Abstract
Multidrug resistance proteins contribute to chemotherapy resistance in various cancers. Among them, the P-glycoprotein (P-gp) has been the most investigated. These works aim to evaluate the efficacy of the new fluorescent tracer LightSpot-FL-1 for quantifying P-gp in CCRF-CEM and KG-1a acute leukemia cell lines, as well as in blood samples from healthy donors and leukemic patients. First, The P-gp quantity in CCRF-CEM and KG-1a cells measured by LightSpot-FL-1, was 7052 ± 2789 FU and 27,666 ± 6706 FU, respectively. Then, cells exposure to 10 µM daunorubicin (DNR) for 3 h reduced P-gp expression by 54% in CCRF-CEM cells and by 62% in KG-1a cells. Moreover, this decrease preceded a dose-dependent reduction in cell viability detected after 24 h of exposure to 10 µM DNR, with 46.6% and 72.2% viable cells for CCRF-CEM and KG-1a, respectively. These findings suggest that P-gp downregulation could serve as a potential biomarker of treatment efficacy. Additionally, LightSpot-FL-1 analysis of six acute myeloid leukemia patient blood samples allowed the identification of 14 distinct blast subpopulations, revealing substantial inter- and intra-individual heterogeneity in P-gp expression. Thus, these findings underscore the potential of LightSpot-FL-1 as a valuable tool for re-evaluating the clinical relevance of P-gp in tumor resistance diagnosis.
Similar content being viewed by others
Introduction
Multidrug resistance (MDR) transporters, which belong to the ATP-binding cassette (ABC) transporter family, contribute to anticancer treatment failure by acting as efflux pumps for a large variety of xenobiotics with varied structures and physiochemical properties. To date, three MDR proteins present a clinical interest i.e., the P-glycoprotein (P-gp), deeply investigated for its role in tumor resistance1,2, the Breast Cancer Resistance Protein (BCRP)3 and the Multi-Drug Resistance-associated Protein 7 (MRP7)4. P-gp plays a major role in the multidrug-resistant phenotype2,5. Indeed, this ubiquitous MDR protein is naturally expressed in a wide variety of living organisms, from prokaryotes to eukaryotes, including cancer cells6,7. At the cellular level, P-gp is expressed at the cell membrane, preventing extended intracellular accumulation of xenobiotics1,3,8. Additionally, P-gp is present on the membranes of intracellular organelles such as vesicles9 and lysosomes6,8, contributing to intracellular drug compartmentalization and further, to the development of cell resistance.
Despite the great interest in using P-gp as a therapeutic target, reliable and effective methodologies for its quantification in clinical practice are missing. In this context, our research group has developed LightSpot-FL-1, a P-gp-targeted fluorescent conjugate. This small molecule enables the specific detection and quantification of P-gp in cells or cellular masses such as spheroids. The P-gp labeling efficiency of LightSpot-FL-1 has already been demonstrated in DU4475 breast cancer cells in suspension, as well as in monolayer and spheroid cell cultures from the SUM1315 triple-negative breast cancer cell line10,11. These advancements make LightSpot-FL-1 an interesting tool, with great potential for evaluating the clinical relevance of P-gp in tumor resistance diagnosis.
Indeed, despite the advancement of new standards and targeted therapies, some cancers, especially Acute Leukemias (AL), remain highly refractory to treatments12. AL are malignant disorders characterized by the clonal expansion of immature and undifferentiated hematopoietic cells called blasts, resulting in their accumulation in the bone marrow. In this malignant transformation process, blast cells progressively replace healthy cells in this compartment and finally lead to depletion of differentiated cells in the blood. AL are classified based on the origin of the clonal cell, distinguishing between lymphoid and myeloid lineages. For several decades, the initial therapeutic approach as involved the concurrent administration of an anthracycline such as daunorubicin (DNR) during the first three days of treatment, and an antimetabolite, cytarabine (Ara-C) for seven days. This therapy is commonly referred to as the “7 + 3 regimen”13. Despite the implementation of this therapeutic approach, the overall survival rate at five years post-treatment averages between 40 and 60%, with a significant decline to 15–20% in patients aged over 6014. These poor survivals reflect the importance of the resistance mechanisms employed by cancer cells in these hematological tumors. MDR proteins in particular are thought to play a critical role in establishing this resistance. Therefore, the direct detection and quantification of P-gp expression level in leukemia cancer cells at diagnosis could help identifying intrinsic drug resistance and provide opportunities for treatment optimization.
Consequently, numerous clinical studies attempted to correlate the P-gp expression level in AL patients to their survival and relapse rates. Many of these studies led to the conclusion that this association lacked clinical relevance15,16,17,18. Other research focused on the most immature blast subpopulation CD34+ cells, identified as leukemic stem cells19. Results suggested that CD34+ blast cells exhibit higher P-gp expression compared to more mature cancer cells and could potentially influence patients outcomes20,21. However, these findings must be interpreted with caution due to small sample sizes and the absence of standardized, clinically adaptable methods for quantifying the P-gp expression in the different blast cell subpopulations.
