Abstract
The VitDPAS study (NCT06104111) was designed as a medical experiment to assess the in vivo effects of vitamin D on immune responses. This study enrolled 45 healthy individuals from Olsztyn, Poland, who received a body weight-adjusted bolus dose of vitamin D3 (1,000 IU/kg). Transcriptome-wide differential gene expression analysis of peripheral blood mononuclear cells, collected before and 24 h after supplementation, identified 758 significantly responsive genes (p < 0.05). By correlating individual gene expression changes with alterations in vitamin D status, participants were categorized into three response groups: 17 high responders, 19 mid responders, and 9 low responders. A comparative analysis with the VitDHiD study (NCT03537027), conducted on a Finnish cohort of 25 healthy participants, revealed 232 overlapping target genes, enabling an integrated assessment of vitamin D responsiveness across all 70 individuals. Applying a more stringent statistical threshold (false discovery rate < 0.05) highlighted 26 shared target genes, demonstrating a consistent in vivo response to vitamin D3 across both cohorts. The modulation of inflammatory processes, mediated primarily via tumor necrosis factor and nuclear factor κB signaling pathways, emerged as a shared effect, highlightening the immunomodulatory potential of vitamin D as a key function of the vitamin in healthy individuals.
Similar content being viewed by others
Introduction
Vitamin D3 is a vital micronutrient that can be synthesized endogenously in human skin when exposed to UV-B light1. Both in vitro studies and animal model experiments suggest that vitamin D plays a significant role in modulating the activity of innate and adaptive immunity2,3,4,5,6. Observational studies comparing individuals with vitamin D deficiency to those with sufficient levels suggest that adequate vitamin D may provide protective benefits against a range of diseases. These include musculoskeletal disorders such as osteoporosis7 and sarcopenia8, autoimmune diseases like multiple sclerosis9, and certain types of cancer, including colon cancer10. In vitamin D deficiency, the underlying issue often relates to insufficient immune modulation5. However, conclusive evidence of the beneficial effects of vitamin D supplementation in healthy, non-deficient individuals is lacking. Large-scale intervention studies with thousands of participants over periods up to five years was unable to demonstrate statistically significant primary benefits from vitamin D supplementation, partly due to ethical constraints against leaving control groups vitamin D deficient for extended periods11,12.
Vitamin D’s molecular mechanisms involve its nuclear receptor, VDR (vitamin D receptor), which binds the active vitamin D metabolite, 1α,25-dihydroxyvitamin D3 (1,25(OH)2D3)13,14, with very high affinity (KD = 0.1 nM). For endocrine signaling, this nuclear hormone is synthesized from 25-hydroxyvitamin D3 (25(OH)D3) in the kidneys but can also be locally produced in immune and skin cells for para- and autocrine functions15. The most abundant and stable metabolite, 25(OH)D3, defines vitamin D status in serum16, with levels above 30 ng/ml (75 nM) considered sufficient and those below 12 ng/ml (30 nM) indicating severe deficiency17.
As a transcription factor, VDR mediates all genomic actions of vitamin D, with 1,25(OH)2D3 being the exclusive activator of VDR at physiological concentrations18. Based on data of the Genotype-Tissue Expression (GTEx) Portal19, VDR is expressed in most tissues, excluding the brain, with hundreds of target genes in each tissue20, many of which exhibit tissue-specific and individual-specific responsiveness12,21. This variation supports the concept of the vitamin D response index, categorizing individuals as high, mid, or low responders to vitamin D supplementation22. Notably, low responders may require higher vitamin D dosages to attain optimal physiological benefits23.
To investigate vitamin D’s molecular effects in healthy individuals under in vivo conditions, we previously conducted studies such as the VitDbol (NCT02063334, ClinicalTrials.gov)24,25 and VitDHiD26,27, involving healthy cohorts in Kuopio, Finland, who received a single 80,000 IU vitamin D3 bolus. In the current study, we expanded on this approach by incorporating the VitDPAS study, which enrolled a cohort of 45 healthy individuals from Olsztyn, Poland. Located more than 1000 km south of Kuopio, the vitamin D winter in Olsztyn is about one month shorter than in Eastern Finland. Transcriptome-wide analysis of peripheral blood mononuclear cells (PBMCs) identified 758 significantly (p < 0.05) responsive vitamin D target genes, of which 232 overlapped with targets observed in the VitDHiD study. A core function of these common in vivo vitamin D target genes was the modulation of inflammatory responses.
Materials and methods
Sample collection
The VitDPAS intervention study commenced in November 2023 in Olsztyn, Poland. Participants eligible for inclusion were between 20 and 65 years old, with a body mass index (BMI) of 18.5–29.9 kg/m2. Exclusion criteria included any disease or infection contracted within the last two weeks before the planned blood draw; severe visual or hearing impairment; gastrointestinal disorders like celiac disease or irritable bowel syndrome; chronic kidney disease; presence of kidney stones; chronic liver diseases; history of pathological bone fractures or fractures due to osteoporosis within the last five years; current diagnosis of cancer; treatment that affects calcium metabolism; diseases with a risk of recurring symptoms, including Parkinson’s disease, post-stroke hemiplegia, epilepsy, recurrent dizziness, and repeated fainting; past or current diseases that may increase serum calcium levels, such as sarcoidosis, lymphoma, and hyperparathyroidism; use of medications that affect 25(OH)D3 levels like phenobarbital, carbamazepine, and phenytoin, Paget’s disease of bone (osteitis deformans), arthritis including rheumatoid arthritis, Reiter’s syndrome, and psoriatic arthritis; participation in research studies within the last 4 months, except for observational studies; known infection with HIV, hepatitis, and other infectious diseases; substance abuse like drug dependence, alcoholism, and tobacco smoking; mental illness; homeostasis disorders that may complicate the blood collection process; current pregnancy, breastfeeding, or planned pregnancy. The study protocol was approved by the Ethics Committee of the Olsztyn Chamber of Doctors (#31/2023/VIII).
