Abstract
This study aims to highlight the effectiveness of computer vision (CV) techniques in classifying brain tumors using a comprehensive dataset consisting of computed tomography (CT) scans. The proposed framework comprises six types of brain tumors, including benign tumors (Meningioma, Schwannoma, and Neurofibromatosis) and malignant tumors (Glioma, Chondrosarcoma, and Chordoma). The acquired images underwent pre-processing steps to enhance the dataset’s quality, including noise reduction through median and Gaussian filters and region of interest (ROIs) extraction using an automated binary threshold-based fuzzy c-means segmentation (ABTFCS) approach. A total of 900 CT-scan images were utilized, 150 images per tumor class, each with a size of 512 × 512 pixels, and 4 ROIs taken per image, so the total dataset size is 3600 (900 × 4) attributes. After pre-processing, the dataset was further analysed to extract 135 statistical multi-features for each ROI. An optimized set of 12 statistical multi-features was selected to identify the most relevant features using a feature selection technique based on correlation. For the classification stage, the optimized statistical multi-feature dataset was evaluated using five computer vision classifiers: multilayer perceptron (MLP), BayesNet, PART, random tree, and randomizable filtered classifier, employing a 10-fold cross-validation method. Among these classifiers, MLP with fine-tuned hyperparameters achieved a promising accuracy rate of 97.83%.
Similar content being viewed by others
Introduction
Technology and computers have recently become an integral part of almost every field. This has significantly improved people’s lives, technological advancements, better healthcare, and more. However, despite these positive changes, we still face several challenges1. Additionally, many diseases continue to affect our society, such as cancer, hepatitis, human immunodeficiency virus (HIV), diabetes, heart disease, tuberculosis, and alzheimer’s. Among these, cancer is the leading cause of death and can affect any body part2. The 2018 International Agency for Research on Cancer (IARC) report found that 9.6 million individuals worldwide succumbed to cancer and that 18.1 million new patients with cancer were diagnosed worldwide. Approximately 20% of males and 16% of females are at risk of cancer3.
The human body has several kinds of cells, around 37.2 trillion. The human brain is the only organ with a high concentration of cells: 100 billion neurons and 1 trillion glial cells. Tumors and neoplasms result from abnormal cell development in a specific area that eventually forms a mass. Tumors generally fall into malignant and benign categories4. In contrast to benign tumors, which stay put and do not spread, malignant ones quickly spread to other organs. The brain is the most critical organ, composed of several layers of soft tissue linked by nerve cells to the spinal cord. It forms a quick and constant transmission network of neurons between the brain and spinal cord, controlling our body’s physical, mental, emotional, and spiritual functions. The central and peripheral nervous systems form the entire nervous system5. Unluckily, tumors potentially threaten such a vital and intricate system. About 700,000 Americans are living with primary brain tumors, according to the 2018 annual study issued by the National Brain Tumor Society (NBTS). According to projections, about 86,000 new instances of brain tumors will be identified in 2019. The effects of a brain tumor on the patient and his or her loved ones are devastating. Brain tumors are the leading cause of death in adults and children of all racial and ethnic backgrounds6.
Primary tumors originating in the brain and secondary tumors spreading from elsewhere in the body are the two main categories. Metastatic tumors are cancer cells originating in one part of the body and spreading to the brain, whereas primary brain tumors begin in the brain and remain there. There are two primary categories of tumors: benign and malignant7. Growth of benign brain tumors is often sluggish, and the tumors’ edges become apparent. Depending on the tumors’ location in the brain, they can be surgically removed. However, malignant brain tumors spread rapidly, causing severe symptoms and even death in some instances. Because their roots spread into neighboring tissues, they lack apparent limits. Malignant brain tumors can spread to other brain and spinal cord regions through the cerebrospinal fluid. In the central brain, about 120 distinct tumors have been identified8. Primary brain tumors include gliomas and meningiomas, while metastatic tumors can originate from anywhere in the body and include melanoma9.
Experts like radiologists, neurologists, and doctors can follow a series of steps to diagnose a brain tumor accurately. Suppose the first radiological or neurological evaluation is inconclusive. In that case, the radiologists or neurologists will move on to the contrast agent procedure, with a biopsy being the only way to conclude. Determining the size, location, and direction of the aberrant tissues is the primary focus of a brain tumor diagnosis. Some people with brain tumors experience no symptoms, while others occasionally experience headaches or seizures. Tumors can be diagnosed using various methods, including imaging tests and neurological exams10. In addition to taking a patient’s medical history and physical exam, neurologists conduct a comprehensive neurological evaluation once a tumor is suspected. To this end, imaging scans are carried out to create digital representations of the brain and spinal cord from various perspectives. The clarity and improvement with contrast agents of a magnetic resonance imaging (MRI) scan make it the preferred method. Computed tomography (CT) can stand in for MRI if the former is unavailable; however, its resolution could be better than MRI, and it is likely unable to effectively diagnose lesions in the posterior fossa and spine. Other scans that may be used for diagnosis and staging include perfusion MRI, functional MRI, and PET11. The various supervised, unsupervised, fully-automatic, and partially-automatic computer vision (CV) algorithms have been developed to diagnose brain tumors. The Artificial neural networks (ANN), support vector machines (SVM), decision trees (DT), k-nearest neighbours (KNN), Random Forest (RF), and convolutional neural networks (CNN) are widely used in medical image classification12.
The study’s contribution can be summarized as follows:
-
An innovative method is deployed to fuse information from six different types of brain tumor CT images to enhance multi-brain tumor classification. Every tumor contributes unique characteristics, resulting in improved segmentation and classification.
-
MRI datasets are mainly used to diagnose brain tumors, and it is a priority for doctors because there is less information in a CT scan. Image pre-processing is an essential aspect of our study to enhance the quality of CT-scan images. In this process, we resized all the images and converted them into grayscale for data standardization. Median and Gaussian filters play a significant role in enhancing the quality of images. In order to improve robustness and reduce overfitting, a pre-processing technique is implemented to equalize the data and remove uninformative images.
