Abstract
Skin cancer, particularly melanoma, remains one of the most life-threatening forms of cancer worldwide, with early detection being critical for improving patient outcomes. Traditional diagnostic methods, such as dermoscopy and histopathology, are often limited by subjectivity, interobserver variability, and resource constraints. To address these challenges, this study proposes a dual-stream deep learning framework that combines histopathological-inherited and vision-based feature extraction for accurate and efficient skin lesion diagnosis. The framework uses the U-Net architecture for precise lesion segmentation, followed by a dual-stream approach: the first stream employs Virchow2, a pretrained model, to extract high-level histopathological embeddings, whereas the second stream uses Nomic, a vision-based model, to capture spatial and contextual information. The extracted features are fused and integrated to create a comprehensive representation of the lesion, which is then classified via a multilayer perceptron (MLP). The proposed approach is evaluated on the HAM10000 dataset, achieving a mean accuracy of 96.25% and a mean F1 score of 93.79% across 10 trials. Ablation studies demonstrate the importance of both feature streams, with the removal of either stream resulting in significant performance degradation. Comparative analysis with existing studies highlights the superiority of the proposed framework, which outperforms traditional single-modality approaches. The results underscore the potential of the dual-stream framework to enhance skin cancer diagnosis, offering a robust, interpretable, and scalable solution for clinical applications.
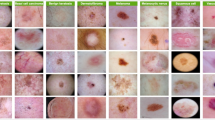
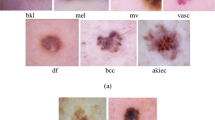
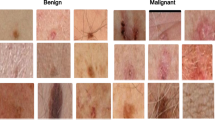
Similar content being viewed by others
Introduction
Skin cancer is one of the most prevalent and life-threatening types of cancer worldwide, with melanoma representing its deadliest form1. Melanoma accounts for the majority of skin cancer-related deaths, despite constituting a smaller proportion of skin cancer cases than nonmelanoma types such as basal cell carcinoma and squamous cell carcinoma2,3.
According to the World Health Organization (WHO), approximately 132,000 new cases of melanoma are diagnosed annually worldwide4,5, and this number continues to rise due to factors such as increased ultraviolet (UV) radiation exposure, aging populations, and improved diagnostic capabilities6. The aggressive nature of melanoma, coupled with its ability to metastasize rapidly, underscores the importance of early detection7.
When diagnosed at an early stage, the five-year survival rate for patients with melanoma exceeds 99%, but this rate decreases to less than 30% for patients with advanced-stage disease. These statistics highlight the critical need for accurate and timely diagnostic tools to improve patient outcomes and reduce mortality rates8,9.
Traditional diagnostic methods for skin cancer rely on dermatologists using tools like dermoscopy and visual inspection10. Dermoscopy, which magnifies and illuminates the skin, improves accuracy but depends heavily on clinician expertise11,12. Histopathological examination remains the “gold standard”, providing definitive diagnostic details through biopsy and microscopic analysis13. However, it is invasive, time-consuming, and inaccessible in resource-limited settings14,15.
Challenges with traditional methods include interobserver variability, leading to misdiagnoses, and subjective reliance on visual interpretation16,17,18. Histopathology can also be error-prone with inadequate samples. These limitations, compounded by a shortage of specialists, highlight the need for objective, scalable, and efficient diagnostic tools19,20.
Whole slide imaging (WSI) has transformed pathology by digitizing glass slides, enabling applications in diagnostics, education, and research21. WSI facilitates telepathology and algorithm-based decision support but faces adoption challenges, including technical issues and diagnostic difficulties22,23. AI integration has expanded WSI’s potential, particularly in cancer diagnosis, where AI algorithms improve accuracy and reduce turnaround times24,25. AI also enhances radiology, pharmacology, and personalized medicine, identifying patterns imperceptible to humans24,25.
Despite its promise, skepticism about AI persists due to interpretability concerns, ethical considerations, and unclear reimbursement opportunities23. Effective communication is needed to build trust and demonstrate AI’s benefits21.
AI, particularly deep learning (DL), offers solutions for skin cancer diagnosis by analyzing dermatoscopic and histopathological images26. Using datasets like HAM10000, AI models identify lesion patterns and assist clinicians, improving efficiency and accessibility in underserved areas27,28,29. This integration represents a paradigm shift, enhancing outcomes30,31. DL automates feature extraction, with CNNs achieving state-of-the-art performance in lesion segmentation and classification32,33,34,35. For example, U-Net captures fine-grained lesion details critical for accurate classification36,37. Challenges remain, such as overfitting, driven by limited, diverse datasets38,39.
To address these challenges, researchers have begun exploring hybrid approaches that combine multiple DL techniques40. For example, pretrained models such as Virchow2, which utilize histopathological data, can extract high-level embeddings that capture intricate patterns in skin lesions. Similarly, vision-based approaches such as Nomic can complement these embeddings by providing additional contextual information.
Therefore, the current study aims to address the challenges associated with skin lesion diagnosis by using advanced DL techniques and a dual-stream methodology to improve skin cancer detection accuracy, efficiency, and interpretability. Using the HAM10000 (HAM10K) dataset, this research employs the U-Net architecture for precise lesion segmentation, ensuring accurate localization of affected areas, critical for subsequent feature extraction and classification.
To enhance diagnostic performance, this study introduces a dual-stream approach: the first track uses Virchow2, a pretrained model that uses dermatoscopic data to extract high-level embeddings capturing intricate patterns in skin lesions, whereas the second track employs a vision-based approach in which Nomic is used to extract spatial and contextual information from these images. This study aims to create a comprehensive diagnostic system to improve classification accuracy and robustness by integrating these methodologies.
The rest of the manuscript is organized in the following manner: Section 2 provides an overview of the relevant research. Section 3 introduces the proposed approach for skin lesion diagnosis. Section 4 evaluates the proposed dual-stream methodology, and its performance was assessed via a comprehensive set of metrics. Finally, Section 5 concludes the work and summarizes potential directions for further research.
