Abstract
Tuberculosis caused by Mycobacterium tuberculosis complex is a significant global health burden, with drug-resistant TB, especially multidrug-resistant TB, causing severe challenges to treatment. In Ethiopia, a high TB-burden country, drug resistance has continued spreading. However, some studies indicate genetic diversity, transmission dynamics, and resistance-conferring mutations by using targeted amplification, there are limited reports of whole genome sequencing analysis to uncover the antimicrobial resistance and virulent genes. Based on that, the objective of this project was to identify antimicrobial resistance regions and characterize virulence factors in M. tuberculosis isolates through in silico whole-genome sequence analysis. A FASTQ file of 45 M. tuberculosis isolates whole genome sequence was downloaded from the SAR database. Following quality control using FASTQC coupled with MultiQC and trimming with Trimmomatic, de novo assembly was conducted using SPAdes. The Burrows-Wheeler Aligner was used for mapping against the M. tuberculosis H37Rv reference genome, followed by variant calling with FreeBayes. In silico spoligotyping was performed using SpoTyping, and drug resistance mutations were identified with TB-Profiler and validated using Mykrobe. Virulence factors were detected through ABRicate and the Virulence Factor Database. STRING was used to network the virulent genes. All statistical analyses were performed using R software. This study revealed the most prevalent TB-lineage in the Amhara region was L4 (58.53%), followed by L3 (34.15%), and L1 (4.88%), and in silico spoligotyping classified 90.24% of the isolates into 12 shared types, with SIT 149 (41.46%) and SIT 21 (14.63%) as the most frequent spoligotypes. Seven major genotypic families were identified, with T3-ETH being the dominant family (48.78%). Drug resistance analysis revealed that 38 isolates (92.7%) were multidrug-resistant, and 1 (2.4%) was pre-extensively drug-resistant. Lineage 4 (59%) and its sub-lineage 4.2.2 (51.3%) show the highest resistance. The most frequent mutations to rifampicin, isoniazid, pyrazinamide, ethambutol, streptomycin, ethionamide, fluoroquinolone, and 2nd-line injectable drugs occurred at rpoB Ser450Leu, katG Ser315Thr, pncA c.-11A > G, embB Gly406Ala, rpsL Lys43Arg, Lys88Thr, ethA Met1, gyrA Ala90Val, Asp94Asn, and rrs 1401A > G, respectively. Additionally, a mutation at the mmpR5 gene for bedaquiline and clofazimine resistance occurred in one isolate. A total of 67 virulence genes were identified and 63 of them occurred in all isolates. The high prevalence of MDR-TB and the detection of resistance to both first- and second-line drugs in this study underscore the urgent need for enhanced TB control measures in the Amhara region.
Similar content being viewed by others
Introduction
Tuberculosis (TB) caused by species within the closely related Mycobacterium tuberculosis complex (MTBC), is an ancient human disease that continues to affect millions annually. MTBC is a group of closely related mycobacterial species that cause TB in humans and animals. The MTBC includes several species that are Mycobacterium tuberculosis, Mycobacterium bovis, Mycobacterium africanum, Mycobacterium microti, Mycobacterium caprae, Mycobacterium pinnipedii, and Mycobacterium canettii. The MTBC members share a high degree of genetic similarity but differ in their host range, pathogenicity, and epidemiological characteristics. Approximately 98% of human TB cases are caused by M. tuberculosis1.
Despite significant advances in diagnostic tools, the availability of effective anti-TB therapy, and extensive global efforts, TB remains a major public health concern worldwide and one of the leading causes of death globally2. In 2023, an estimated 10.8 million people contracted TB, with approximately 1.25 million deaths reported. The highest burden of TB cases was recorded in South-East Asia (45%), Africa (24%), and the Western Pacific (17%), with smaller shares in the Eastern Mediterranean (8.6%), the Americas (3.2%), and Europe (2.1%)3.
The emergence of rifampicin-resistant TB (RR-TB) and multidrug-resistant TB (MDR-TB), as well as resistance to both rifampicin and isoniazid, is a particular concern. The global burden of MDR-TB was estimated at 400,000 new cases in 20233.
Ethiopia is among the 30 high TB-burden countries globally3. There were an estimated 143,000 TB cases in the country in 2021 and an estimated 21,000 people died from TB. Ethiopia reported that 51% of notified individuals with bacteriologically confirmed pulmonary TB were tested for rifampicin resistance (RR-TB)4. The national five-year TB strategic plan (NSP) was revised in 2020, covering the period from July 2021 to June 2026. Within the NSP period, the Ministry of Health (MOH) targets to reduce TB incidence and mortality from 151 per 100,000 population and 22 per 100,000 population, respectively, in 2018 to 91 per 100,000 population and 7 per 100,000 population, by the end of the NSP5.
Drug-resistant tuberculosis (DR-TB) strains, particularly MDR-TB, continue to pose a serious threat to public healthcare systems, mainly in resource-constrained nations such as Ethiopia, where innovative molecular diagnostic technologies and well-equipped laboratory settings are lacking6,7. In Ethiopia, the factors contributing to this rise are not fully understood, but genetic differences among mycobacterial strain lineages, which may contribute to resistance-conferring mutations8,9, coupled with challenges such as population crowding, the HIV/AIDS epidemic, and poor treatment adherence10, play significant roles.
DR-TB usually occurs due to the patient’s delay in early diagnosis and treatment, previous anti-TB drug exposure, inappropriate drug regimens, the patient’s poor adherence to anti-tuberculosis drug regimens, and primary infection with DR-TB strains11. Drug resistance in Mycobacterium tuberculosis is not a product of a single homogeneous genetic unit. Rather it is a result of frequent mutation in various genes that encode for resistance to antibiotics12.
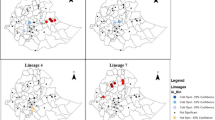
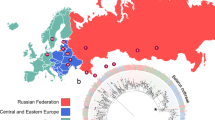
The human-adapted members of MTBC (M. tuberculosis and M. africanum) are classified into nine lineages with distinct geographic structures13. Indo-Oceanic (Lineage 1), East-Asian (Lineage 2), East-African-Indian (Lineage 3), Euro-American (Lineage 4), West-Africa 1 (Lineage 5), West-Africa 2 (Lineage 6), Ethiopian (Lineage 7)14, Lineage 815 and Lineage 913 were reported from the Central and Eastern Africa regions, respectively. Among these, the most common lineage on the planet is lineage 4 (L4)16.
There are first-line and second-line drugs to treat TB. First-line drugs are the most effective and are used as the initial treatment for drug-sensitive TB. These include Isoniazid (INH), Rifampicin (RIF), Ethambutol (EMB), Pyrazinamide (PZA), and Streptomycin (SM). These drugs are typically used in combination to prevent the development of drug resistance. Second-line drugs are used when TB is resistant to first-line drugs (MDR-TB) or when patients cannot tolerate the first-line drugs due to side effects. These include Fluoroquinolones (Levofloxacin, Moxifloxacin), Injectable agents (Amikacin, Kanamycin, Capreomycin), oral bacteriostatic agents (Ethionamide, Cycloserine, Terizidone), Bedaquiline, Delamanid, Linezolid and Clofazimine2,17.
TB drug resistance is classified into five categories. Mono-resistant TB is caused by TB bacteria that are resistant to one first-line anti-TB drug only. Poly-resistant TB is caused by TB bacteria that are resistant to resistance to more than one first-line anti-TB drug, other than both isoniazid and rifampicin. Multidrug-resistant TB (MDR TB) is caused by TB bacteria that are resistant to at least both isoniazid and rifampicin, the most effective first-line TB treatment drugs. Pre-extensively drug-resistant TB (pre-XDR TB) is a type of MDR TB caused by TB bacteria that are resistant to fluoroquinolones in addition to multidrug resistance. Extensively drug-resistant TB (XDR TB) is TB caused by M. tuberculosis strains that fulfill the definition of MDR/RR-TB and are also resistant to any fluoroquinolone and at least one additional Group A drug18,19.
The WHO has listed more than 30,000 variants of MTBC, detailing their frequency and associations with resistance or susceptibility. This includes identified mutations and summaries of key findings for 13 anti-TB drugs, based on the analysis of over 52,000 isolates with matched whole-genome sequencing and phenotypic drug susceptibility testing data from 67 countries20.