The study involved a cohort of 45 healthy participants (23 females and 22 males, aged 24–64, BMI 19.3–29.3, Table 1) who received a single vitamin D3 bolus of 1,000 IU/kg body mass with breakfast. This bolus equates to a monthly dose of vitamin D3, administered only once, whereas other vitamin D interventions, such as the ViDA study28, employed a monthly dose of 100,000 IU of vitamin D3 over a period of more than two years. Therefore, the dose range of 52,000–110,000 IU vitamin D3 used in this study was within safe limits. Blood samples were collected immediately before supplementation (day 0 (d0)) and 24 h after dosing (day 1 (d1)) for serum and PBMC isolation. Based on data collected from previous studies, such as VitDbol24,25, a 24-hour post-supplementation period is considered the optimal time point. Serum 25(OH)D3 levels were assayed using the electrochemiluminescence binding assay on Cobas Pure Immunoassay Analyzer (Roche) in the analytical laboratory of the Municipal Polyclinical Hospital in Olsztyn, Poland. All participants provided written informed consent, and all experimental procedures adhered to relevant guidelines and regulations.
PBMC isolation
PBMCs from the 45 participants of the VitDPAS study were promptly processed without any in vitro culture. Following the collection of 8 ml of peripheral blood on d0 and d1, PBMCs were isolated within one hour using Vacutainer CPT Cell Preparation Tubes with sodium citrate (Becton Dickinson), following the manufacturer’s protocol. After isolation, cells were washed with phosphate-buffered saline, aliquoted at a concentration of 4 million cells per ml, and preserved at -80 °C for future RNA isolation.
Transcriptome analysis
Total RNA was extracted from PBMCs using the High Pure RNA Isolation Kit (Roche) following the manufacturer’s instructions. RNA quality was assessed using an Agilent Tapestation, and library preparation was conducted after rRNA depletion with kits and protocols from New England Biolabs. RNA-seq libraries were sequenced at a read length of 75 bp on a NextSeq2000 system (Illumina) according to standard protocols at the EMBL Gene Core (Heidelberg, Germany). Quality control of the sequencing files was performed using FastQC (version 0.11.9, http://www.bioinformatics.babraham.ac.uk/projects/fastqc) (Supplementary Table S1 online). Reads were aligned to the GRCh38 reference genome (Ensembl version 111.38) using the STAR aligner29 (version 2.7.2a), and quantification was carried out with FeatureCounts30 (version 2.0.1) using default parameters. To standardize gene nomenclature across all datasets, Human Gene Nomenclature Committee (HGNC) symbols were updated using the R package HGNChelper (version 0.8.1, https://CRAN.R-project.org/package=HGNChelper). Annotation, including gene identifiers, descriptions, genomic locations, and biotypes, was added via the Ensembl database (release 100) using the R package BiomaRt31 (version 2.42.1). Entrez Gene identifiers were incorporated using the R package org.Hs.eg.db (version 3.10.0), and any incomplete mappings for target genes were manually verified and retrieved from NCBI (http://www.ncbi.nlm.nih.gov/home/genes). Genes lacking genomic position information or those mitochondrially encoded were excluded from the analysis. This study focused solely on protein-coding genes.
Differential gene expression analysis
Differential gene expression analysis was conducted in R (version 4.3.1) on MacOS 13 (Ventura) using the EdgeR package32 (version 4.0.16) for robust assessment. To reduce transcriptional noise associated with non-coding genes, the analysis focused on 19,287 protein-coding genes. Read counts were normalized to counts per million (CPM) to account for library size differences. Low-expression genes (CPM < 10) were filtered out using the FilterByExpr() function, minimizing the multiple testing burden and optimizing statistical accuracy within the EdgeR framework. Following filtering, library sizes were recalculated, and the trimmed mean of M-values normalization was applied. The transcriptome data structure was explored using multidimensional scaling (MDS) via EdgeR’s plotMDS() function, where distances approximate typical log2 fold changes (FC) between samples. These distances were calculated as the root mean square deviation (Euclidean distance) of log2FC values for genes showing significant changes (p-value < 0.05 or Benjamini-Hochberg adjusted p-value = FDR (false discovery rate) < 0.05)) post-vitamin D3 supplementation. Mean-Difference (MA) plots were generated with vizzy (version 1.0.0, https://github.com/ATpoint/vizzy). Differential expression for each gene was determined through the generalized linear model quasi-likelihood (GLM-QL) pipeline. A trended negative binomial dispersion estimate was calculated using the Cox-Reid profile-adjusted likelihood method. This estimate, along with empirical Bayes-moderated quasi-likelihood gene-wise dispersion estimates33, was employed for GLM fitting, with empirical Bayes shrinkage robustified against outlier dispersions, as recommended. To identify differentially expressed genes with substantial changes, the glmTreat approach33 was applied, setting a threshold for absolute log2FC > 0.25 (Supplementary Table S2 online).