In the present study, we investigate the P-gp quantification using the new P-gp-targeted fluorescent tracer LightSpot-FL-1, and explore its applicability for clinical diagnostics. For this, acute leukemia cell line models (CCRF-CEM and KG-1a) as well as white blood cells (WBCs) from healthy controls and blast cells from acute myeloid leukemia (AML) patients were used. This article is the first to investigate the use of this fluorescent tracer on patient samples.
Results
LightSpot-FL-1 penetration kinetics and P-gp targeting in CCRF-CEM and KG-1a cell lines by flow cytometry analysis
To validate P-gp labeling with LightSpot-FL-1, the penetration and stability of the fluorescent tracer were analyzed over time in the acute leukemia cell lines CCRF-CEM (Fig. 1A) and KG-1a (Fig. 1B).
LightSpot-FL-1 penetration kinetics and P-gp targeting in CCRF-CEM and KG-1a cell lines. CCRF-CEM (A) and KG-1a (B) cells were exposed to 1 µM LightSpot-FL-1 fluorescent probe and the cellular fluorescence intensity was measured at different time points by flow cytometry analysis. After 20 min of exposure, CCRF-CEM and KG-1a cells were washed 3 times with PBS and LightSpot-FL-1 fluorescence intensity in the cells maintained in PBS was monitored over time, after 60, 90, 120 and 180 min. FU: Fluorescence Units. n = 3.
For this, each cell line was exposed to 1 µM LightSpot-FL-1 for 20 min. The results showed that for both cell lines, a rapid penetration of LightSpot-FL-1 was detected within the first 3 min of exposure with 56,433 ± 8717 FU at 3 min and 53,166 ± 14,398 FU at 20 min in CCRF-CEM cells, and with 69,433 ± 10,450 FU at 3 min and 76,833 ± 10,507 FU at 20 min in KG-1a cells.
The cells were then washed three times with PBS to remove excess tracer unbound to the P-gp. After washing, each cell line was incubated in PBS and, from this step, the LightSpot-FL-1 intensity, corresponding to the amount of this tracer fixed on P-gp, was quantified over time. A clear decrease in LightSpot-FL-1 fluorescent signal was detected in both cell lines at 60 min with 7209 ± 2789 FU for CCRF-CEM cells and 27,666 ± 6706 FU for KG-1a cells. LightSpot-FL-1 signal intensity remained stable at 180 min in cells with 7052 ± 2866 FU and 25,066 ± 7720 FU for CCRF-CEM and KG-1a cells, respectively. These results revealed that the P-gp-linked probe remained stable over time. Therefore, the 60-min values were used to compare P-gp expression between the two cell lines, revealing a 3.55-fold higher level in KG-1a.
Intracellular accumulation of DNR in presence or absence of P-gp competitive inhibitor verapamil
In this experiment, both cell lines were exposed to 10 µM DNR, an anticancer therapy, with or without 50 µM verapamil, a P-gp competitive inhibitor. Intracellular DNR accumulation was quantified at 0 h, 3 h, and 6 h by flow cytometry analyses.
In the CCRF-CEM cell line, which has low P-gp expression, the rate of DNR accumulation at 3 h was 11,956 ± 3347 FU, compared to 12,733 ± 1285 FU when the cells were co-treated with verapamil, indicating no statistically significant difference (p value > 0.9999) Similar results were obtained after 6 h of exposure to DNR, with fluorescence intensities of 14,566 ± 829 FU in the DNR alone condition, and 13,088 ± 794 FU in the co-treatment with verapamil (Fig. 2A). These results indicated that the verapamil co-treatment did not affect intracellular DNR accumulation in this cell line.
DNR accumulation with or without verapamil in CCRF-CEM and KG-1a cell lines. Quantification of intracellular DNR levels over time in CCRF-CEM (A) and KG-1a (B) cell lines following exposure to 10 µM DNR, either alone or in co-treatment with 50 µM Verapamil. The analyses were performed using flow cytometry. Significances between conditions were indicated as ns: not significant (p > 0.05), *** (p < 0.001). FU: Fluorescence Units. n = 3.
In contrast, for the KG-1a cell line, which has a higher P-gp expression, the fluorescence intensity measured after 3 h of exposure to DNR alone condition was 6413 ± 2658 FU, whereas it increased to 13,733 ± 1162 FU when the cells were co-exposed to DNR and verapamil. Finally, after 6 h of DNR exposure, fluorescence levels were comparable between both conditions, with 9758 ± 2634 FU for DNR and 13,750 ± 1996 FU for the co-treatment with verapamil (Fig. 2B). These results indicate that for the KG-1a cell line, the presence of verapamil increases the accumulation of DNR in the cells.