Related studies
The field of skin lesion classification has undergone significant advancements in recent years, with DL techniques playing a pivotal role in improving diagnostic accuracy. In 2022, Ali et al. introduced a multiclass skin cancer classification system using EfficientNets, achieving an F1 score of 87% and a top-1 accuracy of 87.91% on the HAM10000 dataset41. Their work demonstrated the effectiveness of transfer learning and fine-tuning in handling imbalanced datasets. However, the study highlighted that increased model complexity does not always lead to better performance, suggesting the need for more efficient architectures.
Similarly, Shetty et al. proposed a machine learning and convolutional neural network (CNN) approach for skin lesion classification, achieving a training accuracy of 95.18% and a testing accuracy of 86.43% with a customized CNN model42. Their work emphasized the importance of data augmentation and k-fold cross-validation to address class imbalance and improve model robustness. Despite these advancements, the reliance on traditional CNNs has limited their ability to capture more complex patterns in skin lesions, and the significant gap between training accuracy and testing accuracy indicates potential overfitting.
In 2024, Adebiyi et al. explored multimodal learning by combining skin lesion images with patient metadata (age, sex, and lesion location) via the ALBEF model43. Their approach achieved an accuracy of 94.11% and an AUCROC of 0.9426 but with a lower recall of 90.19%, outperforming traditional DL models that rely solely on images. This study demonstrated the potential of multimodal learning to improve diagnostic accuracy, particularly in primary care settings where access to expert dermatologists is limited. However, the study was limited by the limited metadata available, suggesting that incorporating additional clinical data could further enhance performance.
Recent advancements in transformer-based models have also contributed to the field. Xin et al. introduced an improved transformer network for skin cancer classification, achieving state-of-the-art performance on the ISIC dataset44. Their work demonstrated the potential of transformers to capture long-range dependencies in dermatoscopic images, but it also highlighted the challenges of training such models on limited medical datasets.
In addition, Mao et al. proposed the medical supervised masked autoencoder (MSMAE), which introduced a better masking strategy and fine-tuning schedule for medical image classification45. Their approach improved the efficiency of transformer-based models, making them more suitable for skin lesion classification tasks.
The reviewed studies collectively highlight several limitations that the current research aims to address. First, while models such as EfficientNets and SBXception have strong performance, they often struggle with class imbalance and computational efficiency. The current study addresses this issue by employing data augmentation and a dual-stream approach that balances accuracy and computational overhead. Second, while multimodal learning, as demonstrated by Adebiyi et al., has shown promise, it is limited by the availability of metadata43.
Third, while transformer-based models proposed by Xin et al.44 and Mao et al.45 have shown potential, they require significant computational resources. The current study integrates efficient feature extraction methods and pretrained models to reduce computational complexity while maintaining high accuracy. The current research aims to create a robust and efficient diagnostic system for skin lesion classification by addressing these limitations.
Methodology
In this section, we introduce the proposed approach for skin lesion diagnosis (see Fig. 1), which uses a dual-stream framework combining histopathological-inherited and vision-based feature extraction. The methodology begins with Data Acquisition, where dermatoscopic images are collected and preprocessed to ensure high-quality input for the model. These images are then passed through a U-Net Model for precise lesion segmentation, generating predicted regions of interest (ROIs) that highlight the affected areas.
The segmented ROIs are processed in parallel by two streams: the embedded stream network using Virchow2 for histopathologically inherited feature extraction and the vision stream network using Nomic for vision-based feature extraction. The features extracted from both streams are fused and integrated to create a comprehensive representation of the lesion, which is then used for classification. The model is trained via a carefully designed loss function and evaluated via a set of performance metrics to ensure robust and accurate diagnosis.
Graphical representation of the proposed dual-stream framework for skin lesion diagnosis. It consists of: (1) Data Acquisition, where dermatoscopic images are collected and preprocessed; (2) U-Net Model, which performs lesion segmentation to generate predicted ROIs; (3) Embedded Stream Network, which uses Virchow2 to extract histopathologically inherited features; (4) Vision Stream Network, which uses Nomic to extract vision-based features; (5) Feature Fusion and Integration, where features from both streams are combined; and (6) Performance Evaluation, where the model’s performance is assessed via metrics such as accuracy and F1 score.
Preprocessing of the dataset
The HAM10K dataset, consisting of 10,015 dermatoscopic images across seven common skin lesion types, was preprocessed to ensure consistency and compatibility with the dual-stream framework. The preprocessing pipeline included several key steps to enhance data quality and facilitate robust model training.
First, all images were resized to a uniform resolution of \(224 \times 224\) pixels to standardize input dimensions while preserving critical details for feature extraction. This resizing step also ensured computational efficiency during training and inference. To address class imbalance, a combination of undersampling and oversampling techniques was applied, ensuring that each skin lesion type was adequately represented in the training set.
Next, data augmentation was performed to simulate real-world variations in lesion appearance and improve the model’s generalizability. Augmentation techniques included random rotations, horizontal and vertical flipping, scaling, and color jittering. These transformations introduced variability in lesion orientation, size, and color, mimicking the diversity observed in clinical settings.
The dataset was then split into training, validation, and testing subsets using a stratified approach to maintain proportional representation of each lesion type across all splits. Specifically, 70% of the dataset was allocated for training, 15% for validation, and the remaining 15% for testing. This partitioning ensures a robust evaluation of the model’s performance on unseen data, as described in Equation 1 where N represents the total number of images in the dataset.
Finally, pixel intensity normalization was applied to all images to standardize the range of input values. Each image’s pixel values were scaled to the range [0, 1], ensuring consistent input distributions for the deep learning models. This normalization step improved convergence speed during training and enhanced the stability of the model. By implementing these preprocessing steps, the dataset was optimized for use in the dual-stream framework, enabling accurate and efficient training while ensuring the model’s ability to generalize across diverse clinical scenarios.
