Abstract
Sepsis remains a critical global health threat characterized by high mortality and complex immune-metabolic dysregulation. This study identifies ferritinophagy-related prognostic genes (FRGs) that are causally linked to sepsis outcomes using a multi-omics approach integrating bulk and single-cell transcriptomics with Mendelian randomization. Among 1047 candidate genes, UBE2Q1, NEDD4L, and TCP11L2 were selected to build a risk model that effectively stratified sepsis patients and predicted mortality. Functional enrichment analysis revealed associations with oxidative phosphorylation, ribosome function, and Parkinson’s disease pathways. Immune infiltration analysis identified increased γδ T and NK cell levels in high-risk patients, with significant correlations to prognostic genes. Single-cell RNA sequencing further revealed dynamic gene expression shifts across key immune cell types including T cells, monocytes, and platelets. Cell–cell communication and pseudo-time analyses highlighted the roles of UBE2Q1 and NEDD4L in immune regulation and development. This study provides preliminary clues that ferritin and autophagy-related genes may be involved in the pathogenesis of sepsis, and proposes potential molecular targets that can be used for prognostic evaluation and therapeutic intervention.
Similar content being viewed by others
Introduction
Sepsis is a major global health issue characterized by a systemic inflammatory response that leads to organ dysfunction and high mortality1,2. A 2020 study estimated approximately 19 million sepsis cases and 5 million sepsis-related deaths worldwide each year3. Sepsis affects millions of people globally, with the incidence and mortality rates varying by country4. Developed nations, such as the United States and Europe, have reported a steady increase in sepsis cases. The annual standardized incidence of hospitalized sepsis in China has increased from 328.25 421.85 cases per 100,000 people between 2017 and 2019, indicating an rising trend5. Despite advancements in medical technology, the prognosis for patients with sepsis remains poor. Bacterial infections are the most common and can result from viral, fungal, and parasitic infections6. The disease’s diverse and variable presentation, increasing antibiotic resistance, and lack of effective early diagnostic markers pose significant challenges for sepsis management7. Therefore, identifying key genes with diagnostic and prognostic significance in sepsis may play a crucial role in improving the treatment and management of these conditions.
Ferritinophagy is a specific autophagy process that degrades ferritin to release iron ions (Fe2+), thereby regulating the intracellular iron balance. Ferritin stores iron in the cytoplasm to prevent oxidative stress caused by iron accumulation8. Ferritinophagy occurs primarily through the autophagy-lysosome pathway, and is influenced by intracellular iron levels, oxidative stress, and nutritional status. It plays a crucial role in various biological processes, including cell growth and division, oxidative stress responses, and immune response9.
Sepsis caused by infection triggers systemic inflammatory response syndrome (SIRS), which involves inflammation, immune dysregulation, metabolic abnormalities, and organ damage10. Zhang et al. found that by blocking ferritin autophagy, YAP1 reduced the availability of free iron, thereby reducing ferroptosis and related cell damage in lung tissue. This mechanism highlights the potential therapeutic value of the ferritin autophagy-ferroptosis pathway in treating sepsis-induced lung injury11. However, the specific biological role of ferritinophagy in sepsis remains unclear. The critical FRGs involved in sepsis are unclear, and their molecular mechanisms and signaling pathways require further elucidation. Further research is needed to validate the potential applications of ferritinophagy in sepsis prognosis and therapy.
This study employed the MR method to address confounding issues in causal inferences. MR offers evidence levels similar to those of randomized controlled trials without the need for randomization. The method relies on three key assumptions: first, the instrument variable must be strongly associated with the exposure factor; second, the instrument variable should be independent of confounding factors affecting both exposure and outcome; and third, the instrument variable should influence the outcome only through the exposure factor. MR analysis uses natural genetic variations or other reliable instrumental variables to simulate randomized controlled trials, thereby assessing the causal impact of the exposure factor on the outcome and reducing confounding influences12.
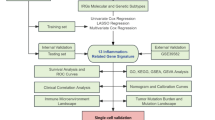
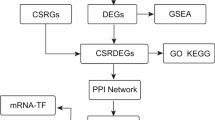
Given the potential importance of ferritinophagy in sepsis, this study aimed to explore new therapeutic targets by identifying the prognostic genes related to ferritinophagy. This study provides new strategies for improving sepsis prognosis and treatment, and utilizes sepsis-related datasets and FRGs from public databases. Differential expression analysis, weighted gene co-expression network analysis (WGCNA), and MR were used to identify prognostic genes related to ferritinophagy. Single-cell RNA sequencing analysis was used to identify the cell types that could potentially affect sepsis. This study aimed to uncover the potential link between ferritinophagy and sepsis, providing new insights and a theoretical foundation for understanding sepsis pathogenesis. Identifying ferritinophagy-related prognostic genes may offer new targeted treatment strategies for sepsis prognosis and therapy.
Materials and methods
Data sources
Sepsis-related datasets GSE65682 (platform: GPL13667) and GSE167363 (platform: GPL24676) were obtained from the Gene Expression Omnibus (GEO, http://www.ncbi.nlm.nih.gov/geo/) database. GSE165682 encompasses 760 whole blood samples from sepsis patients, with survival data available for 479 of these patients, along with 42 blood samples from healthy individuals. The GSE167363 dataset consisted of peripheral blood mononuclear cell (PBMCs) samples from 10 patients with sepsis and two healthy individuals. Twenty ferritinophagy-related prognostic genes (FRGs) were cited in the literature13, including USP24, PCBP1, ATG16L1, ATG7, SNCA, FBXW7, TRIM7, TNF, ATG5, WDR45, CYB561A3, FTH1, NCOA4, HERC2, ALOX15. BECN1, ELAVL1, ZFP36, BCAT2, and FTL. The expression quantitative trait loci (eQTL) of exposure factors and GWAS data for sepsis (ieu-b-4980) were obtained from the Integrative Epidemiology Unit (IEU) Open GWAS database (https://gwas.mrcieu.ac.uk/). The ieu-b-4980 dataset comprised 12,243,539 single nucleotide polymorphisms (SNPs) from 486,484 Europeans. The above GWAS IDs were selected following the principle of the most recent year with the highest number of SNPs, and the data were downloaded on 2 April 2024.