-
The brain CT image is divided into four regions using irregular polygon-shaped seeds to identify distinct areas. These seeds vary in size and are positioned around the image center to maximize the chances of grouping seeds in the same region. The approach accommodates four ROIs with different sizes, dimensions, and shapes. In the post-processing stage, the improved segmented regions were obtained using automated binary threshold-based fuzzy c-means segmentation.
-
A statistical multi-feature dataset was extracted from the segmented regions. The data cleaning and correlation-based feature selection technique was employed to select the 12 optimized statistical multi-features. This technique helps to identify relevant features for classification.
-
The multilayer perceptron (MLP) was trained with hyperparameter tuning, which optimizes the neural network’s performance. The MLP fine-tunes hyperparameters, contributing to the efficiency and effectiveness of the technique.
The second part of this article contains information about related works. The materials and methods are given in “Materials and methods”. Results and discussion are shown in “Results and discussion ” and the conclusion in “Conclusion”, respectively.
Literature review
Despite advances in MRI-based tumor detection, limited research addresses the challenges of CT-based multi-class tumor classification. Most prior works13,14,15,16,17,18,19,20,21 focus on binary classification or MRI modalities, leaving a gap for comprehensive CT-based multi-class systems, especially those integrating segmentation and statistical optimization techniques. The Reference13 proposed a machine learning (ML) based intracranial hemorrhage (brain stroke) classification system utilizing a Computed Tomography (CT) scan dataset. The join feature selection approach was deployed, and discrete cosine (DC), discrete wavelet (DW), and grey-level co-occurrence matrix (GLCM) features were extracted. These features were extracted from two types (Normal and Abnormal) of brain CT images collected from the Kaggle repository. The ML-based Random Forest (RF) classifier obtained the highest accuracy (87.22%). The Reference14 proposed a brain tumor classification system using feature extraction from Magnetic Resonance Imaging (MRI) images. The Grey Level Co-occurrence Matrix (GLCM) was extracted from MRI images. During the feature selection phase, the validity of the extracted features was measured using the ANOVA F-test. Multiple ML classifiers were tested, but the RF classifier showed the highest accuracy (90.41%) in predicting malignant and benign tumors after the hyperparameter tuning process. The Reference15 proposed that brain tumors significantly cause mortality and disability. Early detection and treatment are crucial for improved patient outcomes. Histology, although time-consuming, is commonly used for brain tumor diagnosis. This paper proposes a deep learning-based convolutional neural network (CNN) method for brain tumor (low or high-grade glioma) classification using cellular features and histology images (HI). The model achieved a 91% accuracy with an F1 score of 0.913, which assists human pathologists. The authors believe this approach performed better in the early diagnosis and treatment of brain tumors. The Reference16 Identifying brain tumors using CT scans is challenging due to the complexity of brain tissue and tumor variations. Deep learning, particularly convolutional neural networks (CNNs), holds potential for this task. This study introduces a novel correlation learning mechanism (CLM) with statistical features based on CNNs to enhance neural network architectures. The CLM model achieved an accuracy of 96% on the three types (Meningioma, Glioma, and Pituitary) of the brain tumor Kaggle dataset, outperforming the baseline CNN model. Additionally, it demonstrated faster information acquisition compared to the standard model.
The Reference17 proposed deep learning methods and the Internet of Medical Things (IoMT) for classifying brain tumor CT scan images. The study focuses on remote patient monitoring and diagnosis using internet-integrated medical devices. The dataset, divided into two classes (normal and abnormal), was collected from two online repositories, Kaggle and ISLES-18. A deep learning algorithm, CNN, and a modified U-Net model show 79% accuracy for classifying brain strokes via deep features. They used various methods and techniques to improve the performance of the deep learning model, highlighting the degradation as an issue in image quality for future researchers. The Reference18 proposed a computed tomography (CT) based brain tumor detection system using deep neural networks (DNN). The foundation of the dataset was split into two classes, tumor and non-tumor, collected from the Mansoura University repository. After converting all CT images into digital format, deep features will be extracted via the MobileNet approach. The pre-trained model MobileNet-V2 shows promising accuracy at 97.6%, while its precision, recall, and F1-score values are 96%, 95%, and 96%, respectively. The Reference19 proposed an automated brain tumor detection method using CT images of intensive care unit (ICU) patients. The approach involves pre-processing CT scans, extracting tumor patches through filtering, segmentation, feature extraction, and utilizing a support vector machine (SVM) machine learning algorithm for classification. The system shows promising accuracy and sensitivity in detecting brain tumors, which can help ICU doctors in early diagnosis and treatment. The study also introduces the zoning method and learning vector quantization to categorize normal and abnormal brains with 85% accuracy and improved computational efficiency. The Reference20 aimed to develop a systematic approach for brain tumor diagnosis using deep convolutional neural networks. An MRI dataset with three brain tumor classes (Ependymoma, Medulloblastoma, and Meningioma) and a normal brain was collected from the Radiopaedia database. The dataset was split into training (70%), test (20%), and validation (10%) sets. The study concludes that utilizing deep CNN for feature extraction and classification in MRI-based brain tumor diagnosis is efficient, achieving high accuracy (96%) with 15 epochs. The Reference21 proposed a brain tumor classification system that involves denoising, feature extraction, and classification. BrainWeb MRI dataset, including standard and abnormal types, was utilized for experimentation. The PURE-LET transform is employed for image denoising to address the issue of noise, a common problem in medical images. A combination of the GLCM and Gray Level Run Length Matrix (GLRLM) was extracted from the MRI dataset. Classifiers such as SVM, K Nearest Neighbours (KNN), and Extreme Learning Machine (ELM) are trained with the extracted features to classify the MRI. The results indicate that the SVM achieves the highest accuracy of 95% compared to other classifiers. While previous studies have predominantly used MRI-based datasets and focused on binary classification tasks, a gap exists in the literature concerning multi-class tumor classification using CT scans. Moreover, existing segmentation methods often struggle with CT noise and lack integration of diverse statistical features for robust classification. This study addresses these gaps by proposing an ABTFCS segmentation framework, applying multi-feature fusion, and optimizing feature selection for enhanced classification of six distinct brain tumor types using CT imaging.
