Abstract
This research delved into the molecular mechanism underlying dioctyl terephthalate (DOTP)-related periodontitis (PD) through the application of network toxicology, molecular docking, and molecular dynamics simulations. By leveraging data from SwissTargetPrediction, SuperPred, and GeneCards databases, targets associated with DOTP toxicity and PD were pinpointed, leading to the identification of 37 shared targets through a comprehensive analysis. Enrichment analysis unveiled significant implications in inflammatory responses (e.g., the AGE-RAGE signaling pathway) and immune regulatory pathways (e.g., the C-type lectin receptor pathway). Core targets (PTGS2, MAPK14, NFKB1, STAT1) were pinpointed utilizing Cytoscape and molecular docking techniques. DOTP exhibited robust binding to these targets through hydrogen bonding and hydrophobic interactions, with the DOTP-PTGS2 complex displaying the most favorable binding energy (− 7.1 kcal/mol). Molecular dynamics simulations validated the stability of this complex, demonstrating the lowest root mean square deviation (RMSD) of 0.22 nm and the largest buried solvent-accessible surface area (Buried SASA) of 12 nm2, indicating its superior stability. This investigation elucidates the molecular basis of DOTP-related PD, underscores the efficacy of network toxicology and computational modeling in environmental health risk assessment, and provides a theoretical framework for targeted interventions.
Similar content being viewed by others
Introduction
The advent of industrialization has given rise to a plethora of concerns regarding food and environmental pollutants, which have emerged as significant hazards to human health. The ramifications of these pollutants are pervasive and far-reaching. For instance, in the food sector, residual organophosphorus pesticides have been shown to enter the human body via the food chain, thereby inhibiting the activity of the human body’s acetylcholinesterase enzyme and interfering with the normal function of the nervous system1. A study on PM2.5 has demonstrated that long-term exposure to elevated levels of PM2.5 can result in a decline in lung function and an increased likelihood of developing chronic obstructive pulmonary disease (COPD)2. This underscores the imperative for a comprehensive understanding of the impact of these pollutants on human health.
Periodontitis (PD) is a chronic inflammatory disease of the oral cavity. Severe periodontitis is considered to be the sixth most common disease worldwide, posing a serious threat to human health3. The disease is characterized by red, swollen, bleeding gums and loose teeth4. Notably, periodontitis has been associated with inflammatory factors and immune system dysregulation5. While substantial evidence points to a potential association between environmental pollutants and PD, the precise targets and mechanisms by which it occurs remain to be fully elucidated.
Dioctyl terephthalate (DOTP) is a prevalent plasticizer that is extensively utilized in polyvinyl chloride (PVC) plastic processing. There are two manufacturing routes to DOTP. One is by direct esterification of terephthalic acid with 2-ethylhexanol (2-EH); the other is by transesterification of dimethyl terephthalate with 2-EH. According to statistics, the global consumption of DOTP exceeded 6.8 million tons in 2017, and this figure is still increasing6,7. It is used in significant quantities in food packaging, plastic products, and children’s toys8. In these products, DOTP exists in a physically dispersed form, devoid of chemical bonds with the polymer. This characteristic renders it susceptible to widespread migration into the surrounding environment, underscoring potential exposure risks. The migration rate of DOTP is crucial for assessing real-world exposure levels. In Europe, the total migration limit for DOTP is 60 mg/kg, with a tolerable daily intake of 1 mg/kg body weight for humans. Research indicates that DOTP migration into olive oil reaches this limit of 60 mg/kg9. DOTP and di (2-ethylhexyl) terephthalate (DEHT) are chemically identical, representing the same compound under different names. A study measured plasticizer levels in foods from American fast-food chains and food-handling gloves. It found a median DOTP/DEHT concentration of 251 μg/kg in food samples (n = 19), with chicken wraps showing significantly higher levels than hamburgers (600 μg/kg vs. 220 μg/kg). In glove samples (n = 3), DOTP/DEHT content ranged from 28 to 37% by weight. These findings suggest notable DOTP exposure in fast-food settings10. Furthermore, a recent review reports that global indoor settled dust samples show median/average DOTP/DEHT concentrations of 19 to 164 μg/g, with levels rising in recent years11. Moreover, a study on the temporal trends (2009–2019) of the concentrations of DOTP/DEHT metabolites in urine showed that the concentrations of DOTP/DEHT metabolites in the urine of the US population increased significantly over time12.
Concerning its effects on human health, long-term exposure to DOTP has the potential to disrupt the endocrine system and affect physiological processes such as reproduction and development13. A recent study suggests that DOTP may induce chronic spontaneous urticaria (CSU) by targeting certain proteins that affect inflammation and immune regulation14. Meanwhile, high concentrations of DOTP have been linked to increased production of reactive oxygen species (ROS)8, which may contribute to PD progression via oxidative stress15. Additionally, studies indicate that DOTP exposure can lower blood glucose levels, reduce white adipose tissue weight, trigger inflammatory responses, and alter gut microbiota composition16. Previous studies consistently demonstrate a strong association between gut microbiota and PD17,18. Although direct evidence connecting DOTP to PD is limited, existing data suggest a potential link. Thus, investigating DOTP’s effects on PD is crucial.
Network toxicology, an emerging interdisciplinary field, employs the "compound-target-gene" network model to investigate the toxicological properties of target entities19. This approach facilitates the identification of potential targets and toxic pathways of food and environmental pollutants, thereby assessing their risks to human health. Consequently, network toxicology holds significant promise for evaluating the toxicity of environmental pollutants. Molecular docking, a computational technique, predicts the optimal binding mode and affinity between a compound and its target. In this study, molecular docking was utilized to forecast the binding capacity of DOTP molecules to PD-related targets20. Molecular dynamics simulation captures the real-time motion trajectory of a molecular system, verifying the stability of the binding mode predicted by molecular docking during dynamic processes21,22. These methods enable researchers to gain deeper insights into how DOTP affects diseases like PD by binding to specific target proteins.