To integrate the gene expression data from the VitDPAS and VitDHiD cohorts, matched common gene identifiers across both datasets were identified, ensuring that only shared genes were included in the combined dataset. To account for differences in sequencing depth, the expression values were normalized using CPM, which adjusts raw counts accordingly. Additionally, a scaling step was applied to harmonize the data across time points (d0 and d1). This involved calculating scaling factors based on the average gene expression levels of both cohorts and adjusting the data to minimize technical discrepancies, thereby facilitating a more accurate comparison.
Classification of study participants
To classify participants as high, mid, or low responders, the average absolute log2FC values of all in vivo vitamin D target genes (or relevant subsets) for each individual (Supplementary Table S2 online) was plotted against the changes in total 25(OH)D3 concentrations (d1/d0) (Table 1). Based on prior experience with the vitamin D intervention studies VitDmet34 and VitDbol24,25, plotting the ratio of 25(OH)D3 values proves to be more effective than using their delta. A trendline was drawn through the 1/0 point to divide the plot, and the orthogonal distance of each data point from this line was calculated. These distances were then ranked in descending order, and clustering of participants into three response groups was performed using the K-means algorithm (KMeans function from the sklearn.cluster library, available at https://pypi.org/project/scikit-learn/).
Functional characterization of vitamin D target genes
The STRING database (http://string-db.org/)35 was used to confirm and visualize the physical and functional interactions among vitamin D target genes. In this network, each node represents a protein, while edges between nodes indicate associations inferred from a combination of experimental, computational, and literature-derived data. Functional annotation of the genes, with an emphasis on their primary roles, was derived from databases like Human Protein Atlas (http://www.proteinatlas.org) and GeneCards (http://www.genecards.org).
Results
The VitDPAS intervention trial
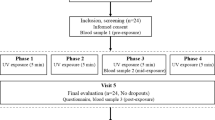
The VitDPAS study design is detailed in Supplementary Figure S1A online. The study began by administering a weight-adjusted vitamin D3 bolus (1,000 IU/kg) to 45 healthy participants (23 females, 22 males; mean age: 39.6 years; mean BMI: 23.8, Supplementary Figure S1B online). Blood samples were collected immediately before supplementation (d0) and 24 h afterward (d1). Baseline serum 25(OH)D3 levels at d0 ranged from 9.0 to 55.9 ng/ml, with an average of 26.5 ng/ml, while levels at d1 increased to a range of 18.6 to 62.6 ng/ml, with an average of 32.9 ng/ml (Table 1). Thus, the vitamin D3 bolus resulted in a significant average increase in vitamin D status by 6.4 ng/ml, or 29% (p = 3.8 × 10− 23, paired T-test, Supplementary Figure S1C online). In summary, a single, weight-adjusted vitamin D3 bolus was sufficient to elevate the vast majority of VitDPAS participants to a level of vitamin D sufficiency.
Vitamin D-induced transcriptomic response in VitDPAS participants
Differential gene expression analysis of RNA-seq data at d1 compared to baseline (d0) revealed that vitamin D3 bolus supplementation significantly impacted the transcriptome of the 45 VitDPAS participants. Principal component analysis, visualized in an MDS plot, showed a notable shift, with d1 data points averaging below the respective d0 points along dimension 2 (Fig. 1A). Out of 8,937 protein-coding genes expressed in PBMCs across all participants (defined as mean CPM > 10 for d0 and d1), 758 genes were significantly (p < 0.05) regulated by vitamin D (Figure S2). Of these, 369 were upregulated and 389 downregulated (Supplementary Table S2 online, Supplementary Figure S2 online). Applying an additional threshold of absolute log2FC > 0.25 narrowed this list to 94 vitamin D target genes, including 39 upregulated and 55 downregulated. A stricter criterion of FDR < 0.05 identified 70 genes, with 15 upregulated and 55 downregulated, and 39 of these also meeting the absolute log2FC > 0.25 threshold (5 upregulated and 34 downregulated) (Fig. 1B). Taken together, vitamin D3 bolus supplementation significantly modulates the transcriptome of circulating immune cells in VitDPAS participants.
Transcriptome-wide analysis of vitamin D3 bolus supplementation in the VitDPAS cohort. (A) The quality of transcriptome datasets (of 758 target genes) from the 45 VitDPAS participants at d0 and d1 was assessed using dimensionality reduction techniques, visualized through a MDS plot. (B) A Volcano plot was employed to illustrate the impact of vitamin D3 supplementation on the expression of 758 significantly regulated genes (p < 0.05). Selected target genes are highlighted.
Segregation of VitDPAS study participants by vitamin D responsiveness
To evaluate the vitamin D responsiveness among the 45 VitDPAS participants, each individual’s average absolute log2FC of all 758 vitamin D target genes was plotted against the ratio of their vitamin D status at d1 relative to baseline (d0) (Fig. 2A). This analysis revealed a spectrum of responsiveness, with high responders clustering at the upper left of the plot (green), mid responders near the trendline (yellow), and low responders at the lower right (red). The distance from the trendline was proportional to each participant’s vitamin D response index, which showed a strong negative correlation (-0.82, Pearson correlation, p = 3.7 × 10− 12, paired T-test) with the d1/d0 vitamin D status ratio. This response index correlated less strongly with baseline vitamin D status (0.61, p = 0.0000089, Supplementary Figure S3A online), and showed no significant correlation with age (0.23, p = 0.12, Supplementary Figure S3B online) or BMI (-0.08, p = 0.58, Supplementary Figure S3C online) of the individuals. K-means clustering further classified the participants into 9 low responders, 19 mid responders, and 17 high responders (Supplementary Figure S4 online). When the analysis was limited to the 94 genes with an absolute log2FC > 0.25, the results were similar; however, participants #9, 14, 16, and 39 were reclassified as mid responders (Fig. 2B). Narrowing the gene list further to 70 targets with FDR < 0.05, individuals #1, 9, 14, 16, 19, and 39 were reclassified as mid responders, while participants #18 and #22 were newly classified as high responders, and #28 as a low responder. Consequently, participants #1 and #19 appear as unstable high responders, while #9, 14, 16, 18, 22, 28, and 39 are unstable mid responders. In summary, VitDPAS participants were stratified into high, mid, and low responders to vitamin D3 supplementation.