Materials and methods
This study involved using a dataset containing CT images, which were used to classify brain tumors into six different classes. The image dataset consisted of two main categories of brain tumors. Benign brain tumor: This category encompasses three sub-categories, namely Meningioma, Schwannoma, and Neurofibromatosis. Malignant brain tumor: This category comprised three sub-categories, namely Glioma, Chondrosarcoma, and Chordoma. Figure 1 provides a visual representation of these six classes of brain tumors.
This study collected a dataset of patients suffering from brain tumors. The foundation of the CT scan dataset is established via a publicly available brain tumor CT scan repository at Radiopaedia22. Radiopaedia provides educational resources and expert-verified radiology case studies, including benign and malignant brain tumor imaging. Tumor labels were derived from accompanying diagnostic reports and verified by a collaborating radiologist. It can be a valuable resource for understanding the imaging characteristics of these tumors. For each type of brain tumor, 150 CT scan images and 4 ROIs were selected, resulting in a total of 3600 attributes balanced dataset (150 CT scan image \(\:\times\:\) 6 types \(\:\times\:\) 4 ROIs) acquired from CT scan images of brain tumor patients. This ensured equal class distribution and avoided any imbalance-related classification bias. An expert radiologist manually examined these images and utilized various biopsy reports for accurate evaluation. A ground truth brain tumor images dataset facilitated the proposed automated binary threshold-based fuzzy c-means segmentation approach.
Proposed methodology
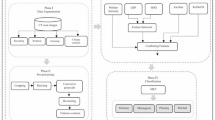
The proposed methodology is outlined in Fig. 2, which describes the main steps of this study as shown in the Algorithm 1.
Now we will explain the proposed methodology steps.
Step 1: In this step, the CT image dataset for the brain tumor is collected. The dataset consists of two main categories: benign and malignant brain tumors.
Step 2: Pre-processing images is the second phase in the process. The first step in the process involves converting the digitized CT scans into a grayscale 8-bit image format. The Second step is to refine the image’s appearance; the “median filter” is utilized throughout the image improvement process. The third step in the process involves the application of “Gaussian filters”23. In conclusion, data cleaning and standardization are done to improve the brain tumor CT Image dataset.
Step 3: During this process stage, segmentation is carried out to remove unnecessary objects, pinpoint the precise location of the lesions, and refine the texture. Extraction of regions of interest (ROI) can be done automatically or semi-automatically using various methods. The proposed method deploys automated binary threshold-based fuzzy c-mean segmentation (ABTFCS) on the CT image dataset of brain tumors to overcome the shortcomings of the two previous approaches24. The demonstration of different types of brain tumors is shown in Fig. 3 to validate the segmentation methodology. This kind of segmentation helps to accomplish accurate segmentation of brain tumors.
Step 4: In the fourth step, statistical multi-features (First-Order and Second-Order) are extracted from the standardized brain tumor CT image dataset. These features include the Co-Occurrence Matrix, which describes the spatial relationship between pixel intensity values25. Spectral features encompass attributes extracted from the frequency domain of a signal26. Histogram Feature: The histogram of pixel intensity values provides statistical information about the image27.
Step 5: The statistical multi-feature dataset is optimized by selecting the 12 best features from the extracted feature dataset. The selection process uses the correlation-based feature selection techniques. These techniques evaluate each feature’s performance and select the most informative based on their discriminative power and correlation with the target variable. The selected features are considered the most relevant and valuable for accurate classification. Table 3 likely presents the feature selection process results and the features’ performance metrics based on their effectiveness in distinguishing brain tumor classes.
Step 6: The final step involves classifying six types of brain tumors using computer vision (CV) classifiers. Five CV classifiers, namely multilayer perceptron, BayesNet, PART, random tree, and randomizable filtered classifier, are employed for this task. The selected optimized statistical multi-feature dataset is used as input to these classifiers. To prevent data leakage, cross-validation with a 10-fold approach is applied to ensure robust evaluation of the classification models. This technique splits the dataset into 10 subsets, performs training and testing on different combinations of the subsets, and provides an averaged assessment of the classifiers’ performance. For the external validation dataset, a 10-fold cross-validation was performed, ensuring no overlap with the primary dataset. This strategy ensures reliable and generalizable performance evaluation across all classifiers.
Image pre-processing
The obtained CT-scan image datasets in 2D format were converted into 8-bit grayscale images to address non-uniformities and enhance contrast28. The grayscale image consists of 256 Gy levels, with the vertical axis representing the pixel values and the horizontal axis ranging from 0 to 255. Equation 1 illustrates the pixel intensity level’s probability density function (p.d.f.).
Here, \(\:0\:\le\:\:\partial\:i\:\le\:\:1\) denotes a pixel’s numeral, \(\:Mi\) represents the pixel intensity, \(\:M\) is the pixel count, and \(\:i\) is 0–255.
While acquisition the brain’s CT image data, speckled noise is detected. This noise can be linked to environmental circumstances impacting the image sensor. In order to solve this problem, a method that eliminates noise is carried out, and it uses Gaussian filters23. This method will resolve the noise issue and the contrast, which will hopefully be improved. The Gaussian filter can be stated mathematically in the following manner:
Let P be a random variable following the Gaussian distribution with a mean of zero and a standard deviation (S.D.) =\(\:\:\omega\:\), i.e., \(\:P\:\sim\text{N}(0,\:{\omega\:}^{2})\). In the unidimensional case, p.d.f. of \(\:P\) is represented by Eq. 2.
Consider \(\:\text{Q}\) as a random variable following the GD with a mean of zero and a standard deviation SD = \(\:{\upomega\:}\), i.e., \(\:\text{Q}\:\sim\:\text{N}(0,\:{{\upomega\:}}^{2})\). In the multidimensional case, the joint the p.d.f of \(\:(\text{P},\:\text{Q})\) is denoted by \(\:{\varrho\:}\left(\text{p},\text{q}\right)\). For the two-dimensional scenario, it is defined in Eq. 3.