Acquisition of differently expressed genes (DEGs)
Initially, the R package ‘limma’ (version 3.54.0)14 was used for differential expression analysis to select DEGs between sepsis patients and controls in the GSE65682 dataset (|Log2fold-change (FC)|> 0.5, p < 0.05)15. All DEGs and the top 10 genes upregulated in the sepsis group are shown using a volcano plot and heat map, respectively. The result was visualized using ‘ggplot2’ (version 3.4.4)16.
Identification of candidate genes
The FRGs scores of the sepsis group and control group in the GSE65682 dataset were evaluated based on the FRGs gene set using the ssGSEA algorithm integrated in the R package “GSVA”17. Sepsis samples in the GSE65682 dataset were split into high- and low-scoring groups in compliance with the median FRGs score. Kaplan–Meier (K-M) analysis was performed using the R package ‘survminer’ (version 0.4.9)18 to assess the survival discrepancies of septic patients in the two scoring groups (p < 0.05). Furthermore, key module genes linked to FRGs score were identified via WGCNA, which was supported by R package ‘WGCNA’ (version 1.7.1)19. Concretely, the median absolute deviation (MAD) of genes in GSE65682 dataset was discovered and removed, and genes with MAD values in the bottom 50% as well as ineligible samples were removed utilizing the ‘goodSamplesGenes’ function. Subsequently, the sepsis samples were subjected to cluster analysis using the ‘hclust’ function to ensure the accuracy of the WGCNA by removing outlier samples. To ensure that the gene interactions closely followed a scale-free distribution, the ideal soft threshold was determined when R2 exceeded 0.85, and the mean connectivity tended to be 0. Dissimilarity coefficients were calculated according to the adjacency and similarity among genes, and a systematic clustering tree was generated. Following the standards of the hybrid dynamic tree cutting algorithm, a minimum of 100 genes per module was established, and a module merging threshold of 0.3 was applied to identify the clustering modules. Using FRGs scores as traits, Pearson’s correlation analysis was performed between the module eigengene (ME) scores of the clustering modules and the FRGs scores (|cor|> 0.3, p < 0.05)20. The clustering modules with the highest positive and negative correlations with FRGs scores were identified as the key modules. Genes included in key modules are called key module genes. The intersecting DEGs and key module genes served as the candidate genes.
Functional exploration of candidate genes
To investigate the functions of the candidate genes, enrichment analyses were conducted using Gene Ontology (GO) (p < 0.05) and the Kyoto Encyclopedia of Genes and Genome (KEGG)21 (p < 0.05). These analyses were performed using the ‘clusterProfiler’ R package (version 4.7.1.003)22. In addition, to explore the interactions between candidate genes at the protein level, a protein–protein interaction (PPI) network was synthesized for the candidate genes using the Search Tool for the Retrieval of Interacting Genes (STRING, https://string-db.org) website (confidence = 0.4).
Screening of instrumental variables (IVs)
A univariate MR (UVMR) study was performed to identify key genes by evaluating the causality between candidate genes (exposure factor) and sepsis (outcome). In this study, SNPs were used as the IVs. Firstly, the ‘extract_instruments’ and ‘extract_outcome_data’ functions from the R package ‘TwoSampleMR’ (version 0.5.7)23 were utilized to determine exposure factors and outcomes, respectively. SNPs related to exposure factors but unrelated to outcomes were ascertained (p < 5 × 10–8, proxies = TRUE, rsq = 0.8). Simultaneously, SNPs with linkage disequilibrium (LD) were excluded (r2 = 0.001, kb = 100)24. Data on exposure factors and outcomes were merged via the ‘ harmonize _data()’ function to generate SNPs for subsequent screening. Additionally, F-statistics of the SNPs were calculated, and weak SNPs with F less than 10 were eliminated. If the exposure factor had fewer than three SNPs, it was excluded. The F-statistic was calculated as the following equation:
The N was sample size for the GWAS, the K was the number of IVs, and R2 indicated the extent to which the IV explained the exposure factors.
UVMR analysis
The effect alleles and effect sizes were standardized using the ‘ harmonize _data’ function. The ‘mr’ function was then invoked to perform UVMR analysis, employing MR Egger25, Weighted median26, Simple mode27, Inverse variance weighted (IVW) method28 and Weighted mode29. The IVW method is the most commonly used. The weighting formula for IVW was calculated as follows:
For the ith IV, βi denoted the estimate of the effect of the IV on the exposure factor, γi denoted the estimate of the effect of the IV on the outcome, and se denoted the standard error. If there were fewer missing values, the observations containing the missing values were deleted directly, and the na.omit function could be used to remove the missing values. If there were more missing values, multiple interpolation would be used to fill in the missing values, with mice package.
Candidate genes were considered to have significant causality with sepsis if the p-value from the IVW method was less than 0.05. Genes with hazard ratios (HRs) greater than 1 were considered risk factors for sepsis, whereas those with HRs less than 1 were considered protective factors. Scatter plots, forest plots, and funnel plots were created to present the UVMR findings.
Sensitivity analysis and Steiger test
Sensitivity analyses including heterogeneity, horizontal pleiotropy, and leave-one-out (LOO) tests were performed to assess the dependability of the UVMR study. A heterogeneity test was conducted using the ‘mr_heterogeneity’ function to determine whether heterogeneity existed among samples. A Q_ p value greater than 0.05 indicated no heterogeneity. To evaluate potential confounding factors in UVMR study, horizontal pleiotropy was assessed with the ‘mr_pleiotropy_test’ function, supplemented by the ‘mr_presso’ function. A P value greater than 0.05, suggested no horizontal pleiotropy. The ‘mr_leaveoneout’ function was employed to perform a LOO test, which examined the impact of each SNP on the outcome. Significant changes in the results upon removal of a variable indicated that the SNP had a substantial impact on the outcome. A Steiger test was employed to confirm the directionality of UVMR analysis. Only candidate genes with a correct causal direction marked as TRUE and a significance level of p < 0.05, were deemed as key genes.