Classification of VitDPAS study participants. (A) The 45 VitDPAS participants were classified into high (green), mid (yellow), and low (red) responders based on the expression profiles of all 758 in vivo vitamin D target genes. (B) Comparison of classification methods utilizing various subsets of vitamin D target genes to evaluate the robustness of the segregation approach.
Joint classification of VitDPAS and vitdhid participants
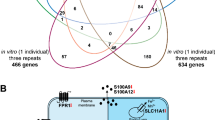
Comparison of the 758 in vivo vitamin D target genes from the VitDPAS study with the 1,654 targets identified in the VitDHiD study (p < 0.05) revealed 232 shared genes (Fig. 3A). The two studies, which both investigated healthy individuals (VitDHiD with 25 participants from Finland), were highly comparable and analyzed using identical methods. The combined transcriptome datasets from all 70 participants were normalized and analyzed using principal component analysis via MDS plots. Based on the 232 common target genes, on average, d1 datasets clustered to the left of d0 datasets along the first dimension, suggesting that vitamin D3 supplementation consistently impacted the PBMC transcriptome across both cohorts (Fig. 3B). Segregation of the 70 individuals by vitamin D responsiveness classified them into 22 high responders, 34 mid responders, and 14 low responders (Fig. 3C). Four VitDPAS participants were reclassified based on this joint analysis: participant #19 shifted from unstable high to mid responder, #16 and #39 from unstable mid to mid responders, and #28 from unstable mid to low responder. Similarly, four VitDHiD participants were reclassified (three from mid to high responders and one from mid to low responder)27. For the remaining 41 VitDPAS and 21 VitDHiD participants (88.6% overall), the joint analysis produced results consistent with each individual study’s classification. In summary, this combined analysis of the VitDPAS and VitDHiD vitamin D intervention studies identified 232 common target genes and yielded consistent response classifications for 62 of the 70 participants.
Overlap and classification of vitamin D target genes between VitDPAS and VitDHiD studies. (A) A Venn diagram highlights the intersection of 758 significant (p < 0.05) in vivo vitamin D target genes identified in the VitDPAS study and 1,654 target genes from the VitDHiD study. (B) The quality of transcriptome datasets for the 232 shared target genes at d0 and d1 was assessed using dimensionality reduction, depicted in a MDS plot. (C) Based on the expression profiles of the 232 common target genes, the 70 participants across both studies were categorized into high (green), mid (yellow), and low (red) vitamin D responders.
Functional impact of joint VitDPAS and vitdhid target genes
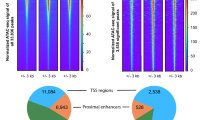
A stringent comparison (FDR < 0.05) between 70 vitamin D target genes identified in the VitDPAS study and 452 targets from the VitDHiD study27 revealed 26 shared genes (Fig. 4A). Notably, of these common genes, only 4 were upregulated by vitamin D3 supplementation, while 22 were downregulated (Supplementary Figure S5 online). The 26 genes were validated by two different approaches. A repeated vitamin D3 bolus supplementation with the same study participants indicated for 80–90% of genes a comparable regulation on the RNA level (Supplementary Figure S6 online). Moreover, an ATAC-seq (assay for transposase accessible chromatin with high-throughput sequencing) analysis for accessible chromatin at TSS (transcription start site) regions confirmed for all downregulated genes a decrease in accessibility (Supplementary Figure S7 online).
Overlap and functional classification of vitamin D target genes between VitDPAS and VitDHiD studies. (A) A Venn diagram illustrates the overlap between 70 highly significant (FDR < 0.05) in vivo vitamin D target genes identified in the VitDPAS study and 452 target genes from the VitDHiD study. (B) The 26 shared vitamin D target genes are grouped into distinct functional categories. Gene interactions, derived from the STRING database (Supplementary Figure S8 online), are depicted with connecting lines. Upregulated genes are shown in green, while downregulated genes are represented in red.