Methods
Access to DOTP toxicity targets
In order to comprehensively assess the toxicity of DOTP, we integrated the targets associated with DOTP exposure from the SwissTargetPrediction23 and SuperPred databases24. The 2D structure and SMILES representation of DOTP were first obtained from the PubChem database25. Potential targets were then identified from SwissTargetPrediction and SuperPred using SMILES. “Homo sapiens” was selected, and the results were cross-referenced to the final toxic target of DOTP. The UniProt database26 was then utilized to standardize the nomenclature of the target proteins.
Disease-related targets
To identify PD-related targets, we obtained the relevant genes from GeneCards27, Online Mendelian Inheritance in Man (OMIM), and Gene Expression Omnibus (GEO) databases28, leveraging comprehensive and accurate data sources. In GeneCards, the keyword “periodontitis” was entered to obtain PD-related targets, and the genes with correlation scores > 1 were filtered from the obtained results to enhance credibility. Concurrently, a PD dataset (GSE10334) comprising 247 samples was selected from the GEO database using the “limma” package29 in R software to screen the characterized genes. The screening criteria included: First, the logFC value had to be greater than 1. Second, the corrected P-value had to be less than 0.05. Consequently, the genes identified through these criteria were then cross-denominated and utilized as the final targets of PD.
GO and KEGG enrichment analysis.
In order to explore the potential mechanism of DOTP-related PD, their cross-targets were imported into the Database for Annotation, Visualization, and Integrated Discovery (DAVID) database30 for gene enrichment analysis. The DAVID database is a widely used bioinformatics tool for identifying biological processes (Gene Ontology, GO) and signaling pathways (e.g., KEGG, Reactome) that are significantly enriched in gene lists. GO analysis evaluates the biological processes (BP), molecular functions (MF), and cellular components (CC) in which the genes are involved. In all screening processes, a P value of less than 0.05 was set to ensure reliable results. Enrichment analysis has been identified as a means of identifying potential pathways of DOTP-related PD at the level of molecular mechanisms, thereby providing further evidence for its possible pathogenicity.
Identify core targets
Protein–protein interaction (PPI) networks of cross-targets were constructed using the Search Tool for the Retrieval of Interacting Genes/Proteins (STRING) database31 with the species parameter set to “Homo”. Nodes with confidence scores exceeding 0.4 were selected, while unconnected nodes were concealed. The ensuing results were then imported into Cytoscape 3.10.2 for core target screening32. Cytoscape is an open-source network analysis and visualization software that supports the construction, analysis, and visualization of complex networks and is particularly adept at dealing with biomolecular interaction networks (e.g., protein–protein interactions and gene regulatory networks, etc.). To accurately identify the core targets, we integrated the results of the “CytoHubba”33 and “MCODE”34 plug-ins in Cytoscape and used the intersection of the two as the core targets. CytoHubba is employed to identify the top 10 nodes that are central in the largest clusters, while MCODE identifies key gene modules. A range of analysis methods were employed to identify the core targets that may play a pivotal role in DOTP-related PD.
Molecular docking of core targets
Molecular docking is a computational simulation technique designed to predict the binding modes and binding strengths of small molecules (ligands) to biological macromolecules (receptors, e.g., proteins, DNA). This method has important applications in the fields of drug discovery, enzyme engineering, and protein–ligand interaction studies. In this study, molecular docking was utilized to assess the binding capacity between DOTP and the core target. To obtain the necessary pre-files for molecular docking, 2D files of DOTP in “SDF” format were obtained from the Public Chemical (PubChem) Database. Concurrently, the identifier of the core target was obtained from the Universal Protein Resource (UniProt), and the corresponding “PDB” format file was obtained from the Protein Data Bank (PDB) database. For small molecule ligands, the minimum binding energy of DOTP was calculated using Chem3D23.1.1 and saved in “MOL2” format. To mitigate the impact of extraneous factors on the ensuing molecular docking, water molecules, and small ligands were eliminated from the protein receptor using PyMOL 4.6.035. AutoDockTools 1.5.7 (ADT) is an open-source tool designed specifically for molecular docking, which simplifies AutoDock’s complex parameter setting and result analysis process36. ADT was employed to hydrogenate protein receptors, identify pocket boxes, and save the structure files of DOTP and target proteins in “PDBQT” format.
AutoDock Vina37 was utilized for the final molecular docking operation, and the results were visualized by GeneCards. Concurrently, the amino acids implicated in the stabilization of small molecules following molecular docking were subjected to two-dimensional interaction analysis using Discovery Studio 2025. This analysis was undertaken with the objective of elucidating the interaction patterns of DOTP with proximate amino acids.
Molecular dynamics simulations
Molecular dynamics simulation (MD) is a computational method based on physical principles that is used to study the dynamic behavior of atomic or molecular systems under specific conditions38. In this study, we employed molecular dynamics simulation to meticulously investigate the interaction mechanism between DOTP and PD-related targets. This approach was undertaken to systematically assess the binding stability of small molecules and proteins.
GROMACS39 is an open-source and efficient MD simulation software that can be used to simulate the motion of atoms/ molecules under specific conditions and analyze the structure, energy, interactions, and dynamic behavior. The optimal binding conformation of DOTP to the target protein, as determined by AutoDock Vina, was employed as the initial conformation for MD. The GROMACS 2022 program was utilized for MD manipulation. The GAFF force field was employed for small molecules, while the AMBER14SB force field and the TIP3P water model were utilized for proteins. The files of protein and small molecule ligands were then merged to construct the simulation system of the complexes. The simulations were performed at constant temperature and pressure, as well as under periodic boundary conditions. During MD simulations, all constraints involving hydrogen bonding were performed using the LINCS algorithm with an integration step of 2 fs. The cutoff value for non-bonding interactions was set to 10 Å and updated at 10-step intervals. The V-rescale temperature coupling method was employed to regulate the simulation temperature to 298 K, and the Berendsen method was utilized to control the pressure to 1 bar. Constant number, volume, and temperature (NVT) and constant number, pressure, and temperature (NPT) equilibrium simulations were carried out at 298 K for 100 ps, and MD simulations were performed for 100 ns for the complex system, with the conformation saved every 10 ps. Following the completion of the simulations, the simulation trajectories were analyzed using VMD and PyMOL. Additionally, the MMPBSA binding free energy analysis between the protein and the small-molecule ligand was performed using the g_mmpbsa program.