Rifampicin-resistant M. tuberculosis isolates have mutations in the 81-bp "hot-spot region" of the rpoB gene, spanning codons 507–533. The most common mutations occur in codons 516, 526, and 531. Mutations in codons like 518 or 529 are linked to low-level rifampicin resistance but remain susceptible to other rifamycins, such as rifabutin or rifalazil. Nearly all rifampicin-resistant strains are also resistant to other drugs, particularly isoniazid, making rifampicin resistance a key marker for MDR-TB21.
Resistance to isoniazid is mainly due to mutations in katG, inhA, ahpC, kasA, mshA, and NDH. Among these, the most common mutations are S315T in katG and 15C/T in inhA. Mutations in dfrA may also contribute to resistance, while mutations in the ahpC promoter can serve as markers for resistance22. Resistance to ethambutol is primarily linked to mutations in the embB gene, codons 306, 406, and 49723,24,25.
Mutations including large deletions in the pncA gene are the most common in pyrazinamide-resistant strains. These mutations are spread throughout the gene, primarily within a 561-bp region of the open reading frame or an 82-bp region of the putative promoter26.
Mutations in rpsL and rrs are the main mechanisms of streptomycin resistance. In rpsL, a common mutation is a lysine to arginine substitution at codon 43. In rrs, mutations frequently occur near nucleotides 530 and 915. Additionally, mutations in gidB, which encodes a 7-methylguanosine methyltransferase for 16S rRNA, result in low-level resistance to streptomycin27.
Fluoroquinolone resistance in M. tuberculosis is primarily associated with mutations in the gyrA and gyrB genes, which encode the subunits of DNA gyrase, a critical enzyme in DNA replication. Mutations in gyrA and gyrB disrupt the action of fluoroquinolones like ofloxacin, levofloxacin, moxifloxacin, and ciprofloxacin, leading to resistance28.
Virulent genes of M. tuberculosis (MTB) are crucial for the bacteria’s ability to cause disease. These genes help MTB evade the host immune system, survive within host cells, and cause damage to host tissues. Some well-known virulence genes and factors in MTB are ESX-1 Secretion System (esp and esx genes), PPE and PE-PGRS gene families, PhoP-PhoR regulatory system, KatG (Catalase-Peroxidase), mce Operons (mce1 and mce4), PknG (Protein Kinase G), LpqH (19-kDa Lipoprotein), HbhA (Heparin-Binding Hemagglutinin Adhesin), Mas and pks Genes (Polyketide Synthases) and Rv1411c (VirS)29,30.
According to Comas et al. (2015), L4 predominates across Ethiopia, L3 is widespread but more common in the north, and L7 is mostly found in the northern Ethiopian highlands. Numerous studies have identified spoligotypes SIT 149 and SIT 53 as major clades circulating in Ethiopia (Merid et al., 2021).
In different parts of Ethiopia, few studies have focused on whole-genome sequencing (WGS) of MTB isolates to analyze their genetic diversity, transmission dynamics, and drug resistance patterns. Studies in Northwest Ethiopia and the Tigray region reported L 4 was predominant followed by L3 and L1, L2 and L7 occurred rarely24,31. From Mycobacterium tuberculosis isolates from the central, eastern, and southeastern Ethiopia six major lineages L4, L3, L2, L1, L5, L6, and L7 were identified32. CAS was the most frequent sub-lineage in the Tigray region, followed by Ural and Haarlem24. Similarly, Delhi-CAS and EA.ETH (L4.2.2) were the predominant sub-lineages in Northwest Ethiopia31. In-silico whole genome sequence analysis of Mycobacterium tuberculosis sub-lineage 4.2.2/SIT149 as Dominant Drug-Resistant Clade in Northwest Ethiopia 2020–2022 that showed that L4.2.2.ETH was the leading drug-resistant sub-lineage showed extensive mutations against first-line anti-TB drugs. Ser450Leu/(tcg/tTg) for Rifampicin, Ser315Thr/(agc/aCc) for Isoniazid, Met306Ile/(atg/atA(C)) for Ethambutol, and Gly69Asp for Streptomycin33.
Drug-resistant tuberculosis poses significant challenges due to the increased difficulty and cost of treatment. Antimicrobial resistance in TB leads to heightened risks of disease transmission, more severe illness, disability, prolonged and complex treatment regimens, higher healthcare costs, and increased mortality rates.
Several methods have been used in previous studies to investigate resistance genes in Mycobacterium tuberculosis (MTB) strains. These methods include targeted gene sequencing of known resistance markers, such as the rpoB gene for rifampicin resistance, and the katG and inhA genes for isoniazid resistance, as well as molecular assays like the GeneXpert MTB/RIF test. Phenotypic drug susceptibility testing (DST) has also been widely employed to determine the resistance profiles of clinical isolates. While effective, these methods may fail to detect novel mutations and genetic factors contributing to drug resistance.
Whole-genome sequencing (WGS) offers a more comprehensive approach by providing detailed insights into both known and novel mutations associated with drug resistance. Although studies in Ethiopia have begun to explore the molecular epidemiology of MTB using WGS, the primary focus has been on lineage distribution and transmission dynamics. Comprehensive studies targeting specific virulence factors and antimicrobial resistance (AMR) regions in M. tuberculosis remained limited.
In different parts of Ethiopia, researchers reported the presence of diverse M. tuberculosis genotypes using WGS analysis. In Tigray region L4 was predominant, followed by L3. The most frequent mutations to RIF, INH, EMB, SM, PZA, ETH, FLQs, and 2nd-line injectable drugs were also reported24. Another comparative whole-genome sequence analysis of Mycobacterium tuberculosis isolated from pulmonary tuberculosis and tuberculous lymphadenitis patients in Northwest Ethiopia revealed L4 followed by L3 and then L731. Whole-genome sequencing-based analysis of Mycobacterium tuberculosis isolates from extrapulmonary tuberculosis patients in western Ethiopia showed that the majority of the isolates belonged to Lineage 4 (L4), with L4.6.3 and L4.2.2.2 emerging as the predominant sub-lineages34.
This study aimed to address this gap by identifying virulence factors and AMR regions in MTB using WGS data from previously published works on the molecular epidemiology and transmission dynamics of MDR-TB strains in the Amhara region of Ethiopia. Identification of these virulence genes and AMR regions is crucial for addressing the public health threat posed by MDR-TB, as WGS provides a detailed understanding of the genetic mutations responsible for resistance to both first-line and second-line anti-TB drugs.
A deeper understanding of the AMR-conferring mutations in MTB, as well as the associated molecular processes, is essential for improving rapid detection methods, discovering new drug targets, and developing more effective treatments and vaccines to reduce the global burden of TB24,35.
Based on the above facts, this study aimed to identify antimicrobial resistance gene mutations and characterize the virulent genes from M. tuberculosis isolates in the Amhara region, Ethiopia.
Methodology
Whole genome sequence data source
The present study used raw whole-genome sequence data of M. tuberculosis in the Amhara region which is available online from an open-source database. A FASTQ file of M. tuberculosis whole genome sequence was downloaded from SRA database. The WGS data were from 45 isolates, and the sequencing was performed using a NextSeq 550 desktop sequencer (Illumina, San Diego, CA, USA) to study the molecular epidemiology and transmission dynamics of MDR-TB strains in the Amhara region, Ethiopia by Shibabaw et al. (2023). Genomic DNA from individual strains was prepared for sequencing using Illumina Nextera XT library preparation kits according to the manufacturer’s instructions (Illumina, San Diego, CA, USA). Each isolate was sequenced using paired-end sequencing with 4 replicates for each fastq read file, resulting in two sequencing read files (designated by R1 and R2). The raw sequence data are publicly available under the project accession number PRJNA935744 (https://www.ncbi.nlm.nih.gov/sra/PRJNA935744).
