Abstract
Aspergillus fumigatus, which causes aspergillosis, has developed resistance to azole antifungal agents in recent years. As only three main classes of antifungal drugs are available, the development of novel therapeutic strategies is crucial. We aimed to control the expression of virulence factors by introducing microRNAs (miRNAs) into fungi as an innovative therapeutic approach. To test our hypothesis, we selected miRNA mimics targeting alb1, which is involved in the synthesis of 1,8-dihydroxynaphthalene (DHN)-melanin, a virulence factor of A. fumigatus, and transfected them into the protoplast of the fungus, resulting in a two-fold reduction in alb1 expression. Next, we created a 3×HA-tagged Alb1 protein (Alb1-HAp)-expressing strain and confirmed the regulation of translation using western blotting with an anti-HA antibody. The protein amount of Alb1-HAp was reduced by one-third after the introduction of the miRNA. Moreover, the reduction in melanin after miRNA transfection promoted the killing of fungus by hydrogen peroxide-induced oxidative stress and sensitised the fungus to neutrophil attack. Additionally, by loading miRNAs into a fungus-targeted delivery system, we demonstrated the potential of transferring miRNAs into intact fungal cells in vitro. These results indicate the potential of miRNAs to regulate target virulence factors in fungi, leading to the development of novel therapies.
Similar content being viewed by others
Introduction
Aspergillus spp. are widespread in the environment, with Aspergillus fumigatus being the primary pathogen that causes invasive aspergillosis. Invasive aspergillosis is estimated to occur annually in > 2.1 million patients with underlying diseases, with a mortality rate of > 80%1. However, the available antifungal drugs have moderate efficacy, and A. fumigatus is resistant to azoles2. The importance of this issue has recently been recognised after the World Health Organization listed A. fumigatus as the most important fungal pathogen that poses a public health threat in the fungal priority pathogen list3. Therefore, there is an urgent need to develop treatments based on mechanisms that differ from those of the existing drugs.
MicroRNAs (miRNAs) are short RNAs of approximately 20–23 nucleotides that control gene expression in many cellular processes4,5. miRNAs are involved in RNA interference (RNAi), and various therapeutic developments that use them for cancer and infectious diseases are underway6,7. RNAi treatments are attractive therapeutic options for fungi8. For example, RNAi systems are present in the closely related species A. nidulans and A. flavus9,10,11. Furthermore, in A. fumigatus, double-stranded RNA reduced the expression of target genes, suggesting the potential use of RNAi systems in this fungus12,13. In the cross-talk between fungi and other species, fungal gene expression is regulated by the transfer of miRNAs through extracellular vesicles released by other organisms14,15,16,17. Although exosomes have been found to move between species of A. fumigatus, there have been no reports indicating that their miRNAs affected the regulation of gene expression18. Although RNAi mechanisms are powerful tools that are thought to be able to suppress fungal growth and have potential therapeutic applications, the delivery of RNA to the fungus is a challenge.
A drug-delivery system (DDS) is a promising carrier for RNA transport. Liposomes and PEG-poly(amino acid)-based micelles are DDS platforms that have long been clinically investigated19. Additionally, these DDS materials have good biocompatibility, low immunogenicity, and can encapsulate medical compounds to enhance their in vivo performance19,20,21. We have successfully constructed micelles based on PEG-poly(amino acid) copolymers to create DDS devices comparable to the size of fungal exosomes. These devices have the property of high fungus-specific accumulation and accessibility to fungi owing to the flexibility of chemical modification of the surface structure and the binding of the dectin-1 molecule22. Additionally, the devices can be loaded with nucleic-acid molecules used for RNAi. Taking advantage of these properties, DDS devices can be loaded with nucleic acids, enabling the transport of nucleic acid molecules to fungi, which is expected to regulate fungal gene expression.
Herein, we introduced miRNA mimics targeting alb1, which is involved in 1,8-dihydroxynaphthalene (DHN)-melanin synthesis, a virulence factor of A. fumigatus, and has been employed as a target for RNAi in previous reports and confirmed that it suppresses target expression23,24,25,26. Furthermore, we confirmed the reduction of resistance to hydrogen peroxide H2O2 stress and the improvement of elimination ability by neutrophils due to the decreased expression of the target gene by miRNA introduction. Finally, we attempted to introduce miRNA mimics into the intact fungi with cell walls using our created DDS devices and observed phenotypic changes in which miRNAs were introduced, and alb1 expression was suppressed.