Results
Toxicity targets of DOTP
In this study, a total of 149 toxicity targets associated with DOTP exposure were obtained after cross-denigration through SwissTargetPrediction and SuperPred databases (Fig. 1A). Similarly, a total of 1618 PD-associated targets were identified by conducting an in-depth analysis of GeneCards, OMIM, and GEO databases (Fig. 1B). The integration of these two sets of data yielded a total of 37 genes that have the potential to serve as targets for PD triggered by DOTP exposure (Fig. 1C). A comprehensive list of these data can be found in the supplementary material.
A Cross-removal of duplicates for DOTP-related toxicity targets; B Cross-removal of duplicates for PD-related targets; C Cross-targets between DOTP and PD; D Gene Ontology (GO) enrichment analysis of 37 shared targets, encompassing Biological Processes (BP), Cellular Components (CC), and Molecular Functions (MF), with presentation limited to the top 10 outcomes; E Presentation of the top 10 pathways derived from Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis of shared targets, along with their associations with relevant genes. Bubble size indicates the number of genes enriched in each pathway, while bubble color reflects the statistical significance of enrichment, with darker shades indicating lower P-values and greater significance.
Enrichment analysis
The present study employed GO and KEGG enrichment analysis on 37 targets. GO enrichment analysis (comprising BP, MF, and CC) identified a total of 165 statistically significant items. Of these, 139 were found to be BP items, 23 were MF items, and 23 were CC items. Figure 1D provides a visual representation of the 10 items with the lowest false discovery rate (FDR). The findings suggest that certain targets may play a role in the metabolic process of reactive oxygen species and the regulation of inflammatory responses, which are closely related to the pathogenesis of PD15.
KEGG enrichment analysis was designed to explore the potential pathways involved in PD by the targets. The analysis yielded a total of 77 statistically significant KEGG items. To elucidate the relationship between these pathways and targets, the top 10 pathways were illustrated using mulberry diagrams (Fig. 1E). Of particular relevance is the AGE-RAGE signaling pathway, which plays a pivotal role in diabetic complications and is a significant risk factor for PD40. Furthermore, cancer-related pathways and receptor-related pathways (e.g., C-type lectin receptor signaling pathway) were enriched. This finding suggests a potential link between the pathogenesis of PD and the tumor microenvironment and cell signaling.
Core cross-targets
A protein–protein interaction (PPI) network was constructed from the STRING database using the cross-targeting of DOTP and PD (Fig. 2A). The network contains 37 nodes and 130 edges, with an average node degree of 7.03. The target network constructed by STRING was systematically analyzed using Cytoscape 3.10.2. The “MCODE” plugin identified key modules containing 11 targets (Fig. 2B). The “CytoHubba” plugin identified 10 nodes with centrality (Fig. 2C). The 10 genes obtained after crossover were finally used as core targets for subsequent analysis (Fig. 2D). Table 1 provides the relevant details of these targets in Cytoscape.
A PPI network of 37 overlapping targets obtained from the STRING platform; B Key gene modules obtained by analyzing the PPI network of overlapping targets using the “MCODE” plugin in Cytoscape software; C The top 10 genes obtained by calculating the maximum clique centrality of overlapping targets using the “CytoHubba” plugin in Cytoscape software. The darker the color, the more or larger the maximum cliques the node participates in, indicating that it may be a core member of the functional module; D Overlapping targets from the two analysis methods.
Molecular docking
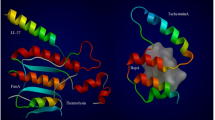
In this study, the binding mode between DOTP and core targets was evaluated using a molecular docking technique. Lower binding energy between the target and DOTP indicates stronger binding strength, with hydrogen bond energy substantially enhancing the complex’s binding ability. Consequently, PTGS2, MAPK14, NFKB1, and STAT1, which form hydrogen bonds with DOTP during molecular docking and exhibit low binding energies, were identified as targets with significant binding potential (Fig. 3). Table 2 provides detailed information on the binding energies and interaction forces of these targets during molecular docking.
Molecular docking results of DOTP with key binding targets (target proteins were selected as receptors, and DOTP was selected as the ligand). In the figures in the upper right corner, the blue parts represent the amino acid residues involved in hydrogen bond formation, and the red parts represent the DOTP molecules. The figures in the lower right corner show the 2D interaction analysis, which refers to the interactions between DOTP and the surrounding amino acids during the molecular docking process, including hydrogen bonds and hydrophobic interactions. More information is provided in Table 2.
The results of the two-dimensional interaction analysis demonstrated that DOTP formed van der Waals interactions with nearby amino acids (Fig. 3). In addition to hydrogen bonding interactions, hydrophobic amino acids such as leucine, valine, and alanine contribute to the stable binding of DOTP in the hydrophobic pocket of the protein. Furthermore, a variety of non-covalent synergistic interactions, including alkyl interactions, π-alkyl interactions, and π-stacking, have been observed to facilitate the formation of relatively stable complexes between DOTP and proteins. The findings of this study provide substantial evidence in support of the hypothesis that DOTP exhibits stable binding to target proteins.