Automated binary threshold-based fuzzy c-mean segmentation (ABTFCS)
There are many different methods for image segmentation, most of which depend heavily on human expertise and manual interpretation. While effective in clinical contexts, such approaches can be time-consuming, subjective, and inconsistent. In contrast, the Automated Binary Threshold-based Fuzzy C-Means Segmentation29 (ABTFCS) technique provides a fully automated pipeline without manual supervision. This framework is particularly suited for processing brain CT images to detect regions of interest (ROIs). The ABTFCS method begins with a comprehensive preprocessing stage. The gray-level CT images are first transformed into their binary form using an automated thresholding method. This step distinguishes foreground structures, typically high-intensity tissue regions, from the background. The input image undergoes median and Gaussian filtering to enhance visibility and suppress noise. These filters improve contrast and edge preservation, which are essential for precise segmentation. After preprocessing, the automated binary threshold guides the fuzzy clustering process by narrowing the segmentation domain. Only the regions identified as foreground in the binary mask are subjected to further analysis. This step reduces unnecessary computation and enhances the focus on clinically relevant structures. The core of the segmentation relies on a modified fuzzy C-means clustering algorithm30. In this approach, the dataset is partitioned into two groups based on the minimization of the following objective function:
In the provided equation, where \(\:1\le\:\text{p}\le\:{\infty\:}\), and, \(\:\:{{\upeta\:}}_{\text{a}\text{b}}\)is the degree of membership of \(\:{\text{y}}_{\text{a}}\)in cluster \(\:b\), where \(\:{\text{y}}_{\text{a}}\) is the \(\:ith\) dimensional measured data, \(\:{{\upzeta\:}}_{\text{b}}\) is the dimensional center of the cluster, and \(\:\left|\right|\text{*}\left|\right|\) is any average representing the similarity between measured data and the center. Fuzzy partitioning is achieved through iterative updates of the objective function (O.F.), involving the renewal of membership \(\:{{\upeta\:}}_{\text{a}\text{b}}\) and the cluster centers \(\:{{\upzeta\:}}_{\text{b}}\) as follows:
,
The repetition process halts when the maximum absolute difference between \(\:{\text{m}\text{a}\text{x}}_{\text{a}\text{b}}\left\{\left|{{\upeta\:}}_{\text{a}\text{b}}^{(\text{h}+1)}\:-{{\upeta\:}}_{\text{a}\text{b}}^{\left(\text{h}\right)}\:\right|\right\}<\:{\Xi\:}\), where \(\:{\Xi\:}\) is an elimination criterion between 0 and 1. The variable \(\:h\) represents the repetition steps. This method converges to a local minimum of \(\:{{\upxi\:}}_{\text{p}}\). To avoid manual intervention, threshold values in ABTFCS are selected automatically using global methods (e.g., Otsu). Thresholding serves only as a pre-processing filter to enhance tumor localization, and is followed by FCM-based soft segmentation. This hybrid approach overcomes individual limitations of both methods and achieves robust performance on CT images with varying intensity and noise levels.
The ABTFCS method differs from standard fuzzy C-means segmentation in three significant ways: (1) it incorporates an automated binary thresholding step to define the segmentation scope, (2) it includes contrast-enhancing and noise-reducing filters in the pre-processing phase, and (3) it restricts the clustering operation to a reduced domain defined by the binary mask. These enhancements lead to faster convergence, improved computational efficiency, and increased segmentation accuracy, especially in noisy and low-contrast medical images31. –32 The ABTFCS algorithm was applied to a dataset of brain CT images with tumors. The results, shown in Fig. 3, illustrate the successful delineation of tumor regions, validating the effectiveness of the proposed segmentation framework.
Feature extraction
For this study, a statistical multi-feature brain tumor dataset was obtained using CT-scan images. The statistical dataset includes various features: first-order histogram, second-order co-occurrence matrix (COM) with laws, including five average texture values in all four dimensions (0o, 45o, 90o, 135o) and spectral features. Selecting a series of feature extraction methods is justified due to their ability to capture diverse information types. The histogram reveals overall pixel intensity distributions, spectral features. It offers multi-resolution analysis for detecting patterns at different scales, and COM provides insights into spatial relationships, particularly beneficial for texture analysis. Combining these methods yields complementary features, enhancing the comprehensive representation of image characteristics and potentially improving discrimination between different tumor tissues.
Each region of interest (ROI) has 135 extracted multi-features. Therefore, the total feature vector space (FVS) for the acquired CT-scan image dataset is 486,000 (135 × 3600). All these features were extracted using CVIP Tools version 5.9 h28. The experiments were conducted on a computer system with an Intel® Core i7 processor operating at 4.40 GHz Turbo, 32 GB of RAM, and running Windows 11, 64-bit.
Histogram features
By choosing the object based on rows and columns, histogram features are used in the research context27. The original image is then processed to extract features using this object as a mask. The intensity values of each pixel that is a part of the chosen item are considered while computing the histogram’s characteristics. These features, also known as first-order histograms or statistical features, are obtained from the image’s histogram. Equation 7 describes the first-order histogram probability, denoted as \(\:{\uppsi\:}\)(\(\:\epsilon\)).
N is the number of pixels in the image, and \(\:\kappa\:\left(\epsilon\right)\) is the number of grayscale values \(\:\epsilon\). The Mean feature shows the average image brightness or darkness. It is calculated according to Eq. 8.
The following values of \(\:i\:\left(rows\right)\) and \(\:j\:\left(columns\right)\) indicate the pixel positions in the image. The Standard Deviation (SD) measures the image’s contrast. It is calculated according to Eq. 9.
Skewness (γ) measures the degree of asymmetry in the distribution of values around the central value (Mean, Median, or Mode). It quantifies the departure from symmetry. It is calculated according to Eq. 10.
The gray levels distribution, which represents the overall intensity distribution in the image, is referred to as Energy (ϑ). It is calculated according to Eq. 11.
Entropy (ν) quantifies the level of randomness or uncertainty in the image data. It offers a measurement of the amount of information that is included inside the image. It is calculated according to Eq. 12.
Co-occurrence matrix features (COM)
The COM features, second-order statistical characteristics, are derived from the grey-level co-occurrence matrix (GLCM)25, also known as texture features. These features with laws are formed by considering the distance and angle between individual pixels in the image. Energy (ζ), which represents the calculated energy, is one of these second-order COM features defined by Eq. 13.
Correlation is a second-order COM feature that measures the similarity between pixels at a specific distance. It quantifies the linear relationship between the gray level values of pixels. Correlation is described by Eq. 14.