Lesion segmentation using U-Net
Accurate lesion segmentation is a critical step in skin lesion diagnosis, as it enables precise localization of affected areas, which is essential for subsequent feature extraction and classification46. This study employs the U-Net architecture for lesion segmentation because of its proven effectiveness in medical image analysis tasks.
U-Net is a convolutional neural network (CNN) designed explicitly for biomedical image segmentation and is characterized by its encoder-decoder structure with skip connections that facilitate the preservation of spatial information47. The encoder captures contextual features through a series of convolutional and pooling layers. In contrast, the decoder reconstructs the spatial resolution of the feature maps to produce a pixel-wise segmentation mask48.
The U-Net architecture is defined by the following key components: an encoder, a decoder, and a loss function. The encoder consists of a series of convolutional layers followed by max-pooling layers49. Each convolutional layer applies a set of filters to the input image, extracting hierarchical features. The output of the i-th convolutional layer can be expressed as in Equation 2, where \(F_i\) represents the feature maps at layer i, \(W_i\) and \(b_i\) are the learnable weights and biases, \(*\) denotes the convolution operation, and \(\sigma\) is the activation function (e.g., ReLU). The max pooling operation reduces the spatial dimensions of the feature maps, enabling the network to capture broader contextual information.
The decoder pathway reconstructs the spatial resolution of the feature maps through upsampling and convolutional layers. The upsampling operation is followed by concatenation with the corresponding feature maps from the encoder pathway via skip connections50. This process can be described as in Equation 3, where \(F_j\) represents the feature maps at the j-th decoder layer, and \(\text {Upsample}(\cdot )\) denotes the upsampling operation. The skip connections ensure that fine-grained details from the encoder are preserved, enabling precise localization of lesion boundaries.
The utilized U-Net model (see Fig. 2) is trained via the Dice loss function, which is particularly suitable for segmentation tasks with imbalanced class distributions51. The Dice loss is defined as in Equation 4, where \(y_p\) and \(\hat{y}_p\) represent the ground truth and predicted segmentation masks, respectively, for pixel p. Dice loss minimizes the discrepancy between the predicted and ground truth masks, ensuring accurate lesion segmentation.
One of the key advantages of U-Net is its reduced complexity compared with other segmentation models, such as DeepLab or Mask R-CNN. This reduction in complexity not only simplifies the training process but also reduces latency, making the model more suitable for real-time applications52. Furthermore, the lesions in the HAM10000 dataset are generally large enough to be effectively analyzed via a moderately complex model such as U-Net53.
Employing an overly complex and advanced model for this task would be computationally inefficient and unnecessary, as the additional complexity would not yield significant improvements in segmentation accuracy. Instead, U-Net strikes an optimal balance between performance and computational efficiency, making it an ideal choice for this study.
The output of the U-Net model is a binary segmentation mask that precisely delineates the lesion region. This mask then isolates the lesion from the background, enabling focused feature extraction and classification in subsequent pipeline stages. By utilizing U-Net’s ability to capture fine-grained details while maintaining computational efficiency, this study ensures accurate and efficient lesion segmentation, laying the foundation for robust skin lesion diagnosis.
Dual-stream feature extraction
Recent advancements in transformer-based architectures, such as Vision Transformers (ViTs) and Swin Transformers54,55,56, have demonstrated superior performance in capturing long-range dependencies within images, making them particularly well-suited for medical image analysis57. In line with this, the proposed dual-stream framework utilizes transformer-based models for both feature extraction streams.
Specifically, Virchow2, a self-supervised Vision Transformer pretrained on 3.1 million whole-slide histopathology images, is employed to extract high-level histopathological features. Its architecture inherently excels at capturing intricate patterns and subtle morphological characteristics of skin lesions, enabling state-of-the-art performance in various computational pathology tasks58,59. Similarly, Nomic, the second stream’s feature extractor, utilizes a vision-based transformer architecture to analyze spatial and contextual information from dermatoscopic images60.
By integrating these transformer-based models, the framework ensures optimal extraction of both global and local features, addressing the reviewer’s recommendation to enhance the capture of subtle lesion characteristics. This design choice underscores the robustness of the proposed approach in overcoming the challenges associated with skin lesion diagnosis.
Virchow2 incorporates a multilayer perceptron (MLP) with gated linear unit (GLU) style gating and sigmoid linear unit (SiLU) activation functions. The GLU mechanism allows for dynamic feature selection, enhancing the model’s ability to focus on relevant patterns in the histopathological data. Register tokens are ignored during processing, ensuring that only meaningful patch and class tokens contribute to the final embeddings.
The embeddings are derived from both class tokens \(\textbf{c}\) and patch tokens \(\textbf{P}\), which are concatenated to form a comprehensive feature representation as in Equation 5, where \(\textbf{c} \in \mathbb {R}^d\) is the class token, \(\textbf{P}_i \in \mathbb {R}^d\) represents the i-th patch token, N is the number of patches, and \(\Vert\) denotes concatenation. This ensures that the model captures both the global and local features critical for accurate diagnosis.
The second track employs Nomic, a vision-based feature extraction model, to analyze spatial and contextual information from the same set of images. Nomic processes the images through a transformer-based architecture, extracting embeddings \(\textbf{E}_{\text {Nomic}} \in \mathbb {R}^d\) that encode visual patterns such as texture, color, and structural irregularities. The Nomic model uses a Nomic Processor to preprocess the input images, followed by a transformer backbone that generates a last hidden state. The class token from this hidden state is extracted and normalized via L2 normalization, as defined in Equation 6, to ensure consistent scaling of the feature vectors where \(\textbf{h}_{\text {cls}} \in \mathbb {R}^d\) represents the class token extracted from the last hidden state of the transformer.
Combining these embeddings with the histopathological-inherited features from Virchow2 provides the model with a more holistic understanding of the lesion characteristics, enhancing its diagnostic capabilities. The fusion process integrates global and local histopathological features with spatial and contextual visual patterns, enabling the model to achieve superior performance in skin lesion classification.
