Abstract
Accurate and early diagnosis of malaria from peripheral blood smear images remains a critical challenge in healthcare, particularly in resource-limited settings. In this work, we propose an optimized convolutional neural network (CNN) framework enhanced by Otsu thresholding-based image segmentation for improved detection of malaria-infected cells. A dataset of 43,400 blood smear images was utilized, divided into a 70:30 ratio for training and testing. A baseline 12-layer CNN achieved 95% accuracy, which improved to 97% with the integration of EfficientNet-B7 through a hybrid parallel feature-fusion model. Further enhancement was achieved using Otsu-based segmentation, where preprocessing emphasized parasite-relevant regions while retaining morphological context in the RGB images. This approach yielded the highest accuracy of 97.96%, reflecting a ~ 3% gain over the baseline CNN. To ensure the reliability of the segmentation step, we created a manually annotated subset of 100 images and computed quantitative segmentation metrics by comparing Otsu-generated masks with reference masks. The method achieved a mean Dice coefficient of 0.848 and Jaccard Index (IoU) of 0.738, confirming that Otsu segmentation effectively isolates parasitic regions despite its simplicity. Five-fold cross-validation was also performed, yielding consistent results (94.8%, 96.9%, and 97.8%), thereby supporting the robustness of the framework. The proposed pipeline demonstrates that simple yet effective preprocessing can significantly boost CNN-based classification while maintaining interpretability and computational feasibility. These findings suggest that segmentation-driven deep learning frameworks can play a vital role in developing reliable, scalable, and cost-effective malaria diagnostic tools.
Similar content being viewed by others
Introduction
The severe infectious illness known as malaria, which is caused by parasites belonging to the genus Plasmodium, continues to be a serious concern to the public health of the whole world. There are about 200 million cases of malaria that are recorded each year, and it is estimated that 400,000 people die from the disease each year. The majority of these fatalities occur in children under the age of five in sub-Saharan Africa1. It is of the utmost importance to make a timely and correct diagnosis in order to effectively treat and control the condition. Therefore, developments in diagnostic procedures are essential in order to battle the broad effects of the disease.
Over the course of medical history, the manual microscopic inspection of stained blood smears has been considered the gold standard for diagnosing malaria. This method relies on the visual identification of Plasmodium parasites that are present inside red blood cells. However, making use of this technology requires a significant amount of manual labour, a significant amount of time, and the skills of professional microscopists. As a consequence of this, there has been an increasing focus on the development of automated image analysis tools in order to improve the effectiveness and precision of the screening process for malaria. Deep learning, and more specifically convolutional neural networks (CNNs), has emerged as a strong tool in the field of medical image analysis in recent years. It has shown great effectiveness in a variety of applications, including the identification of anomalies and illnesses from medical pictures. The purpose of this study is to contribute to the current efforts to automate the diagnosis of malaria by focusing on the use of a unique CNN-EfficientNet hybrid model for the segmentation and classification of blood cells that have been infected with malaria.
Background
Malaria may present itself in a number of different phases, and the life cycle of the parasite involves both the human host and the Anopheles mosquito, which is the vector of the disease. Within the human circulation, the parasites infect red blood cells, generating specific morphological changes that can be visually detected under a microscope. These alterations may be seen by the naked eye. Having the capacity to precisely recognise and categorise these morphological changes is very necessary in order to diagnose the existence of the infection as well as the severity of the sickness2. When it comes to this particular scenario, automated image analysis techniques play a crucial role since they provide an alternative to manual inspection that is both more efficient and perhaps more reliable. The CNN-EfficientNethybrid model that has been presented takes use of the benefits that are associated with both conventional CNN architectures and the most advanced EfficientNet, therefore optimising the balance between the complexity of the model and the computing efficiency of the system. The purpose of this hybridization is to improve the model’s capacity to extract significant characteristics from pictures of blood cells that have been infected with malaria, which will eventually lead to advancements in classification accuracy.
Problem statement
Malaria, an infectious illness caused by Plasmodium parasites and spread by infected female Anopheles mosquitoes, is a major worldwide health concern. Precise and prompt diagnosis is essential for efficient disease control and prevention. Traditional diagnostic techniques often depend on the manual inspection of blood smears, a process that may be time-consuming, demanding in terms of labor, and susceptible to human mistakes. Consequently, there is an urgent want for automated and very precise diagnostic techniques that may speed up the detection of blood cells infected with malaria. Previous attempts in utilizing deep learning models for computer-aided diagnosis of malaria have shown potential; however, challenges remain in attaining the utmost precision and comprehensibility. The key problem addressed by this study is the need for a comprehensive and dependable classification system for blood cells infected with malaria. The study seeks to address the drawbacks of existing diagnostic approaches by introducing a hybrid deep learning technique that integrates a 12-layer CNN model with the efficientNet-b7 model. In addition, the research focuses on the need for enhanced image segmentation methods, particularly employing Otsu’s threshold method to increase feature extraction and further boost the diagnostic accuracy of the model. The study addresses these issues, making a valuable contribution to the improvement of malaria diagnoses. This enables more efficient disease control and eventually aids worldwide initiatives to lessen the impact of malaria.
Objectives
This research aims to accomplish several crucial goals:
-
To develop and evaluate a Convolutional Neural Network (CNN) model for accurate classification of malaria-infected and uninfected red blood cells.
-
To enhance the model’s discriminative capability by applying Otsu’s thresholding technique for segmentation of blood smear images prior to CNN training.
-
To analyze the impact of segmentation on classification performance by comparing CNN results on original versus Otsu-segmented datasets.
-
To validate the effectiveness of the segmentation approach using visual inspection and Canny edge detection, in the absence of ground-truth annotations.
-
To perform a comparative analysis between the proposed CNN model and other transfer learning-based models (e.g., EfficientNet, Inception-V3) to highlight the effectiveness of segmentation-based preprocessing in improving malaria detection performance.
Contribution of the paper
This study significantly contributes to the field of automated malaria diagnostics by demonstrating the effectiveness of Otsu’s thresholding-based image segmentation in improving the classification performance of a CNN model. The primary contributions of this work can be categorized into three main areas: segmentation-based preprocessing, enhanced classification accuracy, and model interpretability. First, the study proposes the application of Otsu’s thresholding as a preprocessing step to segment blood smear images. This technique enables the extraction of key parasitic regions from raw microscopic images, thereby reducing background noise and enhancing the visibility of relevant features. Second, a standard 12-layer CNN was trained on both the original and the Otsu-segmented datasets. While the CNN trained on the original dataset achieved an accuracy of 95%, the model trained on the segmented dataset achieved an accuracy of 97.96%, representing a 2.96% absolute improvement. This improvement also slightly surpasses the CNN–EfficientNet-B7 hybrid model (97.0%), underscoring that segmentation plays a more decisive role in boosting performance than architectural complexity alone. Third, the study conducts visual inspection and edge detection-based validation of the segmentation process to qualitatively assess the effectiveness of Otsu’s method in isolating parasitized regions in the absence of pixel-wise ground truth annotations. Additionally, the proposed CNN model is compared with other well-known deep learning approaches (e.g., EfficientNet-B7, Inception-V3) and lightweight architectures (e.g., SqueezeNet variants), demonstrating that Otsu-segmentation combined with a simple CNN architecture achieves competitive or superior results compared to more complex or computationally heavy models, with clear quantitative performance gains.