Entropy is a second-order COM feature that quantifies the total content or information in the image. It captures the level of randomness or uncertainty in the distribution of gray-level values. Entropy is described by Eq. 15.
The Inverse Difference, a second-order COM feature, characterizes the local homogeneity of the image. It measures the similarity or uniformity of neighboring pixel pairs in terms of their gray level values. Equation 16 defines Inverse Difference.
Contrast, or inertia, is a second-order COM feature representing the amount of local variation or contrast in the image. It measures the differences between neighboring pixel pairs regarding their gray level values. Mathematically, inertia is shown in Eq. 17.
Spectral features
Frequency domain-based features, known as spectral features, are precious for texture-based image classification. These features, called rings and sectors, are calculated regarding power within different regions26. The image’s frequency content is examined in spectral feature analysis to extract relevant information. The image is divided into rings and sectors, and the power within each region is computed. These spectral features capture specific frequency characteristics that can aid in texture-based image classification.
Feature selection
In computer vision (CV), feature selection is a significant step. Its primary goal is to determine which elements of a dataset are the most important while simultaneously eliminating those that are not relevant33. According to the findings of this study, different extracted features contributed different amounts to the classification of brain tumors. Because the acquired dataset had many features, the Feature Vector Space (FVS) was 486,000 (135 × 3600), making it challenging to work with. So, we deployed a data cleaning approach and selected only features with 90% or above unique values and observed the FVS 241,200 (67 × 3600). As stated, the study’s objective was to make the most of the capabilities of the dataset in order to guarantee an accurate representation of all the data and accomplish accurate classification while maintaining a low error rate. In order to conquer these obstacles, a supervised feature selection approach known as Correlation-Based Feature Selection (CFS)34 was utilized on the statistical multi-feature dataset. The CFS is intended to extract the essential features of a dataset by applying information theory, which is mainly focused on entropy and is described in Eq. 19.
Equation 20 defines the entropy of the variable A after values of another variable X have been observed.
Here, Q\(\:\left({z}_{j}\right)\) represents the primary probabilities for all possible values of \(\:Z\), while Q\(\:\left(\raisebox{1ex}{${z}_{j}$}\!\left/\:\!\raisebox{-1ex}{${w}_{k}$}\right.\right)\) represents the secondary probabilities of \(\:Z\) when \(\:Y\) is supplied in its various forms. Information extra refers to the value at which the entropy of \(\:Z\) decreases, indicating that \(\:Y\) is providing more information about \(\:Z\) than was previously available. This value is defined by Eq. 21.
If the following inequality holds, then the Y feature has a stronger correlation with feature Z than with feature A:
The symmetrical uncertainty (SUN), which is an expression of the correlation between characteristics, has to be quantified by us. Equation 23 describes it.
In general, entropy-based quantification calls for standard features. However, in this application, it is used to calculate correlations between continuous and discretized data reliably. In this study, the initial dataset contains a sample feature vector space that has been reduced to a smaller size of FVS 43,200 (12 × 3600) using the CFS approach. The optimized statistical multi-feature dataset, as shown in Table 1, is ready for further analysis and classification for six types of brain tumors.
To validate the selection of 12 features, we conducted a comparative analysis using k-fold cross-validation (k = 10). We evaluated the classification accuracy for different feature subset sizes (ranging from 5 to 50). The accuracy significantly improved with the first 10 features, plateauing around 12 features. Beyond this, performance gains were marginal. Thus, 12 features were selected as the optimal subset to balance performance and computational efficiency.
Classification
In this study, five computer vision (CV) classifiers, namely the multilayer perceptron (MLP), BayesNet (BN), PART, Random Tree (RT), and Randomizable Filtered Classifier (RFC), were employed on the brain tumor statistical multi-features dataset, using the WEKA tool version 3.8.6. A Bayesian network classifier for classification tasks is termed a BayesNet (BN) classifier within probabilistic graphical models35. Constructed by specifying variables and their probabilistic dependencies, the Bayesian network captures correlations among features and the target variable. Inference is facilitated by assigning probabilities to each node, parameterized based on training data. When presented with new cases, the classifier employs Bayes’ theorem to compute posterior probabilities for multiple classes, ultimately predicting the class with the highest likelihood. The PART (Partial, pruned decision trees) classifier36 combines rule induction and decision tree techniques for classification applications. Initially, when building a decision tree through methods like C4.5, PART employs a pruning approach to halt tree development when further expansion no longer significantly enhances accuracy. Subsequently, the decision tree is transformed into a set of rules, further refined to enhance readability and simplicity. These streamlined rules constitute the final PART classifier, balancing the transparency of rule-based models and the accuracy of decision trees. The RT Classifier36 is a distinctive decision tree incorporating randomization into its training approach. Unlike a typical decision tree, it selects a random subset of features at each split, reducing overfitting and enhancing generalization. The infusion of randomization contributes to heightened robustness in the tree, diminishing its vulnerability to specific features and yielding a more resilient classifier. The RFC37 integrates a filtering process with a classification method. Intentionally introducing controlled randomness during the learning process contributes to heightened robustness. This classifier employs a filtering mechanism to pre-process the data before engaging in the classification process. Incorporating the “randomizable” attribute enables the adjustment of randomness levels, providing increased adaptability in model behaviour for improved overall performance. Among these classifiers, the MLP classifier demonstrated the best performance. MLP excels in handling noisy, large, and complex datasets.
The MLP classifier38, operates by calculating the weighted sum of the inputs and biases using the summation function \(\:{\theta\:}_{n}\), as defined in Eq. 24. The specific equation is not provided in the given context, but it represents the mathematical operation used in the MLP classifier to calculate the weighted sum of inputs and biases.
In the given equation, ‘k’ represents the number of inputs, \(\:{I}_{j}\) denotes the input variable ‘I’, \(\:{\theta\:}_{j}\) represents the bias term, and \(\:{\eta\:}_{mj}\) denotes the weight. Multiple activation functions exist for Multilayer Perceptron (MLP), one of which is provided below.
The output of neuron j can be obtained as,
Table 2 provides the fine-tuned hyperparameter settings for the MLP classifier, while Fig. 4 illustrates the optimized statistical multi-feature analysis MLP framework, including all the regulatory parameters39.