P-gp expression levels and cell viability over time, after exposure to increasing concentrations of DNR
For these experiments, CCRF-CEM and KG-1a cell lines were exposed to increasing concentrations, i.e., 0, 0.1, 1, or 10 µM of DNR, for 3, 6 or 24 h. At each drug exposure time point, the P-gp expression level in cells was quantified and normalized relative to the untreated condition by using the LightSpot-FL-1 tracer. Cell viability was also assessed, under the same conditions through a LIVE/DEAD assay (Fig. 3).
Kinetics of P-gp expression levels and cellular viability in KG-1a and CCRF-CEM cell lines after exposure to increasing concentrations of DNR. CCRF-CEM and KG-1a cells were exposed to 0, 0.1, 1 or 10 µM DNR for 3, 6, or 24 h. The P-gp quantification was performed by flow cytometry after CCRF-CEM (A) and KG-1a cells (C) labeling with LightSpot-FL-1. For each drug exposure concentration, the quantified P-gp expression levels in cells was systematically normalized relative to an untreated control. LIVE/DEAD cell staining was performed on CCRF-CEM (B) and KG-1a (D) cells after exposure to 0, 0.1, 1 or 10 µM DNR for 3, 6 or 24 h. The cells were imaged with BioTek Cytation 3 MV (Agilent) (M = 10). Live and dead cells count was performed using Gen5 3.14 software (Agilent). Statistical significances between conditions were indicated as follow: ns: not significant (p > 0.05), *(p < 0.05), **(p < 0.01), ***(p < 0.001) and ****(p < 0.0001). n = 3.
For the CCRF-CEM cells, the P-gp expression level remained stable for 6 h in the presence of 0.1 µM and 1 µM DNR, with 1.03 ± 0.40 and 0.93 ± 0.49, respectively and it was close to the control cells with 1.00 ± 0.02. In contrast, in cells exposed to 1 µM DNR for 24 h, the P-gp expression decreased significantly to 0.38 ± 0.16 compared to the control condition (1.00 ± 0.07). In the presence of 10 µM DNR, P-gp expression in the CCRF-CEM cells dropped to 0.46 ± 0.35 after 3 h of exposure and remained low with 0.13 ± 0.02 after 24 h of exposure compared to the control condition (1.00 ± 0.05 after 3 h and 1.00 ± 0.06 after 24 h of exposure, respectively) (Fig. 3A). Regarding the cell viability analyses, all tested DNR concentrations showed values close to the control condition for up to 6 h of exposure: 97.4 ± 1.4% at 0 µM, 95.7 ± 1.3% at 0.1 µM, 94.9 ± 1.7% at 1 µM and 90.1 ± 4.0% at 10 µM. In contrast, after 24 h of exposure, the cell viability strongly decreased in a dose-dependent manner: 96.9 ± 1.4% at 0 µM, 70.4 ± 7.1% at 0.1 µM, 61.7 ± 8.5% at 1 µM and 46.6 ± 13.4% at 10 µM DNR (Fig. 3B).
For the KG-1a cells, the expression level of P-gp did not significantly change in presence of 0.1 µM and 1 µM DNR over 24 h of exposure, with 1.46 ± 0.74 and 0.98 ± 0.23, respectively, compared to the control condition at 1.00 ± 0.15. However, with 10 µM DNR, a significant decrease in P-gp expression was measured with 0.38 ± 0.13 from 3 h of exposure reaching 0.21 ± 0.02 after 24 h of treatment (Fig. 3C). In the cell viability study, no significant changes were observed during the first 6 h of the experiment at any tested dose, with values of 98.5 ± 1.0% at 0 µM and 87.5 ± 10.4% at 10 µM. In contrast, cell viability decreased slightly but significantly after 24 h of DNR exposure, reaching 88.7 ± 11.5% and 72.2 ± 12.5% in the 1 µM and 10 µM conditions, respectively, whereas cell viability in the control condition remained at 96.1 ± 0.7% (Fig. 3D).
P-gp expression quantification with LightSpot-FL-1 and anti-P-gp immunostaining on white blood cells (WBCs) from healthy donors
LightSpot-FL-1 was then used to study P-gp staining in primary blood cells i.e., monocytes, neutrophils, and lymphocytes from three healthy donors (HDs). The results were expressed as the ratio of fluorescence intensities between the cells stained with LightSpot-FL-1 and the unstained control cells.
In healthy donors HD#1, HD#2, and HD#3, the quantification of P-gp by using LightSpot-FL-1 (Fig. 4A,B,C) revealed respective normalized fluorescence intensities of 3609, 4658 and 550 for monocytes. In contrast, the P-gp expression levels measured by LightSpot-FL-1 in neutrophils and lymphocytes were lower with 1591 and 2029 for HD#1, 1600 and 2796 for HD#2, 45 and 204 for HD#3, respectively. Similarly, the P-gp quantification obtained by immunostaining revealed the higher P-gp expression for monocytes with 32, 43 and 15 for HD#1, HD#2 and HD#3, respectively. Conversely, neutrophils and lymphocytes express much lower P-gp levels with, 2 and 3 for HD#1, 2 and 3 for HD#2, 3 and 1 for HD#3 (Fig. 4D,E,F).