Finally, this work highlights the practical value of segmentation-enhanced deep learning in medical imaging. The methodology improves not only classification accuracy but also model interpretability, providing clearer visual cues about infection regions that can aid in clinical decision-making and further research. By emphasizing segmentation as a performance-enhancing preprocessing strategy, this paper offers a computationally efficient, interpretable, and scalable solution for automated malaria detection using deep learning.
Paper’s structure
-
i.
Section 2 comprises a Literature Review. It investigates current studies on automated malaria detection, with a specific emphasis on image analysis methods and the use of deep learning technologies.
-
ii.
Section 3 presents a Methodology that describes the structure and training procedure of the CNN-EfficientNet hybrid model, along with the use of Otsu’s thresholding technique for cell segmentation.
-
iii.
Section 4 illustrates the findings and displays the outputs of the trained model, including the accuracy of classification and the results of segmentation, which have been verified by Visual Inspection and Edge Detection. Evaluates the performance of the hybrid model concerning transfer-learning models often used in medical image analysis.
-
iv.
Finally, Sect. 5 concludes the research findings. Provides a concise overview of the research’s accomplishments and highlights its importance in furthering the field of automated malaria detection.
Literature review
Malaria, a potentially fatal communicable illness widespread in tropical and subtropical areas, is a significant worldwide health issue. Due to the growing need for prompt and precise diagnosis, there has been a transition in the area towards automated image processing methods. This literature review presents a comprehensive examination of current studies on automated malaria diagnosis, with a specific emphasis on image analysis techniques, deep learning implementations, and the difficulties related to segmentation and classification.
Traditional diagnostic methods
In the past, the main technique used for diagnosing malaria has been the manual microscopic inspection of stained blood smears. This laborious technique, which depends on the knowledge and ability of experienced microscopists, has inherent limitations, such as subjectivity and the possibility of human mistakes. The need for more effective and dependable diagnostic techniques has stimulated the investigation of automated methodologies3.
Image analysis techniques
The use of automated image analysis has become more important in the field of malaria detection because it can overcome the constraints associated with human approaches. Several image processing approaches, such as segmentation and feature extraction, have been used to detect malaria parasites in blood cell pictures. These strategies are crucial in improving the precision of later classification algorithms4.
Deep learning in malaria diagnosis
Deep learning, specifically Convolutional Neural Networks (CNNs), has shown exceptional achievements in the field of medical picture processing in recent times. Convolutional Neural Networks (CNNs) are very proficient in acquiring hierarchical characteristics, making them highly suitable for identifying intricate patterns in pictures of blood cells infected with malaria. Scientists have investigated several structures, such as VGG, RestNet, and Inception, to enhance the accuracy of categorization5,6,7,8.
Transfer learning in medical image analysis
Transfer learning, a method that uses pre-trained models on extensive datasets, has been extensively utilized in medical image processing applications. Transferring information from models trained on broad-picture datasets to specialized medical domains, such as malaria diagnosis, has shown the potential to enhance classification accuracy. Nevertheless, it is crucial to thoroughly evaluate the significance of pre-trained models and their capacity to adjust to various datasets9.
EfficientNet architecture
EfficientNet, a breakthrough in neural network topologies, incorporates a compound scaling technique to achieve a harmonious balance between model depth, breadth, and resolution. This methodology attains cutting-edge performance while using much fewer parameters, making it attractive for applications that prioritize the effective use of resources. The literature review examines the use of EfficientNet in medical image analysis, emphasizing its potential benefits in terms of computing efficiency and model performance10,11.
Challenges in segmentation and classification
Although there have been improvements, difficulties still exist in accurately dividing and categorizing blood cells infected with malaria. Challenges arise from factors such as differences in cell shape, variations in picture clarity, and the lack of standardized datasets with well-labeled reference information, which provide substantial obstacles12. The examination of current research emphasizes the need to tackle these obstacles to improve the dependability of automated approaches for diagnosing malaria13,14,15,16.
Gap in existing research
Although prior research has achieved significant advancements in automated malaria detection, there is still a need to investigate hybrid models that use the advantages of conventional CNN architectures while also including the efficiency of models such as EfficientNet. Moreover, the lack of accurate reference labels for certain datasets presents a distinct difficulty that necessitates inventive techniques for validating segmentation. This study aims to bridge this gap17,18,19.
Summary
The literature review traces the progression of malaria detection techniques from manual microscopic inspection to computerized image analysis. The text emphasizes the significance of deep learning, transfer learning, and innovative architectures like as EfficientNet in enhancing the accuracy of categorization. The difficulties in segmenting and classifying data highlight the need for creative methods, which serve as the basis for using the CNN-EfficientNet hybrid model and segmentation validation methodologies in this study. The next parts will provide a comprehensive explanation of the approach and results of the planned study, which will contribute to the progress of automated malaria diagnosis. Some of the previous studies are listed in Table 1.
Methodology
The given methodology improves the diagnosis of malaria through the combination of Otsu threshold-based segmentation of images and a typical Convolutional Neural Network (CNN). First, the RGB microscopic images of the blood smear will be readjusted to the size of 224 × 224 × 3, then augmented with the rotation, flipping, shifting, and zooming operations. Otsu thresholding is then used to segment the parasitized areas with an aim to provide more concentration on the infected areas and minimized background noise. Such segmented images are fed into a two-convolutional layers (32 and 64 filters), two max-pooling layer, flattening layer, two dense layers (128 and 2 neurons) of an 12-layered CNN architecture to generate final binary classification (parasitized vs. un-parasitized). Segmented data can be used to train a CNN, which enhances the quality of the model in terms of accuracy and its interpretability because the model can focus on biologically meaningful features. Contrary to hybrid or deeper models, this effective method shows better performance due to the fact that it uses segmentation to enhance learning performance. Figure 1 shows the whole process which includes preprocessing, segmentation, training the model, and the classification part.
Dataset
The careful selection and curation of a broad and representative dataset are crucial for the success and generalization capacity of any machine learning model. The dataset-gathering strategy in this work involves a rigorous and comprehensive approach to guarantee the inclusion of varied photos that capture distinct phases of malaria infection. The main goal was to generate a dataset that accurately reflects the intricate and diverse challenges experienced in real-life clinical situations.
The collection comprises approximately 43,400 high-resolution (Fig. 2) photos of blood cells infected with malaria. These images were obtained from reliable medical databases, research organizations, and public sources. Particular emphasis was placed on including photos that covered the whole range of malaria infection phases produced by several Plasmodium species. By including a variety of photos, the model is exposed to a broad range of morphological variances, hence improving its ability to effectively apply learned knowledge to new and unfamiliar data. Every picture went through meticulous preparation procedures to standardize the format, resolution, and color channels. To mitigate possible biases and improve the model’s resilience, deliberate attempts were made to include photos from a wide range of geographical locations, people, and laboratory settings. The variety included in the dataset is of utmost importance, particularly when considering the worldwide occurrence of malaria and the intrinsic variances in physical characteristics associated with distinct Plasmodium species.