Results
Human miRNA candidate against alb1 of A. fumigatus
Candidate human miRNAs used were selected using miRbase27, based on the alb1 exon-only sequence information of strain Af293, a genome strain of A. fumigatus in the FungiDB (https://fungidb.org/fungidb/app). The selected candidate miRNAs are shown in Table 1. Two candidate miRNAs, hsa-miR-3667-5p and hsa-miR-4738-3p, were predicted to have the highest complementarity with the alb1 exon-only sequence (Fig. 1).
Confocal microscopic image of miRNA uptake in A. fumigatus
Next, fluorescence-labelled miRNA mimics of hsa-miR-3667-5p and hsa-miR-4738-3p were prepared to confirm whether the candidate miRNAs had been transfected, and their introduction into the protoplasts was observed. We confirmed that miRNAs were introduced into the protoplasts, and miRNA fluorescence was observed in protoplasts or mycelia 3–24 h after miRNA introduction. As similar fluorescence images were obtained in both cases, results are only shown for the conditions in which miRNA hsa-miR-3667-5p was used as transfection (Fig. 2).
Confocal microscopic image of miRNA uptake in Aspergillus fumigatus. (A) Three hours after introducing fluorescence-labeled miRNAs into the protoplasts, the cells were fixed and photographed using a confocal laser microscope. Bar: 20 µm. (B) Mycelia were photographed 24 h after the introduction of fluorescence-labelled miRNAs. miRNAs indicated fluorescence images of fluorescein, and nuclei indicated fluorescence images of 4′,6-diamidino-2-phenylindole. Bar: 10 µm.
miRNA mimics regulated targeted gene expression and transcription
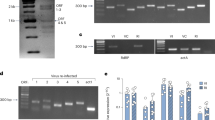
The target gene expression in the mycelium was evaluated using reverse transcription-PCR 3 h and 12 h after the introduction of the two miRNAs and negative-control miRNA. At 3 h after transfection, alb1 expression was unchanged compared with that in the untransfected strain. At 12 h after miRNA transfection, the expression of target alb1 decreased 3.7-fold at 10 nM and 2.1-fold at 40 nM of hsa-miR-3667-5p and decreased 1.1-fold at 10 nM and 1.9-fold at 40 nM of hsa-miR-4738-3p but was not affected by the negative-control miRNA (Fig. 3A). Additionally, we evaluated whether miRNA transfection can lead to the translational regulation of the target gene. To quantify the protein translated from the target gene, we generated an Alb1_HA strain in which a 3×HA-tag was fused to the target protein (Supplementary Methods). The strains were compared to each condition using western blotting with an anti-HA antibody. Mycelium collected after 24 h of incubation following 24 h post-transfection of the Alb1-HA strain with 40 nM hsa-miR-3667-5p showed a 0.3-fold decrease in Alb1-Hap expression, confirming translational regulation (Fig. 3B).
miRNA mimics regulated targeted gene expression and transcription. (A) Expression of alb1 was examined at 12 h after the introduction of negative-control miRNA, hsa-miR-3667-5p or hsa-miR-4738-3p. The graph represents mean ± SEM. Statistical analysis: one-way ANOVA, ns = not significant, asterisk * or *** = p < 0.05 or p < 0.001, respectively. (B) At 24 h after the introduction of negative-control miRNA, hsa-miR-3667-5p and hsa-miR-4738-3p, Alb1-HAp was evaluated using western blotting with the anti-HA antibody; relative ratios are shown.
miRNA mimic-regulated conidia promoted decreased resistance to hydrogen peroxide and elimination by neutrophils
To confirm the quality of the formed conidia, we evaluated their antioxidant capacity in response to 1.1 mM hydrogen peroxide (H2O2) and their fungus-killing ability via neutrophils isolated from mice. Conidia harvested from the miRNA-untransfected AfS35 strain and the negative-control miRNA-transfected AfS35 strain showed no effect on CFUs, with or without exposure to 1.1 mM H2O2. In conidia harvested from AfS35 cells transfected with the respective miRNAs, a 20–25% reduction in CFUs was observed upon exposure to H2O2 (Fig. 4A). The CFUs of harvested conidia from the AfS35 strain without miRNA decreased by 38% when co-cultured with neutrophils. When the AfS35 strain was transfected with hsa-miR-3667-5p and hsa-miR-4738-3p, and the harvested conidia were co-cultured with neutrophils for 3 h, the CFUs decreased by 58% and 59%, respectively (Fig. 4B). This suggests that partial and complete regulation (knockout) of alb1 reduces the intrinsic antioxidant capacity and enhances antifungal killing by mouse neutrophils. In other words, miRNA transfection regulates the quality of conidiation by decreasing the expression and translation of target factors.
