Abstract
To address the challenge of medical image segmentation caused by blurred edges, some researchers have leveraged edge information to improve segmentation performance. However, the current mainstream edge enhanced medical image segmentation networks are limited by the unidirectional information flow mechanism of the encoder decoder architecture, which may affect the network’s inference accuracy for complex anatomical structures. In this paper, we propose a novel edge guided bidirectional iterative network in medical image segmentation (EGBINet), which adopts a cyclic architecture to enable bidirectional flow of edge information and region information between the encoder and decoder, thereby enhancing segmentation performance. Specifically, complementary information is generated by fusing edge features with multi-level region features, constructing an enhanced feedforward information pathway from the encoder to the decoder. Within the feedback mechanism from the decoder to the encoder, region feature representations and edge feature representations are reciprocally propagated, enabling iterative optimization of hierarchical feature representations. This bidirectional flow allows the encoder to dynamically respond to the decoder’s requirements. Furthermore, to improve the aggregation quality of local edge information and multi-level global regional information, we introduce a transformer-based multi-level adaptive collaboration module (TACM). TACM groups local information and multi-level global information, adaptively adjusts their weights according to the aggregation quality, significantly improving the feature fusion quality. Experimental results on multiple medical image segmentation datasets demonstrate that our proposed EGBINet achieves remarkable performance advantages compared to state-of-the-art methods, particularly in edge preservation and complex structure segmentation accuracy, validating the superiority of our proposed network architecture.
Similar content being viewed by others
Introduction
In recent years, deep learning technology has demonstrated remarkable application potential in the field of medical image processing1,2,3, with a particular focus on achieving precise automatic localization and edge delineation of pathological regions. Affected by multiple factors such as biological tissue diversity, pathological structural complexity, and imaging noise interference, target organs and lesion areas often encounter technical challenges during segmentation, including ambiguous edge definition and insufficient data integrity4,5,6.
The UNet architecture7 and its variants have become the backbone of medical image segmentation8,9,10. UNet’s contracting path for feature extraction and expanding path for localization set a foundational framework, but its unidirectional information flow (encoder to decoder) limits dynamic adaptation to segmentation demands. Subsequent improvements can be categorized by their attempts to address specific limitations, though none fully resolve the lack of bidirectional interaction: In network architecture optimization, Res-UNet11 introduced residual learning to alleviate vanishing gradients, enhancing deep network stability. However, it retained the one-way information flow, leaving encoders unaware of decoder needs. Attention U-Net12 embedded attention gates to focus on salient regions, improving feature discrimination but still operating within a unidirectional framework. UNet++13 used nested skip connections and deep supervision to reduce parameters, while DRU-Net14 combined dense connections and residuals for efficiency-both optimized feature propagation but lacked mechanisms for encoder-decoder feedback, a limitation our model directly overcomes.
Edge enhancement has seen significant advances, yet most methods remain constrained by unidirectional inference. ET-Net15 learned edge attention and fused low-level edge features with decoded features, but its encoder could not refine representations based on decoder-derived edge demands. MSEF-Net16 integrated Sobel operators and edge-aware attention to improve MRI edge quality, while FFS-Net17 leveraged spatial-frequency-edge interplay for gland boundaries–both enhanced edge extraction but relied on static encoder outputs. EMidDiff18 integrated diffusion models with edge information, yet its encoder-decoder interaction remained one-way.
While existing studies have advanced network architecture, edge enhancement, and anatomical modeling, they share a critical limitation: reliance on unidirectional, single-pass information flow from encoder to decoder. Encoders lack real-time feedback from decoders, preventing adaptive refinement of features to match decoding demands, and constraining decoder performance.