Feature fusion and integration
The features extracted from both tracks are integrated through a fusion mechanism that combines the strengths of histopathologically inherited and vision-based data. The embeddings from Virchow2 and Nomic are normalized and concatenated, ensuring that the model retains the discriminative power of both feature sets. The fusion process can be expressed as in Equation 7, where \(\text {Norm}(\cdot )\) denotes L2 normalization.
This fusion process is followed by a series of fully connected (FC) dense layers, which further refine the combined feature representation. Each dense block consists of an FC layer, a LeakyReLU activation function, and a dropout layer to prevent overfitting. Integrating these features is critical for capturing the complex interplay between morphological and visual characteristics, enabling the model to make more informed decisions during classification.
Classification model
The classification model is built upon a deep neural network architecture incorporating fused feature representations. The model consists of a multilayer perceptron (MLP) with dropout and batch normalization layers to prevent overfitting and improve generalizability. The output logits \(\textbf{z}\) are computed via Equation 9, where \(\textbf{W}_1, \textbf{W}_2\) are weight matrices, \(\textbf{b}_1, \textbf{b}_2\) are bias terms, and ReLU is the activation function.
The final probabilities \(\textbf{p}\) are obtained via the Softmax function presented in Equation 10, where C is the number of classes.
The model is trained via a cross-entropy loss function, as expressed in Equation 11, where \(y_i\) is the ground truth label. The inclusion of both histopathologically inherited and vision-based features allows the model to achieve high classification accuracy while maintaining robustness to variations in the input data.
To ensure efficient training and convergence, the learning rate was dynamically adjusted using a ReduceLROnPlateau scheduler. This scheduler reduces the learning rate when the validation loss plateaus, as defined in Equation 12, where \(\eta _t\) is the learning rate at epoch t, \(\eta _{t-1}\) is the learning rate at the previous epoch, and \(\alpha\) is the reduction factor applied when the validation loss stops improving.
This approach effectively balances computational efficiency and model performance, ensuring stable convergence without excessive computational overhead. While techniques such as Cosine Annealing or OneCycleLR can be potential alternatives for improving convergence speed and avoiding local minima, the current strategy already demonstrates robust performance across multiple trials. Nevertheless, future suggestions is to explore the integration of these advanced learning rate schedules to further optimize the model’s training dynamics and enhance its adaptability to complex datasets.
Model evaluation and validation
The performance of the proposed model is evaluated via a comprehensive set of metrics, including precision, recall, F1 score, accuracy, and specificity. These metrics are computed via weighted averaging techniques to account for class imbalance in the dataset61. For example, weighted precision \(\text {precision}_{\text {weighted}}\) is calculated via Equation 13, where \(w_i = \frac{n_i}{N}\) is the weight for class i, \(n_i\) is the number of samples in class i, and N is the total number of samples.
Similarly, the weighted F1 score is computed via Equation 14, where \(\text {precision}_i\) and \(\text {recall}_i\) are the precision and recall for class i, respectively62.
Weighted accuracy is defined as in Equation 15, where \(\text {TN}_i\) and \(\text {FN}_i\) are the true negatives and false negatives for class i, respectively.
The model is validated through a series of experiments, including k-fold cross-validation and independent testing on unseen data. Additionally, dimensionality reduction techniques such as PCA, t-SNE, and UMAP are employed to visualize the feature embeddings, providing insights into the model’s ability to separate different classes in the latent space. For example, PCA reduces the dimensionality of the embeddings \(\textbf{E}_{\text {fused}} \in \mathbb {R}^d\) to \(\textbf{E}_{\text {PCA}} \in \mathbb {R}^2\) via Equation 16, where \(\textbf{U}\) and \(\Sigma\) are the eigenvectors and eigenvalues of the covariance matrix of \(\textbf{E}_{\text {fused}}\).
The confusion matrix is used to analyze the model’s performance at the granular level, identifying potential areas for improvement. The results demonstrate that the dual-stream approach significantly enhances diagnostic accuracy compared to single-track models, making it a promising solution for detecting skin cancer.
To illustrate the feature extraction and fusion process, consider a skin lesion image \(\textbf{I}\). Virchow2 first processes the image to extract histopathologically inherited features \(\textbf{E}_{\text {Virchow2}}\), and Nomic is used to extract vision-based features \(\textbf{E}_{\text {Nomic}}\). These features are normalized and concatenated to form the fused embedding \(\textbf{E}_{\text {fused}}\). The fused embedding is then passed through the classification model to produce the final probabilities \(\textbf{p}\), as shown in the following flow:
-
1.
Input Image: \(\textbf{I}\)
-
2.
Feature Extraction:
-
Virchow2: \(\textbf{E}_{\text {Virchow2}} = \left[ \textbf{c} \, \Vert \, \frac{1}{N} \times \sum _{i=1}^{N} \textbf{P}_i \right]\)
-
Nomic: \(\textbf{E}_{\text {Nomic}}\)
-
-
3.
Feature Fusion: \(\textbf{E}_{\text {fused}} = \text {Norm}(\textbf{E}_{\text {Virchow2}}) \, \Vert \, \text {Norm}(\textbf{E}_{\text {Nomic}})\)
-
4.
Classification: \(\textbf{z} = \textbf{W}_2 \times \text {ReLU}(\textbf{W}_1 \times \textbf{E}_{\text {fused}} + \textbf{b}_1) + \textbf{b}_2\)
-
5.
Output Probabilities: \(p_i = \frac{\exp (z_i)}{\sum _{j=1}^C \exp (z_j)}\)
The proposed framework pseudocode
The pseudocode for the proposed framework is outlined in Algorithm 1. The algorithm provides a step-by-step description of the dual-stream deep learning framework for skin lesion diagnosis, encompassing data preprocessing, feature extraction using Virchow2 and Nomic, feature fusion, model training, and performance evaluation. Each step is designed to ensure robustness, generalizability, and interpretability of the model while addressing challenges such as class imbalance and complex lesion characteristics. The modular structure of the pseudocode reflects the key components of the framework, enabling reproducibility and facilitating future enhancements.