miRNA mimic-regulated conidia promoted decreased resistance to hydrogen peroxide and elimination by neutrophils. (A) Conidia harvested from each strain were treated with or without 1.1 mM H2O2 for 24 h, incubated on PDA, and CFUs were measured. The graph represents mean ± SEM. Statistical analysis: two-way ANOVA, ns = not significant, asterisk * or ** = p < 0.05 or p < 0.01, respectively. (B) CFUs were measured when each harvested conidia under each condition were co-incubated with or without neutrophils isolated from mice. The percentage decrease in CFU due to neutrophil co-culture is shown as the percent killing. The graph represents mean ± SEM. Statistical analysis: one-way ANOVA.
miRNA introduction into intact fungi using a Dectin1-installed micelle-loaded miRNA
However, to apply RNAi technology for the treatment of fungal diseases, it is necessary to introduce miRNAs into intact fungi, where the cell wall is present. We have developed polyethylene glycol (PEG)-poly(amino acid)-based micelles, including PEG-poly(L-lysine), as effective DDS nanocarriers. Additionally, we recently developed Dectin-1-installed-polymeric micelles based on PEG-amino acid block copolymers; this carrier has the property of specifically binding to the surface of the mycelium22. Therefore, we aimed to introduce miRNAs into intact fungi using Dectin-1-installed micelle-loaded miRNAs of comparable size to extracellular vesicles. Using this device, miRNAs were detected in the cytoplasm of A. fumigatus with cell walls (Fig. 5A). We found that the introduction of miRNAs by this device reduced CFUs of harvested conidia by approximately 50% after exposure to 1.1 mM H2O2 compared with the untransfected strain and Dectin-1-installed micelle-unloaded miRNA (Fig. 5B).
miRNA introduction into intact fungi using a Dectin1-installed micelle-loaded miRNA. (A) Mycelia were photographed 24 h after the introduction of fluorescence-labeled miRNA-loaded nanodevices. miRNA indicates fluorescence images of fluorescein, and the cell wall indicates fluorescence images of Calcofluor White stain. Bar: 10 μm. (B) Conidia harvested from each strain were treated with or without 1.1 mM H2O2 for 24 h, incubated on PDA, and CFUs were measured. The graph represents mean ± SEM. Statistical analysis: two-way ANOVA, ns = not significant, asterisks * or ** = p < 0.05, or p < 0.01, respectively. Abbreviations: ANOVA, analysis of variance; SEM, standard error of the mean, H2O2, hydrogen peroxide; PDA, potato dextrose agar.
Discussion
Currently, antifungal drugs utilised in the treatment of aspergillosis comprise three classes of compounds: two that target ergosterol (azoles and polyenes) and another (echinocandins) that inhibits the synthesis of β-1,3 glucan, an important component of the cell wall. Furthermore, in recent years, azole-resistant Aspergillus spp. have emerged from the environment and patients2. Hence, there is a concern about being further limited to a few antifungal drugs. To resolve this problem, we attempted to control the expression of virulence factors by using RNAi as a new candidate for therapeutic agents. By introducing two miRNA mimics, we were able to reduce the expression level of the target and obtained results that support the existence of an RNAi system in this fungus, which has been confirmed previously. However, although the mechanism of RNAi has been confirmed and RNAi-based expression control has been shown to be a powerful tool for Aspergillus spp. infection, the challenge has been to enable RNA delivery8. Herein, we showed that polymeric micelles enable the introduction of miRNA into the intact cell wall of A. fumigatus.
Because alb1 was reported as a target for gene suppression by the RNAi system in A. fumigatus, we also selected candidate miRNA mimics targeting alb1 and introduced them into protoplasts25,26. Both miRNA mimics reduced the expression of target genes, while the negative-control miRNA had no effect on transfection (Fig. 3A). Notably, only 40 nM hsa-miR-3667-5p demonstrated translational regulation, despite both miRNA mimics showing transcriptional effects. This suggests that high hsa-miR-3667-5p is a particularly effective interfering molecule, as it achieved transcriptional regulation even at 10 nM(Fig. 3B). Due to sequence differences between miRNA mimics despite targeting the same gene, optimising both concentration and timing is essential to accurately assess their effects. The miRNA-transfected strains did not change to white colonies, a phenotype similar to that of Δalb1 gene deletion (Supplementary Fig. 1A), but instead showed a phenotype involving reduced resistance to hydrogen peroxide, increased susceptibility to neutrophil exclusion and decreased melanin function, which are properties associated with resistance to oxidative stress (Fig. 4). The introduced miRNA mimics potentially cause partial expression of targeted gene expression, differing from the phenotype of complete deletant strain. Since conidia formed after miRNA mimic introduction exhibited qualitative changes, the melanin-containing fraction was extracted from each conidiospore using alkali and acid treatment and quantified relative to controls by UV. The results showed a slight decrease in melanin content in each fraction compared to the WT (Supplementary Fig. 1B, C), although the absolute quantification of 1,8-dihydroxynaphthalene (DHN)-melanin and its association with oxidative stress remains to be assessed.