The selection of isolates for WGS analysis was originally based on multiple criteria to ensure a representative and meaningful dataset. The selection criteria were based on the patient’s admission/ data collection year, molecular drug resistance patterns, MDR/RR-TB treatment center hospitals, geographical location of patients, and Lowenstein-Jensen (LJ) culture-positive isolates. A total of 45 isolates were included for WGS analysis. Demographic and clinical characteristics of patients showed that among 45 study participants, the median age was 29 years25,26,27,28,29,30,31,32,33,34,35,36,37, 60% were male and 20% were HIV-positive. Most of the study participants (67%) had a previous history of anti-TB treatment, the literacy rate was 64% and 51% were urban dwellers. In addition, 18% had a history of contact with MDR/RR-TB patients, 18% had a family history of TB and 91% were smear-positive at the time of diagnosis. The samples were collected from MDR/RR-TB treatment center hospitals in Amhara region, University of Gondar Hospital 19 (42.2%), Boru Meda Hospital 6 (13.3%), Woldia Hospital 6 (13.3%), Ataye District Hospital 5 (11.1%), Finote Selam Hospital 2 (4.4%), Metemma Hospital 3 (6.7%), Debre Birhan Hospital 2 (4.4$), Debre Tabor Hospital 1 (2.2%) and Debre Markos Hospital 1 (2.2%)36.
Sequence read quality check and de novo assembly
The quality assessment of WGS data was checked using FastQC v0.11.937 in aggregation with the MultiQC v1.24.138. Paired-end short reads were trimmed for quality using a flexible read trimming tool for Illumina NGS data Trimmomatic v.0.36 software (sliding-window trimming with a window size of 4 and a read quality threshold of 30), adapter, and other Illumina-specific sequences39. The reads were assembled using SPAdes v3.13.040 and the quality of genome assembly was assessed using QUAST v5.2.0 software41. WGS of Isolates with coverage less than 30X were considered poor coverage42.
Mapping and variant calling
Mapping was done using Burrows-Wheeler Aligner-Maximal Exact Match (BWA-MEM) algorithm using the Mycobacterium tuberculosis H37Rv reference genome. Sequence Alignment/Map tools (SAM tools)43 were used for sorting, indexing, removing putative PCR duplicates, and removing temporary files. Then, variant calling was performed using FreeBayes v0.9.2144. VCF tools v0.1.1645 were used to extract INDELs and SNPs from a VCF file generated from variant calling. The counting of SNPs and Indels from a VCF (Variant Call Format) file was performed using Python, pandas, and seaborn in a Jupyter notebook.
Spoligotyping
While WGS provides precise mutation data, spoligotyping enhances lineage-level insights, historical comparisons, and epidemiological tracking. In silico spoligotyping was conducted using the SpoTyping program version v2.046 with default parameters. The SITVIT2 server was then utilized, based on the identified spoligotypes, to determine the lineage14. Isolates exhibiting a similar pattern to those in the SITVIT database were assigned a Spoligo International Type (SIT) number. Isolates that did not match any SIT numbers were categorized as “Orphan” spoligotypes.
Lineage typing and antimicrobial resistance gene identification
To identify the MTB lineages, sub-lineages, and drug resistance mutations (SNPs, indels, and frameshifts), the isolated strains were analyzed using TB-Profiler v6.2.2. By default, it uses Trimmomatic to trim the reads, BWA to align to the reference genome, and GATK (open source v4) to call variants. This involved aligning raw paired-end illumine sequenced reads against the reference genome MTB H37Rv. To predict resistance the tool uses the curated tbdb database (Phelan et al., 2019). The resistance mutations predicted by TB-Profiler were further validated using mykrobe (v0.10.0)48, which provides a list of mutations in genes associated with antimicrobial resistance for each processed strain. Using Mykrobe to validate drug resistance (DR) mutations identified by TBProfiler is crucial due to its independent algorithmic approach, which relies on k-mer detection rather than traditional variant calling. This allows Mykrobe to detect mutations even in poorly mapped or complex genomic regions, potentially identifying low-frequency or novel resistance mutations that TBProfiler may miss. The inclusion of multiple tools for validation enhances the reliability of the results.
Identification of virulence factors
The tool ABRicate v1.0.1 was used to identify the virulence factor genes of MTB in the Virulence Factor Database (VFDB)49,50 of which the threshold for virulence-gene identification using the VFDB was set at a minimum of 80% coverage and identity. The network analysis of the identified virulent gene’s interaction was conducted using STRING v12.0. A High confidence level (0.700) was used as a minimum required interaction score. The network was downloaded in TSV format and visualized using Cytoscape v3.10.2. Markov Clustering (MCL) was used to identify clusters of functionally related proteins. PlasmidFinder-2.0 Server of the Center for Genomic Epidemiology was used to identify plasmids in the assembled sequence of M. tuberculosis.
Statistical analysis
Data were entered using Microsoft Excel, saved as CSV, and imported into R version 4.4.1 for analysis. Data completeness and consistency were assessed by checking for missing values and running frequency distributions for each variable. Descriptive statistics, including frequency and percentage calculations, were conducted as part of the analysis.
Results
Whole genome sequence data quality
A FASTQ file of M. tuberculosis WGS downloaded from the SAR database was subjected to FASTQC and MultiQc to check its quality. All isolates had mean phred quality > 30. The average read length varied between 227 and 292 bp (mean 282), while the median read length ranged from 127 to 150 bp (mean 149). The mean and median sequencing coverage were 91 and 88 times, respectively.
Out of the 45 sequences three had poor coverage (accession numbers SRR23497702, SRR23497703, and SRR23497959) and one isolate was not identified as M. tuberculosis (SRR23497700). These four isolates were excluded and WGS from 41 isolates were used for the downstream analysis. A total of 4291 SNP sites were identified. There were also 26 Indels (8 insertions and 18 deletions).
Lineage and sub-lineages of M. tuberculosis in Amhara Region
The most frequent lineage was L4 (58.53%), followed by L3 (34.15%), and L1 (4.88%) (Table 1). The most common sub-lineages identified were L4.2.2 (26.1%) followed by L4.2.2.2 (25%) and L4.2 (23.9%) (Fig. 1).
In silico spoligotyping
According to the spoligotyping findings, 90.24% (37 out of 41) of the isolates were classified into 12 shared types (SIT numbers), while the remaining four isolates (9.76%) were categorized as orphans. Among L3 strains, the most prevalent spoligotype was SIT 25 and SIT 21, while among L4 strains, SIT 149 was the dominant spoligotype (Table 2).
Moreover, the SITVIT analysis facilitated the identification of seven major genotypic families, with T3-ETH representing the predominant family at 48.78% (20 out of 41), followed by the CAS1-Delhi family comprising 19.51% (8 out of 41) and CAS1-Kili family, comprising 14.63% (6 out of 41) of the isolates. Interestingly, 9.76% (4 out of 41) of the strains corresponded to spoligotypes not previously documented in the SITVIT2 database (Table 2).
Genetic determinants of drug-resistant tuberculosis
Among 41 M. tuberculosis isolates, 38(92.7%; CI 76.9–97.3%) were MDR-TB, one (2.4%; CI 0.1–12.9%) were Pre-XDR-TB and two isolates (4.9%; CI 0.6–16.5%) were susceptible. The highest frequency of drug resistance was recorded in lineage 4 (23; 59%), followed by lineage 3 (14; 35.9%) and lineage 1 (2; 5.1%). Among the sub-lineages of lineage 4, lineage 4.2.2 has the highest frequency of drug resistance, with 20 isolates, making up 51.3% of the total, followed by lineage 4.1 (2; 5.1%) and lineage 4.2 (1; 2.6%).
From the 39 MDR and Pre-XDR-TB isolates, 37 (94.9%) were resistant to rifampicin. All the mutations have occurred at the rpoB gene, and the dominant mutation was at codon Ser450Leu (24; 64.9%). There were 37 isolates that showed isoniazid resistance of which, 34 (91.9%) exhibited mutations in the katG gene, two (5.4%) in the inhA gene, and one (2.7%) in both katG and inhA genes. Among isolates with mutations in the katG gene, 32 (94.12%) had the Ser315Thr mutation, while one had the Ser315Ile mutation. Additionally, one isolate exhibited both Ser315Thr and Ile317Val mutations at genome positions 2,155,168 and 2,155,163, respectively. All mutations in the inhA gene were observed at codon c.-777C > T. In one isolate, mutations were found in both katG gene codon Ser315Thr and inhA codon c.-154G > A alias fabG1p.Leu203Leu. Pyrazinamide resistance was shown in 22 (56.4%) isolates and all the mutations were at the pncA gene. Mutations showed at diverse codons with a higher frequency at codon c.-11A > G.