Development and verification of a risk model
First, in the GSE65682 dataset, univariate Cox regression analysis was used to identify prognostic genes associated with survival in patients with sepsis from key genes (HR ≠ 1, p < 0.01). Prognostic genes that passed the proportional hazards (PH) assumption test (p > 0.05) were selected for least absolute shrinkage and selection operator regression (LASSO) regression analysis using the R package ‘glmnet’ (version. 4.1–4)30. Samples with complete survival information in the GSE65682 dataset were randomly zoned as a training set and a validation set at cut of 7:3. In the training set, a risk model was developed based on the expression levels of prognostic genes and the overall survival (OS) information of patients with sepsis. A risk score equation was determined based on the LASSO coefficient (coef) and prognostic gene expression (expr) as follows: \(\text{risk score}=\sum_{\text{i}=1}^{\text{n}}\text{coef }({\text{gene}}_{\text{i}})\times \text{expr}{ (\text{gene}}_{\text{i}})\). Subsequently, in the training set, patients with sepsis were divided into high- and low-risk groups, according to the median risk score. Ulteriorly, in training and validation sets, K-M curves were created via the R package ‘survminer’ to compare survival discrepancies between two risk groups (p < 0.05). Moreover, the R package ‘survivalROC’ (version. 1.0.3)31 was used to draw receiver operating characteristic (ROC) curves to evaluate the effectiveness of the risk model by outputting the area under the curve (AUC) values for 9, 18, and 27-day. An expression heatmap of prognostic genes in the two risk groups was generated.
Establishment of a nomogram model and analysis of clinical characteristics
A nomogram model was established via the R package’ rms’ (version 6.5–0)31 in compliance with the expression of each prognostic gene. The predictive precision of the nomogram was appraised using calibration curve, decision, and ROC curves. To investigate the relation between risk score and clinical characteristics, in the training set, age, gender, diabetes, endotype class, ICUA, and pneumonia diagnoses were categorized into disparate clinical subgroups. The Wilcoxon rank-sum test was used to distinguish discrepancies in risk scores among these subgroups (p < 0.05). Stacked bar charts were generated to illustrate the proportion of patients in the two risk groups across various clinical characteristics.
Characterization of immune infiltration and immune checkpoint analysis
The occurrence of the disease is usually inseparable from the role of the immune system; therefore, studying the distribution of immune cells in sepsis can improve our understanding of its pathogenesis. First, the relative percentages of 22 immune cells in the high- and low-risk groups of the training set were estimated using the CIBERSORT algorithm32. Discrepancies in immune cell infiltration between the two risk groups were compared via Wilcoxon rank-sum test (p < 0.05). Discrepancies in the expression levels of 28 immune checkpoints between the two risk groups were estimated using the Wilcoxon rank-sum test (p < 0.05). Spearman’s correlation analysis, supported by the R package” Psych ‘ (version 2.2.9)33 was used to explore the correlations between prognostic genes and discrepant immune cells as well as between prognostic genes and immune checkpoints.
Functional exploration of prognostic genes and molecular network construction
To delve deeper into the signaling pathways associated with prognostic genes and the potential mechanisms of prognostic genes influencing sepsis, a Spearman correlation analysis was conducted on prognostic genes and all genes in the training set through the ‘psych’ package in R. These genes were then ranked based on their correlation coefficients from high to low, and GSEA was conducted on the sorted genes using the R package ‘clusterProfiler’ (p < 0.05). To understand the expression patterns of prognostic genes, upstream molecules targeting these genes were identified. Key miRNAs targeting prognostic genes were obtained by crossing the miRNAs predicted by the miRanda (http://www.miranda-im.pl/) and miRDB (https://mirdb.org/ mining.html) databases. Subsequently, the molecules upstream of these key miRNAs and lncRNAs were predicted using the starBase database (https://starbase.info/). Additionally, the NetworkAnalyst database (https://www.networkanalyst.ca/) was used to retrieve transcription factors (TFs) target prognostic genes. Cytoscape software (version 3.7.2)34 was used to construct the mRNA-miRNA-lncRNA and TF-mRNA networks. To further understand the suborganelles of prognostic genes, the UniProt database (https://www.uniprot.org/) was used for subcellular mapping of proteins coded by prognostic genes.
Disease association analysis, protein phosphorylation analysis and drug prediction
Disease association analysis aims to identify the causes and factors influencing diseases through systematic research. In this study, diseases associated with prognostic genes were explored using the DisGeNet database (https://www.disgenet.org/) and a prognostic gene-disease co-expression network was visualized using Cytoscape software. To investigate the phosphorylation sites of prognostic genes at the protein level, the PhosphoSitePlus database (https://www.phosphosite.org/homeAction.action) was utilized for analyzing the phosphorylation sites of the corresponding proteins. The findings were presented visually on a Laplace plot. The Drug Signatures Database (DsigDB, https://ngdc.cncb.ac.cn/databasecommons/database/id/4603) was used to predict targeted drugs that were likely to act on prognostic genes, and a prognostic gene-drug network was built using Cytoscape software.
Quality control (QC) and cell annotation
Using the GSE167363 dataset, the ‘Seurat’ package in R (version 4.3.0)35 was used for the data filtering. Specifically, cells with fewer than 200 genes, genes present in fewer than three cells, cells expressing fewer than 350 genes or more than 3,000 genes, and genes with counts ≥ 15,000 were excluded. Additionally, the proportion of mitochondrial genes was limited to less than 15%. After data normalization, the ‘vst’ method was used to extract the top 2,000 highly variable genes (HVGs) between cells for further analysis. Furthermore, the samples were subjected to principal component analysis (PCA), the principal components (PCs) were extracted via the ‘JackStrawPlot’ and ‘JackStraw’ functions, and a PCA inflection plot was drawn to show the percentage of variance represented by each PC. The ‘FindNeighbors’ and ‘FindClusters’ functions were executed to do an unsupervised cluster analysis of the cells. The Uniform Manifold Approximation and Projection (UMAP) method was used to cluster the cells, resulting in cell clusters (resolution = 0.3). Cell types were obtained based on the expression of marker genes in cell clusters, which were obtained from a previous study36. Simultaneously, the proportion of each cell type in the discrepant samples from the GSE167363 dataset was determined.
Cell communication and pseudo-time analyses
The expression of prognostic genes in different cell types was estimated, and key cell types with significantly different expression of prognostic genes in sepsis patients and controls were identified using the Wilcoxon rank-sum test (p < 0.05). To assess the patterns of interactions between cell types, cell communication analysis was performed using the R package ‘CellChat’ (version 1.6.1)37. In addition, the patterns of receptor-ligand interactions between different cell types were revealed. Unk ‘Monocle’ (version 2.26.0)38 package in R was used to conduct a pseudo-time analysis of key cell types to reveal the dynamic perspective of key cell differentiation state transformation and prognostic gene expression.