The therapeutic application of RNAi systems against Aspergillus spp. has been avoided because of the difficulties in transporting bare nucleic acids into the cytoplasm of Aspergillus8. However, despite the presence of cell walls, tracking of miRNAs through exosome-like vesicles has been observed in some plants and some fungi15,16,28. There have also been reports suggesting that exosomes released by fungi are transferred between A. fumigatus18. We prepared micelles comparable to the size of exosomes released by fungi29 and loaded labeled miRNA mimics into the micelles22,30,31, and we investigated whether it is possible to introduce miRNA into Aspergillus with intact cell walls by loading miRNA mimics into micelles. Green fluorescence in the cytoplasm was not observed in the fungi un-transfected or transfected only with the miRNA unloaded DDS device; fluorescence in the cytoplasm was observed only in the fungi treated with labeled miRNA mimics in the DDS device (Fig. 5). In other words, the possibility that this was a stress response (autofluorescence) due to the treatment herein was ruled out; indeed, the fluorescence of the RNA fluorescently labeled using vesicles was confirmed in the compartment inside the cell wall, suggesting that the nucleic acid molecules passed through the cell wall.
One limitation of this study is that off-target effects must be considered for RNAi-based methods. We observed phenotypes associated with reduced gene expression and translation upon introducing the miRNA against alb1. However, there was a concern about the off-target effect of miRNA introduction. Therefore, we attempted the method reported by Halder et al.16, which has been reported to demonstrate direct binding of the introduced miRNA to the target mRNA; however, we were unable to replicate this result (data not shown). For effective treatment using RNAi, it is necessary to study the turnover of miRNAs and the appropriate dosage and frequency of administration in vivo. The pharmacokinetics of the micelle itself used herein are well understood and its safety is assured, making it an excellent carrier option; however, understanding the development of toxicity to the host due to the introduction of miRNAs is necessary19,20,21. A limitation of our study is the incomplete in vivo approach. While we observed indicative changes in pathogenic factors in vitro, therapeutic experiments in animal models of infections remain a future challenge. Addressing these issues is expected to bring us closer to therapeutic applications; however, clarification of the mechanisms, such as how nucleic acids are taken up into the fungi by using vesicles, is likely to be the focus of basic biology in the future.
In conclusion, we attempted to regulate the expression of fungal virulence factors using microRNAs (miRNAs). We confirmed that miRNAs can be introduced into intact fungi using DDS-based carriers. This research provides a new therapeutic concept by controlling the expression of pathogenic factors in fungi using miRNAs.
Methods
Ethics
All animal experiments were approved by the Animal Experiment Committee of Tokyo Medical University and were conducted in accordance with animal experiment regulations.
Strains and culture conditions
Strains and primers used in this study are listed in Tables 2 and 3, respectively. The construction of the deletion and complementation strains as well as conditions of the media used in this study are described in the Supplementary Material.
Plasmid preparation and construction
The preparation and construction of plasmids are described in the Supplementary Material.
Conidia preparation
Conidia from each strain were stored in a deep freezer at − 80 °C in phosphate-buffered saline (PBS) containing 20% glycerol. Fresh conidia were harvested after 4 days of culturing in conidial suspensions at 37 °C in PDA medium with PBS containing 0.1% Tween 80. The conidia were passed through a 40-µm EASYstrainer (Greiner Bio-One Co. Ltd., Tokyo, Japan) to remove hyphae. Conidia concentrations in the prepared suspensions were measured using a hemocytometer.
miRNA mimics and fluorescence labeling
The selection of miRNAs was performed using miRbase (miRbase.org)27, based on the alb1 sequence information of strain Af293, the A. fumigatus genome standard strain in the FungiDB (https://fungidb.org/fungidb/app). Two human miRNAs with low e-values were selected based on their high index of complementation and binding. The mirVana™ miRNA mimic of hsa-miR-3667-5p 5′-UGAAACUGGAGCGCCUGGAGGA-3′, hsa-miR-4738-3p 5′-AAAGACCCAUUGGAGGAGAAGGU-3′ and Negative Control #1 were purchased from Thermo Fisher Scientific. We added 50 µL of Tracker Reconstitution Solution, supplied with the kit, to Label IT Tracker Reagent (Takara Bio Inc., Shiga, Japan), and the suspension was vortexed to dissolve completely. Then, miRNA was added to Label IT Reagent and incubated at 37 °C for 1 h. After ethanol precipitation treatment, the fluorescence-labeled nucleic acids were collected.