The STRING database was utilized to explore potential functional relationships among the proteins encoded by the 26 shared genes (Supplementary Figure S8 online). This analysis, combined with a review of publicly available databases such as Human Protein Atlas and GeneCards, facilitated the classification of these genes/proteins into five distinct functional categories: (i) “inflammatory response and TNF (tumor necrosis factor)/NFκB (nuclear factor κB) signaling includes TNFAIP3 (TNF alpha-induced protein 3), TNFSF14 (TNF superfamily member 14), NFKBIA (NFκB inhibitor alpha), NFKBIZ (NFκB inhibitor zeta), PELI1 (pellino E3 ubiquitin protein ligase 1), and ETS2 (ETS proto-oncogene 2, transcription factor); (ii) “immune cell migration and survival” comprises S1PR1 (sphingosine-1-phosphate receptor 1), CXCR4 (C-X-C motif chemokine receptor 4), CX3CR1 (C-X3-C motif chemokine receptor 1), and GIMAP6 (GTPase, IMAP family member 6); (iii) “transcriptional regulation” contains PER1 (period circadian regulator 1), CSRNP1 (cysteine and serine-rich nuclear protein 1), KLF10 (KLF transcription factor 10), KLF11, and HIF1A (hypoxia-inducible factor 1 subunit alpha); (iv) “stress response and apoptosis” consists of RBM7 (RNA binding motif protein 7), TSC22D3 (TSC22 domain family member 3), CDKN1A (cyclin-dependent kinase inhibitor 1A), HERPUD1 (homocysteine-inducible ER protein with ubiquitin-like domain 1), and GADD45B (growth arrest and DNA damage-inducible beta); (v) “metabolism and energy homeostasis” embraces GLUL (glutamate-ammonia ligase), NAMPT (nicotinamide phosphoribosyltransferase), IRS2 (insulin receptor substrate 2), ACSL1 (acyl-CoA synthetase long-chain family member 1), PFKFB3 (6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 3), and SLC2A3 (solute carrier family 2 member 3) (Fig. 4B). Interestingly, based on EnrichR36 analysis of the 232 common target genes between VitDPAS and VitDHiD (Fig. 3A), TNF signaling is the top ranking pathway in the database KEGG (Kyoto Encyclopedia of Genes and Genomes)37. Furthermore, according to STRING analysis (Supplementary Figure S8 online), NFKBIA, HIF1A, TNFAIP3, GADD45B, CXCR4, NFKBIZ, CDKN1A, and CSRNP1 represent the most well-characterized core within this protein set. NFKBIA, NFKBIZ, and TNFAIP3 function downstream of stress sensors, including GADD45D (responsive to DNA damage), CDKN1A (cell cycle regulation), HIF1A (hypoxia response), and CXCR4 (immune activation) (Fig. 4B). Interestingly, in the majority of healthy individuals across both studies (84.3%), the genes encoding these proteins are downregulated in response to vitamin D3 supplementation, a trend that also holds for the average expression of the full set of 26 genes (Supplementary Figure S5 online). However, in a subset of 7 individuals, this gene set is upregulated, indicating an opposite effect of vitamin D3 for these persons. In summary, the functional classification of 26 common highly significant target genes emphasizes a range of biological roles influenced by vitamin D3 supplementation. A central outcome of this regulation is the downregulation of vitamin D target genes involved in inflammatory responses, primarily through TNF and NFκB signaling pathways.
Discussion
The primary aim of this study was to validate and expand the concept of the vitamin D response index in a European cohort distinct from the one where it was originally developed22. Unlike other studies that examined gene expression changes following vitamin D3 supplementation over an 8-week period38, the VitDPAS study employed a different approach: administering a single vitamin D3 bolus and taking a second measurement just one day later. This methodology facilitates direct comparison with in vitro studies, which traditionally use 24-hour stimulations with 1,25(OH)₂D₃39. By bridging these experimental paradigms, the VitDPAS findings offer valuable insights into vitamin D’s gene-regulatory effects on immune function in healthy individuals, potentially advancing our understanding in this area.
The VitDPAS study described here, conducted with a Polish cohort, represents an advanced extension of our previous VitDHiD study26 performed in Kuopio, Finland. Both studies included healthy participants; however, the 45 VitDPAS participants were, on average, 12.1 years older and had a 1.1 kg/m2 higher BMI compared to the 25 VitDHiD participants. Despite these differences, the distribution of high, mid, and low vitamin D responders was similar across both studies, with rates of 37.8%, 42.2%, and 20% in VitDPAS, closely aligning with 36%, 48%, and 16% in VitDHiD, respectively. Interestingly, when combining data from the 70 participants of both studies, the overall response rates were 40% high responders, 38.6% mid responders, and 21.4% low responders. For comparison, a similarly designed study conducted with 50 male and 50 female participants in Jeddah, Saudi Arabia (21°N)40,41, identified 22% high responders, 39% mid responders, and 39% low responders42. This indicates that the proportion of low responders in a population living closer to the equator is nearly double that observed in Northern or Central European populations.
The regions of the two vitamin D intervention studies, Olsztyn (53°N) and Kuopio (63°N), experience extended “vitamin D winters”43,44 lasting approximately 4 and 5 months, respectively. These are periods during which endogenous vitamin D3 production is not possible. This is particularly critical for individuals classified as low vitamin D responders, who require adequate vitamin D3 supplementation during the winter months to prevent health issues associated with vitamin D deficiency45. Although, daily vitamin D3 supplementation of up to 4000 IU is considered safe46, we recommend VitDPAS participants to adjust their daily intake during the winter months according to their body weight, ensuring it does not exceed 40 IU of vitamin D3/kg. The VitDPAS study is currently ongoing, with participants being supplemented and monitored over a total duration of two years.
Previous reports have highlighted significant interindividual variability in the sets of in vivo vitamin D target genes26. The transcriptome analysis of the 45 participants in the VitDPAS study supports this finding. Out of the 8,397 protein-coding genes expressed at a reasonable level (CPM > 10) in the PBMCs of all participants, 758 (9.0%) exhibited a significant (p < 0.05) response to vitamin D. However, only 232 of these vitamin D target genes (30.6%) overlapped with those identified in the VitDHiD study. While the overlap increases to 37.1% when using more stringent statistical criteria (FDR < 0.05), this still suggests that only a small subset of in vivo vitamin D targets (26 genes) is consistently identified across both studies with high confidence.