Comparison of P-gp level quantification after co-staining white blood cells (WBCs) with LightSpot-FL-1 and anti-P-gp immunostaining. Three blood samples from healthy donors were stained with anti-CD45 and LightSpot-FL-1 (A–C) to quantify P-gp in different cell populations. The samples were also stained with an anti-P-gp antibody, and the quantification of P-gp obtained through immunostaining was compared with that obtained using LightSpot-FL-1 (D–F). Fluorescence intensity was measured by flow cytometry (FITC filter for LightSpot-FL-1; APC-A filter for APC mouse anti-human CD243/P-gp antibody). Data were analyzed using FlowJo V10.8.1 software.
Quantification of P-gp levels in blast cells from acute myeloid leukemia patients
At diagnosis, six blood samples from patients with acute myeloid leukemia were stained with the CD45 antibody to distinguish CD45low blast cells from WBCs. A panel of multiple antibodies was also used to identify distinct blast subpopulations within the CD45low blast cell population. Additionally, samples were stained with LightSpot-FL-1 to quantify P-gp expression in the identified cell populations (Figure S1).
The normalized expression levels of P-gp within the CD45low blast population varied significantly among the six patients, ranging from 259 in AL#1 to 7317 in AL#6 (Fig. 5A). The quantification of P-gp expression in identified blast subpopulations revealed intra-individual variability among the different subpopulations, and also between individuals. For example, in the HLA-DR-/CD64- subpopulation, P-gp expression levels ranged from 435 in AL#2 to 2.411 in AL#4. In contrast, the CD34-/CD117- subpopulation exhibited a P-gp expression ranging from 108 in AL#2 to 7410 in AL#4. Regarding the inter-individual variability, in the CD34-/HLA-DR- subpopulation for instance, measured values for the P-gp expression levels were 23 in AL#2, 1029 in AL#1, 2612 in AL#3, 3243 in AL#5, 4028 in AL#4, and 6113 in AL#6 (Fig. 5B).
Quantification of P-gp in blast subpopulations from patients with acute myeloid leukemia. Six different blood samples from patients diagnosed with acute myeloid leukemia were stained with anti-CD45 antibody to identify blast cells and with 1 µM LightSpot-FL-1 to quantify P-gp in these cells (A). P-gp expression was quantified with 1 µM LightSpot®-FL-1 in every blast subpopulation identified by immunostaining using anti-CD34-A700; anti-CD64-APC-H7; anti-HLA-DR-APC and anti-CD117-BV711 antibodies (B). The raw values were normalized relative to the values of the unstained blast cells. Fluorescence intensity for each marker was measured by flow cytometry and data were analysed using FlowJo V10.8.1 software.
Discussion
LightSpot-FL-1 is an innovative fluorescent tracer, recently developed by our research group to allow the direct localization and quantification of P-gp in breast cancer cell line models10,11. The quantification of P-gp at diagnosis and/or at the time of relapse is of particular interest for cancers that are highly refractory to treatment. Despite the development of novel therapies, including immunotherapy and molecular targeted therapies, the standard regimen of DNR combined with Ara-C remains the first-line treatment for acute leukemia. However, this regimen is associated with suboptimal outcomes, particularly in elderly patients13.
The phenomenon of Multi-Drug Resistance is complex and can arise from various mechanisms. The overexpression of the P-glycoprotein is one of the currently explored options, with a trend suggesting the involvement of leukemic stem cells (LSCs) in P-gp-mediated resistance15,16. However, the majority of studies on this subject arrive at different conclusions and no study investigated the expression levels of P-gp across a range of more mature blast subpopulations. This lack of consensus may be attributed to several factors, including the absence of a standardized, cost-effective and easily implementable methodology for efficient detection and quantification of P-gp in clinical practice.
In this context, our research has focused on the application of the P-gp-targeted fluorescent tracer LightSpot-FL-1 on two acute leukemia cell line models i.e., CCRF-CEM and KG-1a, as well as on white blood cell populations from healthy donors and blast cells from patients diagnosed with AML. This represents the first reported application of LightSpot-FL1 on clinical samples.
We first checked the ability of LightSpot-FL-1 to specifically bind P-gp in CCRF-CEM and KG-1a cells by flow cytometry analyses. The lipophilic property of the tracer permits a rapid cellular penetration within the first minutes of exposure. A part of LightSpot-FL-1 was persistently retained in cells up to 2 h after washing. This part corresponds to 13% and 33% of the signal measured before the washing step for CCRF-CEM and KG-1a cells, respectively. These results show that LightSpot-FL-1 specifically binds P-gp over time and reveal a differential P-gp expression between the two cell line models, suggesting that KG-1a cells express approximately 4 times more P-gp than CCRF-CEM cells.