Experiments and discussion
The HAM10K dataset, which consists of 10,015 dermatoscopic images across seven common skin lesion types, was utilized in this study. To evaluate the proposed dual-stream framework, 70% of the dataset was used for training, 15% for validation, and the remaining 15% for testing. This split ensures a robust assessment of the model’s generalization capabilities on unseen data. The performance was assessed via a comprehensive set of metrics, including precision, recall, F1 score, accuracy, and specificity.
Segmentation performance
The U-Net architecture was employed for precise lesion segmentation, ensuring accurate localization of affected areas. Figure 3 shows the learning and validation curves for segmentation, including metrics such as loss, accuracy, Dice coefficient, intersection over union (IoU), recall, precision, and other key performance indicators. The curves indicate stable convergence during training, with minimal overfitting, demonstrating the model’s ability to generalize well to unseen data.
Quantitative results from the segmentation process are summarized in Table 1, showcasing the model’s robust performance across multiple evaluation metrics. Notably, the model achieved a Dice coefficient of 0.9169 , an IoU of 0.8493 , and a mean accuracy of 96.79% , reflecting its effectiveness in delineating lesion boundaries even in challenging cases with irregular shapes and textures. Additionally, the precision and recall values of 0.9146 and 0.9225 , respectively, highlight the model’s balanced performance in identifying both positive and negative regions. The Focal Tversky Loss and BCELoss were minimized to 0.1514 and 0.0754 , respectively, further confirming the model’s strong optimization during training.
Figure 4 shows three prediction samples from the U-Net model. The columns represent the original dermatoscopic images, ground truth masks, and predicted masks, respectively. The visual results highlight the model’s ability to accurately segment skin lesions, even in cases with complex boundaries and varying textures. For example, the model successfully identifies lesions with heterogeneous pigmentation and irregular edges, which are often challenging for traditional segmentation methods. This level of precision is crucial for downstream tasks such as feature extraction and classification, as accurate segmentation ensures that the extracted features represent the lesion’s true characteristics.
Classification performance
The classification performance of the proposed dual-stream approach is summarized in Table 2. The results of over 10 trials were reported, with metrics including precision, recall, F1 score, accuracy, and specificity. The maximum, mean, standard deviation (Std), and confidence interval (CI) values are also provided. The model achieves a mean accuracy of 96.25% and a mean F1 score of 93.79%, demonstrating its robustness and consistency across trials. The high specificity values (mean of 91.82%) further indicate the model’s ability to correctly identify negative cases, which is critical for reducing false positives in skin cancer diagnosis.
The precision and recall values, which are particularly important for medical applications, are consistently high across all trials. This indicates that the model strikes a balance between minimizing false positives (high precision) and false negatives (high recall), ensuring reliable diagnostic outcomes. The low standard deviation and narrow confidence intervals for all the metrics further underscore the stability and reproducibility of the proposed approach.
Ablation studies
Ablation studies were conducted to evaluate the contribution of each component in the dual-stream framework. Table 3 presents the performance metrics when either the embedding branch (Virchow2) or the vision branch (Nomic) is removed. The results show a significant decrease in performance when the embedding branch is removed, with the precision and F1 score decreasing to 61.33% and 64.01%, respectively. This highlights the importance of histopathologically inherited features in capturing domain-specific patterns for accurate diagnosis. The embedding branch, which uses Virchow2, provides high-level morphological information critical for distinguishing between different skin lesions.
Similarly, removing the vision branch results in a noticeable decline in performance, with precision and the F1 score decreasing to 85.51% and 85.50%, respectively. This underscores the complementary role of vision-based features in enhancing the model’s diagnostic capabilities. The vision branch, powered by Nomic, captures spatial and contextual information such as texture, color, and structural irregularities, which are essential for identifying subtle differences between lesion types. The ablation studies demonstrate that both branches contribute uniquely to the model’s performance, and their integration is key to achieving state-of-the-art results.
Figure 5 visualizes the dimensionality reduction of the Virchow2 features via principal component analysis (PCA). The plot reflects the complexity of the problem and provides insights into the separability of different lesion classes in the feature space. The PCA visualization shows distinct clusters for different lesion types, indicating that the Virchow2 embeddings effectively capture discriminative features. However, some overlap between classes is observed, highlighting the challenges of distinguishing between visually similar lesions. This further emphasizes the importance of combining histopathologically inherited and vision-based features, as the dual-stream approach utilizes complementary information to improve class separability.
Dimensionality reduction of Virchow2 embeddings via principal component analysis (PCA). The plot illustrates the separability of different lesion classes in the feature space, highlighting the complexity of the problem and the importance of combining histopathologically inherited and vision-based features.
Comparative analysis
The proposed dual-track approach outperforms traditional single-track methods, which rely solely on either extracted or vision-based features. Integrating both feature types enables the model to capture a more comprehensive representation of skin lesions, improving diagnostic accuracy. For example, while extracted features effectively identify morphological patterns, they may struggle with lesions that exhibit similar structures but differ in texture or color. Conversely, vision-based features excel at capturing visual patterns but may lack the domain-specific knowledge required for precise diagnosis. By combining these two modalities, the proposed approach addresses the limitations of single-track methods and achieves superior performance.
To contextualize the performance of the proposed dual-stream approach, a comparative analysis with existing studies using the HAM10000 dataset is presented in Table 4. It highlights the methodologies, results, and key metrics of recent studies alongside the performance of the proposed framework.
The proposed dual-stream approach achieves an accuracy of 96.25%, outperforming several recent studies. For example, Chaturvedi et al.63 employed MobileNet for skin lesion classification, achieving an accuracy of 83.1%. While their approach is computationally efficient, it lacks the ability to integrate domain-specific features, which limits its ability to distinguish between visually similar lesions. Similarly, Xin et al.44 proposed an improved transformer network, achieving an accuracy of 94.3%. Although their method utilizes advanced transformer architectures, it does not explicitly incorporate embeddings, which are critical for capturing morphological patterns.