Transfection of MiRNA mimics and microscopy
The miRNA mimics were co-incubated with the protoplasts in PEG 4000. The Dectin-1 device-loaded miRNA was co-incubated with mycelium in RPMI 1640 medium (Invitrogen Japan, Tokyo, Japan) for 6–12 h. The protoplasts or mycelia were grown in µ-Dish 35 mm (ibidi GmbH, Gräfelfing, Germany). The cell wall was stained with 2 µL of Calcofluor White stain (product code 18909, Sigma-Aldrich Japan, Tokyo, Japan) diluted in 1000 µL of PBS for 30 min at 15–25 °C. Subsequently, the cells were washed thrice using PBS. The samples were embedded in SlowFade Diamond Antifade Mountant (Invitrogen, Thermo Fisher Scientific, Waltham, MA, USA) with added 4′,6-diamidino-2-phenylindole to counterstain the nuclei. Observations were captured using a confocal laser microscope LSM700 (Carl Zeiss, Oberkochen, Germany).
RNA extraction from A. fumigatus
Mycelia were grown in RPMI 1640 medium at 37 °C under 5% CO2 conditions. After incubation, mycelia were homogenized using zirconia beads. Total RNA was extracted from the homogenized solution using TRIzol Reagent (Invitrogen, Thermo Fisher Scientific, Waltham, MA, USA), according to the manufacturer’s protocol. The extracted RNA was purified using the RNeasy Mini Kit (QIAGEN) and treated with DNase using an RNase-free DNase Set (QIAGEN).
Reverse transcription-PCR analysis of mRNA expression
cDNA was synthesized by reverse transcription reaction of 500 ng of total RNA using the SuperScript VILO cDNA Synthesis Kit (Invitrogen, Thermo Fisher Scientific, Waltham, MA, USA), according to the manufacturer’s instructions. The synthesized cDNA was used as a template, and a combination of primers RT-alb1FW and RT-alb1REV26 (Table 3) was used with TB Green Premix Ex Taq II (Tli RNaseH Plus) (Takara Bio Inc., Shiga, Japan) for reverse transcription-PCR of the target gene using Mx3000P qPCR System (Agilent Technologies Japan, Ltd., Tokyo, Japan). Relative quantification of the gene of interest was performed using beta-tubulin as an endogenous control and the ΔΔCT algorithm.
Protein extraction and Western blot analysis
After miRNA transfection, the Alb1_HA strain was cultured for 24 h at 37 °C under shaking conditions in YG liquid medium. The mycelium, including conidiophores in the liquid and on the surface, was then collected. The mycelium was lyophilised, and Sample Buffer Solution with Reducing Reagent (6×) for sodium dodecyl-sulphate polyacrylamide gel electrophoresis (SDS-PAGE) (NACALAI TESQUE, Inc., Kyoto, Japan) was used. The sample buffer was heated at 100 °C for 5 min, and SDS-PAGE was performed using SuperSep Ace, 10–20% (FUJIFILM Wako Pure Chemical Corporation, Osaka, Japan). After transfer to Immun-Blot PVDF Membrane (Bio-Rad Laboratories, Inc., Hercules, CA, USA), the membrane was blocked with PVDF Blocking Reagent for Can Get Signal (TOYOBO Co., Ltd., Osaka, Japan). Then, the membrane was washed with TBS-T (0.01% Tween 20) and incubated with an anti-HA-tag antibody (sc-7392; Santa Cruz Biotechnology, TX, USA) at a final concentration of 1 µg/mL in Can Get Signal solution 1 (TOYOBO Co., Ltd., Osaka, Japan) at 4 °C overnight. After washing thrice with TBS-T, the secondary antibody (anti-mouse IgG, HRP-linked antibody #7076; Cell Signalling Technology, Danvers, MA, USA) was diluted 15,000-fold using Can Get Signal solution 2 (TOYOBO Co., Ltd., Osaka, Japan) and reacted for 1 h at room temperature. Subsequently, the membrane was washed thrice with TBS-T and reacted with ImmunoStar LD (FUJIFILM Wako Pure Chemical Corporation, Osaka, Japan). The chemiluminescence signals were detected using C-DiGit western blot imaging (LI-COR Corporate, Lincoln, NE, USA). A relative comparison of signals was performed using Image Studio for the C-DiGit software.