The “green” colour represents the first layer of the MLP framework and consists of 12 features in the input layer. The second layer, depicted in “red,” represents the hidden layer with 14 neurons. The third layer, highlighted in “yellow,” corresponds to the output layer and comprises six nodes representing the weights of the hidden layers. The regulatory parameters and their respective values are displayed above the framework.
Results and discussion
To classify brain tumors utilizing chosen optimized statistical multi-features, we used five computer vision (CV) classifiers in this study: the multilayer perceptron (MLP), BayesNet (BN), PART, random tree (RT), and randomizable filtered classifier (RFC). The co-occurrence matrix, histogram, and spectral features extracted from CT-scan images were combined to create the statistical multi-feature dataset. These five classifiers had the best accuracy and processing speed performance among the several CV classifiers. Initially, experimentation was performed on a statistical multi-feature dataset without deploying a feature optimization technique. The conclusions derived from this dataset were encouraging regarding the classifiers’ performance. The observed values for MLP, BN, PART, RT, and RFC overall classification accuracy were 97.66%, 96.94%, 96.83%, 96.27%, and 93.36%, respectively. Other performance evaluation factors, including precision (degree of reproducibility and repeatability), true positive (TP) rate, false positive (FP) rate, and kappa statistics (which compares observed accuracy to expected accuracy), were also taken into account in the analysis. The precision, which quantifies the consistency of repeated measurements under constant conditions, is defined by Eq. 27.
The recall is defined in Eq. 28 as the proportion of retrieved relevant instances to all appropriate instances.
Equation 29 shows how to calculate the f-measure derived from recall and precision, commonly known as the f-score.
Table 3 presents various metrics and concepts related to classifier performance, including the Receiver Operating Characteristic (ROC) curve. The ROC curve illustrates the relationship between true and false positive rates at different classifier thresholds. Additionally, the table includes the Root Mean Squared Error (RMSE), which estimates the standard deviation of the discrepancies between expected and observed values, and the Mean Absolute Error (MAE), which measures the proximity of predicted values to actual outcomes. Time Complexity is also provided.
Figure 5 demonstrates that the MLP classifier achieved a higher classification accuracy of 97.66% when applied to a statistical multi-feature dataset, outperforming other classifiers in this study.
The statistical multi-feature confusion matrix in Table 4 illustrates the classification results. The off-diagonal elements represent instances classified into different classes, while the diagonal elements indicate the classification accuracy within each class. The matrix showcases the performance of the MLP classifier, which exhibited higher overall accuracy compared to other classifiers, using actual and predicted data.
The six types of brain tumor for which classification accuracy results were seen to be 94.16%, 98.83%, 99.00%, 97.50%, 98.50%, and 98.00%, respectively. The graphical representation of these accuracy findings is shown in Fig. 6.
Preliminary studies have shown that the results obtained are not promising. The MLP classifier requires much time to build a classification model. The RFC classifier performs poorly due to the noisy and large size of the statistical multi-feature dataset. We deployed the same approach on an optimized statistical multi-feature dataset with 10-fold cross-validation to improve the performance, as shown in Table 5. The conclusions derived from the optimized dataset were encouraging regarding the classifiers’ performance. The observed values for MLP, BN, PART, RT, and RFC overall classification accuracy were 97.83%, 97.63%, 96.94%, 96.91%, and 96.22%, respectively. To statistically validate the superior performance of the MLP classifier, paired t-tests were conducted against all other classifiers using 10-fold cross-validation results. All tests yielded p-values less than 0.05, confirming the statistical significance of the observed accuracy differences. Additionally, 95% confidence intervals were computed for mean accuracy to assess consistency across folds. These results further support the robustness and reliability of the proposed MLP-based classification framework.
Figure 7 demonstrates that the MLP classifier achieved a significantly higher classification accuracy of 97.83% with selected modal parameters, when applied to an optimized statistical multi-feature dataset based on CT scans, outperforming other classifiers used in the study, and also reducing the time to build a classification model.
The optimized statistical multi-feature confusion matrix in Table 6 illustrates the classification results. The off-diagonal elements represent instances classified into different classes, while the diagonal elements indicate the classification accuracy within each class. The matrix showcases the performance of the MLP classifier, which exhibited higher overall accuracy compared to other classifiers, using actual and predicted data.
The six types of brain tumor for which classification accuracy results were seen to be 94.66%, 98.33%, 99.16%, 98.00%, 98.83%, and 98.00%, respectively. The graphical representation of these accuracy findings is shown in Fig. 8. The lower classification accuracy for certain tumors, such as Chondrosarcoma, compared to other tumors, is attributed to overlapping features in their CT images, including similar textures, shapes, and densities. These shared characteristics can make it challenging for the classifier to differentiate between the tumors effectively.
Figure 9 shows the comparative analysis between the statistical multi-features and the optimized statistical multi-features dataset for brain tumor classification using CT-scan. It has been observed that the feature selection approach improves classification accuracy and reduces the time needed to build a classification model.
The comparative analysis between the statistical multi-feature dataset and the optimized statistical multi-feature dataset demonstrates a clear improvement in classification performance and computational efficiency after applying feature selection. Among the five classifiers examined, the multilayer perceptron (MLP) consistently outperformed others in accuracy, precision, and other evaluation metrics, achieving an impressive accuracy on the optimized dataset. The application of feature selection enhanced classifier performance and significantly reduced the model-building time, addressing limitations such as noise and high dimensionality in the original dataset. These findings highlight the potential of optimized statistical multi-feature approaches in improving the diagnostic accuracy of brain tumor classification from CT scans, offering a promising direction for future research and clinical applications. RFC performed comparatively worse due to its sensitivity to high-dimensional, potentially redundant features and lack of internal feature selection. In contrast, MLP handled feature interactions more effectively. Stability was confirmed through multiple seeds and 10-fold cross-validation, showing consistent performance across all experiments.