Molecular dynamics simulation
In order to determine further the binding ability of DOTP with the four core targets, we performed molecular dynamics simulations to validate the results of molecular docking. The Root Mean Square Deviation (RMSD) metric was used to track changes in molecular structure relative to the initial structure during the course of the simulation. This permitted observation of whether the magnitude of changes tended to stabilize. As illustrated in Fig. 4, the RMSD value of the complex structure underwent a gradual stabilization after 75 ns, indicating that DOTP exhibited a stable binding with the four proteins. It is noteworthy that the lower the RMSD, the more stable the complex structure. Notably, the DOTP-PTGS2 complex exhibited the lowest RMSD (~ 0.22 nm), suggesting highly stable binding.
RMSD values of the complexes as a function of simulation time. RMSD is used to track the changes in the molecular structure relative to the initial structure during the simulation process and observe whether the amplitude of the changes tends to be stable. A lower RMSD indicates that the two structures are closer. The target protein is represented by a purple color, the small molecule ligand (DOTP) by a yellow color, and the complex by a blue color. As can be seen from Figures (A) to (D), the RMSD values of the four complexes gradually stabilize as the simulation progresses.
Principal Component Analysis (PCA) was employed to distill the complex high-dimensional molecular motion data into a few principal components and to identify the most significant motion patterns of the molecules during the simulation (Fig. 5). As illustrated in the figure, the conformational distribution of DOTP in MAPK14 and STAT1 is more dispersed, while the conformational distribution in PTGS2 and NFKB1 is more concentrated, exhibiting a higher frequency conformation. This observation indicates that DOTP exhibits enhanced conformational stability in the presence of PTGS2 and NFKB1. The Radius of Gyration (Rg) was employed as a metric to assess the compactness of the overall small molecule-protein structure. As the simulation progressed (particularly after 50 ns), the Rg values of the four complexes remained largely stable, suggesting that there were no evident structural alterations in the complexes (Fig. 6). During this period, the DOTP-MAPK14 and DOTP-NFKB1 complexes exhibited lower Rg values and more compact structures.
PCA analysis of the molecular trajectories of DOTP in the four complexes. Through PCA analysis, complex high-dimensional molecular motion data can be simplified into a few principal components, and the most significant motion patterns of molecules during the simulation process can be captured. (A) The conformational distribution of DOTP is relatively concentrated, with one conformation having a relatively high occurrence frequency, indicating that its conformation is relatively stable; (B) The conformational distribution of DOTP is relatively dispersed, and there are two conformations with relatively high frequencies simultaneously; (C) There is one conformation of DOTP with a relatively high occurrence frequency, indicating that the conformation of the small molecule is relatively stable; (D) There is one conformation of DOTP with a relatively high frequency, but the conformational distribution is relatively dispersed, indicating that the structure of the small molecule is highly variable.
The RMSD and Rg of the complexes were utilized as the free energy topography of the complexes to demonstrate the possible conformations of the molecules or systems and their relative stability (Fig. 7). The results indicated that all four complexes existed in a lower energy state, suggesting that the overall structure of the complexes was more stable. Of particular interest is the observation of two energy wells in close proximity to the DOTP-PTGS2 complex (Fig. 7A). This observation suggests that the molecules may undergo frequent conformational transitions between these two energy minima to establish a dynamic equilibrium. Concomitantly, the transition between these two stable states is facilitated by the relatively modest energy barriers that must be surmounted for interconversion between the two conformations. This flexibility in conformation enables the complexes to respond adaptively to environmental changes while maintaining stability. The Buried Solvent Accessible Surface Area (Buried SASA) is a crucial metric for assessing the surface exposure of proteins. The results demonstrated that the Buried SASA values of the three complexes, with the exception of DOTP-STAT1, exhibited stability. This finding suggests that the contact area between small molecules and proteins remained stable, with no substantial expansion or contraction occurring subsequent to binding (Fig. 8). Among the complexes studied, DOTP-PTGS2 exhibited the most significant Buried SASA value, approximately 12 nm2, suggesting that this complex displays the strongest intermolecular interactions and the largest contact area (Fig. 8A).
The free energy landscape of the complex plotted using RMSD and Rg of the complex is used to display various possible conformations of the molecule or system and their relative stabilities. As can be seen from Figures A to D, all four complexes exhibit states with relatively low energy, indicating that the overall structures of the complexes are relatively stable.
Buried SASA values of the complexes over time. A larger value of the Buried SASA indicates stronger intermolecular interactions and a larger contact area. Except for Figure D, the Buried SASA values in the other three groups of complexes remain basically stable, indicating that the contact area between the small molecule and the protein remains stable, and their binding remains stable. Among them, Figure A shows a higher Buried SASA value.
In summary, in addition to STAT1, PTGS2, MAPK14, and NFKB1, all of which demonstrated stable binding to DOTP, the complex exhibited notable characteristics. It is noteworthy that the DOTP-PTGS2 complex demonstrated the lowest RMSD value and the highest Buried SASA value. In comparison with the other three complexes, the complete system of DOTP-PTGS2 demonstrated enhanced stability. Consequently, PTGS2 target proteins may play a pivotal role in the induction of PD by DOTP.
Discussion
The effects of the food and environmental pollutant DOTP, a plasticizer, on inflammatory and immune pathways constitute a potential threat to PD. This study identified MAPK14, NFKB1, STAT1, and PTGS2 as primary binding targets of DOTP influencing PD, through the integration of data from multiple databases and the application of network toxicology and molecular docking techniques. Furthermore, MAPK14, NFKB1, and PTGS2 demonstrated remarkable stability in molecular dynamics simulations. Given the established roles of these targets in the pathogenesis of PD, we have placed particular emphasis on them in our discussion.
Mitogen-activated protein Kinase 14 (MAPK14) is a pivotal component of the MAPK signaling pathway family. It plays an important role in cellular stress response and inflammation regulation41. During the pathogenesis of PD, host recognition of cellular components (including lipopolysaccharide, etc.) induces activation of the MAPK14 pathway. The activated MAPK14 subsequently promotes the expression of inflammatory factors, such as IL-6 and MMP-13, which in turn induce further inflammatory responses in the gingiva42.