Hydrogen peroxide oxidative susceptibility assay
Approximately 1.0 × 104 conidia were suspended in 10 mL of 1.1 mM hydrogen peroxide (FUJIFILM Wako Pure Chemical Corporation, Osaka, Japan) and rotated at room temperature for 24 h. Subsequently, 10 µL of the diluted solution was inoculated in triplicate onto PDA plates and incubated at 37 °C for 24 h. The colonies were visually measured.
Neutrophil killing assay
Neutrophils were collected using the Neutrophil Isolation Kit (Funakoshi Co., Ltd., Tokyo, Japan). We injected 1 mL of 7.5% Sodium Caseinate Solution Assay Reagent into the abdominal cavity of C57/B6 J mice. After 24 h, 5 mL of PBS was injected into the peritoneal cavity of the mice, and the peritoneal fluid was collected. The collected peritoneal fluid was treated with Neutrophil Isolation Medium and Red Blood Cell Lysis Buffer using Percoll Gradient Assay Reagent to isolate neutrophils. Subsequently, 1.0 × 105 aliquots were prepared using RPMI liquid medium and co-cultured for 3 h with an equal number of neutrophils. After co-cultivation, colonies were inoculated onto PDA plates and incubated at 37 °C for 24 h; colonies were visually measured. The sterilization rate was expressed as the percentage decrease in CFU in the presence of neutrophils, with CFU in the absence of neutrophils as the control. The mice used were C57BL/6J, male, 5 weeks old, purchased from CLEA Japan, Inc. All mice used for the treatment and other experiments were acclimated for at least 1 week.
Dectin-1-installed micelle-loaded miRNA Preparation
Dectin-1 Recombinant Human Dectin-1/CLEC7A Protein (hFc Tag) (Sino Biological Japan Inc., Kanagawa, Japan) was conjugated to a DBCO-PEG4-NHS linker for attachment to N3-PEG-poly(L-lysine) (pLL). Dectin-1 (100 µg) was dissolved in 50 mM NaHCO3 buffer (pH 8.6), which was exchanged twice in a 10k vivaspin tube. Ultrafiltration was conducted at 830×g for 4 min. The DBCO-PEG4-NHS linker (10 mg/mL, 5 µg) was added to the solution and incubated at 4 °C for 12 h. Then, the solution was purified with HEPES buffer in a 10k vivaspin tube. Ultrafiltration was conducted at 3000 rpm for 4 min for purification. To prepare miRNA-loaded nanocarriers, N3-PEG(12k)-pLL was dissolved in pH 8.5 HEPES buffer at 2 mg/mL concentration. Then, 140 µL of the solution was added to the miRNA solution (10 µg of miRNA was dissolved in 200 µL of nuclease-free water) and incubated for an hour at 25 °C. After incubation, a total of 300 µL of MeO-PEG-pLL(CAA) solution (2 mg/mL) was added; 15 µL was added every 5 min. After 12 h of incubation, the micelle solution was filtered through a 0.22-µm PVDF syringe filter.
Statistical analysis
Statistical analyses of biological replicates of reverse transcription-PCR data and percent killing data were performed using one-way analysis of variance (ANOVA), followed by Sidak/Dunnett’s multiple comparison tests. Analysis of hydrogen peroxide oxidative susceptibility assay data was performed using a two-way ANOVA, followed by Sidak’s multiple comparison tests. All analyses were performed using the GraphPad Prism 7 software (GraphPad Software Inc., San Diego, CA, USA).
Data availability
All data generated or analysed during this study are included in this published article and its supplementary information files.
References
Denning, D. W. Global incidence and mortality of severe fungal disease. Lancet Infect. Dis. 24, e428–e438 https://doi.org/10.1016/S1473-3099(23)00692-8 (2024).
Burks, C., Darby, A., Gómez Londoño, L., Momany, M. & Brewer, M. T. Azole-resistant Aspergillus fumigatus in the environment: Identifying key reservoirs and hotspots of antifungal resistance. PLOS Pathog. 17, e1009711. https://doi.org/10.1371/journal.ppat.1009711 (2021)
WHO fungal priority pathogens list to guide research, development, and public health action. (2022). Available from: https://www.who.int/publications/i/item/9789240060241
Bartel, B. MicroRNAs directing SiRNA biogenesis. Nat. Struct. Mol. Biol. 12, 569–571. https://doi.org/10.1038/nsmb0705-569 (2005).