In addition, to mitigate overfitting and guarantee unbiased assessments, the dataset was divided into three distinct subsets: a training set used for optimizing the model, a validation set used for fine-tuning hyperparameters and avoiding overfitting, and a testing set utilized for unbiased evaluation of the trained model (The class balance and split are summarized in Fig. 3). Data distribution and split. Bar chart showing the number of Parasitized and Uninfected images and the 70:30 train–test split used in all experiments. Values indicate counts per class and per split. This partitioning technique follows established principles in machine learning to guarantee that the model’s performance measures precisely represent its capacity to generalize to novel, unseen data. Aside from guaranteeing diversity, ethical issues were of utmost importance in the dataset-gathering process. The photos were anonymized to safeguard patient confidentiality, and strict safeguards were implemented to adhere to relevant data protection legislation. Ensuring ethical management of medical data is crucial for building confidence and promoting responsible use in scientific research. The carefully selected dataset, which contains a wide range of morphological variants and different representations of malaria infection phases, is essential for training, verifying, and assessing the proposed hybrid model, which combines an EfficientNet with the Convolutional Neural Network (CNN) architecture. The model’s comprehensive design allows it to acquire intricate patterns related to various phases of malaria infection, hence enhancing its resilience and practical use. The dataset, which takes into account ethical issues and emphasizes variety, serves as the basis for the succeeding stages of model building and assessment in this study.
Preprocessing
Preprocessing plays a vital role in creating strong and precise models, particularly in the field of medical picture analysis. Important factors to consider for preprocessing are target_size, batch_size, subset, and class_mode. The target size parameter is a resizing parameter that optimizes the trade-off between computational efficiency and the preservation of spatial information. Batch size refers to the amount of samples that are processed in each iteration during a computational task. It is used to optimize the utilization of computing resources and to mitigate problems such as overfitting or underfitting. A subset is selected for the training phase when the pictures are subjected to essential modifications and augmentations to improve the model’s capacity to generalize to unfamiliar data. The class mode is set to ‘binary’ and is specifically intended for instances involving binary classification, such as distinguishing between infected and uninfected blood cells. Image augmentation methods are often used to increase the variety of the training dataset.
The selection of the preprocessing configuration is crucial for the efficacy of machine learning models in properly categorizing blood cells infected with malaria. The preprocessing pipeline establishes a strong and adaptable training process by downsizing pictures to a standardised dimension, optimizing the batch size, prioritizing the training subset, and employing a binary class mode. Preprocessing plays a crucial role not only in numerical aspects, but also in the ability of models to identify important traits, make accurate predictions on new data, and help to automated malaria diagnosis.
Model architecture: 12-Layer CNN
This research utilized a 12-layer Convolutional Neural Network (CNN) model to classify images of blood cells infected with malaria. The architecture begins with an initial layer that resizes grayscale images to a standardized format, followed by convolutional layers that capture low-level information. The convolutional operation can be represented as shown in Eq. ()1)
where:
-
yi, j,k is the output feature map at position (i, j) for the k-th feature map,
-
x is the input feature map,
-
w is the filter/kernel of size M×N,
-
b is the bias term.
Activation functions (ReLU) are used to introduce non-linearity, defined as shown in Eq. 2
Batch normalization stabilizes and accelerates the training process by normalizing the inputs to each layer, which can be represented as Eq. (3) and Eq. (4)
where:
-
µ and σ2 are the mean and variance of the batch,
-
γ and β are learnable parameters,
-
\( \in \) is a small constant for numerical stability.
Max pooling layers reduce spatial dimensions while preserving crucial information. The max pooling operation can be represented as shown in Eq. (5)
where:
-
R represents the pooling region,
-
x and y are the input and output feature maps.
The flattened layer converts the 3D output into a 1D vector, which prepares the data for input into dense layers. If the input feature map has dimensions h×w×d (height, width, depth), the flatten operation reshapes it into a vector of size h×w×d.
Dense layers are used to extract complex features and can be represented as shown in Eq. (6)
where:
-
x is the input vector,
-
W is the weight matrix,
-
b is the bias vector,
-
f is the activation function (e.g., ReLU, sigmoid).
Dropout layers mitigate overfitting by randomly deactivating a fraction of neurons during training. The final output layer, typically using a sigmoid or softmax activation function, generates the model’s output. The model undergoes training using optimization methods such as stochastic gradient descent (SGD) or Adam optimization. Hyperparameters are fine-tuned to balance convergence speed and overfitting prevention. The training process involves multiple epochs, allowing iterative adjustment of parameters and improving classification performance. The design of this 12-layer CNN architecture is optimized for analyzing malaria-infected blood cell images, balancing model complexity and effective feature extraction. Layer-wise description is shown in Table 2, and the Schematic diagram is shown in Fig. 4.
Model architecture: CNN-efficientnet hybrid
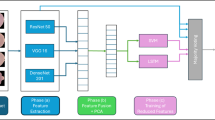
The CNN-EfficientNet-B7 hybrid approach is implemented as a parallel feature-fusion system that integrates the advantages of a traditional 12-layer CNN and the current-state-of-the-art system EfficientNet-B7. CNN Branch: 12-layer CNN takes the passed blood smear images and extracts low-level and morphological feature as cell boundaries, textures, and fine structural variations. EfficientNet-B7 Branch: Parallel to this, EfficientNet-B7 is a high-capacity feature extractor with high level semantic representations and contextual information of the identical input image. Its principle of scaling compound scaling enables extraction of features deeper, wider and with higher-resolution, yet with computational efficiency. Feature Fusion: At the dense layer phase the feature vectors of the two branches are combined to create a single feature. This integration is an effective way to integrate fine-grained localization (CNN comes up with) and global discriminative features (EfficientNet-B7 comes up with). Head: The fused one is fed through fully connected dense layers with dropout regularization to avoid overfitting, and a sigmoid activation is applied to classify it as binary (infected vs. uninfected).
As a result of this integration strategy, the hybrid model will be capable of leveraging the strengths of the two architectures: the CNN, and its capacity to extract localized low-level features, and EfficientNet-B7 that will be able to extract deeper contextual information. The hybrid method is also better in classification accuracy and retains strong performance when compared to using either branch alone, especially on diverse blood smear samples25. The overall architecture of the hybrid framework is shown in Fig. 5, where the CNN and EfficientNet-B7 branches process the input in parallel, and their feature vectors are concatenated for final classification. Unlike sequential pipelines or simple ensembles, the proposed parallel fusion strategy (Fig. 5) integrates complementary low- and high-level features, which improves classification accuracy while keeping the architecture interpretable.