Resistance-conferring mutations in 31 (79.5%) ethambutol-resistant isolates occurred in the embB and embA genes. In 27 (87.1%) isolates, resistance mutations were identified at embB codons Gly406Ala (11; 40.7%), Met306Ile (6; 22.2%), Gly406Ser (2; 7.4%), Asp328Gly (1; 3.7%), Asp328Tyr (1; 3.7%), and Gln497Arg (1; 3.7%). Double mutations were detected at embB codons Met306Ile and Asp1024Asn (1; 3.7%), as well as Met306Val and Gln497Arg (1; 3.7%). In four (12.9%) isolates, mutations occurred in both the embA codons c.-16C > T (3; 75%) and c.-12C > T (1; 25%), as well as in the embB codon Met306Ile.
Mutations of streptomycin resistance were identified in 34 isolates. Mutations in the rpsL gene occurred in 10 isolates (29.4%) at codon Lys43Arg (8; 80%), Lys88Thr (1; 10%), and Lys88Gln (1; 10%). Mutations were also observed in both the rpsL and gid genes in 10 isolates (29.4%), involving codon Lys88Thr in rpsL and Gly69Asp in gid. Additionally, in 9 isolates (26.5%), resistance mutations were found at the gid codon Gly69Asp. Moreover, streptomycin resistance mutations were observed in the rrs gene in 2 isolates (5.9%) at codon n.888G > A, as well as in both the rrs and gid genes in 2 isolates (5.9%) at codons 799C > T and Gly69Asp, respectively. In 1 isolate (2.94%), mutations were identified in all three genes rpsL, rrs, and gid at codons Lys88Thr, 799C > T, and Gly69Asp, respectively.
Ethionamide resistance was observed in 27 isolates (69.2%), with mutations in the ethA gene found in 23 isolates (85.2%). Of these, a mutation at codon p.Met1? was identified in 17 isolates (73.9%), while the c.859_999del mutation appeared in 2 isolates (8.7%). Additionally, mutations in the inhA gene were present in 4 isolates (14.8%), with the c.-777C > T mutation detected in 3 isolates (75%) and the c.-154G > A mutation, also known as fabG1 Leu203Leu, found in one isolate. The drug resistance-conferring mutation for 2nd-line anti-TB drugs occurred at gyrA codon Ala90Val and Asp94Asn for fluoroquinolone (levofloxacin and moxifloxacin) resistance in one isolate and rrs gene codon 1401A > G, 1402C > A, 1484G > T, noncoding transcript exon variant, for 2nd-line injectable drugs amikacin, kanamycin, and capreomycin in four isolate for each. Additionally, a mutation at mmpR5 gene codon c.338_339insC and c.273_274insG, frameshift mutation, for bedaquiline and clofazimine resistance, respectively in one isolate (Table 3).
Varying levels of drug resistance across different sublineages were found. The most common form of drug resistance observed is MDR-TB, which is present in all sublineages, with the highest frequency found in sublineages L 4.2, L 4.2.2, and L 4.2.2.2, each exhibiting 21 cases. Pre-XDR TB was detected in sublineages L 3.1 and L 3.1.1, each showing one case. Mono-resistant TB (streptomycin resistance) was found in L 3, with one case (Table 4).
Virulence genes of Mycobacterium tuberculosis isolates
A total of 66 virulence genes were identified, with 63 virulent genes (95.45%) found in all isolates. The identified genes had a coverage range of 81.4% to 100% (mean 99.94%) and an identity range of 83.1% to 100% (mean 99.34%) when compared to the virulence factor database. The gene espB was present in 9 isolates, espK in 2 isolates, and eccA1 in 1 isolate (Table 5). Virulent genes esxN and esxM were found frequently in all isolates of M. tuberculosis with a total frequency of 126 and 125, respectively.
Network analysis of identified virulence genes of MTB isolates
In the network analysis, the identified virulent genes were recognized as nodes and there were 242 edges connecting the nodes. The average node degree was 7.68 and the average local clustering coefficient was 0.639. The network has significantly more interactions than expected, such an enrichment indicates that the proteins are at least partially biologically connected, as a group (Fig. 2). There were five isolated genes (relA, lipF, icl1, mgtC, and ideR) from the network.
Markov clustering of the network grouped the genes into seven distinct clusters based on functional classifications. Cluster 1 (Red), 22 genes involved in protein secretion by the type VII secretion system. Cluster 2 (Yellow), 13 genes related to biosynthesis of siderophore group nonribosomal peptides. Cluster 3 (Orange), 11 genes classified as mixed, including organelle lumen and zinc ion homeostasis. In cluster 4 (Green), 5 genes were classified as mixed, including EccD-like transmembrane domain and peptidase S8, subtilisin, and Asp-acti. Cluster 5 (Light green), 3 genes involved in catechol-containing compound metabolism and long-chain fatty acid metabolism. Cluster 6 (Light blue), 2 genes linked to the two-component regulatory system. In cluster 7 (Purple), 2 genes are classified as mixed, including the EspG family and PPE family (C-terminal) (Fig. 2). There was no plasmid found in all isolates.
Discussion
The current study highlights the genetic diversity, AMR, and virulence genes of MTB strains circulating in the Amhara region of Ethiopia.
Among 41 M. tuberculosis isolates 39 had drug resistance mutations. In most discordant cases, isolate pairs harbored variants that could cause low- or moderate-level resistance or were previously associated with variable minimum inhibitory concentrations (MICs)51. Additionally, resistance detection through WGS relies on known databases of resistance-conferring mutations, and some isolates may carry novel or rare variants that are not yet well characterized52. The simultaneous performance of phenotypic DST and WGS-based resistance profiling provides the most accurate assessment of drug resistance in M. tuberculosis isolates51.
Among the identified lineages, L4 (58.53%) was the most frequent followed by L3 (36.59%). This finding is consistent with previous studies conducted across various regions of Ethiopia, from Tigray region24 Northwest Ethiopia31 St.Peter’s TB Specialized Hospital, Ethiopia53, central, eastern, and southeastern Ethiopia32, and a nationwide review in Ethiopia54. Afro-TB dataset of MTB in Africa also showed L4 and L3 were the predominant lineages in Ethiopia55. Ethiopia’s location as a crossroads between Africa, the Middle East, and Europe has contributed to the introduction and spread of different MTB lineages. Historically, migration and trade routes likely facilitated the spread of L4 (originating from Europe) and L3 (originating from the Indian subcontinent) into Ethiopia56. Nevertheless, a lower proportion (40.1%) of lineage 4 has been documented in Northwest Ethiopia57. However, another study conducted among refugees residing in refugee camps in Ethiopia reported that lineage 3 (52, 77.6%) was the prevalent lineage. This could be due to that people in the refugee camps were from different countries (Eritrea, Somalia, Sudan, and South Sudan), which contributed to the dominance of this lineage in the camp58.
In terms of sub-lineages, L4.2.2 (26.1%) was the most frequent, followed by L4.2.2.2 (25.0%) and L4.2 (23.9%). Similarly, a study by31 reported that L4.2.2 was the predominant sub-lineage in Northwest Ethiopia. The high prevalence of sub-lineages within L4 was also reported by previous studies in different parts of Ethiopia34,59,60. However, these previous studies did not report L4.2.2 and L4.2.2.2 in high frequencies as observed in the present study.
Genotypes of lineage 3 of M. tuberculosis infection showed a predominance of the CAS1-Dehli family; this agrees with other studies57,61,62. The other dominant family was CAS1-Kili, which was also reported previously in Ethiopia24 and was not reported in a previous study in Ethiopia60. In lineage 4, T3-ETH was the most prevalent. Another study in Northwest Ethiopia also reported that L4-T3-ETH (32.0%), L3-CAS1-Delhi (22.7%), and L3-CAS1-Killi (14.8%) families were the most common63. The dominantly identified Spoligo SITs were SIT25, T3-ETH and SIT149. The finding is in agreement with previous reports64,65. The highest frequency of drug resistance was recorded in lineage 4 (59%), Among the sub-lineages, sub-lineage 4.2.2 had the highest frequency (51.3%) of drug resistance. This finding is comparable with previous reports in different parts of Ethiopia33,63. Prior studies in Ethiopia showed that these spoligotypes were more often linked with drug resistance-conferring mutations and clonal expansion in Ethiopia66. Perhaps, the connection of certain MTBC genotypes with MDR could be attributed to their genetics and enhanced intrinsic ability to acquire resistance to anti-TB drugs67.