Bartel, D. P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 116, 281–297 https://doi.org/10.1016/S0092-8674(04)00045-5(2004).
Borna, H., Imani, S., Iman, M. & Azimzadeh Jamalkandi, S. Therapeutic face of RNAi: In vivo challenges. Expert Opin. Biol. Ther. 15, 269–285. https://doi.org/10.1517/14712598.2015.983070 (2015).
Sayyed, A. A. et al. Role of MiRNAs in cancer diagnostics and therapy: A recent update. Curr. Pharm. Des. 28, 471–487 https://doi.org/10.2174/1381612827666211109113305(2022).
Bruch, A., Kelani, A. A. & Blango, M. G. RNA-based therapeutics to treat human fungal infections. Trends Microbiol. 30, 411–420. https://doi.org/10.1016/j.tim.2021.09.007 (2022).
Khatri, M. & Rajam, M. V. Targeting polyamines of Aspergillus Nidulans by SiRNA specific to fungal ornithine decarboxylase gene. Med. Mycol. 45, 211–220 https://doi.org/10.1080/13693780601158779 (2007).
Nami, S. et al. The utilization of RNA Silencing technology to mitigate the voriconazole resistance of Aspergillus flavus; Lipofectamine-based delivery. Adv. Pharm. Bull. 7, 53–59. https://doi.org/10.15171/apb.2017.007 (2017).
Raruang, Y. et al. Targeting the Aspergillus flavus p2c gene through host-induced gene Silencing reduces A. flavus infection and aflatoxin contamination in Transgenic maize. Front. Plant. Sci. 14, 1150086. https://doi.org/10.3389/fpls.2023.1150086 (2023).
Mouyna, I., Henry, C., Doering, T. L. & Latgé, J. P. Gene Silencing with RNA interference in the human pathogenic fungus Aspergillus fumigatus. FEMS Microbiol. Lett. 237, 317–324. https://doi.org/10.1016/j.femsle.2004.06.048 (2004).
Jöchl, C., Loh, E., Ploner, A., Haas, H. & Hüttenhofer, A. Development-dependent scavenging of nucleic acids in the filamentous fungus Aspergillus fumigatus. RNA Biol. 6, 179–186. https://doi.org/10.4161/rna.6.2.7717 (2009).
Croston, T. L., Lemons, A. R., Beezhold, D. H. & Green, B. J. MicroRNA regulation of host immune responses following fungal exposure. Front. Immunol. 9, 170. https://doi.org/10.3389/fimmu.2018.00170 (2018).
Das Gupta, M. et al. Aspergillus fumigatus induces microRNA-132 in human monocytes and dendritic cells. Int. J. Med. Microbiol. 304, 592–596. https://doi.org/10.1016/j.ijmm.2014.04.005 (2014).
Halder, L. D. et al. Candida albicans induces cross-kingdom miRNA trafficking in human monocytes to promote fungal growth. mBio 13, e0356321 https://doi.org/10.1128/mbio.03563-21 (2021).
Wang, Y., Cui, C., Wang, G., Li, Y. & Wang, S. Insects defend against fungal infection by employing microRNAs to silence virulence-related genes. Proc. Natl Acad. Sci. U. S. A. 118, e2023802118 https://doi.org/10.1073/pnas.2023802118 (2021).
Bitencourt, T. A. et al. Fungal extracellular vesicles are involved in intraspecies intracellular communication. mBio 13, e0327221 https://doi.org/10.1128/mbio.03272-21 (2022).
Cabral, H. & Kataoka, K. Progress of drug-loaded polymeric micelles into clinical studies. J. Control Release. 190, 465–476. https://doi.org/10.1016/j.jconrel.2014.06.042 (2014).
Petrini, M. et al. Effects of surface charge, pegylation and functionalization with dipalmitoylphosphatidyldiglycerol on liposome-cell interactions and local drug delivery to solid tumors via thermosensitive liposomes. Int. J. Nanomed. 16, 4045–4061. https://doi.org/10.2147/IJN.S305106 (2021).
Kataoka, K. et al. Doxorubicin-loaded poly(ethylene glycol)-poly(beta-benzyl-L-aspartate) copolymer micelles: their pharmaceutical characteristics and biological significance. J. Control Release. 64, 143–153. https://doi.org/10.1016/s0168-3659(99)00133-9 (2000).