Hyper-parameter tuning
The grid search method: To maximize the performance of the model, a grid search was conducted looking at a range of values for significant hyper-parameters. This approach changed the learning rate, batch size, number of filters, and kernel width systematically. The learning rate was investigated across a range from 0.1 to 0.0001 while batch sizes of 16, 32, and 64 were evaluated. The convolutional layers ranged in number from 32 to 128; kernel sizes of (3,3) and (5,5) were looked at. This all-encompassing search aimed to identify the optimal hyper-parameter configuration that would mix training economy with model complexity and generalization capacity.
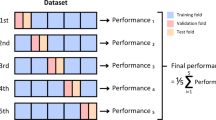
Cross-validation: Cross-validation was applied to ensure a robust evaluation of every hyper-parameter value. Particularly k-fold cross-valuation was used with k = 5 to divide the dataset into five equal halves. The model was tested on one part after being trained on four sections in every fold. This method lets one more regularly assess the performance of the model by lowering any deviations or prejudices in the data split. Cross-valuation turned out to be fairly crucial in preventing overfitting as it provided a fuller evaluation of the model’s potential to generalize to unknown data.
Additional methods: Additional methods like early stopping and dropout were applied to improve the model’s resilience and stop overfitting. Early halting was based on tracking training validation accuracy. Should the validation accuracy remain unchanged after a set number of epochs (patience = 3), the training was stopped to stop the model from overfitting to the training data. Applying a dropout rate of 0.5, Dropout was applied to the thick layers randomly deactivating 50% of the neurons throughout training. This method serves to lessen the dependence on any one neuron, therefore enhancing the generalization of the model by increasing its resilience to fluctuations in the input data.
Feature extraction of CNN-EfficientNet hybrid model
The visualization shows feature maps from the first layer of the hybrid model (Fig. 6), highlighting the model’s ability to identify basic structural elements within the input image at the beginning. The distinct shapes and edges differentiate key features, such as cell boundaries and regions of interest, which are critical in identifying malaria-infected blood cells. The feature maps reveal that the model is effectively focusing on significant areas, which supports the model’s robustness in early-stage feature extraction.
Cell segmentation
Automated image analysis relies heavily on accurate segmentation, which is especially important in medical diagnostics due to the critical nature of accurately delineating areas of interest. Identifying and isolating individual cells is critical for further categorization and diagnosis in the field of malaria-infected blood cell analysis. Using Otsu’s Thresholding, a powerful and extensively used approach in image processing, this section explores the techniques used for cell segmentation. Cell segmentation is the process of dividing a picture into distinct sections, specifically individual blood cells. Precise segmentation is crucial in malaria diagnosis to correctly locate and separate infected cells, allowing subsequent classification algorithms to accurately detect and analyze the morphological features linked to various infection phases. Accurate segmentation is especially difficult when cell borders are not well defined and when working with large and diverse information. One notable and computationally efficient approach to picture segmentation is Otsu’s Thresholding, which was created by Nobuyuki Otsu in 1979. The idea behind it is histogram-based thresholding, where the goal is to choose the best threshold for achieving maximum class separation in a grayscale picture. To isolate cells from background pixels in blood cell pictures, Otsu’s approach finds the intensity threshold that maximizes inter-class variation while minimizing intra-class variance. The first step in using Otsu’s Thresholding is to create a histogram that shows how the image’s pixel intensities are distributed (Fig. 7)26. The weighted total of variances for the classes that come from the method iteratively selecting candidate thresholds follows. The best segmentation threshold is the one that produces the lowest intra-class variation. When dealing with photographs of blood cells infected with malaria, where there are frequent changes in cell shape and staining intensity, Otsu’s Thresholding is useful. By adjusting to the dataset’s natural variability, it successfully extracts areas of interest and delineates cell borders. When looking at pictures of blood cells, Otsu’s Thresholding is an effective method for identifying malaria. Effective segmentation over a wide range of picture quality is guaranteed by its adaptability to different staining intensities. By eliminating the need for human interaction during parameter tuning, Otsu’s Thresholding improves scalability and repeatability due to its unsupervised nature. The algorithm’s processing efficiency makes it a good fit for real-time applications and massive picture collections. But in dynamic and complicated biological pictures, it encounters problems such artefacts, uneven lighting, and varying staining quality. To get around these problems, segmentation pipelines may include pre-processing procedures like noise reduction and contrast normalization, while post-processing methods like morphological operations can take the segmented sections and make them even better27. This study employed Otsu’s thresholding segmentation method to identify critical areas within the images of infected blood cells. This technique helps to separate regions of interest by maximizing inter-class variance while minimizing intra-class variance. The segmented images highlight important features such as parasitic regions, which are essential for subsequent analysis by the hybrid deep learning model. By integrating Otsu’s thresholding, we improved the model’s interpretability and accuracy in detecting malaria-infected cells. The segmented dataset generated using this method significantly enhanced the model’s capacity to focus on relevant features, thus improving the overall classification performance.
Otsu’s thresholding is a well-established method in image processing used to determine the optimal threshold value for separating foreground and background elements in an image. The method aims to maximize inter-class variance while minimizing intra-class variance, which is particularly effective in cases where there is a significant difference in pixel intensities, such as in medical images. The algorithm works by evaluating the histogram of pixel intensities to find the threshold T that minimizes the weighted within-class variance σ2w, defined as shown in Eq. (7).
where:
\(\:{\omega\:}_{1}\left(T\right)\) and \(\:{\omega\:}_{2}\left(T\right)\)are the probabilities of the two classes separated by threshold T,
\(\:{\sigma\:}_{1}^{2}\left(T\right)\) and \(\:{\sigma\:}_{2}^{2}\left(T\right)\)are the variances of the pixel values in the two classes.Otsu’s method effectively distinguishes malaria-infected areas in blood cell images, enhancing subsequent feature extraction and classification steps. We have included these equations and relevant studies in the revised manuscript to further substantiate the effectiveness of this method.
Segmented dataset Preparation and segmentation validation
In this work, the term segmented dataset refers to images processed through Otsu’s thresholding. Two distinct formats were used depending on the task: For segmentation validation, Otsu’s method was applied to grayscale images to generate 1-channel binary masks (foreground = parasitic region, background = non-parasitized area). These binary masks were used exclusively for quantitative validation of segmentation performance against manually annotated masks using Dice and IoU metrics. For CNN training, the Otsu thresholding was applied directly to the original 3-channel RGB images, allowing the model to preserve important color and morphological information while emphasizing parasite-relevant regions. The CNN models were therefore trained on these Otsu-processed RGB images, not on pure binary masks.
To strengthen the validation of the Otsu segmentation step, we created a small manually annotated subset of 100 blood smear images. For this, we used the open-source tool LabelMe to draw polygonal masks around visible parasitic regions. Each image was visually examined, and boundaries were delineated based on characteristic features of malaria-infected cells such as chromatin dots, ring structures, and staining intensity differences. To reduce subjectivity, each annotation was reviewed twice by the authors, and ambiguous cases were cross-checked against published examples of parasitized and non-parasitized cells. The final masks were exported in standard formats (JSON/PNG) and used as ground truth for quantitative evaluation of the Otsu segmentation method. Figure 8 shows representative reference masks.
Results
This section provides the experimental results obtained by assessing the model’s performance under various parameter settings. Additionally, graphical representations are included to visually illustrate and enhance the comprehension of the results.