An intriguing observation from the VitDPAS study is that, although the initial list of 758 vitamin D target genes included an approximately equal number of upregulated and downregulated genes, only 4 of the 26 genes in the final shortlist were upregulated. This finding suggests that the most critical targets of vitamin D in healthy individuals are more likely to be downregulated than elevated. This aligns with previous findings from the VitDHiD study47, which indicated that vitamin D-triggered pathways are predominantly downregulated or maintained in equilibrium, rather than being actively upregulated.
In a previous study utilizing VitDHiD data, the genes NFKBIA, NFKBIZ, and TNFAIP3 were identified as top-ranking players in eight signaling pathways related to innate immunity48. Interestingly, the downregulation of NFKBIA and NFKBIZ, both inhibitors of NFκB, promotes the translocation of the transcription factor NFκB from the cytoplasm to the nucleus, thereby activating multiple proinflammatory target genes. Moreover, downregulation of TNFAIP3, an upstream inhibitor of TNF-mediated inflammatory pathways, may further amplify the inflammatory response. These findings suggest that vitamin D enhances the sensitivity of inflammatory signaling pathways to external triggers, thereby facilitating a more efficient response when necessary.
In conclusion, the VitDPAS study validated and expanded upon the principles of the vitamin D response index. It underscored the need for special attention to low vitamin D responders within this Polish cohort, ensuring adequate vitamin D3 supplementation during the winter months. The combined analysis of the VitDPAS and VitDHiD studies identified 26 high-confidence vitamin D target genes, whose core function is the regulation of inflammatory responses in response to external stress sensors.
Data availability
Fastq files of the raw data can be found at Gene Expression Omnibus (GEO, http://www.ncbi.nlm.nih.gov/geo) with accession number GSE278885.
Abbreviations
- 1,25(OH)2D3 :
-
1α,25-dihydroxyvitamin D3
- 25(OH)D3 :
-
25-hydroxyvitamin D3
- ACSL1:
-
Acyl-CoA synthetase long chain family member 1
- ATAC-seq:
-
assay for transposase accessible chromatin with high-throughput sequencing
- BMI:
-
Body mass index
- CDKN1A:
-
Cyclin dependent kinase inhibitor 1 A
- CPM:
-
Counts per million
- CSRNP1:
-
cysteine and serine rich nuclear protein 1
- CX3CR1:
-
C-X3-C motif chemokine receptor 1
- CXCR4:
-
C-X-C motif chemokine receptor 4
- ETS2:
-
ETS proto-oncogene 2, transcription factor
- FC:
-
Fold change
- FDR:
-
False discovery rate
- GADD45B:
-
Growth arrest and DNA damage inducible beta
- GIMAP6:
-
GTPase, IMAP family member 6
- GLUL:
-
Glutamate-ammonia ligase
- HERPUD1:
-
Homocysteine inducible ER protein with ubiquitin like domain 1
- HGNC:
-
Human genome nomenclature committee
- HIF1A:
-
Hypoxia inducible factor 1 subunit alpha
- IRS2:
-
Insulin receptor substrate 2
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- KLF:
-
KLF transcription factor
- MDS:
-
Multidimensionality scaling
- NAMPT:
-
Nicotinamide phosphoribosyltransferase
- NFκB:
-
Nuclear factor κB
- NFKBI:
-
NFkB inhibitor
- PBMC:
-
Peripheral blood mononuclear cell
- PELI1:
-
Pellino E3 ubiquitin protein ligase 1
- PER1:
-
Period circadian regulator 1
- PFKFB3:
-
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 3
- RBM7:
-
RNA binding motif protein 7
- RNA-seq:
-
RNA sequencing
- S1PR1:
-
Sphingosine-1-phosphate receptor 1
- SLC2A3:
-
Solute carrier family 2 member 3
- TNF:
-
Tumor necrosis factor
- TNFAIP3:
-
TNF alpha induced protein 3
- TNFSF14:
-
TNF superfamily member 14
- TSC22D3:
-
TSC22 domain family member 3
- TSS:
-
Transcription start site
- VDR:
-
Vitamin D receptor
References
Holick, M. F., MacLaughlin, J. A. & Doppelt, S. H. Regulation of cutaneous previtamin D3 photosynthesis in man: skin pigment is not an essential regulator. Science 211, 590–593. https://doi.org/10.1126/science.6256855 (1981).
Martens, P. J., Gysemans, C., Verstuyf, A. & Mathieu, A. C. Vitamin D’s effect on immune function. Nutrients 12 https://doi.org/10.3390/nu12051248 (2020).
Verway, M. et al. Vitamin D induces interleukin-1beta expression: paracrine macrophage epithelial signaling controls M. tuberculosis infection. PLoS Pathog. 9, e1003407. https://doi.org/10.1371/journal.ppat.1003407 (2013).
Dimitrov, V. et al. Vitamin D-regulated gene expression profiles: species-specificity and cell-specific effects on metabolism and immunity. Endocrinology 162 https://doi.org/10.1210/endocr/bqaa218 (2021).
Bishop, E., Ismailova, A., Dimeloe, S. K., Hewison, M. & White, J. H. Vitamin D and immune regulation: antibacterial, antiviral, anti-inflammatory. JBMR Plus. 5, e10405. https://doi.org/10.1002/jbm4.10405 (2021).
Harrison, S. R. et al. Autoimmune disease and rheumatoid arthritis. Calcif Tissue Int. 106, 58–75. https://doi.org/10.1007/s00223-019-00577-2 (2020).
Sunyecz, J. A. The use of calcium and vitamin D in the management of osteoporosis. Ther. Clin. Risk Manag. 4, 827–836. https://doi.org/10.2147/tcrm.s3552 (2008).