Resistance of MTB strains to RIF is mainly due to canonical mutations in the hot-spot region of the rpoB gene (HSRrpoB). However, there are also disputed rpoB mutations that confer RIF-resistance, and their occurrence is not rare68. The association of these mutations with RIF resistance is endorsed by WHO in the updated catalog of mutations in MTBC and their association with DR20. Rifampicin works by binding to this subunit, inhibiting RNA synthesis and effectively killing Mycobacterium tuberculosis69. Specific mutations on the rpoB gene, alter the binding site of rifampicin on RNA polymerase, preventing the drug from binding effectively. This allows the bacteria to continue transcribing DNA into RNA, despite the presence of rifampicin70,71.
According to the WGS analysis, 91.9% of mutations that confer resistance to isoniazid occurred at the katG codon Ser315Thr, which is the most commonly reported mutation associated with high-level isoniazid resistance24,72,73. The KatG gene encodes the enzyme catalase-peroxidase and mutation in this gene decreases or blocks the enzyme activity. Mutation in the katG gene is the main mechanism of INH resistance in most strains74. The inhA mutations with or without a katG mutation were detected in the present study. There are similar reports in previous studies75,76.
In addition to rifampicin and isoniazid resistance, pyrazinamide resistance was detected in 56.4% of the isolates, with mutations in the pncA gene being responsible for the resistance. The diversity of mutations observed in the pncA gene highlights the complexity of pyrazinamide resistance mechanisms which has been reported in other studies24,33. The gene pncA is one such gene that encodes for pyrazinamidase (PZAse) and helps in the activation of pyrazinamide (PZA) to its active form pyrazinoic acid (POA). Thus, a mutation in this gene is linked to PZA resistance77.
Ethambutol (EMB) resistance was found in 79.5% of the isolates, with mutations in the embB and embA genes. The most common mutations of embB were at codons 406 and 306, which are considered as hotspot resistance codons78. There were also mutations at codons 328 and 497. Previous reports showed that codon 306 was shown to be directly involved in EMB binding while codons 406 and 497 were not directly involved78,79. Nevertheless, mutations at codon 497 cause conformational changes that affect codon 327, one of the EMB binding sites. Codon 406 mutations may also affect drug binding by causing protein conformation changes78. Mutations at the embA gene also confer resistance to ethambutol. This result aligns with the principle that resistance to EMB is caused by mutation of the embCAB operon (embC, embA, and embB) that encodes membrane-associated arabinosyltransferases involved in the synthesis of cell wall arabinogalactan80.
In our study, mutations that confer SM resistance were observed at the rpsL, gid, and rrs genes. Previous studies have indicated that most SM-resistance Mycobacterium tuberculosis isolates can be determined by mutations in rpsL, rrs, and gid27. The most prevalent mutation associated with SM-resistance was at the rpsL gene codon Lys43Arg. Although the proportion of mutations that confer resistance to SM at Lys43Arg varied geographically, this finding is concordant with an earlier study that reported its dominance across the world and its association with a high drug resistance level81. Similar findings were also reported by24. The frequently detected mutation at K43R highlighted its importance as a surrogate marker for rapid detection of SM-resistance. The present findings indicated that 29.4% of isolates showed co-existing mutations at both genes, rpsL codon Lys88Thr in rpsL and gid codon Gly69Asp.
Ethionamide resistance-conferring mutations occurred at ethA gene codon M1 (85.2%), inhA and inhA alias with fabG1 gene (14.8%). The present finding revealed that all isolates resistant to ETH were co-resistant to INH. This finding is supported by prior reports that showed the isolation of Mtb strains co-resistant to INH and ETH from TB patients previously treated with INH but never treated with ETH24,82. The fabG1 gene codon C-154G > A mutation, which conferred resistance to both INH and ETH, was detected from all MDR-TB. This is consistent with other study reports24,83.
The study also detected resistance-conferring mutations for second-line anti-TB drugs, including fluoroquinolones (levofloxacin and moxifloxacin) and injectable drugs (amikacin, kanamycin, and capreomycin). According to the WHO Consolidated Guidelines (2022), second-line injectable drugs are no longer recommended in the treatment regimen for drug-resistant TB (DR-TB) due to their lower efficacy and higher toxicity84. In one isolate mutations in the gyrA gene codons Ala90Val and Asp94Asn conferring fluoroquinolone resistance were detected. This finding is supported by previous reports85. However, the mutation sites and frequencies of gyrA varied across studies86,87,88,89, which may be attributed to the difference in detection techniques, breakpoint concentrations, epidemic strains, research population, and medical history. Four isolates had mutations in the rrs gene at codons 1401A > G, 1402C > A, and 1484G > T, which are associated with resistance to aminoglycosides and capreomycin. This finding agrees with previous reports90,91. These findings underscore the potential emergence of XDR-TB in the region, which could complicate treatment options and outcomes.
The detection of mutations in the mmpR5 gene (Rv0678) has been associated with resistance to bedaquiline and clofazimine, two essential drugs in the treatment of rifampicin-resistant tuberculosis (TB). Previous studies (Nimmo et al., 2024) highlighted similar findings, suggesting that baseline resistance due to mmpR5 mutations increases the risk of treatment failure in an all-oral 6-month regimen that includes bedaquiline, linezolid, and moxifloxacin (Timm et al., 2023). Importantly, some mmpR5 mutations, like Met146Thr, emerged before the introduction of bedaquiline, conferring cross-resistance to both bedaquiline and clofazimine. This mutation is now recognized in the WHO’s updated catalogue of resistance mutations (Beckert et al., 2020; WHO, 2023). The emergence of such resistance poses a challenge for the treatment of multidrug-resistant (MDR) and extensively drug-resistant (XDR) TB, threatening the efficacy of future regimens. It is worth noting that mutations in the ethA, fabG1, and mmpR5 genes cannot be detected by LPA.
Several virulent genes associated with the MTB strain were identified. The ecc genes (eccA, eccB, eccC, eccCa, eccCb, eccD, eccE) are part of the ESX (Type VII secretion) systems that play an important role in the secretion of virulence factors, which are crucial for the pathogenicity of M. tuberculosis92.
The identified esp and esx genes (espA, espB, espC, espD, espG3, espK, esxA, esxB, esxG, esxH, esxN, and esxM) are part of the Type VII Secretion Systems (T7SS) in Mycobacterium tuberculosis and are essential for its virulence. These genes encode proteins that help secrete virulence factors, allowing the bacterium to survive and evade the host immune system. Their identification highlights the high virulence potential of these genes, given their crucial roles in intracellular survival, immune evasion, and nutrient acquisition93. About 20.7% of L4 and 33.3% of L3 have only espB gene. The ESX-1 system, with key components such as EsxA and EsxB, allows M. tuberculosis to escape from the phagosome and survive inside macrophages. Meanwhile, the ESX-3 system, supported by EsxN and EsxM, enables the bacterium to acquire essential metals (such as iron and zinc) from the host, which is crucial for its growth and persistence. Furthermore, the secretion of proteins like EspA, EspB, and EspC helps M. tuberculosis evade the host immune response, creating a more favorable environment for its replication93,94,95.
The genes fbpA, fbpB, and fbpC in Mycobacterium tuberculosis encode proteins that are part of the antigen 85 complex (Ag85), which plays a crucial role in the virulence and pathogenesis of M. tuberculosis. This complex is composed of three main proteins Ag85A, Ag85B, and Ag85C encoded by fbpA, fbpB, and fbpC, respectively. These proteins are involved in key biological processes that contribute to the bacterium’s ability to establish infection and persist within the host96.
The mbtA, mbtB, mbtC, mbtD, mbtE, mbtF, mbtG, mbtH, mbtI, mbtJ, mbtK, mbtL, mbtM, and mbtN genes in Mycobacterium tuberculosis comprise the mycobactin biosynthetic gene cluster. Their primary function is to facilitate the synthesis of mycobactins, which are siderophore-like molecules crucial for sequestering iron from the host environment97. Iron is vital for bacterial growth and metabolism, particularly during infection. The ability to acquire iron is directly linked to the virulence of M. tuberculosis, and the mbt gene cluster is essential for establishing infection and persistence in the host98. Understanding the mechanisms of mycobactin biosynthesis and regulation can inform vaccine development and novel treatment strategies against tuberculosis.