Subsequently, the uptake of DNR, known for being a P-gp substrate22, was investigated in the presence and absence of verapamil, a competitive P-gp inhibitor23. The presence of verapamil had no impact on the intracellular accumulation of DNR in low P-gp expressing CCRF-CEM cells. In contrast, in the KG-1a cells, which exhibit a higher P-gp expression, DNR accumulation increased by 52% in the presence of verapamil. Thus, the level of P-gp expression in cells influence the intracellular accumulation of anticancer agents. Therefore, we subsequently investigated the effect of exposure to increasing concentrations of DNR on both P-gp expression and cell viability over time.
The relationship between P-gp expression levels, i.e., cellular drug resistance and treatment efficacy through cell viability analysis was investigated. Interestingly, a decrease in P-gp levels was detected in both cell lines after high DNR concentration exposure that systematically precede a decrease in cell viability. Indeed, the treatment with 10 µM DNR for 3 h resulted in 55% and 62% decrease in P-gp levels for CCRF-CEM and KG-1a cells, respectively. After 24 h of 10 µM DNR exposure, the P-gp expression rate decreased dramatically by 87% and 80% for CCRF-CEM and KG-1a cells, respectively. Regarding the cell viability, after 24 h of 10 µM DNR exposure this parameter dropped to 47% and 72% in CCRF-CEM and KG-1a cells, respectively. These results suggest that a drop in P-gp expression level following anticancer therapy may serve as an early indicator of treatment efficacy.
After validating the efficacy of LightSpot-FL-1 on in vitro cellular models of AL, we extended our study to assess the effectiveness of P-gp labeling in primary cells from healthy donors and patients with AML. Subsequently, LightSpot-FL-1 P-gp labeling and immunostaining were compared in monocytes, neutrophils, and lymphocytes from three healthy donors. The P-gp expression level was much higher in monocytes than in neutrophils and lymphocytes, regardless of the methodology. Indeed, the P-gp expression in monocytes measured with LightSpot-FL-1 was 5.79 and 2.04 times higher on average than in neutrophils and lymphocytes, respectively. Regarding immunostaining, the P-gp expression in monocytes was found 14.85 and 12.29 times higher on average than in neutrophils and lymphocytes, respectively. Despite similar quantification profiles, differences can be detected between the two techniques. This highlights the ability of the LightSpot-FL-1 tracer to target both P-gp expressed at the cell membrane and intracellular P-gp, unlike immunostaining which only targets an external epitope. This is a clear advantage of the LightSpot technology, as intracellular P-gp was shown to be a significant factor in overall drug resistance, particularly in AL models9. Inter-individual variability in the WBCs P-gp expression of the three healthy donors was also detected. This natural variability results from the combined influence of genetic and environmental factors on the expression of MDR proteins, particularly P-gp24,25. Thus, the total quantification of cell membrane and intracellular P-gp by LightSpot-FL-1 underscores its advantages in predicting global cell resistance, thereby demonstrating its applicability and relevance for clinical P-gp investigation.
Finally, LightSpot-FL-1 was used in combination with a panel of clinically relevant membrane antibodies to quantify P-gp in blast subpopulations from six patients diagnosed with AML. The collected data revealed that significant variations can be detected by using LightSpot-FL-1, both in the average P-gp expression levels in blast cells among patients and within fourteen distinct blast subpopulations of a given patient. These encouraging preliminary results pave the way to investigating inter and intra-individual P-gp expression variabilities with LightSpot-FL1 on a broader sample. If confirmed, such variabilities could partially explain the inconsistencies observed in studies attempting to predict tumor resistance based on the average P-gp expression level of blast cells15,16,17,18. In the context of recent studies aimed at stratifying patients based on P-gp expression levels in their blast cells to evaluate responses to treatment combining standard chemotherapy with a P-gp inhibitor26,27 LightSpot-FL-1 emerges as a powerful tool for accurate and reliable patient screening. In conclusion, LightSpot-FL-1 appears as a new P-gp-targeting tracer capable to overcome technical and financial limitations, thereby enabling a reevaluation of the clinical significance of P-gp in tumor resistance.
Material and methods
Cell lines culture
The human T lymphoblast CCRF-CEM (CCL-119) and the promyeloblasts macrophage KG-1a (CCL-246.1) cell lines were purchased from ATCC and were tested for the absence of mycoplasma contamination (MycoplasmaCheck test, Eurofins Genomics, Nantes, France). These two cell lines have been authenticated by DNA comparison with known databases (Eurofins Genomics, Nantes, France). CCRF-CEM cells were cultured in RPMI 1640 medium (Gibco, Dublin, Ireland) supplemented with 5% heat-inactivated fetal calf serum (Dominique Dutscher, Bernolsheim, France) 100 µg/mL Penicillin Streptomycin (Gibco, Dublin, Ireland) and 2 mM L-glutamine (Gibco, Dublin, Ireland). KG-1a cells were cultured in IMDM medium (Gibco, Dublin, Ireland) containing L-glutamine (Gibco, Dublin, Ireland) and 25 mM HEPES (Gibco, Dublin, Ireland) supplemented with 5% heat-inactivated fetal calf serum and 100 µg/mL Penicillin–Streptomycin (Gibco, Dublin, Ireland).