Lastly, to validate this study, we deployed the proposed model on a privately collected brain tumor dataset from Bahawal Victor Hospital, Bahawalpur, Punjab, Pakistan. The validation dataset, ethical approval was obtained under institutional guidelines, and all patient data was anonymized. For this purpose, 150 attributes were obtained for each of the following types of brain tumors: benign and malignant. The total size of the validation dataset is 300 attributes, with an optimized statistical multi-feature dataset utilized for validation. The proposed method ABTFCS was utilized while employing the CV classifiers shown in Table 7.
As shown in Fig. 10, the evaluation produced promising results, with the MLP, BN, PART, RT, and RFC classifiers reaching classification accuracies of 96.33%, 97.33%, 97.33%, 96.66% and 95%, respectively.
Due to discrepancies in the multi-institutional brain tumor image databases, the experimental results indicated variations. It is essential to promote the creation of a worldwide platform for medical patient data to resolve these variations and enhance the precision with which medical health issues are addressed. Such a platform would allow for more precise analysis and solutions because it would include various patient modes, areas, demographics, geographies, and medical histories. A comparison diagram between the proposed and validation optimized statistical multi-feature CT-scan datasets is shown in Fig. 11, illustrating the variances and differences between the two datasets.
The previous studies have shown that most research has been done on MRI datasets to classify brain tumors. Researchers conducted experiments on different datasets and obtained promising results. Fewer researchers have proposed the CT scan dataset for brain tumor classification, and mainly used two classes (normal and abnormal) just for diagnosis. Considering all these facts, we concluded that our proposed methodology would open a new horizon in multi-brain tumor classification using CT scans. Table 8 compares the proposed methodology (PM) and the current state-of-the-art techniques used in this study.
Conclusion
This research study primarily focuses on classifying six types of brain tumors: meningioma, schwannoma, neurofibromatosis, glioma, chondrosarcoma, and chordoma. The classification uses CT scan images and texture analysis, employing a statistical multi-feature approach. The study’s main objectives include ABTFCS, selection of optimized statistical multi-features, and identifying the best classifiers for efficient classification. Firstly, five computer vision classifiers, namely MLP, BN, PART, RT, and RFC, were utilized with the statistical multi-feature dataset and observed 97.66%, 96.94%, 96.83%, 96.27%, and 93.36%, respectively. Secondly, the same classifiers were deployed in an optimized statistical multi-feature dataset; the observed values for MLP, BN, PART, RT, and RFC overall classification accuracy were 97.83%, 97.63%, 96.94%, 96.91%, and 96.22%, respectively. While all classifiers yielded satisfactory results, the MLP classifier with fine-tuned hyperparameters demonstrated exceptionally high accuracy compared to the others. Overall, an accuracy of 97.66% was achieved across all six types of brain tumors using a multi-feature dataset, but it takes much time to build a model. While using an optimized multi-feature dataset, a model was built quickly, and overall, an accuracy of 97.83% was achieved across all six types of brain tumors. The individual accuracies obtained by the MLP classifier for the six classes were as follows: chondrosarcoma (94.66%), chordoma (98.33%), glioma (99.16%), meningioma (98.00%), neurofibromatosis (98.83%), and schwannoma (98.00%).
Strength of work
Diagnosing brain tumors is of great importance in modern times. In previous research, most researchers used CT scan datasets to diagnose brain tumors, divided into normal and abnormal. Our proposed methodology is innovative from other studies because we have selected six types of brain tumors. After pre-processing and segmentation, optimized statistical multi-features were selected for classification. The MLP classifiers with fine-tuned hyperparameters show excellent accuracy, which will be helpful for future researchers and radiologists to classify and detect brain tumors efficiently.
Future work
In subsequent research, this study will be conducted with various other types and modalities of brain tumors. While this study focuses on optimized statistical multi-feature classification using computer vision, future work will include benchmarking against deep learning methods such as CNNs and MobileNet on the same CT dataset to assess comparative strengths and generalization performance further.
Data availability
The CT-scan dataset is publicly available at Radiopaedia (https://radiopaedia.org), Brain tumor CT-scan repository. The optimized hybrid features datasets used and analyzed during the current study available from the corresponding author on reasonable request.
References
Senbekov, M. et al. The recent progress and applications of digital technologies in healthcare: A review. Int. J. Telemed. Appl. (2020).
Tiwari, S. & Talreja, S. Do you think disease and disorder are same? –Here is the comparative review to brash up your knowledge. J. Pharm. Sci. Res. 12 (4), 462–468 (2020).
Motlana, M. K., Ginindza, T. G., Mitku, A. A. & Jafta, N. Spatial distribution of cancer cases seen in three major public hospitals in KwaZulu-Natal, South Africa. Cancer Inform. 20, 11769351211028194 (2021).
Vilella, F. & Simon, C. Reproductive medicine, as seen through single-cell glasses. Fertil. Steril. 115 (2), 296–297 (2021).
Nersesjan, V. et al. Central and peripheral nervous system complications of COVID-19: a prospective tertiary center cohort with 3-month follow-up. J. Neurol. 268, 3086–3104 (2021).
Miller, K. D. et al. Brain and other central nervous system tumor statistics, 2021. Cancer J. Clin. 71 (5), 381–406 (2021).
Mukhtar, M. et al. Nanomaterials for diagnosis and treatment of brain cancer: Recent updates. Chemosensors 8(4),117z (2020).
Hamblin, M. R. Photodynamic therapy for cancer: what’s past is prologue. Photochem. Photobiol. 96 (3), 506–516 (2020).
Biratu, E. S., Schwenker, F., Ayano, Y. M. & Debelee, T. G. A survey of brain tumor segmentation and classification algorithms. J. Imaging. 7 (9), 179 (2021).
Ithayan, J. V. et al. Machine learning approach for brain tumor detection. J. Surv. Fisheries Sci. 10 (4S), 793–802 (2023).
Abedalthagafi, M., Mobark, N., Al-Rashed, M. & AlHarbi, M. Epigenomics and immunotherapeutic advances in pediatric brain tumors. NPJ Precision Oncol. 5 (1), 34 (2021).
Barragán-Montero, A. et al. Artificial intelligence and machine learning for medical imaging: A technology review. Phys. Med. 83, 242–256 (2021).
Gudadhe, S., Thakare, A. & Anter, A. M. A novel machine learning-based feature extraction method for classifying intracranial hemorrhage computed tomography images. Healthc. Analytics. 3, 100196 (2023).