As demonstrated in previous studies, NFKB1 has been identified as a core transcription factor associated with immunity and inflammation43. Furthermore, the NFKB1 signaling pathway is induced by PD pathogens, which in turn activates the expression of inflammatory factors, such as TNF-α and IL-6, leading to exacerbated periodontal tissue loss. Furthermore, the NFKB1 signaling pathway may interact with other signaling pathways, such as Wnt/β-catenin, which collectively regulate the inflammatory response and cellular function of periodontal tissues44. Notably, the cryotherapy approach proposed by Lin et al.45 has been shown to effectively downregulate the expression of multiple proinflammatory factors, including NFKB1, thereby attenuating inflammation in periodontal ligament cells. This finding provides a novel approach to the treatment of PD.
Signal Transducer and Activator of Transcription 1 (STAT1) is a pivotal pathway that regulates immune and inflammation-related diseases. STAT1 expression has been observed to be significantly increased in cases of chronic and aggressive periodontitis in comparison to healthy subjects46. Furthermore, studies have demonstrated that STAT1 plays a pivotal role in PD with hypertension, and the inhibition of STAT1 in mice has been shown to reduce the expression of pro-inflammatory factors (e.g., IL-6, etc.) and macrophage infiltration, thereby attenuating bone resorption and periodontal destruction in PD lesion areas47. Consequently, STAT1 may play an important role in the development, progression, and regression of PD by regulating the expression of inflammatory factors.
Prostaglandin-Endoperoxide Synthase 2 (PTGS2), also known as Cyclooxygenase-2 (COX-2), is activated by neutrophil extracellular traps via Toll receptors48. During periods of inflammation, there is a significant increase in PTGS2 expression. Prostaglandin E2 (PGE2), catalyzed by PTGS2, is a pivotal inflammatory mediator that instigates vasodilatation and elevates vascular permeability, consequently amplifying the inflammatory response49. A recent study demonstrated that PTGS2 is an important factor in PD that mediates the inflammatory response in periodontal tissues and shows excellent diagnostic efficacy50. Furthermore, molecular dynamics simulations revealed that PTGS2 formed complexes with DOTP, exhibiting optimal stability, thereby underscoring the significance of PTGS2 in these processes. The collective evidence from these studies suggests a collaborative mechanism involving the four key binding targets, working in concert to promote PD progression through a series of interconnected inflammatory pathways.
PD is a complex inflammatory disease whose pathogenesis involves multiple inflammatory and immune pathways. PTGS2, MAPK14, NFKB1, and STAT1, as key binding targets of DOTP-related PD, play important roles in the regulation of these pathways. The results of the KEGG enrichment analysis emphasized that PD-L1 expression and the PD-1 checkpoint pathway in cancer (P = 1.18E-06) are also implicated. It is noteworthy that the expression of PD-1 and PD-L1 was higher in periodontitis patients than in the healthy population, and this was associated with the regulation of the key pathogen Porphyromonas gingivalis (P. gingivalis)51. This finding suggests the potential for therapeutic interventions targeting the inflammatory response of periodontal tissues by modulating the PD-1/PD-L1 pathway regulated by P. gingivalis. Additionally, the AGE-RAGE signaling pathway (P = 2.33E-06) in diabetic complications is closely associated with the progression of PD. It has been demonstrated that hyperglycemia increases the expression of AGEs in periodontal tissues and promotes the release of inflammatory factors by binding to RAGE on the surface of immune cells, thereby exacerbating periodontal tissue destruction52. Concurrently, we observed a significant enrichment of genes such as PTGS2 in the C-type lectin receptors (CLRs) signaling pathway (P = 5.18E−05). This finding suggests a potential association between the CLR signaling pathway and the differentiation of immune cells during PD progression. A recent study demonstrated that the over-expression of Dectin-2 in the CLRs family enhances osteoclast-mediated bone resorption, leading to alveolar bone resorption and promoting PD progression53. The significant enrichment of these pathways underscores the potential impact of DOTP-associated targets on PD, as well as providing a foundation for the future development of preventive and therapeutic strategies.
Despite being an alternative to phthalate plasticizers, DOTP still poses toxicity concerns. There is currently no direct evidence of toxic concentrations of DOTP in periodontal tissues, and research in this area is virtually nonexistent, highlighting the urgency of this study. L Ball et al. suggest an oral reference dose limit for DOTP/DEHT at 0.2 mg/kg/day, which could serve as a guideline54. Although general population exposure to DOTP is low, localized exposure in periodontal tissues may occur under specific conditions, such as prolonged contact with dental materials and medical devices containing DOTP. Future research should focus on the effects of DOTP in these specific scenarios and assess the relationship between local exposure levels and periodontal disease.
This study demonstrates the considerable potential of network toxicology in the assessment of environmental pollutants’ toxicity. In contrast to conventional toxicology, which focuses on a single pollutant or a single toxicity pathway, network toxicology employs multifactorial integration to systematically localize key toxicity drivers. It also utilizes methods such as KEGG and network construction to comprehensively identify cross-pathway nodes interfered with by pollutants. The limitations of conventional toxicology studies, which depend on animal models, become evident when attempting to accurately assess the toxicity of pollutants to humans. Additionally, static experimental models are ill-equipped to capture the dynamic evolution of diseases induced by pollutants55. Conversely, network toxicology facilitates rapid assessment of key toxicity targets of pollutants by constructing molecular networks and molecular docking models. Molecular dynamics simulation has played a pivotal role in this study, as it has been instrumental in overcoming the static nature and high-cost limitations of traditional experimental methods. By dynamically resolving the binding mechanisms of toxic small molecules and targets, molecular dynamics simulation has emerged as a crucial tool in network toxicology. Notably, it is indispensable in elucidating the metathesis effect, solvation effect, and mutation tolerance of the complexes. The integration of network toxicology with these approaches furnishes a formidable instrument for the analysis of health hazards posed by complex environmental exposures. The advancements in this field are poised to exert a substantial influence on environmental regulation, precision preventive medicine, and chemical safety design.