To address this, we propose a novel edge guided bidirectional iterative network in medical image segmentation (EGBINet), which adopts a cyclic architecture to enable bidirectional flow of edge information and region information between the encoder and decoder, thereby enhancing segmentation performance. Specifically, complementary information is generated by fusing edge features with multi-level region features, constructing an enhanced feedforward information pathway from the encoder to the decoder. Within the feedback mechanism from the decoder to the encoder, region feature representations and edge feature representations are reciprocally propagated, enabling iterative optimization of hierarchical feature representations. This bidirectional flow allows the encoder to dynamically respond to the decoder’s requirements. Furthermore, to improve the aggregation quality of local edge information and multi-level global regional information, we introduce a transformer-based multi-level adaptive collaboration module (TACM). TACM groups local information and multi-level global information, adaptively adjusts their weights according to the aggregation quality, significantly improving the feature fusion quality. Experimental results on multiple medical image segmentation datasets, including automated cardiac diagnosis challenge (ACDC)19, medical image computation and computer-assisted intervention atrial segmentation challenge (ASC)20 and infrapatellar fat pad (IPFP)21, demonstrate that our proposed EGBINet achieves remarkable performance advantages compared to state-of-the-art methods, particularly in edge preservation and complex structure segmentation accuracy, validating the superiority of our proposed network architecture.
Contributions:
-
1.
We propose a novel edge guided bidirectional iterative network in medical image segmentation. This network adopts a cyclic architecture that enables bidirectional flow of edge information and region information between the encoder and decoder, thereby enhancing the segmentation performance of medical images.
-
2.
We propose a transformer-based multi-level adaptive collaboration module, which groups local information and multi-level global information, adaptively adjusts their weights according to the aggregation quality, significantly improving the feature fusion quality.
-
3.
Compared with the current state-of-the-art medical image segmentation networks on medical image segmentation datasets (ACDC, ASC and IPFP), our method achieves superior performance.
Related work
Medical image segmentation
Medical image segmentation focuses on dividing medical images into clinically meaningful sub-regions to support diagnosis and treatment planning. Its technological development has gradually evolved from traditional methods relying on hand-crafted features to deep learning methods with end-to-end learning as the core22. Among these, the UNet architecture and its variants, leveraging an encoder-decoder structure with skip connections, effectively balance high-level semantic feature extraction and low-level spatial detail preservation, thus becoming the fundamental models in this field7. The Transformer architecture23, by virtue of its self-attention mechanism, can accurately capture long-range anatomical correlations across regions in medical images, making up for the limitations of local receptive fields in convolution-based models such as UNet. The Segment Anything Model (SAM)24, through large-scale general image pre-training and a flexible prompt interaction mechanism, provides stronger generalization and adaptability for medical image segmentation, enabling it to quickly respond to the segmentation needs of different modalities and different organs. However, most current frameworks still have key limitations: on the one hand, they adopt a unidirectional information flow from the encoder to the decoder, lacking a feedback mechanism from the decoder to the encoder, which prevents adaptive refinement of features according to segmentation requirements; on the other hand, they often treat regional segmentation and edge delineation in isolation, ignoring the complementary correlation between the two, which restricts the further improvement of segmentation accuracy.
Edge enhancement
Edge enhancement is crucial for medical image segmentation, as clear boundary information can effectively distinguish adjacent tissues and directly improve the clinical usability and accuracy of segmentation results. Deep learning-based edge enhancement methods significantly optimize edge representation by deeply integrating edge learning into the entire segmentation workflow. Specifically, ET-Net15 learned edge attention and fused low-level edge features with decoded features, but its encoder could not refine representations based on decoder-derived edge demands. EANet25 designs a dedicated edge-aware attention module to enhance feature responses for low-contrast boundary regions in medical images while suppressing noise interference. ECM-TransUNet26 combines edge-enhanced multi-scale transposed attention and multi-scale convolutional state space modules to improve feature extraction and spatial consistency modeling. However, existing edge enhancement frameworks still have two key drawbacks: First, they rely on a unidirectional, single-pass information flow from the encoder to the decoder. The encoder lacks real-time feedback from the decoder, which hinders the adaptive refinement of features to match decoding requirements and limits the performance of the decoder; Second, there is edge-region disconnection–some methods insufficiently design the interactive fusion of edge features and regional features, leading to inconsistencies between the predicted edge masks and the final regional segmentation results at the boundary positions, which restricts the further improvement of segmentation accuracy.