Agyenta et al.64 utilized DenseNet201, reporting a training accuracy of 99.12% but a testing accuracy of only 86.91%. This significant drop in performance suggests potential overfitting, highlighting the challenges of generalizing single-modality approaches to unseen data. In contrast, the proposed dual-stream approach demonstrates robust generalization, as evidenced by its consistent performance across multiple trials (see Table 2).
Ali et al.41 explored EfficientNets for multiclass skin lesion classification, achieving an F1 score of 87% and a top-1 accuracy of 87.91%. While their approach is effective, it relies solely on vision-based features, which may not fully capture the complexity of skin lesions. The proposed dual-stream framework addresses this limitation by integrating histopathologically inherited features, resulting in higher accuracy and F1 scores.
Adebiyi et al.43 introduced a multimodal learning approach, achieving an accuracy of 94.11% and an AUCROC of 0.9426. Their method combines multiple data modalities, demonstrating the benefits of integrating diverse feature types. However, their approach does not explicitly utilize pretrained models such as Virchow2, which are specifically designed for dermatoscopic data. The proposed dual-stream approach builds on this idea by incorporating Virchow2 embeddings, further enhancing diagnostic accuracy.
Clinical implications
The high accuracy and robustness of the proposed framework have significant implications for clinical practice. Skin cancer diagnosis often requires expert dermatologists to analyze complex visual and morphological patterns, which can be time-consuming and prone to human error. The proposed dual-stream approach provides a reliable and efficient tool for assisting clinicians in diagnosing skin lesions, reducing the burden on healthcare systems and improving patient outcomes. The interpretability of the model, as demonstrated by PCA visualization and ablation studies, further enhances its clinical utility, as it allows clinicians to understand the basis for the model’s predictions.
Moreover, the model’s ability to generalize across different trials and datasets suggests that it can be adapted to various clinical settings and populations. This is particularly important for skin cancer diagnosis, as lesion characteristics can vary significantly depending on factors such as skin type, age, and geographic location. The robustness of the proposed framework to such variations makes it a promising candidate for widespread deployment in dermatology.
Limitations
Despite its strong performance, the proposed approach has several limitations. First, the model relies on high-quality dermatoscopic images, which may not always be available in resource-limited settings. Future work could explore alternative imaging modalities or techniques for enhancing low-quality images. Second, while ablation studies demonstrate the importance of both feature types, further research is needed to optimize the fusion mechanism and explore alternative architectures for integrating histopathologically inherited and vision-based features.
Additionally, the model’s performance could be further improved by incorporating additional data sources, such as patient metadata (e.g., age, sex, medical history) or multispectral imaging data. These extra inputs could provide valuable context for improving diagnostic accuracy and personalizing treatment plans. Finally, future work should validate the model on more extensive and diverse datasets to ensure its generalizability across different populations and clinical scenarios.
Conclusions and future directions
This study presents a dual-stream DL framework for skin lesion diagnosis that combines histopathological-inherited and vision-based feature extraction to achieve state-of-the-art performance. The proposed approach uses advanced DL techniques to address key limitations of traditional diagnostic methods, such as subjectivity, interobserver variability, and resource constraints. The U-Net architecture ensures precise lesion segmentation, whereas the dual-stream feature extraction mechanism, which uses Virchow2 and Nomic, captures both the morphological and contextual information critical for accurate diagnosis. The fusion of these features enables the model to achieve a mean accuracy of 96.25% and a mean F1 score of 93.79% on the HAM10000 dataset, demonstrating its robustness and generalizability across multiple trials. Ablation studies highlight the complementary roles of histopathological and vision-based streams, with the removal of either stream leading to significant performance degradation. Comparative analysis with existing studies further validates the superiority of the proposed framework, which outperforms traditional single-modality approaches and recent multimodal methods. The interpretability of the model, as demonstrated by dimensionality reduction techniques such as PCA, provides valuable insights into the feature space, enhancing its clinical utility. The proposed framework has significant implications for clinical practice, offering a reliable and efficient tool for assisting dermatologists in diagnosing skin lesions. Its ability to generalize across diverse datasets and populations makes it a promising candidate for widespread deployment, particularly in resource-limited settings.
Future directions
Future work will focus on key enhancements to strengthen the framework. First, the fusion mechanism will be optimized, potentially incorporating attention mechanisms or multi-scale feature fusion to better capture local and global spatial information. Second, additional data sources, such as patient metadata (e.g., age, sex, medical history) and advanced imaging modalities like thermal or multispectral data, multispectral or 3D imaging, will be integrated to enrich contextual information and enable personalized predictions. Third, the model will be validated on larger, diverse datasets, expanding the evaluation to include datasets that explicitly represent underrepresented populations, varied skin phototypes (e.g., Fitzpatrick skin types I-VI), and regional differences in lesion presentation. These improvements aim to revolutionize skin cancer diagnosis, enhance accuracy, improve outcomes, and reduce mortality, paving the way for clinical adoption.
Data Availability
This study used the HAM10000 dataset, a large collection of multi-source dermatoscopic images of common pigmented skin lesions, which is available at: https://dataverse.harvard.edu/dataset.xhtml?persistentId=doi:10.7910/DVN/DBW86T
References
Jindal, M. et al. Skin cancer management: Current scenario and future perspectives. Curr. Drug Saf. 18, 143–158 (2023).
Papadopoulos, O., Karantonis, F.-F. & Papadopulos, N. A. Non-melanoma skin cancer and cutaneous melanoma for the plastic and reconstructive surgeon. Non-Melanoma Skin Cancer and Cutaneous Melanoma: Surgical Treatment and Reconstruction 153–239 (2020).