Despite the encouraging potential of network toxicology and molecular dynamics simulations, it is imperative that enhanced sampling techniques (e.g., Metadynamics) are employed to expedite the acquisition of rare events and that dedicated force field parameters are developed for novel contaminants. Meanwhile, previous studies have shown that factors such as smoking, diet, genetics, and alterations in the microbiome also have potential impacts on the progression of periodontitis. Therefore, future research should rely on standardized epidemiological data, multi-omics analysis, intelligent algorithms, and relevant dynamic experimental models to comprehensively analyze how DOTP jointly affects the progression of PD through combined effects with other factors. This systematic research strategy will promote the precision of cyber-toxicology and safe chemical design, making network toxicology a core tool for next-generation environmental health risk management.
Conclusion
In this study, we systematically revealed the relevant targets and potential molecular mechanisms of periodontitis (PD) induced by DOTP, a food and environmental pollutant, by integrating network toxicology, molecular docking, and molecular dynamics simulation. Through multi-database cross-tabulation analysis, 37 targets shared by DOTP and PD were identified in this study, among which PTGS2, MAPK14, NFKB1, and STAT1 were identified as core targets in molecular docking. Enrichment analysis revealed that these targets were significantly enriched in inflammatory responses (e.g., AGE-RAGE signaling pathway, C-type lectin receptor pathway) and immune regulation-related pathways, suggesting that DOTP may synergistically drive inflammatory injury in periodontal tissues through activation of pro-inflammatory factors (e.g., IL-6, TNF-α) and inhibition of anti-inflammatory mechanisms. Molecular dynamics simulations further confirmed that PTGS2, MAPK14, and NFKB1 were stably bound to DOTP. Of particular note, the DOTP-PTGS2 complex demonstrated optimal binding stability, suggesting that PTGS2 may be a core driver target of DOTP-related PD. These findings provide a theoretical basis for mechanism resolution and precise intervention in environmental toxicology and highlight the unique value of network toxicology, an interdisciplinary approach, in the study of complex diseases.
Data availability
Original research data are provided in the supplementary material. Other data in the course of the study can be obtained from the corresponding author upon reasonable request.
Abbreviations
- PD:
-
Periodontitis
- DOTP:
-
Dioctyl terephthalate
- PVC:
-
Polyvinyl chloride
- CSU:
-
Chronic spontaneous urticaria
- GO:
-
Gene ontology
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- BP:
-
Biological processes
- MF:
-
Molecular functions
- CC:
-
Cellular components
- PPI:
-
Protein–protein interaction
- ADT:
-
AutoDockTools
- MD:
-
Molecular dynamics
- FDR:
-
False discovery rate
- RMSD:
-
Root mean square deviation
- PCA:
-
Principal component analysis
- Rg:
-
Radius of gyration
- Buried SASA:
-
Buried solvent accessible surface area
- MAPK14:
-
Mitogen-activated protein kinase 14
- STAT1:
-
Signal transducer and activator of transcription 1
- PTGS2:
-
Prostaglandin-endoperoxide synthase 2
- PGE2:
-
Prostaglandin E2
References
Xu, W. et al. Biomimetic single Al–OH site with high acetylcholinesterase-like activity and self-defense ability for neuroprotection. Nat. Commun. 14(1), 6064. https://doi.org/10.1038/s41467-023-41765-x (2023).
Lo, W. C. et al. Long-term exposure to ambient fine particulate matter (PM2.5) and associations with cardiopulmonary diseases and lung cancer in Taiwan: A nationwide longitudinal cohort study. Int. J. Epidemiol. 51(4), 1230–1242. https://doi.org/10.1093/ije/dyac082 (2022).
Kassebaum, N. J. et al. Global burden of severe periodontitis in 1990–2010: A systematic review and meta-regression. J. Dent. Res. 93(11), 1045–1053. https://doi.org/10.1177/0022034514552491 (2014).
Papapanou, P. N. et al. Periodontitis: Consensus report of workgroup 2 of the 2017 world workshop on the classification of periodontal and peri-implant diseases and conditions. J. Periodontol. 89(Suppl 1), S173–S182. https://doi.org/10.1002/JPER.17-0721 (2018).
Hajishengallis, G. Periodontitis: From microbial immune subversion to systemic inflammation. Nat. Rev. Immunol. 15(1), 30–44. https://doi.org/10.1038/nri3785 (2015).
Geyer, R., Jambeck, J. R. & Law, K. L. Production, use, and fate of all plastics ever made. Sci. Adv. 3(7), e1700782. https://doi.org/10.1126/sciadv.1700782 (2017).
Tran, H. T. et al. Bacterial community progression during food waste composting containing high dioctyl terephthalate (DOTP) concentration. Chemosphere 265, 129064. https://doi.org/10.1016/j.chemosphere.2020.129064 (2021).
Mohamed, D. F. M. S., Tarafdar, A., Lee, S. Y., Oh, H. B. & Kwon, J. H. Assessment of biodegradation and toxicity of alternative plasticizer di(2-ethylhexyl) terephthalate: Impacts on microbial biofilms, metabolism, and reactive oxygen species-mediated stress response. Environ. Pollut. 355, 124217. https://doi.org/10.1016/j.envpol.2024.124217 (2024).
Harmon, P. & Otter, R. A review of common non-ortho-phthalate plasticizers for use in food contact materials. Food Chem. Toxicol. 164, 112984. https://doi.org/10.1016/j.fct.2022.112984 (2022).