Feature fusion
Feature fusion is a core component in image processing27,28, which aims to integrate complementary information from multi-scale and multi-modal features to address the limitations of single-feature representations. In the early stage, fusion methods centered on Convolutional Neural Network (CNN) dominated this field7. However, the local receptive field of CNN makes it difficult to model long-range anatomical correlations. With the rise of Transformer, its self-attention mechanism has provided a new paradigm for global dependency modeling, and related fusion methods have developed rapidly: Swin-UNet29 achieves multi-scale fusion through hierarchical window-based attention, while TransUNet23 combines CNN and Transformer to balance local details and global context. Nevertheless, existing methods still have shortcomings: some overlook the fine-grained fusion of edge and regional features, and others rely on fixed structures, lacking anatomical adaptability. Thus, there is an urgent need for a Transformer-based feature fusion strategy that is more suitable for the needs of medical segmentation.
Methods
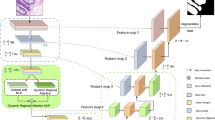
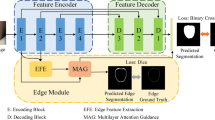
The overall architecture of the EGBINet proposed in this article is shown in Figure 1. In the first stage, the input image is processed by the encoder (Such as VGG19, ResNet50, Vision transformer (ViT), et al. In this paper, we adopt the VGG19) to extract five encoded features at different scales, denoted as \(E^{1}_{i}\) (i = 1, 2, 3, 4, 5). These initial multi-scale encoded features provide abundant semantic information for the subsequent iterative optimization process.
The overall architecture of the EGBINet. \(E^{i}_{1}\), \(E^{i}_{2}\), \(E^{i}_{3}\), \(E^{i}_{4}\) and \(E^{i}_{5}\) denotes extracting encoding regional features of different scales in the i stage, \(D_{edge}^i\) denotes extracting edge features in the i stage, \(D^{i}_{1}\), \(D^{i}_{2}\), \(D^{i}_{3}\) and \(D^{i}_{4}\) denotes decoding regional features of different scales in the i stage.
To implement the inference in the first stage, we adopt a progressive decoding strategy inspired by UNet7 for processing regional features. Specifically, multi-layer convolutional blocks are utilized to achieve cross-layer fusion between different feature hierarchies,
Here, \(D_i^1\) denotes the decoded regional features at the i-th layer in the first stage, and Con represents the multi-layer convolutional blocks. In the second stage, these decoded regional features will be fed back to the encoder.
For edge feature, we aggregate both local edge information (\(E^{1}_{2}\)) and global positional information (\(E^{1}_{5}\)) to extract edge features. Subsequently, multi-layer convolutional blocks are employed to achieve multi-scale feature fusion.
Here, \(D_{edge}^1\) denotes edge feature in the first stage. In the second stage, edge features will be fed back to the encoder.
In the second stage, the edge feature \(D_{edge}^1\) and the decoded multi-level regional features \(D_i^1\) are fed back to the encoder to emphasize target-related information. This process can be regarded as introducing prior knowledge of the target to guide the encoder’s feature extraction. We first introduce transformer-based multi-level adaptive collaboration module (TACM) to achieve high-quality aggregation of local edge information and multi-level global regional information,
Here, \(R_i^1\) denotes the aggregated features combining the edge features from the first stage and the decoded regional features at the i-th layer. TACM represents the TACM module. Down indicates the down sampling operation. Subsequently, \(R_i^1\) is fed back to the encoder and fused with the corresponding encoder features,
Here, \(E_i^2\) denotes the encoder regional features in the second stage. \(\otimes\) and \(\oplus\) denote element-wise multiplication and addition operations, respectively. Subsequently, the encoder regional features from the second stage are decoded,
Here, \(D_i^2\) represents the decoder regional features at the i-th layer in the second stage. The edge features in the second stage are as follows,
Here, \(D_{edge}^2\) denotes edge feature in the second stage. By iteratively performing the aforementioned steps, we establish a bidirectional flow framework between encoder and decoder features. Finally, we select the decoder regional features from the last iteration and fuse them with edge features to generate the final prediction,
Here, \(P_i^j\) denotes the decoder regional prediction results at the i-th layer in the j-th stage, and \(D_i^j\) represents the decoder regional features at the i-th layer in the j-th stage.