To improve understanding, the model is trained utilizing two unique architectures: a Convolutional Neural Network (CNN) and the novel CNN-EfficientNet-B7. The proposed models were evaluated on a dataset of 43,400 malaria blood smear images using a 70:30 training–testing split. The baseline 12-layer CNN achieved an accuracy of 95%, while the hybrid CNN–EfficientNet-B7 model improved performance to 97%. Incorporating Otsu-based segmentation further enhanced classification accuracy to 97.96%, demonstrating the benefit of preprocessing in highlighting parasite-relevant regions. To ensure robustness, a five-fold cross-validation was also conducted, yielding mean accuracies of 94.8%, 96.9%, and 97.8% for the CNN, hybrid, and Otsu-CNN models, respectively. The close agreement between the single split and cross-validation results confirms the reliability and stability of the proposed framework.
CNN model
The mentioned model is a convoluted 12-layer, which consists of convolutional, pooling, and fully connected layers, to precisely categories blood cells infected with malaria. The architecture is specifically developed to achieve a harmonious equilibrium between intricacy and computing efficiency, so guaranteeing optimal training effectiveness and resilient performance. The Convolutional Neural Network (CNN) undergoes 20 iterations on a well-selected dataset, enabling it to progressively acquire knowledge and fine-tune its parameters for optimal performance. Ensuring this is of utmost importance to effectively apply learned knowledge to new and unfamiliar medical images. Following the completion of the training procedure, the Convolutional Neural Network (CNN) demonstrates an impressive accuracy rate of 95%, thereby demonstrating its capability in automated malaria detection (Shown in Fig. 9). The performance parameters (Table 3) and confusion matrix (Fig. 10) provide a thorough evaluation of the CNN’s performance, including accuracy, recall, and F1 score. These metrics provide valuable information about the model’s capacity to accurately detect true positives, false positives, false negatives, and false negatives, enabling a comprehensive assessment of its classification performance. The confusion matrix provides a visual representation of the classification findings of the model, demonstrating the relationship between the anticipated and real labels.
CNN-efficientnet-B7 model
A hybrid model that combines the strong feature extraction skills of a conventional CNN with the efficiency and optimization capabilities of the cutting-edge EfficientNet-B7 architecture is used in this study. The CNN module captures complex hierarchical characteristics, whereas EfficientNet-B7 achieves efficiency improvements via compound scaling. The training regimen consists of 20 epochs, which enables the model to adjust and optimize its parameters. The extended length of training enables the model to acquire a deep understanding of intricate patterns and intricate connections inside pictures of malaria-infected blood cells. After the training process, the hybrid model has an exceptional accuracy rate of 97% (Fig. 11), which signifies its remarkable precision in accurately categorizing blood cells as either infected or uninfected with malaria. The performance parameters table (Table 4) and confusion matrix (Fig. 12) provide a thorough assessment of the model’s skills, including accuracy, recall, and F1 score. The confusion matrix provides a visual representation of the categorization outcomes, clarifying the relationship between the expected and actual labels. The amalgamation of precision, performance indicators, and the confusion matrix establishes a resilient framework for assessing and comprehending the efficacy of the hybrid model in segmenting and classifying malaria-infected blood cells.
Segmentation
Segmentation is a crucial component in the complex field of medical image analysis. It serves as a fundamental step that enables following tasks like classification and diagnosis. This work uses a robust segmentation method to accurately detect and isolate areas of parasitic infection within pictures of blood cells infected with malaria. The segmentation process comprises a series of meticulously coordinated stages, with each one playing a role in enhancing the exactness and correctness of identifying the regions of interest.
Conversion to grayscale
The first step of the segmentation process is converting the original colour pictures into grayscale. Grayscale conversion streamlines the next processing stages by reducing the picture to a solitary channel, hence removing colour data (The preprocessing pipeline is illustrated in Fig. 13,14,15,16).
Figure 13 shows Preprocessing-grayscale conversion. Example RGB blood smear converted to grayscale as the first step in the pipeline prior to Otsu thresholding. This stage is especially important in medical imaging, since it emphasises structural characteristics rather than fluctuations in colour. Grayscale pictures are used as a basis for future thresholding and edge detection operations, making the study of morphological traits more efficient28.
Thresholding techniques
Thresholding is a basic method used in image processing to differentiate between pixels that belong to the foreground and those that belong to the background in grayscale photographs. More precisely, a mixture of thresholding methods, such as Otsu’s thresholding, is used. Figure 4 is the representative visualization of segmentation and preprocessing steps. (a) Otsu-thresholded binary mask highlighting the parasitic region in green. (b) Canny edge detection overlay on the Otsu mask, showing detected boundaries in white/yellow. (c) Segmented blood smear image with parasite region preserved and background suppressed, used as input for CNN training. Otsu’s technique calculates the most suitable threshold by maximizing the variation between different classes, successfully distinguishing parasitized areas from the background. Otsu’s thresholding is very valuable in situations where there may be variations in staining intensity across photos since it guarantees a strong and reliable segmentation process29.
Representative visualization of segmentation and preprocessing steps. (a) Otsu-thresholded binary mask highlighting the parasitic region in green. (b) Canny edge detection overlay on the Otsu mask, showing detected boundaries in white/yellow. (c) Segmented blood smear image with parasite region preserved and background suppressed, used as input for CNN training.
Canny edge detection
To improve the clarity of the borders inside the areas affected by parasites, the segmentation process includes the integration of Canny edge detection. The Canny edge detection algorithm detects sudden variations in pixel intensity, accurately delineating the boundaries of objects in the image (Fig. 15). The integration of thresholding and Canny edge detection enhances the segmentation process by adding a higher level of accuracy, enabling the capture of delicate morphological characteristics related to parasitized blood cells. This hybrid method guarantees that the model can accurately distinguish between textural and structural alterations that indicate infection30.
Detection and highlighting
The last stage of the segmentation procedure is identifying and emphasizing the areas within the pictures that are infected with parasites. This is accomplished by using contour detection methods, notably the “findContours” and “drawContours” functions (Fig. 16). These algorithms detect and delineate the borders of parasitized blood cells by identifying continuous patches within the segmented picture. The identified outlines are then emphasized, offering a visual depiction of the regions of significance. This process is crucial for further research, as it helps to extract distinct properties from parasitized cells, which are then used for categorization reasons31.
Histogram plotting
Finally, a histogram is generated to visually represent the distribution of pixel intensities in the segmented images (Fig. 17). Histograms provide a means to analyse the distribution of pixel values in grayscale, enabling a qualitative evaluation of the segmentation outcomes. Deviation or anomalies in the histogram may indicate segmentation difficulties, offering useful insights for refining the parameters of the segmentation procedure32.
The segmentation method is consistently applied to the complete collection of images of blood cells infected with malaria. By using the segmentation process in a methodical and consistent manner, a uniform methodology to accurately detecting and isolating areas can be established that are affected by parasites.
