019).
Peng, H. et al. General improved SSD model for picking object recognition of multiple fruits in natural environment. Nongye Gongcheng Xuebao/Transactions Chin. Soc. Agricultural Eng. 34, 155–162. https://doi.org/10.11975/j.issn.1002-6819.2018.16.020 (2018).
Saedi, S. I. & Rezaei, M. A Modified xception deep learning model for automatic sorting of olives based on ripening stages. In Inventions 9,1. (2024). https://doi.org/10.3390/inventions9010006
Chollet, F. Deep Learning with Python 1st edn (Manning Publications Co., 2017).
Saedi, S. I. & Khosravi, H. A deep neural network approach towards real-time on-branch fruit recognition for precision horticulture. Expert Syst. Appl. 159, 113594 (2020).
Khosravi, H., Saedi, S. I. & Rezaei, M. Real-time recognition of on-branch Olive ripening stages by a deep convolutional neural network. Sci. Hort. 287, 110252. https://doi.org/10.1016/j.scienta.2021.110252 (2021).
Ren, S., He, K., Girshick, R. & Sun, J. Faster R-CNN: Towards Real-Time Object Detection with Region Proposal Networks. In C. Cortes, N. Lawrence, D. Lee, M. Sugiyama, & R. Garnett (Eds.), Advances in Neural Information Processing Systems 28. Curran Associates, Inc. (2015). https://proceedings.neurips.cc/paper_files/paper/2015/file/14bfa6bb14875e45bba028a21ed38046-Paper.pdf
Sa, I. et al. DeepFruits: A fruit detection system using deep neural networks. In Sensors16 8. (2016). https://doi.org/10.3390/s16081222
Salim, F., Saeed, F., Basurra, S., Qasem, S. N. & Al-Hadhrami, T. DenseNet-201 and xception pre-trained deep learning models for fruit recognition. In Electronics 12,14. (2023). https://doi.org/10.3390/electronics12143132
Saedi, S., Iman, Rezaei, M. & Khosravi, H. Dual-path lightweight convolutional neural network for automatic sorting of Olive fruit based on cultivar and maturity. Postharvest Biol. Technol. 216, 113054. https://doi.org/10.1016/j.postharvbio.2024.113054 (2024).
Upadhyay, A. et al. Advances in ground robotic technologies for site-specific weed management in precision agriculture: A review. Comput. Electron. Agric. 225, 109363. https://doi.org/10.1016/j.compag.2024.109363 (2024).
Quan, L. et al. Maize seedling detection under different growth stages and complex field environments based on an improved faster R–CNN. Biosyst. Eng. 184, 1–23. https://doi.org/10.1016/j.biosystemseng.2019.05.002 (2019).
Fan, X., Chai, X., Zhou, J. & Sun, T. Deep learning based weed detection and target spraying robot system at seedling stage of cotton field. Comput. Electron. Agric. 214, 108317 (2023).
Koparan, G. C. S. et al. A novel automated cloud-based image datasets for high throughput phenotyping in weed classification. Data Brief. 111097. https://doi.org/10.1016/j.dib.2024.111097 (2024).
Nasiri, A., Omid, M., Taheri-Garavand, A. & Jafari, A. Deep learning-based precision agriculture through weed recognition in sugar beet fields. Sustainable Computing: Inf. Syst. 35, 100759. https://doi.org/10.1016/j.suscom.2022.100759 (2022).
Subeesh, A. et al. Deep convolutional neural network models for weed detection in polyhouse grown bell peppers. Artif. Intell. Agric. 6, 47–54. https://doi.org/10.1016/j.aiia.2022.01.002 (2022).
Gao, J. et al. Deep convolutional neural networks for image-based convolvulus sepium detection in sugar beet fields. Plant. Methods. 16 (1), 29. https://doi.org/10.1186/s13007-020-00570-z (2020).
Rai, N. & Sun, X. WeedVision: A single-stage deep learning architecture to perform weed detection and segmentation using drone-acquired images. Comput. Electron. Agric. 219, 108792. https://doi.org/10.1016/j.compag.2024.108792 (2024).
Shao, Y. et al. GTCBS-YOLOv5s: A lightweight model for weed species identification in paddy fields. Comput. Electron. Agric. 215, 108461. https://doi.org/10.1016/j.compag.2023.108461 (2023).
GC, S., Zhang, Y., Howatt, K., Schumacher, L. G. & Sun, X. Multi-Species weed and crop classification comparison using five different deep learning network architectures. J. ASABE. 67 (2), 275–287. https://doi.org/10.13031/ja.15590 (2024).
Zhang, G. C. S., Koparan, Y., Ahmed, C., Howatt, M. R., Sun, X. & K., & Weed and crop species classification using computer vision and deep learning technologies in greenhouse conditions. J. Agric. Food Res. 9, 100325. https://doi.org/10.1016/j.jafr.2022.100325 (2022).
Makarian, H. & Saedi, S. I. Automated classification of saffron and broadleaf weeds of flixweed and hoary cress using deep learning and color images. Crop Prot. 106750. https://doi.org/10.1016/j.cropro.2024.106750 (2024).
Funding
This research has been financially supported by the Saffron Institute, University of Torbat Heydarieh, Iran. The grant number was P/179506.
Author information
Authors and Affiliations
Contributions
H. Makarian conceived and designed the research, prepared the images, and contributed to writing the manuscript. S. I. Saedi analyzed the images and data using modeling and software, and also contributed to writing the manuscript. H. Sahabi assisted in image preparation and revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Makarian, H., Saedi, S.I. & Sahabi, H. Smart weed recognition in saffron fields based on an improved EfficientNetB0 model and RGB images. Sci Rep 15, 15412 (2025). https://doi.org/10.1038/s41598-025-00331-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-00331-9