Transformer-based multi-level adaptive collaboration module
Traditional fixed weight fusion applies a uniform proportion for fusion regardless of the utility of feature subgroups at different positions, which may give rise to several issues. For instance, it might cause the coarse-grained features of regional subgroups to overshadow the details of edge subgroups, leading to confusion between lesion boundaries and normal tissues. We introduce a transformer-based multi-level adaptive collaboration module (TACM). TACM customizes weights for each spatial position and each feature subgroup: subgroups that are more useful for the current position are assigned higher weights to dominate the fusion result, while useless or interfering subgroups have their weights reduced to minimize negative impacts. This is illustrated in Fig. 2. Given two input feature sets, \(S \in R^{H \times W \times C}\) and \(T \in R^{H \times W \times C}\) (H, W, and C denote the height, width, and number of channels of the image, respectively), we first divide them into multiple subgroups along the channel dimension: \(S = \{ S_1, S_2, S_3,..., S_N \}\), \(T = \{ T_1, T_2, T_3,..., T_N \}\). Subsequently, we perform fusion between corresponding subgroups of \(S_N\) and \(T_N\) based on Swin transformer29,
Here, \(F_n\) denotes the fused subgroups features. represents the Swin transformer operation. indicates the concatenation operation. Second, we compute the maximum response at each position across all subgroups features,
Here, P(x, y, c) denotes the maximum response map. (x, y) represent the spatial coordinates, while c denotes the channel. Third, we compute the dice similarity coefficient between each subgroup \(F_n(x,y,c)\) and P(x, y, c),
Here, \(Dice_n\) denotes the dice similarity score between the n-th subgroup \(F_n(x,y,c)\) and the maximum response P(x, y, c). Fourth, assign weights to each subgroup based on their dice similarity coefficient with P(x, y, c),
Here, FNN denotes feedforward neural network. Finally, the N subgroups are concatenated and processed through a convolutional block to output the final fused result \(F_{TACM},\)
Loss function
The loss function \(L_{total}\) in this work consists of two components: the edge loss \(L_{edge}\) and the regional loss \(L_{region},\)
Here, the regional loss \(L_{region}\) combines binary cross-entropy (BCE) loss with dice loss, while the edge loss \(L_{edge}\) adopts a weighted BCE loss.
Experiments
Datasets, preprocessing, implementation details and evaluation metrics
Datasets and preprocessing
We evaluated our network on ACDC19, ASC20 and IPFP21 datasets. The ACDC dataset contains 100 3D cardiac magnetic resonance imaging (MRI) cases with physician-annotated ground truths: right ventricle (RV), myocardium (Myo), and left ventricle (LV). All slices were uniformly cropped to 224\(\times\)224 pixels. The ASC dataset includes 154 3D cardiac MRI cases with physician-annotated ground truths: left atrial (LA). We used 100 cases for training and 54 for testing. All slices were uniformly cropped to 224\(\times\)224 pixels. The IPFP dataset includes 195 3D infrapatellar fat pad MRI cases, and doctors have annotated the basic fact: infrapatellar fat pad (IPFP). We used 138 cases for training and 57 cases for testing. All slices are uniformly cropped to 224 \(\times\) 224 pixels. All datasets have not undergone data augmentation. All slices underwent normalization (0-1) prior to network training and testing.