Edwards, L. et al. Phthalate and novel plasticizer concentrations in food items from U.S. fast food chains: A preliminary analysis. J. Expo Sci. Environ. Epidemiol. 32(3), 366–373. https://doi.org/10.1038/s41370-021-00392-8 (2022).
Wei, W. et al. Plasticizer sources and concentrations in indoor environments in Europe: A systematic review of existing data. Sci. Total Environ. 972, 179080. https://doi.org/10.1016/j.scitotenv.2025.179080 (2025).
Domínguez-Romero, E. et al. Time-trends in human urinary concentrations of phthalates and substitutes DEHT and DINCH in Asian and North American countries (2009–2019). J. Expo Sci. Environ. Epidemiol. 33(2), 244–254. https://doi.org/10.1038/s41370-022-00441-w (2023).
Lyche, J. L. et al. Reproductive and developmental toxicity of phthalates. J. Toxicol. Environ. Health B Crit. Rev. 12(4), 225–249. https://doi.org/10.1080/10937400903094091 (2009).
Cao, F., Guo, C. & Guo, J. Deciphering CSU pathogenesis: Network toxicology and molecular dynamics of DOTP exposure. Ecotoxicol. Environ. Saf. 291, 117864. https://doi.org/10.1016/j.ecoenv.2025.117864 (2025).
Sczepanik, F. S. C. et al. Periodontitis is an inflammatory disease of oxidative stress: We should treat it that way. Periodontol. 2000 84(1), 45–68. https://doi.org/10.1111/prd.12342 (2020).
Teng, M. et al. Effects of BBIBP-CorV vaccine on gut microbiota and short-chain fatty acids in mice exposed to bis (2-ethylhexyl) phthalate and dioctyl terephthalate. Environ. Int. 190, 108851. https://doi.org/10.1016/j.envint.2024.108851 (2024).
Ye, X. et al. Genetic evidence strengthens the bidirectional connection between gut microbiota and periodontitis: Insights from a two-sample Mendelian randomization study. J. Transl. Med. 21(1), 674. https://doi.org/10.1186/s12967-023-04559-9 (2023).
Di Stefano, M. et al. Impact of oral microbiome in periodontal health and periodontitis: A critical review on prevention and treatment. Int. J. Mol. Sci. 23(9), 5142. https://doi.org/10.3390/ijms23095142 (2022).
Huang, S. Efficient analysis of toxicity and mechanisms of environmental pollutants with network toxicology and molecular docking strategy: Acetyl tributyl citrate as an example. Sci. Total Environ. 905, 167904. https://doi.org/10.1016/j.scitotenv.2023.167904 (2023).
Pinzi, L. & Rastelli, G. Molecular docking: Shifting paradigms in drug discovery. Int. J. Mol. Sci. 20(18), 4331. https://doi.org/10.3390/ijms20184331 (2019).
Hu, X. et al. Molecular dynamics simulation of the interaction of food proteins with small molecules. Food Chem. 405(Pt A), 134824. https://doi.org/10.1016/j.foodchem.2022.134824 (2023).
Zhu, J., Wang, J., Han, W. & Xu, D. Neural relational inference to learn long-range allosteric interactions in proteins from molecular dynamics simulations. Nat. Commun. 13(1), 1661. https://doi.org/10.1038/s41467-022-29331-3 (2022).
Daina, A., Michielin, O. & Zoete, V. SwissTargetPrediction: Updated data and new features for efficient prediction of protein targets of small molecules. Nucleic Acids Res. 47(W1), W357–W364. https://doi.org/10.1093/nar/gkz382 (2019).
Gallo, K., Goede, A., Preissner, R. & Gohlke, B. O. SuperPred 3.0: Drug classification and target prediction-a machine learning approach. Nucleic Acids Res. 50(W1), W726–W731. https://doi.org/10.1093/nar/gkac297 (2022).
Kim, S. et al. PubChem in 2021: New data content and improved web interfaces. Nucleic Acids Res. 49(D1), D1388–D1395. https://doi.org/10.1093/nar/gkaa971 (2021).
UniProt Consortium. UniProt: The universal protein knowledgebase in 2023. Nucleic Acids Res. 51(D1), D523–D531. https://doi.org/10.1093/nar/gkac1052 (2023).
Safran, M. et al. GeneCards Version 3: The human gene integrator. Database (Oxford). 2010, baq020. https://doi.org/10.1093/database/baq020 (2010).
Barrett, T. et al. NCBI GEO: Archive for functional genomics data sets-update. Nucleic Acids Res. 41(Database issue), D991–D995. https://doi.org/10.1093/nar/gks1193 (2013).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43(7), e47. https://doi.org/10.1093/nar/gkv007 (2015).
Sherman, B. T. et al. DAVID: A web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res. 50(W1), W216–W221. https://doi.org/10.1093/nar/gkac194 (2022).
Szklarczyk, D. et al. The STRING database in 2023: Protein–protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res. 51(D1), D638–D646. https://doi.org/10.1093/nar/gkac1000 (2023).
Shannon, P. et al. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 13(11), 2498–2504. https://doi.org/10.1101/gr.1239303 (2003).
Yu, Z. et al. Identification of hub genes and key pathways in arsenic-treated rice (Oryza sativa L.) based on 9 topological analysis methods of CytoHubba. Environ. Health Prev. Med. 29, 41. https://doi.org/10.1265/ehpm.24-00095 (2024).
Bader, G. D. & Hogue, C. W. An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinform. 4, 2. https://doi.org/10.1186/1471-2105-4-2 (2003).
Seeliger, D. & de Groot, B. L. Ligand docking and binding site analysis with PyMOL and Autodock/Vina. J. Comput. Aided Mol. Des. 24(5), 417–422. https://doi.org/10.1007/s10822-010-9352-6 (2010).