Gene set variation analysis (GSVA)
To explore changes in key cell-enriched pathways, GSVA was proceeded via the ‘GSVA’ package (version 1.42.0)17, with the reference gene set h.all.v2023.2. Hs.symbols.gmt, from the Molecular Signatures Database ( https://www.gsea-msigdb.org/gsea/msigdb).
Statistical analysis
Bioinformatics analysis was performed using the R software (version 4.2.2). Differential expression analysis was performed using limma (version 3.54.0) to identify differential genes between disease and normal samples; a co-expression network was constructed via WGCNA and key modules were screened; clusterProfiler (version 4.7.1.003) was used to conduct GO/KEGG functional enrichment analysis on candidate genes; TwoSampleMR was employed to perform Mendelian randomization analysis for inferring the causal relationship between genes and sepsis; the construction of a prognostic model using survival (version 3.5–3) combined univariate Cox regression and glmnet (version 4.1–4) LASSO regression, and rms (version 6.5–0) and nomogramFormula (version 1.2.0.0) were utilized to construct and validate the nomogram model; immune infiltration analysis was carried out to evaluate cell composition through CIBERSORT; for single-cell data analysis, Seurat (version 4.3.0) was used for quality control, standardization and clustering, CellChat (version 1.6.1) was applied to analyze intercellular communication, and monocle (version 2.26.0) was used to perform pseudotime analysis; in addition, GSVA (version 1.42.0) and enrichplot (version 1.18.3) were also used to conduct pathway enrichment analysis. Discrepancies between the two groups were evaluated using the Wilcoxon rank-sum test, with a p-value of less than 0.05 deemed statistically significant.
Results
A total of 1047 candidate genes linked to sepsis and ferritinophagy were obtained
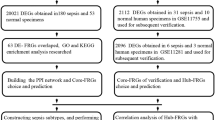
In the GSE65682 dataset, differential expression analysis identified 3,920 sepsis-associated DEGs, comprising 1,245 upregulated and 2,675 downregulated genes (Fig. 1A, Table S1). A heatmap was generated to display the expression patterns of the top 10 genes with the largest absolute log2 fold change values (Fig. 1B). The FRGs score was higher in sepsis group (p < 0.001). K-M analysis revealed that patients with sepsis in the high-scoring cohort maintained a lower survival probability (p = 0.00029) (Fig. S1A, B). No outliers were observed in the GSE65682 dataset (Fig. S1C). The ideal soft threshold was determined to be 8, with an R2 value approaching 0.85 and a mean connectivity near 0 (Fig. S1D). Twelve clustering modules were generated in the co-expression network (Fig. S1E). Among these, the MEblue module showed the strongest positive correlation with the FRGs scores (cor = 0.93, p < 0.001), whereas the MEturquoise module exhibited the highest negative correlation with the FRGs score (cor = −0.63, p < 0.001). Consequently, they serve as the key modules. A set of 3,06 key module genes was obtained by merging the module genes from the two key modules (Fig. 1C). Subsequently, 1,047 genes intersecting DEGs and key module genes were identified as candidate genes (Fig. S1F, Table S2).
Candidate genes related to ferritinophagy in sepsis were obtained. (A) DEGs between sepsis and control samples in the GSE65682 dataset; (B) Heatmap of expression of up-regulated DEGs in the sepsis group; (C) Correlation of modules with FRGs scores; (D) The top 5 entries for BPs, CCs, and MFs based on enrichment analyses of candidate genes; (E) The top 10 KEGG pathways21,39,40; (F) PPI network of Candidate genes.
Candidate genes might be involved in protein regulation and modification, DNA maintenance and repair, energy metabolism
Further enrichment analyses of candidate genes revealed that they were involved in 758 biological processes (BPs), 124 cellular components (CCs), 135 molecular functions (MFs) and 38 KEGG pathways (Table S3). The top five BPs, CCs, and MFs entries were presented in Fig. 1D, such as ATP hydrolysis activity and ubiquitin-like protein transferase activity and so on. KEGG enrichment analysis identified a total of 38 pathways (Table S4). The top 10 KEGG pathways39,40 included nucleocytoplasmic transport, protein processing in the endoplasmic reticulum, and the cell cycle (Fig. 1E). Candidate genes may be involved in protein regulation and modification, DNA maintenance and repair, energy metabolism, and cell cycle regulation. The PPI network revealed 8,167 interaction pairs between 1047 candidate genes, including ATP10D-TMEM86B, GDE1-NAPEPLD, and B3GALNT2-XYLT1. (Fig. 1F).
A number of 95 candidate genes retained significant causal relationships with sepsis
To clarify the potential causal relationship between candidate genes and sepsis, we employed a univariate Mendelian randomization approach for analysis. The results showed that among the 1047 candidate genes, 95 genes had a significant causal relationship with sepsis. Among them, 47 candidate genes are risk factors for sepsis, such as XBP1 (HR = 1.060, p < 0.001, 95% confidence interval [CI]: 1.04–1.081), FAM117B (HR = 1.025, p < 0.001, 95% CI: 1.013–1.038) and MYBPC3 (HR = 1.038, p < 0.001, 95% CI: 1.024–1.052), while 48 candidate genes were protective factors for sepsis, such as DPCD (HR = 0.803, p < 0.001, 95% CI: 0.742–0.869) and CTCF (HR = 0.805, p < 0.001, 95% CI: 0.735–0.882) (Fig. 2A). Details of the 95 candidate genes are presented in Supplementary Table S1. Using FAM117B as an example, the IVW method identified it as a risk factor for sepsis owing to its positive slope, and its effect on sepsis was almost unaffected by confounding effects (Fig. 2B). In addition, the MR effect size for FAM117B in sepsis was greater than zero in the forest plot, suggesting that FAM117B may increase the risk of sepsis (Fig. 2C). The evenly distributed points on the funnel plots indicate that the causal effect of FAM117B on sepsis adhered to Mendel’s second law (Fig. 2D). Both MR-Egger and IVW methods showed Q_pval values above 0.05, confirming no heterogeneity (Table S5). The horizontal pleiotropy test suggested no influence of confounding factors (p > 0.05) (Tables S6, S7). In addition, none of the SNPs was sensitive to sepsis upon sequential elimination, indicating that individual SNP effects were unlikely to affect the causality of FAM117B in sepsis (Fig. 2E). The Steiger test further confirmed the causal direction of these genes on sepsis (p < 0.05), ruling out the possibility of reverse causality (Table S8). The results for the top 10 genes ranked by p-values are presented in Supplementary File 1.