Implementation details
Our network is implemented in PyTorch and runs on an RTX 3090 card. All networks in this paper adopt the same training and preprocessing methods. All networks were trained using the Adam optimizer with a batch size of 8 and a learning rate of \(5 \times 10^{-4}.\)
Evaluation metrics
We employed the 95% Hausdorff distance30 (HD95) and dice similarity coefficient to evaluate the segmentation performance,
Here, A and B denote the prediction result and ground truth, respectively.
Ablation experiments and analyses
Impact of TACM on segmentation performance
We conducted experiments on the ASC dataset to evaluate the impact of TACM on segmentation performance. We compared TACM with other region and edge feature aggregation strategies: (a) convolutional neural network (CNN), (b) transformer (Transformer), (c) TACM. The results are presented in Table 1. As shown in Table 1, the segmentation performance of the TACM-based regional and edge aggregation approach outperforms both CNN-based and T-based approaches, demonstrating the superiority of the proposed TACM method. Figure 3 shows the feature map of TACM. As can be seen from Fig. 3, the fused image regional and edge are clearer, which further proves the progressiveness of TACM proposed in this paper.
Impact of backbone on segmentation performance
Experiments were conducted on the ASC dataset to evaluate the impact of backbone networks on segmentation performance. We compared different backbone networks, including (a) VGG19, (b) ResNet50, and (c) ViT. The results are presented in Table 2. As indicated in Table 2, VGG achieved the optimal performance. This may be attributed to the fact that VGG captures richer edge information. Medical images, especially MRI images, often have indistinct edges, and the inclusion of rich edge information facilitates improving segmentation performance.
Impact of the number of iterative feedback steps on segmentation performance
Experiments were conducted on the ASC dataset to evaluate the impact of the number of iterative feedback steps on segmentation performance. We compared different number of iterative feedback steps, including (a) No feedback, (b) Feedback iteration once (Once), (c) Feedback iteration twice (Twice) and (d) Feedback iteration three times (Three Times). The results are presented in Table 3. As indicated in Table 3, the segmentation performance improves progressively with the increase in the number of iterations; however, when the number of iterations is greater than or equal to Twice, the segmentation performance no longer significantly improves with further increases in the number of iterations. Since a larger number of iterations leads to a larger number of network parameters, we thus select Twice as the number of iterations in this paper.
Comparison with the state-of-the-art
We compared the proposed EGBINet with state-of-the-art medical image segmentation networks on both ACDC, ASC and IPFP datasets. The results are presented in Tables 4, 5 and 6. The proposed EGBINet achieved optimal performance on all three datasets, demonstrating that the proposed EGBINet is advanced. Figures 4, 5 and 6 visualize the segmentation results of the proposed EGBINet and other networks. As shown in Figs. 4, 5 and 6, under conditions of unclear edges, the proposed EGBINet exhibits superior segmentation performance compared to other networks.
The number of parameters average (Params), floating point operations (FLOPs), Frames Per Second (FPS) of our method are shown in Table 7. As presented in Table 7, the Params and FLOPs of our method are relatively large, while the FPS is relatively slow. However, medical image segmentation has relatively low requirements for high real-time performance. Our method achieves a significant improvement in segmentation performance, thus demonstrating a certain degree of practicality.
Discussion
Medical image segmentation is divided into traditional methods and deep learning methods. Currently, due to the superior performance of deep learning methods, traditional methods receive less attention, yet they still hold reference value. Statistical shape models (SSMs) are based on prior statistical knowledge: they are constructed using feature points on tissue edges, with parameters adjusted to gradually approximate the tissue, and essentially model edge information–this aligns with the edge feature extraction goal of our method. The strength of SSMs lies in their ability to utilize prior knowledge of tissue shapes, while their weakness is that constructing feature points and initial positioning of the model may require manual operation. Combining traditional and deep learning methods may be a future research direction.