There was no plasmid in all isolates and this finding is supported by lacking previous evidence of the presence of plasmids in M. tuberculosis99. These finding is a good insight that M. tuberculosis does not rely on plasmids for survival or virulence, but its genome is rich in virulence factors and complex regulatory systems encoded on its chromosome.
Limitations
While this study provides valuable insights into the genetic diversity, antimicrobial resistance (AMR) patterns, and virulence factors of M. tuberculosis strains in the Amhara region, it has limitations. The reliance on whole-genome sequencing (WGS) without phenotypic drug susceptibility testing (DST) limits the validation of resistance profiles. Additionally, the study lacked clinical and epidemiological data, which would have provided a more comprehensive understanding of the factors driving drug-resistant TB spread. Resource constraints also limited the depth of genomic analysis.
Conclusion and recommendations
This study demonstrated that TB in the Amhara region is caused by a wide diversity of MTB strains belonging to lineages L1, L3, and L4 with a predominance of L4-T3-ETH, L3-CAS1-Delhi and L3-CAS1-Killi families. Overall, L4 was the most frequently observed MTB genotype and was associated with the highest proportion of drug resistance. The study also highlighted the usefulness of mutations at rpoB, katG, embB, rpsL, pncA, ethA, gyrA, and rrs genes as molecular markers for the rapid detection of resistance for RIF, INH, EMB, SM, PZA, ETH, FLQs, and SLIDs, respectively. This study also provided valuable insights into the virulence genes of Mycobacterium tuberculosis isolates from the Amhara region of Ethiopia. Notably, the virulent genes esxN and esxM were frequently present in all isolates. The high prevalence of MDR-TB and the detection of resistance to second-line drugs, including bedaquiline and clofazimine, underscore the urgent need for advanced molecular diagnostic tools. Establishing a robust surveillance system to monitor the spread of resistant strains and their genetic evolution can inform public health interventions and prevent the further spread of resistant TB. Tailoring treatment based on the genetic profile of the infecting strain can improve treatment outcomes and reduce the risk of further resistance development.
Data availability
All data generated or analyzed during this study are available upon the request of the corresponding author.
Abbreviations
- AMR:
-
Antimicrobial resistance
- MTB:
-
Mycobacterium tuberculosis
- MDR-TB:
-
Multidrug-resistant tuberculosis
- FLQ:
-
Fluoroquinolones
- SLID:
-
Second line injectable drugs
- WGS:
-
Whole genome sequencing
References
Lin, S. Y. G. & Desmond, E. P. Molecular diagnosis of tuberculosis and drug resistance. Clin. Lab. Med. 34(2), 297–314 (2014).
Alsayed, S. S. R. & Gunosewoyo, H. Tuberculosis: pathogenesis, current treatment regimens and new drug targets. Int. J. Mol. Sci. 24(6), 5202 (2023).
WHO. Global tuberculosis report 2024. Geneva: World Health Organization; 2024. Licence: CC BY-NC-SA 3.0 IGO [Internet]. https://www.who.int/teams/global-tuberculosis-programme/tb-reports/global-tuberculosis-report-2024 (2024).
WHO Africa. Country Disease Outlook Ethiopia, 55–58. https://www.afro.who.int/sites/default/files/2023-08/Ethiopia.pdf (2023).
US Agency for International Development. Ethiopia tuberculosis roadmap overview, Fiscal Year 2023. https://www.usaid.gov/sites/default/files/2024-02/Ethiopia_TB_roadmap_narrative_22_508.pdf (2023).
Onyedum, C. C., Alobu, I. & Ukwaja, K. N. Prevalence of drug-resistant tuberculosis in Nigeria: A systematic review and meta-analysis. PLoS ONE 12(7), e0180996 (2017).
Saravanan, M. et al. Review on emergence of drug-resistant tuberculosis (MDR & XDR-TB) and its molecular diagnosis in Ethiopia. Microb. Pathog. 117, 237–242 (2018).
Alelign, A. et al. Molecular detection of Mycobacterium tuberculosis sensitivity to rifampicin and isoniazid in South Gondar Zone, northwest Ethiopia. BMC Infect. Dis. 19(1), 343 (2019).
Tessema, B. et al. Molecular epidemiology and transmission dynamics of Mycobacterium tuberculosis in Northwest Ethiopia: new phylogenetic lineages found in Northwest Ethiopia. BMC Infect. Dis. 13(1), 131 (2013).
Mesfin, Y. M., Hailemariam, D., Biadglign, S. & Kibret, K. T. Association between HIV/AIDS and multi-drug resistance tuberculosis: a systematic review and meta-analysis. PLoS ONE 9(1), e82235 (2014).
Bedewi, Z. et al. Mycobacterium tuberculosis in central Ethiopia: drug sensitivity patterns and association with genotype. New Microbes New Infect. 17, 69–74 (2017).
Abraham, A. O. Mechanism of drug resistance in Mycobacterium Tuberculosis. Am. J. Biomed. Sci. Res. 7(5), 378–383 (2020).
Coscolla, M. et al. Phylogenomics of Mycobacterium africanum reveals a new lineage and a complex evolutionary history. Microb. Genom. https://doi.org/10.1099/mgen.0.000477 (2021).
Couvin, D., Segretier, W., Stattner, E. & Rastogi, N. Novel methods included in SpolLineages tool for fast and precise prediction of Mycobacterium tuberculosis complex spoligotype families. Database 2020, baaa108 (2020).
Ngabonziza, J. C. S. et al. A sister lineage of the Mycobacterium tuberculosis complex discovered in the African Great Lakes region. Nat. Commun. 11(1), 2917 (2020).
Stucki, D. et al. Mycobacterium tuberculosis lineage 4 comprises globally distributed and geographically restricted sublineages. Nat. Genet. 48(12), 1535–1543 (2016).
Jhun, B. W. & Koh, W. J. Treatment of isoniazid-resistant pulmonary tuberculosis. Tuberc. Respir. Dis. 83(1), 20 (2020).
Centers for Disease Control and Prevention. Clinical overview of drug-resistant tuberculosis disease. https://www.cdc.gov/tb/hcp/clinical-overview/drug-resistant-tuberculosis-disease.html (2024).
WHO. Meeting report of the WHO expert consultation on the definition of extensively drug-resistant tuberculosis. [cited 2025 Mar 3]. https://www.who.int/publications/i/item/9789240018662.
Catalogue of Mutations in Mycobacterium Tuberculosis Complex and Their Association with Drug Resistance, 2nd ed. (World Health Organization, 2023).
Muthaiah, M. et al. Prevalence of mutations in genes associated with rifampicin and isoniazid resistance in Mycobacterium tuberculosis clinical isolates. J. Clin. Tuberc. Mycobact. Dis. 8, 19–25 (2017).
Cao, B. et al. Genetic characterization conferred co-resistance to isoniazid and ethionamide in Mycobacterium tuberculosis isolates from Southern Xinjiang, China. Infect. Drug Resist. 16, 3117–3135 (2023).
Bakuła, Z. et al. Mutations in the embB gene and their association with ethambutol resistance in multidrug-resistant Mycobacterium tuberculosis clinical isolates from Poland. BioMed. Res. Int. 2013, 1–5 (2013).
Welekidan, L. N. et al. Whole genome sequencing of drug resistant and drug susceptible Mycobacterium tuberculosis isolates from Tigray Region, Ethiopia. Front. Microbiol. 6(12), 743198 (2021).
Bwalya, P. et al. Characterization of embB mutations involved in ethambutol resistance in multi-drug resistant Mycobacterium tuberculosis isolates in Zambia. Tuberculosis 133, 102184 (2022).
Kim, N. Y., Kim, D. Y., Chu, J. & Jung, S. H. pncA large deletion is the characteristic of pyrazinamide-resistant Mycobacterium tuberculosis belonging to the East Asian Lineage. Infect. Chemother. 55(2), 247 (2023).
Wang, Y. et al. The roles of rpsL, rrs, and gidB mutations in predicting streptomycin-resistant drugs used on clinical Mycobacterium tuberculosis isolates from Hebei Province, China. Int. J. Clin. Exp. Pathol. 12(7), 2713–2721 (2019).
Farhat, M. R. et al. Gyrase mutations are associated with variable levels of fluoroquinolone resistance in Mycobacterium tuberculosis. J. Clin. Microbiol. 54(3), 727–733 (2016).