Zhang, Y. & Sanner, M. F. AutoDock CrankPep: Combining folding and docking to predict protein-peptide complexes. Bioinformatics 35(24), 5121–5127. https://doi.org/10.1093/bioinformatics/btz459 (2019).
Eberhardt, J., Santos-Martins, D., Tillack, A. F. & Forli, S. AutoDock Vina 1.2.0: New docking methods, expanded force field, and python bindings. J. Chem. Inf. Model. 61(8), 3891–3898. https://doi.org/10.1021/acs.jcim.1c00203 (2021).
Wu, X., Xu, L. Y., Li, E. M. & Dong, G. Application of molecular dynamics simulation in biomedicine. Chem. Biol. Drug Des. 99(5), 789–800. https://doi.org/10.1111/cbdd.14038 (2022).
Yu, D. et al. Application of the molecular dynamics simulation GROMACS in food science. Food Res. Int. 190, 114653. https://doi.org/10.1016/j.foodres.2024.114653 (2024).
Polak, D. & Shapira, L. An update on the evidence for pathogenic mechanisms that may link periodontitis and diabetes. J. Clin. Periodontol. 45(2), 150–166. https://doi.org/10.1111/jcpe.12803 (2018).
Madkour, M. M., Anbar, H. S. & El-Gamal, M. I. Current status and future prospects of p38α/MAPK14 kinase and its inhibitors. Eur. J. Med. Chem. 213, 113216. https://doi.org/10.1016/j.ejmech.2021.113216 (2021).
Kirkwood, K. L. & Rossa, C. Jr. The potential of p38 MAPK inhibitors to modulate periodontal infections. Curr. Drug Metab. 10(1), 55–67. https://doi.org/10.2174/138920009787048347 (2009).
Tak, P. P. & Firestein, G. S. NF-kappaB: A key role in inflammatory diseases. J. Clin. Investig. 107(1), 7–11. https://doi.org/10.1172/JCI11830 (2001).
Milward, M. R. et al. Differential activation of NF-kappaB and gene expression in oral epithelial cells by periodontal pathogens. Clin. Exp. Immunol. 148(2), 307–324. https://doi.org/10.1111/j.1365-2249.2007.03342.x (2007).
Lin, C., Liu, M., Guo, J. & Jia, R. Cryotherapy attenuates inflammation via the lncRNA SNHG1/miR-9-5p/NFKB1 regulatory axis in periodontal ligament cells. Int. J. Mol. Sci. 24(15), 12097. https://doi.org/10.3390/ijms241512097 (2023).
Haftcheshmeh, S. M., Mohammadi, A., Soltani, A., Momtazi-Borojeni, A. A. & Sattari, M. Evaluation of STAT1 and Wnt5a gene expression in gingival tissues of patients with periodontal disease. J. Cell Biochem. 120(2), 1827–1834. https://doi.org/10.1002/jcb.27487 (2019).
Wei, W. et al. Activation of the STAT1 pathway accelerates periodontitis in Nos3-/- mice. J. Dent. Res. 98(9), 1027–1036. https://doi.org/10.1177/0022034519858063 (2019).
Schneider, A. H. et al. Neutrophil extracellular traps mediate joint hyperalgesia induced by immune inflammation. Rheumatology (Oxford) 60(7), 3461–3473. https://doi.org/10.1093/rheumatology/keaa794 (2021).
Martín-Vázquez, E., Cobo-Vuilleumier, N., López-Noriega, L., Lorenzo, P. I. & Gauthier, B. R. The PTGS2/COX2-PGE2 signaling cascade in inflammation: Pro or anti? A case study with type 1 diabetes mellitus. Int. J. Biol. Sci. 19(13), 4157–4165. https://doi.org/10.7150/ijbs.86492 (2023).
Qiu, W. et al. Single-cell atlas of human gingiva unveils a NETs-related neutrophil subpopulation regulating periodontal immunity. J. Adv. Res. https://doi.org/10.1016/j.jare.2024.07.028 (2024).
Liu, X., Yang, L. & Tan, X. PD-1/PD-L1 pathway: A double-edged sword in periodontitis. Biomed. Pharmacother. 159, 114215. https://doi.org/10.1016/j.biopha.2023.114215 (2023).
Blasco-Baque, V. et al. Periodontitis induced by Porphyromonas gingivalis drives periodontal microbiota dysbiosis and insulin resistance via an impaired adaptive immune response. Gut 66(5), 872–885. https://doi.org/10.1136/gutjnl-2015-309897 (2017).
Ye, W. et al. Dectin-2 depletion alleviates osteoclast-induced bone loss in periodontitis via Syk/NOX2/ROS signaling. Free Radic. Biol. Med. 229, 13–29. https://doi.org/10.1016/j.freeradbiomed.2025.01.011 (2025).
Ball, G. L., McLellan, C. J. & Bhat, V. S. Toxicological review and oral risk assessment of terephthalic acid (TPA) and its esters: A category approach. Crit. Rev. Toxicol. 42(1), 28–67. https://doi.org/10.3109/10408444.2011.623149 (2012).
Robinson, N. B. et al. The current state of animal models in research: A review. Int. J. Surg. 72, 9–13. https://doi.org/10.1016/j.ijsu.2019.10.015 (2019).
Funding
This work was supported by the Xinjiang Uygur Autonomous Region Natural Science Foundation Upper-level Program (Grant No. 2023D01C116).
Author information
Authors and Affiliations
Contributions
L.Q conceptualized the idea and was responsible for the design of the study and the accuracy of the data analysis. JJ.W and QG.D performed the statistical analyses and wrote and revised the manuscript. All authors have read and approved the submitted version to be published.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, J., Deng, Q. & Qi, L. Exploring the toxicological effects of DOTP exposure on periodontitis by combining molecular docking and molecular dynamics simulations. Sci Rep 15, 20915 (2025). https://doi.org/10.1038/s41598-025-05740-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-05740-4