Conclusion
To address the challenge of insufficient segmentation accuracy caused by blurred edges in medical images, this study proposes an innovative edge guided bidirectional iterative network in medical image segmentation. Its core contributions are as follows: First, we construct a bidirectional cyclic information flow mechanism that enables dynamic interactions between the encoder and decoder, achieving multi-level collaborative optimization of edge features and regional features. This bidirectional propagation mode allows the encoder to adaptively adjust feature representations in real-time based on the decoder’s segmentation demands, significantly enhancing the segmentation precision of complex anatomical structures. Second, we introduce a transformer-based multi-level adaptive collaboration module, which efficiently integrates local edge information with global multi-scale contextual information through a dynamic weight allocation mechanism. Experimental results on multiple medical image segmentation datasets demonstrate that our proposed EGBINet achieves remarkable performance advantages compared to state-of-the-art methods, particularly in edge preservation and complex structure segmentation accuracy, validating the superiority of our proposed network architecture.
Data availability
The datasets used and analysed during the current study available from the Shaolong Chen (E-mail: chenshlong@mail2.sysu.edu.cn) on reasonable request.
References
Zhao, Y., Zhou, X., Pan, T., Gao, S. & Zhang, W. Correspondence-based generative bayesian deep learning for semi-supervised volumetric medical image segmentation. Comput. Med. Imaging Graph. 113, 102352 (2024).
Zhang, Z., Li, Y. & Shin, B.-S. Robust color medical image segmentation on unseen domain by randomized illumination enhancement. Comput. Biol. Med. 145, 105427 (2022).
Zhao, J. & Li, S. Evidence modeling for reliability learning and interpretable decision-making under multi-modality medical image segmentation. Comput. Med. Imaging Graph. 116, 102422 (2024).
Wang, R. & Zheng, G. Pfmnet: prototype-based feature mapping network for few-shot domain adaptation in medical image segmentation. Comput. Med. Imaging Graph. 116, 102406 (2024).
Wu, J. et al. Medical sam adapter: adapting segment anything model for medical image segmentation. Med. Image Anal. 102, 103547 (2025).
You, X., He, J., Yang, J. & Gu, Y. Learning with explicit shape priors for medical image segmentation. IEEE Trans. Med. Imaging (2024).
Ronneberger, O., Fischer, P. & Brox, T. U-net: convolutional networks for biomedical image segmentation. In International Conference on Medical Image Computing and Computer-Assisted Intervention 234–241 (Springer, 2015).
Li, X. et al. Dmsa-unet: dual multi-scale attention makes unet more strong for medical image segmentation. Knowl.-Based Syst. 299, 112050 (2024).
Liu, X., Gao, P., Yu, T., Wang, F. & Yuan, R.-Y. Cswin-unet: transformer unet with cross-shaped windows for medical image segmentation. Inf. Fusion 113, 102634 (2025).
Tang, H. et al. Rm-unet: Unet-like mamba with rotational ssm module for medical image segmentation. SIViP 18, 8427–8443 (2024).
Shankaranarayana, S. M., Ram, K., Mitra, K. & Sivaprakasam, M. Joint optic disc and cup segmentation using fully convolutional and adversarial networks. In International Workshop on Ophthalmic Medical Image Analysis 168–176 (Springer, 2017).
Oktay, O. et al. Attention u-net: learning where to look for the pancreas. arXiv:1804.03999 (2023).
Zhou, Z., Siddiquee, M. M. R., Tajbakhsh, N. & Liang, J. Unet++: redesigning skip connections to exploit multiscale features in image segmentation. IEEE Trans. Med. Imaging 39, 1856–1867 (2019).
Jafari, M., Auer, D., Francis, S., Garibaldi, J. & Chen, X. Dru-net: an efficient deep convolutional neural network for medical image segmentation. In 2020 IEEE 17th International Symposium on Biomedical Imaging (ISBI) 1144–1148 (IEEE, 2020).
Zhang, Z. et al. Et-net: a generic edge-attention guidance network for medical image segmentation. In International Conference on Medical Image Computing and Computer-Assisted Intervention 442–450 (Springer, 2019).