A number of candidate genes retained significant causal relationships with sepsis. (A) UVMR analysis; (B) Scatter plot of FAM117B in relation to sepsis; (C) Forest plot of FAM117B in relation to sepsis; (D) Funnel plot with evenly distributed points (FAM117B); (E) Leave-one-out (LOO) tests(FAM117B).
The risk model demonstrated reliable predictive ability on sepsis
Three prognostic genes were selected from 95 candidate genes (UBE2Q1, NEDD4L, and TCP11L2) via univariate Cox regression analysis and PH hypothesis testing (p > 0.05) (Fig. 3A). Three genes were subjected to LASSO regression analysis, and all three were retained in the most accurate model (lambda min = 0.01648664) (Fig. 3B-C). The LASSO regression coefficients for each prognostic gene are shown in Table S9 and are as follows: risk score = UBE2Q1 × (−1.401702039) + NEDD4L × (0.288855044) + TCP11L2 × (0.513506055). In the training set, based on the risk model using these three prognostic genes, the K-M curve confirmed that high-risk patients had a lower survival probability (p = 0.012) (Fig. 3D). The risk model demonstrated credible predictive ability as evidenced by the ROC curve, with AUCs of 0.65, 0.64, and 0.63 at 9, 18, and 27 days, respectively (Fig. 3E). Additionally, the expression of UBE2Q1 was low in the high-risk group, whereas that of NEDD4L and TCP11L2 was high (Fig. 3F). Furthermore, the validation set proved the universality of the risk model, showing outcomes similar to those of the training set (Fig. S2A–C).
The prognostic gene-based nomogram performed well in in predicting the risk of sepsis
A nomogram model was built to evaluate the predictive value of the prognostic genes. The risk of sepsis increased with the increase in the total number of points (Fig. 4A). The calibration curve indicated a high fit for the nomogram, supported by a Hosmer–Lemeshow test p-value of 0.616 (Fig. S3A). The decision curve showed that the nomogram model provided a higher net benefit than any single diagnostic gene (Fig. 4B). Moreover, the AUC value of the nomogram model was higher than that of individual prognostic genes (Fig. 4C). In summary, the nomogram model appeared to be a credible tool for predicting sepsis risk. The samples were classified into four phenotypes using the unsupervised consensus clustering method and named MARS1-441. Subsequent analysis of the differences in risk scores between different clinical subgroups revealed evident differences in risk scores among the endotype class subgroups, but not among other clinical subgroups. The risk score of Mars1 was significantly higher than those of Mars2, Mars3, and Mars4 (p < 0.001) (Fig. 4D, Fig. S3B-F). Additionally, there were significantly more high-scoring patients in the Mars1 subgroup than in the low-scoring group (Fig. 4E, Fig. S4A–E).
Totally 3 discrepant immune cells and 16 discrepant immune checkpoints were identified
The relative percentages of the 22 immune cells in the two risk groups in the training set are shown in Fig. 5A. Among the 22 immune cells, activated natural killer (NK) cells, resting NK cells, and gamma delta (γδ) T cells maintained discrepant infiltration levels between the high- and low-risk groups, with high infiltration of resting NK cells and γδ T cells in the high-risk group (p < 0.05) (Fig. 5B). TCP11L2 was positively correlated with γδ T cells (cor = 0.14, p < 0.01) and resting NK cells (cor = 0.23, p < 0.01), and negatively correlated with activated NK cells (cor = −0.20, p < 0.05). Conversely, UBE2Q1 expression negatively correlated with γδ T cells (cor = −0.14, p < 0.05) (Fig. 5C). The expression of 28 immune checkpoints between the two risk groups was then estimated, identifying 16 discrepant immune checkpoints. CD160, CD40LG, CD70, and ICOSLG were upregulated in the high-risk group, whereas the other discrepant immune checkpoints were upregulated (Fig. 5D). Importantly, among the immune checkpoints that were significantly correlated with prognostic genes, most of the immune checkpoints were significantly negatively correlated with NEDD4L and TCP11L2, whereas UBE2Q1 was significantly positively correlated with most of the immune checkpoints (Fig. 5E).
The differential immune cell infiltration and immune checkpoint expression between high-risk and low-risk groups, as well as their correlations with prognostic genes. (A) Relative percentages of 22 immune cell types in high-risk and low-risk teams from the training set; (B) Differential expression of immune cells in high and low risk groups; (C) Correlation analysis of TCP11L2 and UBE2Q1 with differential immune cells; (D) Expression analysis of 28 immune checkpoints between high-risk and low-risk groups; (E) Correlation between immune checkpoints and prognostic genes.
Prognostic genes might be regulated by multiple miRNAs and TFs when performing functions such as mitochondrial and protein synthesis
In GSEA, prognostic genes were clearly co-enriched in ribosomes, Parkinson’s disease, and oxidative phosphorylation. This suggests that the functions of the prognostic genes might be related to mitochondrial function and protein synthesis (Fig. 6A–C). The mRNA-miRNA-lncRNA network included three mRNA, seven miRNAs, and 45 lncRNAs. AL031282.2 may simultaneously regulate the expression of UBE2Q1 and NEDD4L by influencing hsa-miR-526b-5p (Fig. 6D). Moreover, in sum of 57 TFs targeting prognostic genes were retrieved from the NetworkAnalyst database. As represented in the TF-mRNA network, the expression of UBE2Q1 and TCP11L2 may also be simultaneously regulated by KLF16 and TFAP4, as well as TCP11L2 and NEDD4L by DMAP1 (Fig. 6E). Subcellular localization revealed that NEDD4L was mainly distributed in the cytoplasm and endoplasmic reticulum, whereas UBE2Q1 and TCP11L2 were mainly distributed in the cytoplasm and cytoskeleton, respectively (Fig. 6F–H).
Prognostic gene enrichment, network regulation and cellular localization. (A–C) Gene Set Enrichment Analysis of prognostic genes (A: UBE2Q1 B: NEDD4L C: TCP11L2); (D) mRNA-miRNA-lncRNA network; (E) TF-mRNA network; (F) Subcellular localization of NEDD4L; (G) Subcellular Localization of TCP11L2; (H) Subcellular localization of UBE2Q1.