Inukai, T., Chen, P., Kokuba, H., Cabral, H. & Nakamura, S. A novel fungal-targeted drug delivery system dectin-1-targeted-PEG-amino acid polymer block enhances antifungal activity and reduces cytotoxicity of amphotericin B. J. Drug Deliv Sci. Technol. 100, 106073 https://doi.org/10.1016/j.jddst.2024.106073 (2024).
Tsai, H. F., Chang, Y. C., Washburn, R. G., Wheeler, M. H. & Kwon-Chung, K. J. The developmentally regulated alb1 gene of Aspergillus fumigatus: its role in modulation of conidial morphology and virulence. J. Bacteriol. 180, 3031–3038 (1998). https://doi.org/10.1128/JB.180.12.3031-3038 (1998).
Jahn, B. et al. Isolation and characterization of a pigmentless-conidium mutant of Aspergillus fumigatus with altered conidial surface and reduced virulence. Infect. Immun. 65, 5110–5117. https://doi.org/10.1128/iai.65.12.5110-5117.1997 (1997).
Enayati, S., Azizi, M., Aminollahi, E., Ranjvar Shahrivar, M. & Khalaj, V. T7-RNA polymerase dependent RNAi system in Aspergillus fumigatus: a proof of concept study. FEMS Microbiol. Lett. 363, fnw029. https://doi.org/10.1093/femsle/fnw029 (2016).
Khalaj, V., Eslami, H., Azizi, M., Rovira-Graells, N. & Bromley, M. Efficient downregulation of alb1 gene using an AMA1-based episomal expression of RNAi construct in Aspergillus fumigatus. FEMS Microbiol. Lett. 270, 250–254. https://doi.org/10.1111/j.1574-6968.2007.00680.x (2007).
Kozomara, A., Birgaoanu, M. & Griffiths-Jones, S. MiRBase: From MicroRNA sequences to function. Nucleic Acids Res. 47, D155–D162. https://doi.org/10.1093/nar/gky1141 (2019).
Samuel, M., Bleackley, M., Anderson, M. & Mathivanan, S. Extracellular vesicles including exosomes in cross Kingdom regulation: A viewpoint from plant-fungal interactions. Front. Plant. Sci. 6, 766 (https://doi.org/10.3389/fpls.2015.00766 (2015).
Rizzo, J., Taheraly, A. & Janbon, G. Structure, composition and biological properties of fungal extracellular vesicles. Microlife 2, uqab009. https://doi.org/10.1093/femsml/uqab009 (2021).
Bae, Y. & Kataoka, K. Intelligent polymeric micelles from functional poly(ethylene glycol)-poly(amino acid) block copolymers. Adv. Drug Deliv Rev. 61, 768–784. https://doi.org/10.1016/j.addr.2009.04.016 (2009).
Yang, W., Chen, P., Boonstra, E., Hong, T. & Cabral, H. Polymeric micelles with pH-responsive cross-linked core enhance in vivo mRNA delivery. Pharmaceutics 14, 1205. https://doi.org/10.3390/pharmaceutics14061205 (2022).
Szewczyk, E. et al. Fusion PCR and gene targeting in Aspergillus Nidulans. Nat. Protoc. 1, 3111–3120. https://doi.org/10.1038/nprot.2006.405 (2006).
Verde-Yanez, L. et al. Identification and biosynthesis of DHN-melanin related pigments in the pathogenic fungi monilinia Laxa, M. fructicola, and M. fructigena. J. Fungi (Basel). 9, 138. https://doi.org/10.3390/jof9020138 (2023).
Acknowledgements
We would like to thank the Center for Diversity at Tokyo Medical University for supporting this study (TMUCD-202401).
Funding
This work was supported by JSPS KAKENHI Grant Numbers 24K19271 (TI).
Author information
Authors and Affiliations
Contributions
T.I, R.W, Y.M, H.C: Performed research and analyzed data. T.I, Y.M, H.C, M.K, S.N Conceptualized this study. T.I, H.C, S.N: wrote the main manuscript text and prepared figures . All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Inukai, T., Watanabe, R., Murakami, Y. et al. Fungus-targeted nanomicelles enable microRNA delivery for suppression of virulence in Aspergillus fumigatus as a novel antifungal approach. Sci Rep 15, 17398 (2025). https://doi.org/10.1038/s41598-025-02742-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-02742-0
Keywords
This article is cited by
-
Sustainable nanomaterials for precision dental medicine: green synthesis, therapeutic applications, and future directions
Journal of Nanobiotechnology (2026)