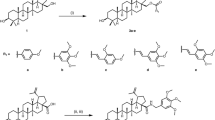
LightSpot-FL-1 chemical synthesis and characterization
LightSpot-FL-1 was synthetized as previously described11. Briefly, an amine bearing P-gp inhibitor derivative was reacted with commercial NHS ester fluorochrome BODIPY FL (Interchim, Montlucon, France) to yield LightSpot-FL-1. After semi-preparative HPLC–UV-DAD purification, LightSpot-FL-1 was fully characterized by 1H NMR, 13C NMR, and high-resolution mass spectrometry, and was stored as dry aliquots at − 20 °C. For biological experiments, LightSpot-FL-1 was dissolved in DMSO (1.0 mM stock solutions) and analysed by analytical HPLC before use for quality control purposes.
LightSpot-FL-1 cellular penetration and P-gp targeting analysis by flow cytometry
CCRF-CEM and KG-1a cell cultures were maintained according to the previously described conditions. For the LightSpot-FL-1 penetration experiments, cell suspensions containing 1,000,000 cells/mL were collected from both cell lines and fixed with 4% paraformaldehyde (PFA) for 10 min. The fixed cells were then incubated with 1 µM LightSpot-FL-1 dissolved in dimethyl sulfoxyde (DMSO) and diluted in phosphate-buffered saline (PBS). Fluorescence intensity in the cells was measured after 3, 10, and 20 min of LightSpot-FL-1 exposure. The cells were then washed three times with PBS and the fluorescence intensity was measured after 60, 90, 120, 180 min using a Luminex Guava easyCyte HT flow cytometer (Cytek Biosciences, Fremont, CA, USA) with a blue excitation laser (488 nm) and a green fluorescent filter (525/30 nm). All data were analyzed using the GuavaSoft 4.0 software.
Daunorubicin (DNR) intracellular accumulation in presence or absence of verapamil
KG-1a and CCRF-CEM cells were seeded in 96-well plates at a concentration of 20,000 cells per well in serum-free medium. Daunorubicin hydrochloride (Merck KGaA, Darmstadt, Germany) and verapamil hydrochloride (Sigma-Aldrich, Saint Louis, MO, USA) were diluted in PBS with 0.1% DMSO and added to the wells at final concentrations of 10 and 50 µM respectively. After 0, 3 or 6 h of treatment, the cells were washed once with serum-free medium and once with Dulbecco’s PBS (Thermo Fisher Scientific, Waltham, MA, USA). Following these washing steps, the cell pellets were resuspended in PBS for analysis using a Luminex Guava® easyCyte™ HT flow cytometer (red excitation laser: 642 nm; red fluorescent filter: 695/50 nm; Cytek Biosciences, Fremont, CA, USA). All data were analyzed using the GuavaSoft 4.0 software.
Impact of daunorubicin (DNR) on P-gp expression quantified by LightSpot-FL-1
CCRF-CEM and KG-1a cells were seeded at a density of 200,000 cells/mL. After 48 h of culture, the cells were exposed to DNR at varying concentrations of 0, 0.1, 1 or 10 µM for 3, 6 or 24 h. For each condition, 1,000,000 cells were collected and fixed with 4% PFA for 10 min. The fixed cells were then incubated with 1 µM LightSpot-FL-1 dissolved in DMSO and diluted in PBS for 20 min. To remove non-specific staining, three successive PBS washes were performed. Fluorescence intensity in the cells was measured using a Luminex Guava easyCyte™ HT flow cytometer (Cytek Biosciences, Fremont, CA, USA) as previously described. The P-gp expression level of cells exposed to anticancer agents was normalized to the basal expression level of the cells. All data were analyzed using the GuavaSoft 4.0 software.
Evaluation of the cytotoxicity of daunorubicin (DNR)
CCRF-CEM and KG-1a cells were seeded in 96-well plates at a concentration of 100,000 cells per well. Daunorubicin hydrochloride (DNR) (Merck KGaA, Darmstadt, Germany) was diluted in PBS with 0.1% DMSO for drugs accumulation experiments. DNR was added to each well at final concentrations of 0, 0.1, 1, and 10 µM. The cells were incubated with these molecules for 3, 6 or 24 h. To determine drug cytotoxicity, cells were stained with LIVE/DEAD™ Viability/Cytotoxicity Kit (calcein/ethidium, Invitrogen, Waltham, MA, USA) for 30 min, following the manufacturer’s instructions. Cells were then imaged using the BioTek Cytation 3 MV (Agilent, Santa Clara, CA, USA) fluorescent microscopy (magnification = 10X, fluorescent filters: Green Fluorescent Protein (GFP) for calcein detection, Iodide Propidium (IP) for ethidium detection). For each field, viability was determined by counting live and dead cells using the cellular analysis primary mask and count tool in the Gen5 3.14 software (Agilent, Santa Clara, CA, USA).