Balaha, H. M. & Hassan, A.E.-S. Skin cancer diagnosis based on deep transfer learning and sparrow search algorithm. Neural Comput. Appl. 35, 815–853 (2023).
Church, K. Early Detection of Melanoma. Master’s thesis, school The College of St. Scholastica (2022).
Shamshad, M. F. et al. A comprehensive review on treatment modalities of malignant melanoma. J. Liaquat Natl. Hosp. 1, 90–102 (2023).
Narayanan, D. L., Saladi, R. N. & Fox, J. L. Ultraviolet radiation and skin cancer. Int. J. Dermatol. 49, 978–986 (2010).
Gosman, L. M., Tăpoi, D.-A. & Costache, M. Cutaneous melanoma: A review of multifactorial pathogenesis, immunohistochemistry, and emerging biomarkers for early detection and management. Int. J. Mol. Sci. 24, 15881 (2023).
Geller, A. C., Swetter, S. M., Brooks, K., Demierre, M.-F. & Yaroch, A. L. Screening, early detection, and trends for melanoma: Current status (2000–2006) and future directions. J. Am. Acad. Dermatol. 57, 555–572 (2007).
Karakousis, G. C. & Czerniecki, B. J. Diagnosis of melanoma. PET Clin. 6, 1–8 (2011).
Heibel, H. D., Hooey, L. & Cockerell, C. J. A review of noninvasive techniques for skin cancer detection in dermatology. Am. J. Clin. Dermat. 21, 513–524 (2020).
Meng, X. et al. Non-invasive optical methods for melanoma diagnosis. Photodiagn. Photodyn. Ther. 34, 102266 (2021).
Leachman, S. A. et al. Methods of Melanoma Detection. Melanoma 51–105 (2016).
Werner, B. Skin biopsy and its histopathologic analysis: Why? what for? how? part i. Anais Bras. Dermatol. 84, 391–395 (2009).
Mogensen, M. & Jemec, G. B. Diagnosis of nonmelanoma skin cancer/keratinocyte carcinoma: A review of diagnostic accuracy of nonmelanoma skin cancer diagnostic tests and technologies. Dermatol. Surg. 33, 1158–1174 (2007).
Dobre, E.-G. et al. Skin cancer pathobiology at a glance: A focus on imaging techniques and their potential for improved diagnosis and surveillance in clinical cohorts. Int. J. Mol. Sci. 24, 1079 (2023).
Bridge, P., Fielding, A., Rowntree, P. & Pullar, A. Intraobserver Variability: Should We Worry? (2016).
Van Den Einden, L. C. et al. Interobserver variability and the effect of education in the histopathological diagnosis of differentiated vulvar intraepithelial neoplasia. Mod. Pathol. 26, 874–880 (2013).
Bajaj, S. et al. The role of color and morphologic characteristics in dermoscopic diagnosis. JAMA Dermatol. 152, 676–682 (2016).
Malik, S. & Dixit, V. V. Skin cancer detection: State of art methods and challenges. In ICCCE 2021: Proceedings of the 4th International Conference on Communications and Cyber Physical Engineering 729–736 (Springer, 2022).
Anand, V., Gupta, S. & Koundal, D. Skin disease diagnosis: challenges and opportunities. In Proceedings of Second Doctoral Symposium on Computational Intelligence: DoSCI 2021 449–459 (Springer, 2022).
Zarella, M. D. et al. A practical guide to whole slide imaging: A white paper from the digital pathology association. Arch. Pathol. Lab. Med. 143, 222–234 (2019).
Rizzo, P. C. et al. Digital pathology world tour. Digit. Health 9, 20552076231194550 (2023).
Rizzo, P. C. et al. Technical and diagnostic issues in whole slide imaging published validation studies. Front. Oncol. 12, 918580 (2022).
Marletta, S. et al. Artificial intelligence-based algorithms for the diagnosis of prostate cancer: A systematic review. Am. J. Clin. Pathol. 161, 526–534 (2024).
Koteluk, O., Wartecki, A., Mazurek, S., Kołodziejczak, I. & Mackiewicz, A. How do machines learn? Artificial intelligence as a new era in medicine. J. Pers. Med. 11, 32 (2021).
Halabi, D. Enhancing skin cancer detection and classification: Exploring the impact of attention mechanisms in transfer learning models. Preprints (2023).
Kushimo, O. O., Salau, A. O., Adeleke, O. J. & Olaoye, D. S. Deep learning model to improve melanoma detection in people of color. Arab. J. Basic Appl. Sci. 30, 92–102 (2023).
Ashfaq, M. & Ahmad, A. Skin cancer classification with convolutional deep neural networks and vision transformers using transfer learning. In Advances in Deep Generative Models for Medical Artificial Intelligence 151–176 (publisherSpringer, 2023).
Jones, O. et al. Artificial intelligence and machine learning algorithms for early detection of skin cancer in community and primary care settings: A systematic review. Lancet Digit. Health 4, e466–e476 (2022).
Takiddin, A., Schneider, J., Yang, Y., Abd-Alrazaq, A. & Househ, M. Artificial intelligence for skin cancer detection: Scoping review. J. Med. Internet Res. 23, e22934 (2021).
Hauser, K. et al. Explainable artificial intelligence in skin cancer recognition: A systematic review. Eur. J. Cancer 167, 54–69 (2022).
Höhn, J. et al. Combining cnn-based histologic whole slide image analysis and patient data to improve skin cancer classification. Eur. J. Cancer 149, 94–101 (2021).
Mampitiya, L. I., Rathnayake, N. & De Silva, S. Efficient and low-cost skin cancer detection system implementation with a comparative study between traditional and cnn-based models. J. Comput. Cogn. Eng. 2, 226–235 (2023).
Shah, A. et al. A comprehensive study on skin cancer detection using artificial neural network (ann) and convolutional neural network (cnn). Clinical eHealth (2023).
Junayed, M. S., Anjum, N., Noman, A. & Islam, B. A deep cnn model for skin cancer detection and classification. International Conference in Central Europe on Computer Graphics, Visualization and Computer Vision (2021).