Uchitomi, R., Oyabu, M. & Kamei, Y. Vitamin D and sarcopenia: potential of vitamin D supplementation in sarcopenia prevention and treatment. Nutrients 12 https://doi.org/10.3390/nu12103189 (2020).
Feige, J. et al. Vitamin D supplementation in multiple sclerosis: a critical analysis of potentials and threats. Nutrients 12 https://doi.org/10.3390/nu12030783 (2020).
Vaughan-Shaw, P. G. et al. The effect of vitamin D supplementation on survival in patients with colorectal cancer: systematic review and meta-analysis of randomised controlled trials. Br. J. Cancer. 123, 1705–1712. https://doi.org/10.1038/s41416-020-01060-8 (2020).
Bouillon, R. et al. The health effects of vitamin D supplementation: evidence from human studies. Nat. Rev. Endocrinol. 18, 96–110. https://doi.org/10.1038/s41574-021-00593-z (2022).
Gospodarska, E., Ghosh Dastidar, R. & Carlberg, C. Intervention approaches in studying the response to vitamin D3 supplementation. Nutrients 15 https://doi.org/10.3390/nu15153382 (2023).
Whitfield, G. K. et al. Cloning of a functional vitamin D receptor from the lamprey (Petromyzon marinus), an ancient vertebrate lacking a calcified skeleton and teeth. Endocrinology 144, 2704–2716. https://doi.org/10.1210/en.2002-221101 (2003).
Haussler, M. R. et al. Vitamin D receptor: molecular signaling and actions of nutritional ligands in disease prevention. Nutr. Rev. 66, 98–112. https://doi.org/10.1111/j.1753-4887.2008.00093.x (2008).
Chun, R. F., Liu, P. T., Modlin, R. L., Adams, J. S. & Hewison, M. Impact of vitamin D on immune function: lessons learned from genome-wide analysis. Front. Physiol. 5, 151. https://doi.org/10.3389/fphys.2014.00151 (2014).
Hollis, B. W. Circulating 25-hydroxyvitamin D levels indicative of vitamin D sufficiency: implications for Establishing a new effective dietary intake recommendation for vitamin D. J. Nutr. 135, 317–322 (2005).
Holick, M. F. et al. Evaluation, treatment, and prevention of vitamin D deficiency: an endocrine society clinical practice guideline. J. Clin. Endocrinol. Metab. 96, 1911–1930. https://doi.org/10.1210/jc.2011-0385 (2011).
Pike, J. W., Meyer, M. B., Lee, S. M., Onal, M. & Benkusky, N. A. The vitamin D receptor: contemporary genomic approaches reveal new basic and translational insights. J. Clin. Invest. 127, 1146–1154. https://doi.org/10.1172/JCI88887 (2017).
GTEx-Consortium. The Genotype-Tissue expression (GTEx) project. Nat. Genet. 45, 580–585. https://doi.org/10.1038/ng.2653 (2013).
Campbell, M. J. Vitamin D and the RNA transcriptome: more than mRNA regulation. Front. Physiol. 5, 181. https://doi.org/10.3389/fphys.2014.00181 (2014).
Seuter, S. et al. Molecular evaluation of vitamin D responsiveness of healthy young adults. J. Steroid Biochem. Mol. Biol. 174, 314–321. https://doi.org/10.1016/j.jsbmb.2016.06.003 (2017).
Carlberg, C. & Haq, A. The concept of the personal vitamin D response index. J. Steroid Biochem. Mol. Biol. 175, 12–17. https://doi.org/10.1016/j.jsbmb.2016.12.011 (2018).
Carlberg, C. & Mycko, M. P. Linking mechanisms of vitamin D signaling with multiple sclerosis. Cells 12 https://doi.org/10.3390/cells12192391 (2023).
Vukic, M. et al. Relevance of vitamin D receptor target genes for monitoring the vitamin D responsiveness of primary human cells. PLoS ONE. 10, e0124339. https://doi.org/10.1371/journal.pone.0124339 (2015).
Seuter, S. et al. Molecular evaluation of vitamin D responsiveness of healthy young adults. J. Steroid Biochem. Mol. Biol. 174, 314–321 (2017).
Hanel, A. et al. Common and personal target genes of the micronutrient vitamin D in primary immune cells from human peripheral blood. Sci. Rep. 10, 21051. https://doi.org/10.1038/s41598-020-78288-0 (2020).
Ghosh Dastidar, R. et al. In vivo vitamin D targets reveal the upregulation of focal adhesion-related genes in primary immune cells of healthy individuals. Sci. Rep. 14, 17552. https://doi.org/10.1038/s41598-024-68741-9 (2024).
Scragg, R. et al. Effect of monthly High-Dose vitamin D supplementation on cardiovascular disease in the vitamin D assessment study: A randomized clinical trial. JAMA Cardiol. 2, 608–616. https://doi.org/10.1001/jamacardio.2017.0175 (2017).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21. https://doi.org/10.1093/bioinformatics/bts635 (2013).
Liao, Y., Smyth, G. K. & Shi, W. FeatureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930. https://doi.org/10.1093/bioinformatics/btt656 (2014).
Durinck, S., Spellman, P. T., Birney, E. & Huber, W. Mapping identifiers for the integration of genomic datasets with the R/Bioconductor package biomart. Nat. Protoc. 4, 1184–1191. https://doi.org/10.1038/nprot.2009.97 (2009).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. EdgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140. https://doi.org/10.1093/bioinformatics/btp616 (2010).