Blood samples from patients with AML and healthy donors
Blood samples from healthy donors and patients with AML were collected in EDTA tubes. We used leftover samples, corresponding to the residue of a sample that had been obtained and used for clinical purposes, and would otherwise have been discarded. These samples could be used for research because the donors were informed and did not express any disagreement, as stipulated by French law.
P-gp expression level quantification in WBCs from healthy controls and blast subpopulations from patients diagnosed with AML
All experiments were carried out with fresh cells, within 24 h of sampling. Blood samples were washed with PBS and the cells were incubated with NH4Cl for 15 min to lyse red blood cells. The white blood cells (WBCs) were then washed again with PBS before being counted using the Sysmex XN-1000 system (Sysmex Corporation, Kobe, Hyogo, Japan). After counting, 1,000,000 WBCs were collected and exposed for 20 min to 1 µM LightSpot-FL-1, or APC mouse anti-human CD243 (1:100, ABCB1, clone UIC2; eBioscience, ThermoFisher Scientific, Waltham, MA, USA) for healthy controls, and anti-CD45-A700 (1:160, clone HI30, BD Biosciences), anti-HLA-DR-APC (1:80, clone L243, BD Biosciences), anti-CD34-BV421 (1:80, clone 8G12, BD Biosciences), anti-CD117-BV711 (1:40, clone 104D2, BD Biosciences), anti-CD64-APC-H7 (1:40, clone 10.1, BD Biosciences), anti-CD14-PE™-Cy7 (1:40, clone MφP9, BD Biosciences) for samples from patients with AML. The cells were washed three times with PBS and the pellets were resuspended in 300 µL of PBS for analysis using BD LSRFortessa cytometer (Beckton Dickinson, Franklin Lakes, NJ, USA). The raw values were normalized relative to the values of the unstained blast cells. All data were analyzed using FlowJo-10.8.1 software.
Statistical analysis
Results were presented as mean ± standard deviation. All experiments were performed independently on three separate occasions. Mann–Whitney test was used to compare the difference between two variables. Kruskal–Wallis test was performed when we compared more than two variables. GraphPad Prism 8.0.2 (GraphPad Software) was used for all statistical analyses. The differences were noted as follows: p < 0.05 (*), p < 0.01 (**), p < 0.001 (***), p < 0.0001(****). Non-significant results were noted as “ns”.
Data availability
The datasets generated and/or analyzed during the current study are available from the corresponding author upon reasonable request.
References
Bosch, I. & Croop, J. P-glycoprotein multidrug resistance and cancer. Biochim. Biophys. Acta BBA Rev. Cancer 1288, 37–54 (1996).
Zheng, H.-C. The molecular mechanisms of chemoresistance in cancers. Oncotarget 8, 59950–59964 (2017).
Doyle, L. A. et al. A multidrug resistance transporter from human MCF-7 breast cancer cells. Proc. Natl. Acad. Sci. 95, 15665–15670 (1998).
Hopper, E. et al. Analysis of the structure and expression pattern of MRP7 (ABCC10), a new member of the MRP subfamily. Cancer Lett. 162, 181–191 (2001).
Silverman, J. A. Multidrug-resistance transporters. Pharm. Biotechnol. 12, 353–386 (1999).
Goldstein, L. J. et al. Expression of multidrug resistance gene in human cancers. JNCI J. Natl. Cancer Inst. 81, 116–124 (1989).
Lubelski, J., Konings, W. N. & Driessen, A. J. M. Distribution and physiology of ABC-type transporters contributing to multidrug resistance in bacteria. Microbiol. Mol. Biol. Rev. MMBR 71, 463–476 (2007).
Juliano, R. L. & Ling, V. A surface glycoprotein modulating drug permeability in Chinese hamster ovary cell mutants. Biochim. Biophys. Acta BBA - Biomembr. 455, 152–162 (1976).
Ferrao, P., Sincock, P., Cole, S. & Ashman, L. Intracellular P-gp contributes to functional drug efflux and resistance in acute myeloid leukaemia. Leuk. Res. 25, 395–405 (2001).
Goisnard, A. et al. LightSpot®-FL-1 fluorescent probe: An innovative tool for cancer drug resistance analysis by direct detection and quantification of the P-glycoprotein (P-gp) on monolayer culture and spheroid triple negative breast cancer models. Cancers 13, 4050 (2021).
Daumar, P. et al. Chemical biology fluorescent tools for in vitro investigation of the multidrug resistant P-glycoprotein (P-gp) expression in tumor cells. RSC Adv. 13, 27016–27035 (2023).
Zhang, J., Gu, Y. & Chen, B. Mechanisms of drug resistance in acute myeloid leukemia. OncoTargets Ther. 12, 1937–1945 (2019).
Döhner, H. et al. Diagnosis and management of AML in adults: 2022 recommendations from an international expert panel on behalf of the ELN. Blood 140, 1345–1377 (2022).
Liu, H. et al. 1- and 5-year survival for adults with acute myeloid leukaemia and 30-day mortality after initial systemic anti-cancer therapy, with an exploration of factors associated with poorer outcomes: Data from a national registry in England, 2013–2020. Blood 142, 2846 (2023).