This segmentation approach is well illustrated by a collection of carefully selected images. It includes the original grayscale image, the segmented image with red parasitic regions emphasized to make them stand out and an affinity associated histogram. In Fig. 18 the image shows the red and blue regions, here the red represents the infected cells (segmented) and blue represents uninfected. Qualitative research such expose if the model can accurately identify parasitized regions in pictures of blood cells, amongst other factors. Accurate segmentation is critical to automated diagnosis of malaria as it separates areas within blood cell pictures that are infected with parasites. This allows classification models to concentrate on certain morphological characteristics of infection, thus improving the precision and sensitivity for diagnosing this disease.
12-Layer CNN model trained on segmented dataset
Using a unique method that uses segmentation as an intermediate stage in the training process, the CNN model has been made more complete. The entire dataset, which contains 43,400 pictures of blood cells infected with malaria, is segmented in order to produce a training dataset designed for this particular purpose. This means converting to grayscale since there is one channel only; distinguishing different areas using a threshold method; detecting contours and then highlighting parasitized regions in the process. This produces an even more focused improved set of training data for the CNN to learn. This newly generated segmentation dataset is then used as input for future CNN model training. It is also consistent with the objective of enhancing network performance, by providing images that emphasize on important areas which are carefully highlighted. The original dataset previously trained CNN model further training on the segmentation dataset for 20 epochs (Fig. 19). This phase allows the model to adjust and incorporate subtle features extracted during segmentation. The learning process is repeated, and the model’s weights become richer and biases greater, so it has capacity to produce accurate prediction based on separate parasitized areas consequently.
The CNN model, which was trained by the segmented dataset, achieved a remarkable accuracy improvement, rising from 95% to 97.96%. This underscores the effectiveness of using segmentation to guide the model toward more precise and directional learning. Evaluation of the CNN model’s performance is carried out by providing a confusion matrix (Fig. 20) and a performance parameter (Table 5). These tools give an all-round analysis of the model’s ability to accurately identify true positives, false positives, true minuses and false minuses, thereby considering the impact of segmentation. The synthesis of segmentation and training improves the overall performance of the CNN model, highlighting the necessity of incorporating domain-specific preprocessing methods into a machine learning pipeline.
Using this manually annotated subset of 100 images, we computed the Dice coefficient and Jaccard Index (IoU) between the Otsu-generated masks and the reference masks. The Otsu method achieved an average Dice score of 0.848 and an IoU of 0.738, demonstrating that it effectively isolates parasite-relevant regions despite its simplicity. The predicted masks are shown in the Fig. 21.
Cross-validation results
In order to assess rigorously the robustness, the stability, and generalizability of the proposed models, we have carried out the cross-validation study on the whole malaria dataset (containing 43,400 images). The method is commonly considered to be a gold standard of the machine learning to give an accurate estimate of the performance of a model on unknown data and reduce the danger of overfitting. During the 5-fold cross-validation trial, the set was divided into five partitions (or folds) of equal size. Each iteration had four model training subsets and one-sub set kept aside as validation. This was used five times with each subset having been used once as the validation set. By taking a mean of comes across all folds we were able to get a more accurate measurement of the generalization capacity of the models, which otherwise would have been biased due to a single random split of the data into train and test. Table 6 shows the average values of accuracy, precision, recall and F1-score over five folds.
All the measurements on the results of the 12-layer CNN have consistent results of an average accuracy of 94.8%, and the standard deviation across the folds is small (the standard deviation is 0.4).
This means that the model generalised well, and the performance on different validation sets was not considerably different. This uniformity implies that the CNN architecture proves to be stable when identifying malaria-infected and uninfected blood cells; even without extra optimization methods. There was a marked improvement in the hybrid model, which was the combination of the CNN and EfficientNet-B7 with an average accuracy of 96.9). Notably, it was also a drop in its precision, recall and F1-score, that appeared rather stable across all folds, having an incredibly low variance. These findings show the advantages of the architectural fusion, in which the compound scaling strategy in EfficientNet can contribute to improved feature extraction performance efficiency and supplement hierarchical learning ability of the CNN. Such synergy leads to the improved overall classifier that can better generalize over all resulting different samples.
The best performance was recorded by the CNN model with the segmented dataset created by Otsu thresholding that scored 97.8 (average accuracy). This model also showed better results even on the other two models since it has the highest average score and the variance (0.2 or 0.3) was also narrower. These results prove quite well how beneficial the process of image segmentation can be added to the flow of preprocessing. The segmentation process eliminates noise and irrelevant background, increasing sensitivity and specificity of occurrences of malaria due to focused concentration of the model in learning the most concerned areas that are parasitized.
Such results of cross-validation are very suggestive that the proposed models, especially the hybrid CNN- EfficientNet-B7 and CNN that was learned on a segmented data, can be called robust and has the potential to be generalized to unseen data. They also confirm the significance of integrating several methods including architectural fusion and enhanced image preprocessing to enhance the performance of classification of medical images. Moreover, the low variance between folds means that the models do not overfit on particular subsets of the data and can stay highly performant on different sets. This strength would be necessary to be deployed in the real world since in real-world deployment what really matters is the reliability of the model across the different populations of the patients, the different staining in the specimens, and different imaging protocols.
Results comparative analysis
This study evaluated three deep learning models for malaria detection using blood smear images, focusing on accuracy improvement and diagnostic robustness. A comprehensive comparison with recent state-of-the-art (SOTA) approaches is also conducted to contextualize the performance of the proposed models.
Performance of proposed models
The baseline 12-layer CNN achieved a robust accuracy of 95%, confirming its strong capacity for malaria-infected vs. uninfected cell classification. However, to further enhance performance, two advanced variations were implemented: A hybrid model combining the 12-layer CNN with EfficientNet-B7, achieving an improved accuracy of 97%. The integration of EfficientNet-B7 added significant representational power due to its compound scaling and lightweight design, contributing to more precise feature extraction. The final model incorporated Otsu-based image segmentation prior to CNN training, enabling focused learning on parasite-relevant regions. This segmented CNN achieved the best result with an accuracy of 97.96%, highlighting the effectiveness of preprocessing in improving discriminative learning. These findings, summarized in Table 7, confirm that architectural enhancements and domain-specific preprocessing contribute substantially to performance improvements.
Comparative analysis with state-of-the-art methods
A comparative study was conducted to benchmark the proposed malaria detection models against recent state-of-the-art (SOTA) deep learning and machine learning approaches. These comparisons not only highlight the improvements in classification accuracy but also provide insights into computational efficiency and practical usability. Suresh Babu Nettur et al.33 demonstrated that MobileNetV2 achieved 97.06% accuracy, showcasing the robustness of lightweight models optimized for deployment on mobile and embedded devices. However, the model requires extensive tuning to generalize across datasets. Similarly, the SqueezeNet1.1 family34 reported 97.12% accuracy, while a smaller variant (Variant 3, four fire modules) achieved 96.55% accuracy with a significantly reduced parameter count, illustrating the trade-off between performance and efficiency for edge applications.