Rahlwes, K. C., Dias, B. R. S., Campos, P. C., Alvarez-Arguedas, S. & Shiloh, M. U. Pathogenicity and virulence of Mycobacterium tuberculosis. Virulence. 14(1), 2150449 (2023).
Ramon-Luing, L., Palacios, Y., Ruiz, A., Téllez-Navarrete, N. & Chavez-Galan, L. Virulence factors of Mycobacterium tuberculosis as modulators of cell death mechanisms. Pathogens. 12(6), 839 (2023).
Mekonnen, D. et al. Comparative whole-genome sequence analysis of Mycobacterium tuberculosis isolated from pulmonary tuberculosis and tuberculous lymphadenitis patients in Northwest Ethiopia. Front. Microbiol. 30(14), 1211267 (2023).
Agonafir, M. et al. Genetic diversity of Mycobacterium tuberculosis isolates from the central, eastern and southeastern Ethiopia. Heliyon. 9(12), e22898 (2023).
Mekonnen, D. et al. Mycobacterium tuberculosis sub-lineage 4.2.2/SIT149 as dominant drug-resistant clade in Northwest Ethiopia 2020–2022: In-silico whole-genome sequence analysis. Infect. Drug Resist. 16, 6859–6870 (2023).
Chekesa, B. et al. Whole-genome sequencing-based genetic diversity, transmission dynamics, and drug-resistant mutations in Mycobacterium tuberculosis isolated from extrapulmonary tuberculosis patients in western Ethiopia. Front. Public Health. 9(12), 1399731 (2024).
Bi, K. et al. The past, present and future of tuberculosis treatment. J. Zhejiang Univ. Med. Sci. 51(6), 657–668 (2022).
Shibabaw, A. et al. Molecular epidemiology and transmission dynamics of multi-drug resistant tuberculosis strains using whole genome sequencing in the Amhara region, Ethiopia. BMC Genom. 24(1), 400 (2023).
Babraham Bioinformatics. FastQC: A quality control tool for high throughput sequence data (Version 0.12.0). https://www.bioinformatics.babraham.ac.uk/projects/fastqc/. 2023. https://www.bioinformatics.babraham.ac.uk/projects/fastqc/
Ewels, P., Magnusson, M., Lundin, S. & Käller, M. MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinformatics 32(19), 3047–3048 (2016).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30(15), 2114–2120 (2014).
Prjibelski, A., Antipov, D., Meleshko, D., Lapidus, A. & Korobeynikov, A. Using SPAdes De Novo Assembler. Curr. Protoc. Bioinform. 70(1), e102 (2020).
Mikheenko, A., Prjibelski, A., Saveliev, V., Antipov, D. & Gurevich, A. Versatile genome assembly evaluation with QUAST-LG. Bioinformatics 34(13), i142–i150 (2018).
Bogaerts, B. et al. Evaluation of WGS performance for bacterial pathogen characterization with the Illumina technology optimized for time-critical situations. Microb. Genom. https://doi.org/10.1099/mgen.0.000699 (2021).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinform. Oxf. Engl. 25(16), 2078–2079 (2009).
Garrison, E., Marth, G. Haplotype-based variant detection from short-read sequencing. arXiv; 2012 [cited 2024 Sep 25]. http://arxiv.org/abs/1207.3907.
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27(15), 2156–2158 (2011).
Xia, E., Teo, Y. Y. & Ong, R. T. H. SpoTyping: fast and accurate in silico Mycobacterium spoligotyping from sequence reads. Genome Med. 8(1), 19 (2016).
Phelan, J. E. et al. Integrating informatics tools and portable sequencing technology for rapid detection of resistance to anti-tuberculous drugs. Genome Med. 11, 41 (2019).
Hunt, M. et al. Antibiotic resistance prediction for Mycobacterium tuberculosis from genome sequence data with Mykrobe. Wellcome Open Res. 2(4), 191 (2019).
Chen, L., Zheng, D., Liu, B., Yang, J. & Jin, Q. VFDB 2016: hierarchical and refined dataset for big data analysis—10 years on. Nucleic Acids Res. 44(D1), D694–D697 (2016).
Seemann, T. ABRicate: mass screening of contigs for antiobiotic resistance genes. (2020). https://github.com/tseemann/abricate.
Sadovska, D. et al. Discordance between phenotypic and WGS-based drug susceptibility testing results for some anti-tuberculosis drugs: A snapshot study of paired Mycobacterium tuberculosis isolates with small genetic distance. Infect. Drug Resist. 17, 3289–3307 (2024).
García-Marín, A. M. et al. Role of the first WHO mutation catalogue in the diagnosis of antibiotic resistance in Mycobacterium tuberculosis in the Valencia Region, Spain: a retrospective genomic analysis. Lancet Microbe. 5(1), e43-51 (2024).
Damena, D. et al. Genetic diversity and drug susceptibility profiles of Mycobacterium tuberculosis obtained from Saint Peter’s TB specialized Hospital, Ethiopia. PLoS ONE 14(6), e0218545 (2019).
Mekonnen, D. et al. Molecular epidemiology of M. tuberculosis in Ethiopia: A systematic review and meta-analysis. Tuberculosis 118, 101858 (2019).
Laamarti, M., El Fathi, L. Y., Elfermi, R., Daoud, R. & El Allali, A. Afro-TB dataset as a large scale genomic data of Mycobacterium tuberuclosis in Africa. Sci. Data. 10(1), 212 (2023).
Reta, M. A. et al. Genetic diversity of Mycobacterium tuberculosis strains isolated from spiritual holy water site attendees in Northwest Ethiopia. A cross-sectional study. New Microbes New Infect. 59, 101235 (2024).
Ejo, M. et al. Genetic diversity of the Mycobacterium tuberculosis complex strains from newly diagnosed tuberculosis patients in Northwest Ethiopia reveals a predominance of East-African-Indian and Euro-American lineages. Int. J. Infect. Dis. 103, 72–80 (2021).
Meaza, A. et al. Genomic transmission clusters and circulating lineages of Mycobacterium tuberculosis among refugees residing in refugee camps in Ethiopia. Infect. Genet. Evol. 116, 105530 (2023).
Worku, G. et al. Molecular epidemiology of tuberculosis in the Somali region, eastern Ethiopia. Front. Med. 14(9), 960590 (2022).
Wondale, B. et al. Molecular epidemiology of clinical Mycobacterium tuberculosis complex isolates in South Omo, Southern Ethiopia. BMC Infect. Dis. 20(1), 750 (2020).
Srilohasin, P. et al. Genetic diversity and dynamic distribution of Mycobacterium tuberculosis isolates causing pulmonary and extrapulmonary tuberculosis in Thailand. J. Clin. Microbiol. 52(12), 4267–4274 (2014).
Hadifar, S. et al. Variation in Mycobacterium tuberculosis population structure in Iran: a systemic review and meta-analysis. BMC Infect. Dis. 21(1), 2 (2021).
Ejo, M. et al. Strain diversity and gene mutations associated with presumptive multidrug-resistant Mycobacterium tuberculosis complex isolates in Northwest Ethiopia. J. Glob. Antimicrob. Resist. 32, 167–175 (2023).
Ayalew, S. et al. Drug resistance conferring mutation and genetic diversity of Mycobacterium tuberculosis isolates in tuberculosis lymphadenitis patients; Ethiopia. Infect. Drug Resist. 14, 575–584 (2021).
Diriba, G. et al. Mycobacterial lineages associated with drug resistance in patients with extrapulmonary tuberculosis in Addis Ababa, Ethiopia. Tuberc. Res. Treat. 2021, 5239529 (2021).
Diriba, B., Berkessa, T., Mamo, G., Tedla, Y. & Ameni, G. Spoligotyping of multidrug-resistant Mycobacterium tuberculosis isolates in Ethiopia. Int. J. Tuberc. Lung Dis. 17(2), 246–250 (2013).
Chihota, V. N. et al. Population structure of multi- and extensively drug-resistant Mycobacterium tuberculosis strains in South Africa. J. Clin. Microbiol. 50(3), 995–1002 (2012).
Shea, J. et al. Low-level rifampin resistance and rpoB mutations in Mycobacterium tuberculosis: an analysis of whole-genome sequencing and drug susceptibility test data in New York. J. Clin. Microbiol. 59(4), e01885-e1920 (2021).