Similar diseases and common drugs targeting prognostic genes were identified
Only the diseases associated with UBE2Q1 and NEDD4L were predicted in the DisGeNet database. Among these, 10 disease types, including breast carcinoma, tumor progression, malignant neoplasm of the breast, and tumor cell invasion, were simultaneously correlated with UBE2Q1 and NEDD4L (Fig. S5A). Protein phosphorylation analysis highlighted that the positions of the phosphorylated mutations in UBE2Q1, NEDD4L, and TCP11L2 were concentrated between 375 and 475 amino acids, 300–500 amino acids and 0–50 amino acids (Fig. S5B–D). In addition, the gene-drug network revealed 60 compounds that target prognostic genes. Recently, anisomycin, VALPROIC, and trichostatin co-targeted NEDD4L and UBE2Q1, while acetaminophen and benzo[a]pyrene co-targeted NEDD4L and TCP11L2 (Fig. S5E).
T cells, B cells, NK cells, monocytes and platelets were regarded as key cell types
The findings in Fig. S6 A-B depicted the scRNA-seq data before and after QC. The red spots in Fig. S6 C represented the top 2,000 (HVGs). Based on the PCA results (p < 0.05), the first 30 PCs were selected for subsequent analysis (Fig.S6 D-E). Additionally, UMAP revealed that 21 cell clusters were combined into seven cell types (T cells, B cells, NK cells, monocytes, platelets, erythroid precursors, and dendritic cells [DCs]) according to the expression of marker genes in each cell clusters (Fig. 7A–C, Fig. S6F, Table S10). In summary, cell types in which prognostic genes were expressed differentially included T cells, B cells, NK cells, monocytes, and platelets, and were regarded as key cell types (Fig. S6 G-K). Cell communication analysis confirmed that in the controls, T cells communicated more closely with monocytes and B cells, platelets with monocytes, monocytes with B cells, B cells with monocytes, and NK cells with monocytes in quantity and strength (Fig. 7D, E). In addition, the close interaction between T cells and B cells is mainly mediated by MIF − (CD74 + CXCR4). NK cells and platelets communicated with monocytes mainly through CCL5-CCR1 (Fig. 7F). Pseudo-time analysis of key cell types indicated that the differentiation of B cells and platelets included five branches (states 1–5) and they were differentiated in order from state 1 (early development) to state 3/4 (late development). NK cells, monocytes, and T cells had 11,3 and 9 branches, respectively. For NK cells and monocytes, state 1 was in the early stage of development, whereas state 8 was in the early stage of T cell development. In sepsis, T cells predominated in stages 1 and 3 of development (Fig. 8A). The expression of prognostic genes significantly increased from early to late development of T cells, and the expression of UBE2Q1 increased with differentiation of each key cell type (Fig. 8B).
Pseudo-timing and enrichment analysis of key cells. (A) Pseudo-temporal analysis of key cells (From left to right are: B cells, monocytes, NK cells, platelets, T cells); (B) Expression of prognostic genes at different time states in key cells(From left to right are: B cells, monocytes, NK cells, platelets, T cells) (C) Analysis of pathways involved in key cells (From left to right are: B cells, monocytes, NK cells, platelets, T cells).
The changes of classical signaling pathways in key cell types were revealed
GSVA revealed changes in classical signaling pathways in key cell types. For instance, in sepsis, Notch signaling pathways are inhibited in NK cells and activated in platelets and T cells. P53 pathway was inhibited in the T cells, B cells, NK cells, and platelets in the sepsis group (Fig. 8C–G).
Discussion
Sepsis remains a major global health issue and is characterized by a systemic inflammatory response that leads to organ dysfunction and high mortality42. Previous studies have highlighted the complexity of sepsis pathogenesis, involving inflammation, immune dysregulation, metabolic abnormalities, and organ damage43. Despite extensive research, the specific biological role of ferritinophagy in sepsis has not been adequately explored44. This study addressed this gap by identifying the prognostic genes related to ferritinophagy and examining their potential impact on sepsis outcomes. In this study, we identified three prognostic genes (UBE2Q1, NEDD4L, and TCP11L2) that are associated with ferritinophagy in sepsis. A prognostic model was developed based on the genes that accurately predicted the outcomes of patients with sepsis. Subsequently, the potential functions of these prognostic genes were explored using GSEA, molecular regulatory network construction, and immune cell infiltration analyses. Additionally, single-cell RNA sequencing revealed dynamic changes in the expression of these genes within the key cell types implicated in sepsis.
This comprehensive approach provides new insights into the role of FRGs in sepsis and offers novel perspectives on the pathogenesis of the disease and potential therapeutic targets. Identifying these prognostic genes not only enhances our understanding of sepsis but also paves the way for improved prognostic assessment and targeted treatment strategies.
Each of these genes plays a crucial role in various diseases; however, their involvement in sepsis has not been previously reported. As a member of the E2 ubiquitin-conjugating enzyme family, UBE2Q1 plays a central role in the ubiquitin–proteasome system (UPS), which is essential for intracellular protein degradation, cell cycle regulation, and cellular growth45,46,47. Aberrant expression of UBE2Q1 has been documented in multiple types of malignancies46. For instance, in hepatocellular carcinoma (HCC), UBE2Q1 is upregulated due to copy number amplification and promotes tumor progression through activation of the β-catenin–EGFR–PI3K–Akt–mTOR signaling pathway, indicating its pivotal role in cell proliferation and oncogenesis48,49. Moreover, abnormal UBE2Q1 expression has also been observed in colorectal and breast cancers50,51.
Sepsis is a life-threatening syndrome characterized by dysregulated host responses to infection, leading to organ dysfunction52. The ubiquitin–proteasome system participates in regulating immune cell function and inflammatory signaling pathways45,53. Among these, the NF-κB signaling pathway plays a crucial role in mediating inflammatory responses during sepsis54,55. Previous evidence suggests that UBE2Q1 may indirectly modulate cytokine production or immune cell activation by affecting the ubiquitination of key signaling molecules. Although UBE2Q1 itself has not been directly investigated in this context, ubiquitination-mediated regulation of the NF-κB pathway has been shown to enhance macrophage glycolysis and M1 polarization, thereby exacerbating sepsis, highlighting the critical role of ubiquitin modification in the inflammatory response during sepsis56.