An ensemble approach integrating ResNet-50, VGG-16, and DenseNet-201 with an SVM and LSTM classification layer35 attained 96.47% accuracy on the NIH Malaria dataset. While effective, this method introduces considerable computational overhead due to the fusion of multiple models and features. In another study36, EfficientNet achieved 97.57% accuracy by leveraging compound scaling and deep feature extraction, though the hardware demands of this architecture may limit real-time clinical deployment.
In contrast, traditional machine learning approaches such as Logistic Regression (65.38%) and SVM (84%) underperformed significantly compared to deep learning models. Even with moderate optimization, their accuracies remain well below those of CNN-based methods. By comparison, Inception-V3 achieved 94.52% accuracy on 27,558 red blood cell images within just five epochs, highlighting the superiority of deep convolutional models in capturing complex morphological patterns relevant for medical diagnostics.
A summary of the comparative results, including accuracy, parameter counts, model size, and inference times, is presented in Table 7. As shown, our Otsu-CNN achieves a strong balance between accuracy and computational efficiency, outperforming lightweight models such as MobileNetV2 and SqueezeNet in accuracy, while requiring fewer parameters than deeper ensembles. The hybrid CNN–EfficientNet-B7 model delivers the highest accuracy, though at the cost of increased complexity and inference time. Table 7 highlights these trade-offs, underscoring that Otsu-CNN provides an effective middle ground for practical deployment in resource-constrained environments.
Future work and limitations
Although the results are promising, several limitations of the present study should be acknowledged. First, the experiments were conducted using publicly available malaria datasets that lack expert-verified pixel-level masks. As a result, the validation of Otsu’s thresholding was limited to qualitative assessment and a small manually annotated subset, which may not capture the full variability present in real clinical samples. Second, the dataset is susceptible to biases introduced by staining variability, illumination differences, and slide preparation, which could affect the model’s generalizability. Third, while the CNN–EfficientNet-B7 hybrid model provided accuracy gains, it also comes with higher computational costs compared to lightweight models such as MobileNetV2 or SqueezeNet, limiting its deployment in resource-constrained environments. Finally, external validation using multi-center datasets and real-world clinical trials remains necessary to confirm the robustness and applicability of the proposed framework. The results are encouraging although there exists a number of areas to carry out future research in. The first contribution to the topic of transfer learning could be an extension to various malaria datasets through fine-tuning of pre-trained networks in order to exploit generalized knowledge43. Second, it is possible that ensemble modelling (a combination of predictions by several neural networks) can further improve diagnostic accuracy44. Third, multimodal data including those of patient clinical records and imaging may also give a more comprehensive picture of the disease45. Besides, the interpretability tools need to be refined to become clinically applicable, and real-time deployment has to be considered by using platform testing and validation in medical facilities35. The methods related to data augmentation need to be extended as well so that the model could withstand different conditions of the images46. Finally, it is proper to test the model with bigger and more diverse sets to ascertain its generalizability and flexibility47.
Conclusions
The study is a significant hit towards automating diagnosis of malaria through deep learning. Inclusion of superior neural structures and smart preprocessing mechanisms have shown dramatic results on the accuracy of the classification task. Hybrid model that implemented a 12-layer CNN and EfficientNet-B7 scored high (97% accuracy compared to 97.96% and thereby surpassing the baseline models) whereas 12-Layer CNN model that was trained using segmented databases scored higher (97.96% compared to 97%). The choice of using the segmentation approach helped the model to concentrate on the most significant areas which were parasitized thus increasing the discriminatory capabilities. Moreover, normalization procedures like the Batch Normalization and size transformations also helped the model to be more stable and have better generalization between the datasets. Such findings confirm the efficiency of the integration of classical CNNs and effective feature extractors and image segmentation in the classification tasks of medical images.
Data availability
All the datasets analyzed during the current study are available from the corresponding author upon reasonable request.
References
Nettur, S. B. et al. UltraLightSqueezeNet: A deep learning architecture for malaria classification with up to 54× fewer trainable parameters for resource constrained Devices, in IEEE access, 13, pp. 89428–89440, (2025).
Murmu, A. & Kumar, P. DLRFNet: deep learning with random forest network for classification and detection of malaria parasite in blood smear. Multimed Tools Appl. 83 (23), 63593–63615 (2024).
Aniket, S. P. Malaria Dataset. (2020).
Yoon, J., Jang, W. S., Nam, J., Mihn, D. C. & Lim, C. S. An automated microscopic malaria parasite detection system using digital image analysis. Diagnostics (Basel). 11 (3), 527 (2021).
Hemachandran, K. et al. Performance analysis of deep learning algorithms in diagnosis of malaria disease. Diagnostics (Basel). 13 (3), 534 (2023).
Vijayalakshmi, A. & Rajesh Kanna, B. Deep learning approach to detect malaria from microscopic images. Multimed Tools Appl. 79, 21–22 (2020).
Jones, C. B. & Murugamani, C. Malaria parasite detection on microscopic blood smear images with integrated deep learning algorithms. Int. Arab. J. Inf. Technol., 20, 2, (2023).
Siłka, W., Wieczorek, M., Siłka, J. & Woźniak, M. Malaria detection using advanced deep learning architecture. Sens. (Basel). 23 (3), 1501 (2023).
Morid, M. A., Borjali, A. & Del Fiol, G. A scoping review of transfer learning research on medical image analysis using imagenet. Comput. Biol. Med. 128 (104115), 104115 (2021).
Van-Thanh, K. H. Practical analysis on architecture of EfficientNet, in 14th International Conference on Human System Interaction (HSI), IEEE, 2021, pp. 1–4., IEEE, 2021, pp. 1–4. (2021).
Sethi, M., Ahuja, S., Singh, S., Snehi, J. & Chawla, M. An Intelligent Framework for Alzheimer’s disease Classification Using EfficientNet Transfer Learning Model, International Conference on Emerging Smart Computing and Informatics (ESCI), https://doi.org/10.1109/ESCI53509.2022.9758195 (2022).
Zhu, Z., Liu, L., Free, R. C., Anjum, A. & Panneerselvam, J. OPT-CO: optimizing pre-trained transformer models for efficient COVID-19 classification with stochastic configuration networks. Inf. Sci.. 680 (2024).
N, M. P. et al. Automated Malaria Parasite Identification and Classification Using Deep Learning Techniques, 2024 International Conference on Science Technology Engineering and Management (ICSTEM), Coimbatore, India, pp. 1–6 (2024).
Sukumarran, D. et al. Automated Identification of Malaria-Infected Cells and Classification of Human Malaria Parasites Using a Two-Stage Deep Learning Technique, In IEEE Access, vol. 12, pp. 135746–135763, (2024).
Zhu, Z., Ren, Z., Lu, S., Wang, S. & Zhang, Y. DLBCNet: A deep learning network for classifying blood cells. Big Data Cogn. Comput. 7 (2), 75 (2023).
Lu, S. Y., Zhu, Z., Tang, Y., Zhang, X. & Liu, X. CTBViT: A novel ViT for tuberculosis classification with efficient block and randomized classifier. Biomed. Signal. Process. Control. 100 (2025).