Palomino, J. & Martin, A. Drug resistance mechanisms in Mycobacterium tuberculosis. Antibiotics. 3(3), 317–340 (2014).
Molodtsov, V., Scharf, N. T., Stefan, M. A., Garcia, G. A. & Murakami, K. S. Structural basis for rifamycin resistance of bacterial RNA polymerase by the three most clinically important RpoB mutations found in Mycobacterium tuberculosis. Mol. Microbiol. 103(6), 1034–1045 (2017).
Patel, Y., Soni, V., Rhee, K. Y. & Helmann, J. D. Mutations in rpoB that confer rifampicin resistance can alter levels of peptidoglycan precursors and affect β-lactam susceptibility. MBio 14(2), e0316822 (2023).
Kigozi, E. et al. Prevalence and patterns of rifampicin and isoniazid resistance conferring mutations in Mycobacterium tuberculosis isolates from Uganda. PLoS ONE 13(5), e0198091 (2018).
Zhang, M. et al. Detection of mutations associated with isoniazid resistance in Mycobacterium tuberculosis isolates from China. J. Clin. Microbiol. 43(11), 5477–5482 (2005).
Moaddab, S. R., Farajnia, S., Kardan, D., Zamanlou, S. & Alikhani, M. Y. Isoniazid MIC and KatG gene mutations among Mycobacterium tuberculosis Isolates in Northwest of Iran. Iran. J. Basic Med. Sci. 14(6), 540–545 (2011).
Niehaus, A. J., Mlisana, K., Gandhi, N. R., Mathema, B. & Brust, J. C. M. High prevalence of inhA promoter mutations among patients with drug-resistant tuberculosis in KwaZulu-Natal, South Africa. PLoS ONE 10(9), e0135003 (2015).
Reta, M. A., Alemnew, B., Abate, B. B. & Fourie, P. B. Prevalence of drug resistance-conferring mutations associated with isoniazid- and rifampicin-resistant Mycobacterium tuberculosis in Ethiopia: a systematic review and meta-analysis. J. Glob. Antimicrob. Resist. 26, 207–218 (2021).
Khan, M. T. et al. Pyrazinamide resistance and mutations in pncA among isolates of Mycobacterium tuberculosis from Khyber Pakhtunkhwa, Pakistan. BMC Infect. Dis. 19(1), 116 (2019).
Zhang, L. et al. Structures of cell wall arabinosyltransferases with the anti-tuberculosis drug ethambutol. Science 368(6496), 1211–1219 (2020).
Xu, Y., Jia, H., Huang, H., Sun, Z. & Zhang, Z. Mutations found in embCAB, embR, and ubiA genes of ethambutol-sensitive and -resistant Mycobacterium tuberculosis clinical isolates from China. BioMed. Res. Int. 2015, 1–8 (2015).
Cui, Z. et al. Mutations in the embC-embA intergenic region contribute to Mycobacterium tuberculosis resistance to ethambutol. Antimicrob Agents Chemother. 58(11), 6837–6843 (2014).
Nhu, N. T. Q. et al. Association of streptomycin resistance mutations with level of drug resistance and Mycobacterium tuberculosis genotypes. Int. J. Tuberc. Lung Dis. 16(4), 527–531 (2012).
Vilchèze, C. & Jacobs, W. R. Resistance to isoniazid and ethionamide in Mycobacterium tuberculosis: Genes, mutations, and causalities. Microbiol. Spectr. 2(4), 2401 (2014).
Walker, T. M. et al. A cluster of multidrug-resistant Mycobacterium tuberculosis among patients arriving in Europe from the Horn of Africa: a molecular epidemiological study. Lancet Infect. Dis. 18(4), 431–440 (2018).
WHO. WHO consolidated guidelines on drug-resistant tuberculosis treatment (2022). https://iris.who.int/bitstream/handle/10665/311389/9789241550529-eng.pdf.
Teng, C. et al. Evaluation of genetic correlation with fluoroquinolones resistance in rifampicin-resistant Mycobacterium tuberculosis isolates. Heliyon. 10(11), e31959 (2024).
Aitkenhead, A. R. et al. International standards for safety in the intensive care unit Developed by the international task force on safety in the intensive care unit. Intensive Care Med. 19(3), 178–181 (1993).
An, Q., Lin, R., Yang, Q., Wang, C. & Wang, D. Evaluation of genetic mutations associated with phenotypic resistance to fluoroquinolones, bedaquiline, and linezolid in clinical Mycobacterium tuberculosis: A systematic review and meta-analysis. J. Glob. Antimicrob. Resist. 34, 214–226 (2023).
Hu, Y. et al. Genotyping and molecular characterization of fluoroquinolone’s resistance among multidrug-resistant Mycobacterium tuberculosis in Southwest of China. Microb. Drug Resist. Larchmt N. 27(7), 865–870 (2021).
Shago, M. et al. Modulation of the retinoic acid and retinoid X receptor signaling pathways in P19 embryonal carcinoma cells by calreticulin. Exp. Cell Res. 230(1), 50–60 (1997).
Sirgel, F. A. et al. Mutations in the rrs A1401G gene and phenotypic resistance to amikacin and capreomycin in Mycobacterium tuberculosis. Microb. Drug Resist. Larchmt N. 18(2), 193–197 (2012).
Via, L. E. et al. Polymorphisms associated with resistance and cross-resistance to aminoglycosides and capreomycin in Mycobacterium tuberculosis isolates from South Korean Patients with drug-resistant tuberculosis. J. Clin. Microbiol. 48(2), 402–411 (2010).
Wagner, J. M. et al. Structures of EccB1 and EccD1 from the core complex of the mycobacterial ESX-1 type VII secretion system. BMC Struct. Biol. 16(1), 5 (2016).
Roy, S., Ghatak, D., Das, P. & BoseDasgupta, S. ESX secretion system: The gatekeepers of mycobacterial survivability and pathogenesis. Eur. J. Microbiol. Immunol. 10(4), 202–209 (2020).
Guo, Q., Bi, J., Wang, H. & Zhang, X. Mycobacterium tuberculosis ESX-1-secreted substrate protein EspC promotes mycobacterial survival through endoplasmic reticulum stress-mediated apoptosis. Emerg. Microbes Infect. 10(1), 19–36 (2021).
Serafini, A., Pisu, D., Palù, G., Rodriguez, G. M. & Manganelli, R. The ESX-3 secretion system is necessary for iron and zinc homeostasis in Mycobacterium tuberculosis. PLoS ONE 8(10), e78351 (2013).
Xu, Z. et al. Generation of monoclonal antibodies against Ag85A antigen of Mycobacterium tuberculosis and application in a competitive ELISA for serodiagnosis of bovine tuberculosis. Front. Vet. Sci. 30(4), 107 (2017).
Sritharan, M. Iron homeostasis in Mycobacterium tuberculosis: Mechanistic insights into siderophore-mediated iron uptake. J. Bacteriol. 198(18), 2399–2409 (2016).
Rodriguez, G. M., Sharma, N., Biswas, A. & Sharma, N. The iron response of Mycobacterium tuberculosis and its implications for tuberculosis pathogenesis and novel therapeutics. Front. Cell Infect. Microbiol. 11(12), 876667 (2022).
Zainuddin, Z. F. & Dale, J. W. Does Mycobacterium tuberculosis have plasmids?. Tubercle 71(1), 43–49 (1990).
Funding
No funding was received for this research.
Author information
Authors and Affiliations
Contributions
Conceptualization and methodology: A.T.G.; Data acquisition and formal analysis: A.T.G.; Software, supervision and validation: A.T.G. and B.K.; Writing-original draft preparation: A.T.G.; Investigation, visualization, and writing-review and editing: A.T.G., T.E., M.Z.K., B.K. All authors approved the final draft of the manuscript prior to submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics statement
Not applicable as there is not any animal or human subject involved directly in the study. The study utilized publicly available raw WGS data. Demographic and clinical data of patients were accessed from an open-source database.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gessese, A.T., Kinde, M.Z., Eshetu, T. et al. Whole-genome sequencing analysis to identify antimicrobial resistance regions and virulence factors in Mycobacterium tuberculosis isolates from the Amhara Region, Ethiopia. Sci Rep 15, 16076 (2025). https://doi.org/10.1038/s41598-025-01241-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-01241-6