NEDD4L, an E3 ubiquitin ligase, plays a critical role in the ubiquitin–proteasome system (UPS) by mediating the specific ubiquitination of target proteins, thereby regulating their degradation, subcellular localization, and activity57,58. The most well-characterized substrate of NEDD4L is the epithelial sodium channel (ENaC), which is predominantly expressed in pulmonary and renal epithelial cells and is closely associated with related diseases59. In addition, NEDD4L promotes polyubiquitination and degradation of Smad2/3, thereby attenuating the TGF-β signaling pathway .
The TGF-β pathway plays a complex and context-dependent role in sepsis, exhibiting both immunosuppressive and proinflammatory effects depending on the disease stage and microenvironment60.
Taken together, NEDD4L, as an E3 ubiquitin ligase, is pivotal in maintaining protein homeostasis and regulating immune responses and intracellular signaling. Given the extensive involvement of the ubiquitin–proteasome system in the multifaceted pathophysiology of sepsis, NEDD4L is highly likely to influence the onset, progression, and prognosis of sepsis through multiple regulatory mechanisms.
T-Complex Protein 11 like 2 (TCP11L2) is less well studied but is known to be involved in spermatogenesis and has been linked to reproductive health61. The role of these genes in other diseases and our findings suggest that their dysfunction may contribute to the complex pathophysiology of sepsis. However, the exact mechanism remains to be elucidated.
The current study is the first to link these three genes to sepsis and develop a prognostic model that accurately predicts patient outcomes. GSEA results showed that these genes were co-enriched in pathways related to ribosomes, Parkinson’s disease, and oxidative phosphorylation. Ribosomes are involved in protein synthesis, which is essential for cellular repair and immune functions in sepsis62. Parkinson’s Disease pathways suggest possible roles in neuroinflammation and mitochondrial dysfunction, which are both relevant to the systemic effects of sepsis63. In addition,oxidative phosphorylation is crucial for energy production and its dysregulation can lead to metabolic abnormalities in sepsis64. The co-enrichment of these genes in Parkinson’s disease and oxidative phosphorylation pathways suggests broader implications beyond sepsis, potentially linking systemic inflammation and metabolic dysregulation observed in other diseases. Investigating these cross-disease pathways may not only enhance sepsis treatment strategies but also contribute to our understanding of other inflammatory and neurodegenerative disorders. Ultimately, the insights gained from this study could pave the way for personalized gene-targeted therapies to improve patient outcomes in sepsis and beyond.
Immune infiltration analysis revealed significant differences in resting NK cells and gamma delta (γδ) T cells between the high- and low-risk groups. Resting NK cells are involved in early immune responses and the control of viral infections, with studies showing that their dysfunction in sepsis contributes to immune paralysis65. Gamma delta T cells play roles in bridging innate and adaptive immunity and promoting inflammatory responses and have been implicated in sepsis pathogenesis through their cytokine production and cytotoxic activity66. In addition, correlation analysis showed a certain correlation between prognostic genes and differential immune cells, suggesting that regulating the differential infiltration of immune cells through genes may have an inhibitory effect on the immune dysregulation of sepsis.
Single-cell RNA sequencing has identified five key cell types: T-cells, B-cells, NK cells, monocytes, and platelets. Based on the role of these immune cells in immune response, they may also be critical for immune dysregulation and inflammation in sepsis. Among them, T and B cells are involved in adaptive immunity, NK cells are involved in innate immunity, the main function of monocytes is phagocytosis and participation in the inflammatory response, and platelets regulate coagulation and inflammation. Recently, Bird et al. found that NOP receptor signaling may adversely affect the outcome of sepsis by regulating the immune response of B cells and T cells67, and Yao et al. found that S100A9 + monocytes exhibited profound immunosuppressive function in the CD4 + T cell immune response68. These results were consistent with our results.
Furthermore,cell communication analysis showed strong interactions between monocytes and other cell types, primarily through the TN-TLR4 and CCL5-CCR1 signaling pathways, which are important for inflammatory responses69. Pseudo-time analysis indicated that the expression of UBE2Q1 increased with differentiation of key cell types, suggesting its role in cell development and function in sepsis. These findings highlight the importance of intercellular communication in sepsis progression and the potential impact of prognostic genes on these processes.
GSVA was used to explore the biological pathways involved in key cells, and the results highlighted classic signaling pathways, such as P53 and Notch. The P53 pathway is a crucial tumor suppressor involved in cell cycle regulation and apoptosis, and its suppression can lead to uncontrolled cell proliferation and impaired immune response in sepsis70. Yu et al. found that P53 affects the occurrence of ferritinophagy by regulating autophagy-related genes, iron metabolism-related proteins, NCOA4, and oxidative stress71. In contrast, the Notch pathway is vital for cell differentiation and immune regulation, and its dysregulation contributes to immune dysfunction observed in sepsis72. However, Notch exists in different states in different cell types. In NK cells in sepsis, Notch is inhibited but is activated in other key cells. This complex regulation suggests that immune cells and pathways are involved in multiple mechanisms of sepsis, which require further exploration.
This study identified UBE2Q1, NEDD4L, and TCP11L2 as key ferritinophagy-related genes involved in the prognosis of sepsis. We not only established a prognostic model capable of effectively stratifying patients according to mortality risk but also, at single-cell resolution, uncovered the dynamic expression patterns of these genes within immune cells and their close associations with specific immune subsets, such as γδ T cells and NK cells. These findings introduce a novel paradigm for understanding the immunometabolic dysregulation of sepsis from the perspective of iron metabolism–immune crosstalk, providing both a theoretical foundation and potential targets for the development of innovative prognostic assessment tools and precision immunotherapeutic strategies for sepsis in the future.
Despite the significant findings of this study, several limitations should be acknowledged. First, although three ferritinophagy-related prognostic genes were identified and a risk prediction model was established, the specific molecular mechanisms through which these genes influence sepsis remain to be elucidated. In particular, their interactions with core molecules within the ferritinophagy pathway, as well as their regulatory roles in immune dysregulation and metabolic abnormalities during sepsis, have not yet been fully clarified.Second, this study primarily relied on bioinformatic analyses of public databases, lacking validation in independent clinical cohorts and functional experiments to confirm the expression patterns, mechanistic roles, and causal relationships of the key genes. Therefore, the generalizability and robustness of the prognostic model require further confirmation through prospective studies.