Wuchter, C. et al. Clinical significance of P-glycoprotein expression and function for response to induction chemotherapy, relapse rate and overall survival in acute leukemia. Haematologica 85, 711–721 (2000).
Olson, D. P. et al. The prognostic significance of P-glycoprotein, multidrug resistance-related protein 1 and lung resistance protein in pediatric acute lymphoblastic leukemia: A retrospective study of 295 newly diagnosed patients by the Children’s Oncology Group. Leuk. Lymphoma 46, 681–691 (2005).
van Grotel, M. et al. CD34 expression is associated with poor survival in pediatric T-cell acute lymphoblastic leukemia. Pediatr. Blood Cancer 51, 737–740 (2008).
Svirnovski, A. I. et al. ABCB1 and ABCG2 proteins, their functional activity and gene expression in concert with drug sensitivity of leukemia cells. Hematol. Amst. Neth. 14, 204–212 (2009).
Lapidot, T. et al. A cell initiating human acute myeloid leukaemia after transplantation into SCID mice. Nature 367, 645–648 (1994).
de Figueiredo-Pontes, L. L. et al. Determination of P-glycoprotein, MDR-related protein 1, breast cancer resistance protein, and lung-resistance protein expression in leukemic stem cells of acute myeloid leukemia. Cytometry B Clin. Cytom. 74, 163–168 (2008).
Suárez, L. et al. CD34+ cells from acute myeloid leukemia, myelodysplastic syndromes, and normal bone marrow display different apoptosis and drug resistance-associated phenotypes. Clin. Cancer Res Off. J. Am. Assoc. Cancer Res. 10, 7599–7606 (2004).
Wang, E., Casciano, C. N., Clement, R. P. & Johnson, W. W. Cooperativity in the Inhibition of P-glycoprotein-mediated daunorubicin transport: Evidence for half-of-the-sites reactivity. Arch. Biochem. Biophys. 383, 91–98 (2000).
Futscher, B. W. et al. Verapamil suppresses the emergence of P-glycoprotein-mediated multi-drug resistance. Int. J. Cancer 66, 520–525 (1996).
Schuetz, E. G., Furuya, K. N. & Schuetz, J. D. Interindividual variation in expression of P-glycoprotein in normal human liver and secondary hepatic neoplasms. J. Pharmacol. Exp. Ther. 275, 1011–1018 (1995).
Prasad, B. et al. Interindividual variability in hepatic organic anion-transporting polypeptides and P-glycoprotein (ABCB1) protein expression: Quantification by liquid chromatography tandem mass spectroscopy and influence of genotype, age, and sex. Drug Metab. Dispos. 42, 78–88 (2014).
Marcelletti, J. F. & Sikic, B. I. A clinical trial of zosuquidar plus gemtuzumab ozogamicin (GO) in relapsed or refractory acute myeloid leukemia (RR AML): Evidence of efficacy based on leukemic blast P-glycoprotein functional phenotype. Cancer Chemother. Pharmacol. 92, 369–380 (2023).
Marcelletti, J. F. & Sikic, B. I. Continuous 72-h infusion of zosuquidar with chemotherapy in patients with newly diagnosed acute myeloid leukemia stratified for leukemic blast P-glycoprotein phenotype. Cancer Chemother. Pharmacol. 93, 595–604 (2024).
Funding
This study was funded by the “Comité du Puy-de-Dôme de la Ligue Contre le Cancer” and the French government under the project “Plan de relance, DRV_PIP_2021-115_UCA_ANR_ANR21-PRRD-0009-01_RESIST”.
Author information
Authors and Affiliations
Contributions
M.B designed and supervised the project, participated in the revision and edition of the manuscript. M.D performed the experiments, analyzed the data, and wrote the manuscript. A.G and C.B, participated in the supervision of the project, validation and edition of the manuscript. A.V performed the experiments. E.G, M.DE, M.R, M.BE and F.P-L participated in the revision and edition of the manuscript. P.D and E.M supervised the project and participated in the revision and edition of the manuscript. All the authors made revisions and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
Our study exclusively involves patient-derived cells without any associated clinical data. According to the ethical regulations of our institution and French law (Article L.1211-2 al.2 du Code de la santé publique), such research does not require Institutional Review Board (IRB) approval. Additionally, as per our institutional policy, all patients systematically sign a non-opposition statement, allowing the use of their biological samples for research purposes. Indeed, all patients included in the study were informed via the CHU Clermont-Fd welcome booklet that the remaining biological samples taken during their diagnosis and follow-up could be used for research unless they objected; no patient expressed their objection. All methods were carried out in accordance with relevant guidelines and regulations.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Dubois, M., Goisnard, A., Bourgne, C. et al. P-gp quantification for cell resistance studies in leukemia cells using the innovative LightSpot-FL-1 fluorescent conjugate. Sci Rep 15, 18173 (2025). https://doi.org/10.1038/s41598-025-03370-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-03370-4