Nawaz, M. et al. Melanoma segmentation: A framework of improved densenet77 and unet convolutional neural network. Int. J. Imaging Syst. Technol. 32, 2137–2153 (2022).
Miradwal, S., Mohammad, W., Jain, A. & Khilji, F. Lesion segmentation in skin cancer detection using unet architecture. In Computational Intelligence and Data Analytics: Proceedings of ICCIDA 2022 329–340 (publisherSpringer, 2022).
Li, Z. et al. Artificial intelligence in dermatology image analysis: Current developments and future trends. J. Clin. Med. 11, 6826 (2022).
Liopyris, K., Gregoriou, S., Dias, J. & Stratigos, A. J. Artificial intelligence in dermatology: Challenges and perspectives. Dermatol. Ther. 12, 2637–2651 (2022).
Sarker, I. H. Deep learning: A comprehensive overview on techniques, taxonomy, applications and research directions. SN Comput. Sci. 2, 420 (2021).
Ali, K., Shaikh, Z. A., Khan, A. A. & Laghari, A. A. Multiclass skin cancer classification using efficientnets-a first step towards preventing skin cancer. Neurosci. Inform. 2, 100034 (2022).
Shetty, B. et al. Skin lesion classification of dermoscopic images using machine learning and convolutional neural network. Sci. Rep. 12, 18134 (2022).
Adebiyi, A. et al. Accurate skin lesion classification using multimodal learning on the ham10000 dataset. MedRxiv 2024–05 (2024).
Xin, C. et al. An improved transformer network for skin cancer classification. Comput. Biol. Med. 149, 105939 (2022).
Mao, J. et al. Medical supervised masked autoencoder: Crafting a better masking strategy and efficient fine-tuning schedule for medical image classification. Appl. Soft Comput. 112536 (2024).
Yacin Sikkandar, M. et al. Deep learning based an automated skin lesion segmentation and intelligent classification model. J. Ambient Intell. Hum. Comput. 12, 3245–3255 (2021).
Neha, F. et al. U-net in medical image segmentation: A review of its applications across modalities. arXiv preprint arXiv:2412.02242 (2024).
Minaee, S. et al. Image segmentation using deep learning: A survey. IEEE Trans. Pattern Anal. Mach. Intell. 44, 3523–3542 (2021).
Sahli, H., Ben Slama, A. & Labidi, S. U-net: A valuable encoder-decoder architecture for liver tumors segmentation in ct images. J. X-ray Sci. Technol. 30, 45–56 (2022).
Azad, R. et al. Medical image segmentation review: The success of u-net. IEEE Trans. Pattern Anal. Mach. Intell. (2024).
Lu, Y., Lin, J., Chen, S., He, H. & Cai, Y. Automatic tumor segmentation by means of deep convolutional u-net with pre-trained encoder in pet images. IEEE Access 8, 113636–113648 (2020).
Bagheri, F., Tarokh, M. J. & Ziaratban, M. Skin lesion segmentation from dermoscopic images by using mask r-cnn, retina-deeplab, and graph-based methods. Biomed. Signal Process. Control 67, 102533 (2021).
Lilhore, U. K. et al. A precise model for skin cancer diagnosis using hybrid u-net and improved mobilenet-v3 with hyperparameters optimization. Sci. Rep. 14, 4299 (2024).
Saleem, S. & Sharif, M. I. An integrated deep learning framework leveraging nasnet and vision transformer with mixprocessing for accurate and precise diagnosis of lung diseases. arXiv preprint arXiv:2502.20570 (2025).
Hafeez, R. et al. Deep learning in early alzheimers diseases detection: A comprehensive survey of classification, segmentation, and feature extraction methods. arXiv preprint arXiv:2501.15293 (2025).
Balaha, H. M., Ali, K. M., Mahmoud, A., Ghazal, M. & El-Baz, A. Integrated grading framework for histopathological breast cancer: Multi-level vision transformers, textural features, and fusion probability network. In International Conference on Pattern Recognition 76–91 (Springer, 2024).
Shah, O. I., Rizvi, D. R. & Mir, A. N. Transformer-based innovations in medical image segmentation: A mini review. SN Comput. Sci. 6, 375 (2025).
Zimmermann, E. et al. Virchow2: Scaling self-supervised mixed magnification models in pathology. arXiv preprint arXiv:2408.00738 (2024).
Vorontsov, E. et al. Virchow: A million-slide digital pathology foundation model. arXiv preprint arXiv:2309.07778 (2023).
Nussbaum, Z., Morris, J. X., Duderstadt, B. & Mulyar, A. Nomic embed: Training a reproducible long context text embedder. arXiv preprint arXiv:2402.01613 (2024).
Balaha, H. M., Hassan, A.E.-S., El-Gendy, E. M., ZainEldin, H. & Saafan, M. M. An aseptic approach towards skin lesion localization and grading using deep learning and Harris Hawks optimization. Multimed. Tools Appl. 83, 19787–19815 (2024).
Abd El-Khalek, A. A. et al. A concentrated machine learning-based classification system for age-related macular degeneration (amd) diagnosis using fundus images. Sci. Rep. 14, 2434 (2024).
Chaturvedi, S. S., Gupta, K. & Prasad, P. S. Skin lesion analyser: an efficient seven-way multi-class skin cancer classification using mobilenet. In Advanced machine learning technologies and applications: proceedings of AMLTA 2020 165–176 (Springer, 2021).
Agyenta, C. & Akanzawon, M. Skin lesion classification based on convolutional neural network. J. Appl. Sci. Technol. Trends 3, 21–26 (2022).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The author declares that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Ethics approval and consent to participate
Not applicable
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Almutairi, S.A. A dual-stream deep learning framework for skin cancer classification using histopathological-inherited and vision-based feature extraction. Sci Rep 15, 32301 (2025). https://doi.org/10.1038/s41598-025-01319-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-01319-1