Zhao, J. et al. Msef-net: multi-scale edge fusion network for lumbosacral plexus segmentation with mr image. Artif. Intell. Med. 148, 102771 (2024).
Luo, Y. B. et al. Ffs-net: fourier-based segmentation of colon cancer glands using frequency and spatial edge interaction. Expert Syst. Appl. 262, 125527 (2025).
Tang, Q., Zhu, Q., Xiong, Y., Xu, Y. & Du, B. Edge-and-mask integration-driven diffusion models for medical image segmentation. IEEE Signal Process. Lett. (2024).
Bernard, O. et al. Deep learning techniques for automatic mri cardiac multi-structures segmentation and diagnosis: is the problem solved?. IEEE Trans. Med. Imaging 37, 2514–2525 (2018).
Xiong, Z. et al. A global benchmark of algorithms for segmenting the left atrium from late gadolinium-enhanced cardiac magnetic resonance imaging. Med. Image Anal. 67, 101832 (2021).
Chen, S., Zhong, L., Qiu, C., Zhang, Z. & Zhang, X. Transformer-based multilevel region and edge aggregation network for magnetic resonance image segmentation. Comput. Biol. Med. 152, 106427 (2023).
Yi, Y. et al. Mfnet: multi-fusion network for medical image segmentation. Digital Signal Process. 163, 105219 (2025).
Chen, J. et al. Transunet: rethinking the u-net architecture design for medical image segmentation through the lens of transformers. Med. Image Anal. 97, 103280 (2024).
Ma, J. et al. Segment anything in medical images. Nat. Commun. 15, 654 (2024).
Wang, K. et al. Eanet: iterative edge attention network for medical image segmentation. Pattern Recogn. 127, 108636 (2022).
Lv, C. et al. Ecm-transunet: edge-enhanced multi-scale attention and convolutional mamba for medical image segmentation. Biomed. Signal Process. Control 107, 107845 (2025).
Talreja, J., Aramvith, S. & Onoye, T. Xtnsr: Xception-based transformer network for single image super resolution. Complex Intell. Syst. 11, 162 (2025).
Talreja, J. & Chauhan, D. Advanced computational techniques: Bridging metaheuristic optimization and deep learning for material design through image enhancement. In Metaheuristics-Based Materials Optimization 197–228 (Elsevier, 2025).
Cao, H. et al. Swin-unet: Unet-like pure transformer for medical image segmentation. In European Conference on Computer Vision 205–218 (Springer, 2022).
Taha, A. A. & Hanbury, A. An efficient algorithm for calculating the exact hausdorff distance. IEEE Trans. Pattern Anal. Mach. Intell. 37, 2153–2163 (2015).
Jagadeesh, B. & Kumar, G. A. Brain tumor segmentation with missing mri modalities using edge aware discriminative feature fusion based transformer u-net. Appl. Soft Comput. 161, 111709 (2024).
Ding, H., Zhang, X., Lu, W., Yuan, F. & Luo, H. Mmaformer: multiscale modality-aware transformer for medical image segmentation. Electronics 13, 4636 (2024).
Acknowledgements
This work was supported in part by Science and Technology Planning Project of the Guangdong Science and Technology Department under Grant Guangdong Key Laboratory of Advanced IntelliSense Technology (2019B121203006), Science and Technology Planning Project of Shenzhen City Polytechnic (2511001).
Funding
This work was supported in part by Science and Technology Planning Project of the Guangdong Science and Technology Department under Grant Guangdong Key Laboratory of Advanced IntelliSense Technology (2019B121203006), Science and Technology Planning Project of Shenzhen City Polytechnic (2511001).
Author information
Authors and Affiliations
Contributions
S.C. conceived the experiments, X.P. conducted the experiment, X.P. and S.C. analysed the results. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Peng, X., Chen, S. Edge guided bidirectional iterative network in medical image segmentation. Sci Rep 15, 39304 (2025). https://doi.org/10.1038/s41598-025-23085-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-23085-w