Mshani, I. H. et al. Key considerations, target product profiles, and research gaps in the application of infrared spectroscopy and artificial intelligence for malaria surveillance and diagnosis. Malar J, 22, 1, (2023).
Islam, M. R. et al. Explainable transformer-based deep learning model for the detection of malaria parasites from blood cell images. Sens. (Basel). 22 (12), 4358 (2022).
Lu, S. Y., Zhu, Z., Zhang, Y. D. & Yao, Y. D. Tuberculosis and pneumonia diagnosis in chest X-rays by large adaptive filter and aligning normalized network with report-guided multi-level alignment. Eng. Appl. Artif. Intell. 158 (2025).
Sarkar, A., Vandenhirtz, J., Nagy, J., Bacsa, D. & Riley, M. Identification of images of COVID-19 from chest x-rays using deep learning: comparing cognex visionpro deep learning 1.0tm software with opensource convolutional neural networks. SN Appl. Sci. 2 (3), 1–18 (2021).
Ramos-Briceño, D. A., Flammia-D’Aleo, A., Fernández-López, G., Carrión-Nessi, F. S. & Forero-Peña, D. A. Deep learning-based malaria parasite detection: convolutional neural networks model for accurate species identification of plasmodium falciparum and plasmodium Vivax. Sci. Rep. 15 (1), 3746 (2025).
Arco, J. E., Górriz, J. M., Ramírez, J., Álvarez, I. & Puntonet, C. G. Digital image analysis for automatic enumeration of malaria parasites using morphological operations. Expert Syst. Appl. 42 (6), 3041–3047 (2015).
Nakasi, R. et al. A new approach for microscopic diagnosis of malaria parasites in Thick blood smears using pre-trained deep learning models. SN Appl. Sci, 2, 7, (2020).
Delgado-Ortet, M., Molina, A., Alférez, S., Rodellar, J. & Merino, A. A deep learning approach for segmentation of red blood cell images and malaria detection. Entropy (Basel). 22 (6), 657 (2020).
Marques, G., Agarwal, D. & de la Díez, I. Automated medical diagnosis of COVID-19 through EfficientNet convolutional neural network. Appl. Soft Comput. 96 (106691), 106691 (2020).
Al-Kofahi, Y., Lassoued, W., Lee, W. & Roysam, B. Improved automatic detection and segmentation of cell nuclei in histopathology images. IEEE Trans. Biomed. Eng. 57 (4), 841–852 (2010).
Zhang, J. & Hu, J. Image segmentation based on 2D Otsu method with histogram analysis, In International Conference on Computer Science and Software Engineering, (2008).
Saravanan, C. Color Image to Grayscale Image Conversion, in Second International Conference on Computer Engineering and Applications, (2010).
Al-amri, S. S. & Kalyankar, N. V. and K. S. D., Image segmentation by using threshold techniques, (2010).
Bao, P., Zhang, L. & Wu, X. Canny edge detection enhancement by scale multiplication. IEEE Trans. Pattern Anal. Mach. Intell. 27 (9), 1485–1490 (2005).
Mahamud, S., Williams, L. R., Thornber, K. K. & Xu, K. Segmentation of multiple salient closed contours from real images. IEEE Trans. Pattern Anal. Mach. Intell. 25 (4), 433–444 (2003).
Alonso-Ramírez, A. A. et al. Malaria cell image classification using compact deep learning architectures on Jetson TX2. Technol. (Basel). 12 (12), 247 (2024).
Nettur, S. B. et al. UltraLightSqueezeNet: A deep learning architecture for malaria classification with up to 54× fewer trainable parameters for resource constrained devices. IEEE Access. 13, 89428–89440 (2025).
Shahin, O. R., Alshammari, H. H., Alabdali, R. N., Salaheldin, A. M. & Saleh, N. Automated multi-model framework for malaria detection using deep learning and feature fusion. Sci. Rep. 15 (1), 25672 (2025).
Mujahid, M. et al. Efficient deep learning-based approach for malaria detection using red blood cell smears. Sci. Rep. 14 (1), 13249 (2024).
Leveraging Machine Learning and Deep Learning for Advanced. Malaria Detection Through Blood Cell Images.
Khan, M. et al. IoMT-Enabled Computer-Aided Diagnosis of Pulmonary Embolism from Computed Tomography Scans Using Deep Learning, Sensors, https://doi.org/10.3390/s23031471 (2023).
Khan, H. H., Shah, P. M., Shah, M. A. & Islam, S. Cascading Handcrafted Features and Convolutional Neural Network for IoT-enabled Brain Tumor Segmentation, Comput. Commun., https://doi.org/10.1016/j.comcom.2020.01.013. (2020).
Shah, P. M. et al. Deep GRU-CNN model for COVID-19 detection from chest X-Rays data. IEEE Access. 10, 35094–35105. https://doi.org/10.1109/ACCESS.2021.3077592 (2022).
Ghaffar, Z. et al. Comparative Analysis of State-of-the-Art Deep Learning Models for Detecting COVID-19 Lung Infection from Chest X-Ray Images. https://doi.org/10.48550/arXiv.2208.01637 (2022).
Hussain, S. S. et al. and others, Classification of Parkinson’s Disease in Patch-Based MRI of Substantia Nigra, Diagnostics 13, no. 17, 2827, https://doi.org/10.3390/diagnostics13172827 (2023).
Hussain, S. S. et al. A Swin Transformer and CNN Fusion Framework for Accurate Parkinson Disease Classification in MRI, Sci. Rep., 15, 15117, https://doi.org/10.1038/s41598-025-93671-5 (2025).
Ikerionwu, C. et al. Application of machine and deep learning algorithms in optical microscopic detection of plasmodium: A malaria diagnostic tool for the future. Photodiagnosis Photodyn Ther. 40 (103198), 103198 (2022).
Nasir, S. M. I., Amarasekara, S., Wickremasinghe, R., Fernando, D. & Udagama, P. Prevention of re-establishment of malaria: historical perspective and future prospects. Malar J, 19, 1, (2020).
Sukumarran, D. et al. Machine and deep learning methods in identifying malaria through microscopic blood smear: A systematic review. Eng. Appl. Artif. Intell. 133 (108529), 108529 (2024).
Jdey, I., Hcini, G. & Ltifi, H. Deep learning and machine learning for Malaria detection: overview, challenges and future directions, (2022).
Chaudhry, H. A. H., Farid, M. S., Fiandrotti, A. & Grangetto, M. A lightweight deep learning architecture for malaria parasite-type classification and life cycle stage detection. Neural Comput. Appl., (2024).
Acknowledgements
The authors acknowledge Organizations for the support provided for carrying out the research work in the stipulated time.
Author information
Authors and Affiliations
Contributions
Retinderdeep Singh: Conceptualization, Methodology, Software. Chander Prabha: Data curation, Writing- Original draft preparation, Writing- Reviewing and Editing; Shahab Abdulla: Visualization, Investigation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Singh, R., Prabha, C. & Abdulla, S. Optimized CNN framework for malaria detection using Otsu thresholding−based image segmentation. Sci Rep 15, 40117 (2025). https://doi.org/10.1038/s41598-025-23961-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-23961-5