Chen, Y., Lun, A. T. & Smyth, G. K. From reads to genes to pathways: differential expression analysis of RNA-Seq experiments using Rsubread and the edger quasi-likelihood pipeline. F1000Research 5, 1438. https://doi.org/10.12688/f1000research.8987.2 (2016).
Carlberg, C. et al. Primary vitamin D target genes allow a categorization of possible benefits of vitamin D3 supplementation. PLoS ONE. 8, e71042. https://doi.org/10.1371/journal.pone.0071042 (2013).
Szklarczyk, D. et al. The STRING database in 2023: protein-protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res. 51, D638–D646. https://doi.org/10.1093/nar/gkac1000 (2023).
Chen, E. Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinform. 14 https://doi.org/10.1186/1471-2105-14-128 (2013).
Kanehisa, M., Furumichi, M., Tanabe, M., Sato, Y. & Morishima, K. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 45, D353–D361. https://doi.org/10.1093/nar/gkw1092 (2017).
Hossein-nezhad, A., Spira, A. & Holick, M. F. Influence of vitamin D status and vitamin D3 supplementation on genome wide expression of white blood cells: a randomized double-blind clinical trial. PLoS ONE. 8, e58725. https://doi.org/10.1371/journal.pone.0058725 (2013).
Hanel, A. & Carlberg, C. Time-resolved gene expression analysis monitors the regulation of inflammatory mediators and Attenuation of adaptive immune response by vitamin D. Int. J. Mol. Sci. 23 https://doi.org/10.3390/ijms23020911 (2022).
AlGhamdi, S. A. et al. A single oral vitamin D3 bolus reduces inflammatory markers in healthy Saudi males. Int. J. Mol. Sci. 23 https://doi.org/10.3390/ijms231911992 (2022).
Alsufiani, H. M. et al. A single vitamin D3 bolus supplementation improves vitamin D status and reduces Proinflammatory cytokines in healthy females. Nutrients 14 https://doi.org/10.3390/nu14193963 (2022).
AlGhamdi, S. A. et al. Evaluation of the vitamin D response index in a Saudi cohort. Saudi Pharm. J. 32, 102137. https://doi.org/10.1016/j.jsps.2024.102137 (2024).
Ohlund, I., Lind, T., Hernell, O. & Silfverdal, S. A. Karlsland Akeson, P. Increased vitamin D intake differentiated according to skin color is needed to Meet requirements in young Swedish children during winter: a double-blind randomized clinical trial. Am. J. Clin. Nutr. 106, 105–112. https://doi.org/10.3945/ajcn.116.147108 (2017).
Khanna, T. et al. Comprehensive analysis of seasonal and geographical variation in UVB radiation relevant for vitamin D production in Europe. Nutrients 14 https://doi.org/10.3390/nu14235189 (2022).
Holick, M. F. Vitamin D deficiency. N Engl. J. Med. 357, 266–281. https://doi.org/10.1056/NEJMra070553 (2007).
Johnson, K. C. et al. Safety and tolerability of high-dose daily vitamin D3 supplementation in the vitamin D and type 2 diabetes (D2d) study-a randomized trial in persons with prediabetes. Eur. J. Clin. Nutr. 76, 1117–1124. https://doi.org/10.1038/s41430-022-01068-8 (2022).
Jaroslawska, J. & Carlberg, C. In vivo regulation of signal transduction pathways by vitamin D stabilizes homeostasis of human immune cells and counteracts molecular stress. Int. J. Mol. Sci. 24, 14632. https://doi.org/10.3390/ijms241914632 (2023).
Jaroslawska, J., Ghosh Dastidar, R. & Carlberg, C. In vivo vitamin D target genes interconnect key signaling pathways of innate immunity. PLoS ONE. 19, e0306426. https://doi.org/10.1371/journal.pone.0306426 (2024).
Acknowledgements
Kind thanks to the Gene Core Facility at the EMBL in Heidelberg, Germany, for massively parallel sequencing services, the Centre of Informatics Tricity Academic Supercomputer and Network of the Technical University of Gdansk (https://task.gda.pl/en) for the use of their the supercomputers for sequence alignment and Procter & Gamble Health Poland for donating Vigantoletten Max 4000 to our study.
Funding
This publication is part of a project that has received funding from the European Union’s Horizon2020 research and innovation program under grant agreement no. 952601, from the David and Amy Fulton Foundation, Seattle, US and from the National Science Centre (Poland), project number 2023/49/B/NZ9/00402.
Author information
Authors and Affiliations
Contributions
The VitDPAS study was designed and conducted by E.G., M.Ra., K.T.M., J.R., and C.C.. Data analysis and visualization were performed by R.G.D., M.Ry. and J.J., while RNA isolation and RNA-seq library preparation were carried out by E.G. and C.C.. C.C. refined the figures and drafted the manuscript, which was reviewed and approved by all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gospodarska, E., Dastidar, R.G., Jaroslawska, J. et al. Transcriptomic profiling of immune modulation induced by vitamin D3 in the VitDPAS and VitDHiD cohort studies. Sci Rep 15, 17334 (2025). https://doi.org/10.1038/s41598-025-02495-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-02495-w
Keywords
This article is cited by
-
25-Hydroxyvitamin D3 metabolism modulates the effect of variable UV exposure on 25-hydroxyvitamin D3 plasma concentrations
Communications Medicine (2026)
-
Chromatin and transcriptional dynamics underlying the immune-modulatory effects of vitamin D3 in vivo
Scientific Reports (2025)