Vijithananda, S. M. et al. Feature extraction from MRI ADC images for brain tumor classification using machine learning techniques. Biomed. Eng. Online 21(1), 52 (2022).
Ker, J., Bai, Y., Lee, H. Y., Rao, J. & Wang, L. Automated brain histology classification using machine learning. J. Clin. Neurosci. 66, 239–245 (2019).
Woźniak, M., Siłka, J. & Wieczorek, M. Deep neural network correlation learning mechanism for CT brain tumor detection. Neural Comput. Appl. (2023) 35:14611–14626, https://doi.org/10.1007/s00521-021-05841-x
Omarov, B., Tursynova, A. & Uzak, M. Deep learning enhanced internet of medical things to analyze brain computed tomography images of stroke patients. Int. J. Adv. Comput. Sci. Appl., 14(8), 668-676, (2023) http://dx.doi.org/10.14569/IJACSA.2023.0140874
Dawood, N. M. et al. Brain tumors detection using computed tomography scans based on deep neural networks. Vol. 12 (2023).
Fahmi, F., Apriyulida, F. & Nasution, I. K. Automatic detection of brain tumor on computed tomography images for patients in the intensive care unit. J. Healthc. Eng. 2020, 13 (2020).
Mohammed, B. A. & Al-Ani, M. S. An efficient approach to diagnose brain tumors through deep CNN. Math. Biosci. Eng. 18, 851–867 (2020).
Devi, M. & Maheswaran, S. An efficient method for brain tumor detection using texture features and SVM classifier in MR images. Asian Pac. J. Cancer Prevention: APJCP. 19 (10), 2789 (2018).
Brain Tumor Repository & Radiopaedia. https://radiopaedia.org/ . Accessed 13 June 2024 (2024).
Zhang, L. et al. Finger vein image enhancement based on guided tri-Gaussian filters. ASP Trans. Pattern Recognit. Intell. Syst. 1 (1), 17–23 (2021).
Cai, L., Gao, J. & Zhao, D. A review of the application of deep learning in medical image classification and segmentation. Ann. Transl. Med. 8(11) (2020).
Hussain, L. et al. Lung cancer prediction using robust machine learning and image enhancement methods on extracted gray-level co-occurrence matrix features. Appl. Sci. 12(13), 6517 (2022).
Bandara, W. G. C. & Patel, V. M. Hypertransformer: A textural and spectral feature fusion transformer for pansharpening, In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition. 1767–1777 (2022).
Girsang, N. D. Classification of Batik images using multilayer perceptron with histogram of oriented gradient feature extraction. In Proceeding International Conference on Science and Engineering. Vol. 4. 197–204 (2021).
Umbaugh, S. E. Digital Image Processing and Analysis: Computer Vision and Image Analysis (CRC, 2023).
Chowdhary, C. L., Mittal, M. P. K., Pattanaik, P. A. & Marszalek, Z. An efficient segmentation and classification system in medical images using intuitionist possibilistic fuzzy C-mean clustering and fuzzy SVM algorithm. Sensors 20 (14), 3903 (2020).
Latif, G., Alghazo, J., Sibai, F. N., Iskandar, D. A. & Khan, A. H. Recent advancements in fuzzy C-means based techniques for brain MRI segmentation. Curr. Med. Imaging Reviews. 17 (8), 917–930 (2021).
Kumar, V. P., Pattanaik, S. R. & Kumar, V. S. An automated brain tumor segmentation and classification using adaptive bayesian fuzzy clustering. Appl. Soft Comput. 175, 113061 (2025).
Aslam, M. A., Munir, M. A. & Cui, D. Noise removal from medical images using hybrid filters of technique. J. Phys. Conf. Ser. 1518(1), 012061 (IOP Publishing, 2020).
Mishra, P., Biancolillo, A., Roger, J. M., Marini, F. & Rutledge, D. N. New data preprocessing trends based on ensemble of multiple preprocessing techniques. TrAC Trends Anal. Chem. 132, 116045 (2020).
Mohamad, M. et al. Enhancing big data feature selection using a statistical correlation-based feature selection. Electronics 10 (23), 2984 (2021).
Albini, E., Rago, A., Baroni, P. & Toni, F. Relation-Based counterfactual explanations for bayesian network classifiers. In IJCAI, pp. 451–457 (2020).
Dou, Y. & Meng, W. Comparative analysis of weka-based classification algorithms on medical diagnosis datasets. Technol. Health Care (preprint). 1–12 (2023).
Mirmozaffari, M., Alinezhad, A. & Gilanpour, A. Data mining classification algorithms for heart disease prediction. Int’l J. Comput. Commun. Instrum. Engg. 4 (1), 11–15 (2012).
Desai, M. & Shah, M. An anatomization on breast cancer detection and diagnosis employing multi-layer perceptron neural network (MLP) and convolutional neural network (CNN). Clin. eHealth 4, 1–11 (2021).
Zhang, J., Li, C., Yin, Y., Zhang, J. & Grzegorzek, M. Applications of artificial neural networks in microorganism image analysis: a comprehensive review from conventional multilayer perceptron to popular convolutional neural network and potential visual transformer. Artif. Intell. Rev. 56 (2), 1013–1070 (2023).
Acknowledgements
I would like to express my sincere gratitude to the editor and the three anonymous reviewers for their insightful comments and constructive suggestions, which greatly contributed to improving the clarity, quality, and rigor of this article. Their valuable feedback helped refine the arguments and strengthen the overall presentation of the research.
Author information
Authors and Affiliations
Contributions
A.A.: Conceptualization, Methodology, Data curation, Investigation, Writing – original draft. X.L.: Project administration, Resources, Validation, Supervision. W.K.M: Data curation, Validation, Writing – review & editing. M.A.: Formal analysis, Funding acquisition, Investigation, Validation. M.T.: Data curation, Writing – review & editing. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ali, A., Li, X., Mashwani, W.K. et al. Computer vision based efficient segmentation and classification of multi brain tumor using computed tomography images. Sci Rep 15, 32198 (2025). https://doi.org/10.1038/s41598-025-16825-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-16825-5