Future research can advance in several directions. (1) In vitro cellular and in vivo animal experiments should be conducted to elucidate the precise roles of these prognostic genes in the ferritinophagy process. (2) Independent, multicenter clinical cohorts of sepsis patients should be collected to evaluate the correlations between the expression of these genes and patient survival, inflammatory markers, and organ function scores, thereby optimizing the risk model and nomogram to enhance clinical applicability.(3) Based on predictions from the DisGeNet and DsigDB databases, drug repurposing studies involving compounds such as anisomycin and valproic acid could be performed to verify their efficacy in improving sepsis outcomes by modulating these prognostic genes. Moreover, integrating single-cell RNA sequencing data could help identify the key cellular subpopulations involved and uncover the roles of these genes in cell–cell communication. Finally, exploring potential cross-disease associations of these prognostic genes with disorders such as Parkinson’s disease or cancer may provide novel insights into shared molecular mechanisms and open new avenues for therapeutic innovation in sepsis.
Conclusion
This study successfully identified three ferritinophagy-related prognostic genes (UBE2Q1, NEDD4L, and TCP11L2) in sepsis using bioinformatic analysis and developed a predictive model based on these genes that accurately forecast patient outcomes. Our findings indicate that these genes are enriched in pathways related to ribosomes, Parkinson’s disease, and oxidative phosphorylation, suggesting that they may affect sepsis progression through regulation of protein synthesis, mitochondrial function, and energy metabolism. Furthermore, immune infiltration analysis revealed high levels of resting NK cells and gamma delta (γδ) T cells in the high-risk group, highlighting their crucial roles in the immune response to sepsis. Single-cell RNA sequencing provided insights into the dynamic expression changes of these prognostic genes in key cell types, offering a new perspective on the cellular and molecular mechanisms underlying sepsis.These findings have significant clinical implications and provide a foundation for future research to further understand and treat sepsis.
Data availability
All data generated in this study are publicly available in the Gene Expression Omnibus (GEO) database (http://www.ncbi.nlm.nih.gov/). Specifically, the datasets GSE154918 and GSE134347. All materials used in this study are commercially available and are described in detail in the Methods section. Additional data supporting the findings of this study are available from the corresponding authors upon request. The R script and all supplementary materials are available at https://pan.baidu.com/s/1aCBDCFEjVMZda-JM0l9ukA Extraction code: 1234.
Abbreviations
- FRGs:
-
Ferritinophagy-related genes
- MR:
-
Mendelian randomization
- GWAS:
-
Genome-wide association studies
- UVMR:
-
Univariable Mendelian randomization
- LASSO:
-
Least absolute shrinkage and selection operator
- GSEA:
-
Gene set enrichment analysis
- NK:
-
Natural killer
- SIRS:
-
Systemic inflammatory response syndrome
- WGCNA:
-
Weighted gene co-expression network analysis
- GEO:
-
Gene expression omnibus
- PBMCs:
-
Peripheral blood mononuclear cells
- IEU:
-
Integrative epidemiology unit
- SNPs:
-
Single nucleotide polymorphisms
- eQTL:
-
Expression quantitative trait loci
- DEGs:
-
Differently expressed genes
- MAD:
-
Median absolute deviation
- ME:
-
Module eigengene
- GO:
-
Gene ontology
- KEGG:
-
Kyoto encyclopedia of genes and genome
- PPI:
-
Protein-protein interaction
- IVs:
-
Instrumental variables
- LD:
-
Linkage disequilibrium
- IVW:
-
Inverse variance weighted
- HRs:
-
Hazard ratios
- LOO:
-
Leave-one-out
- PH:
-
Proportional hazards
- OS:
-
Overall survival
- K-M:
-
Kaplan-Meier
- ROC:
-
Operating characteristic
- AUC:
-
Area under curve
- QC:
-
Quality control
- HVGs:
-
Highly variable genes
- PCA:
-
Principal component analysis
- PCs:
-
Principal components
- UMAP:
-
Uniform manifold approximation and projection
References
Gong, T., Liu, Y. T. & Fan, J. Exosomal mediators in sepsis and inflammatory organ injury: Unraveling the role of exosomes in intercellular crosstalk and organ dysfunction. Mil. Med. Res. 11, 24. https://doi.org/10.1186/s40779-024-00527-6 (2024).
Donnelly, J. P. et al. Impact of reducing time-to-antibiotics on sepsis mortality, antibiotic use, and adverse events. Ann. Am. Thorac. Soc. 21, 94–101. https://doi.org/10.1513/AnnalsATS.202306-505OC (2024).
Rudd, K. E. et al. Global, regional, and national sepsis incidence and mortality, 1990–2017: analysis for the Global Burden of Disease Study. Lancet 395, 200–211. https://doi.org/10.1016/S0140-6736(19)32989-7 (2020).
Li, J., Shen, L. & Qian, K. Global, regional, and national incidence and mortality of neonatal sepsis and other neonatal infections, 1990–2019. Front. Public Health 11, 1139832. https://doi.org/10.3389/fpubh.2023.1139832 (2023).
Weng, L. et al. National incidence and mortality of hospitalized sepsis in China. Crit. Care 27, 84. https://doi.org/10.1186/s13054-023-04385-x (2023).
Guo, Q. et al. Organism type of infection is associated with prognosis in sepsis: An analysis from the MIMIC-IV database. BMC Infect. Dis. 23, 431. https://doi.org/10.1186/s12879-023-08387-6 (2023).
Sweeney, T. E., Liesenfeld, O. & May, L. Diagnosis of bacterial sepsis: why are tests for bacteremia not sufficient?. Expert Rev. Mol. Diagn. 19, 959–962. https://doi.org/10.1080/14737159.2019.1660644 (2019).
Liu, Y. C. et al. Ferritinophagy induced ferroptosis in the management of cancer. Cell. Oncol. 47, 19–35. https://doi.org/10.1007/s13402-023-00858-x (2024).
Kolahdouzmohammadi, M., Kolahdouz-Mohammadi, R., Tabatabaei, S. A., Franco, B. & Totonchi, M. Revisiting the role of autophagy in cardiac differentiation: A comprehensive review of interplay with other signaling pathways. Genes https://doi.org/10.3390/genes14071328 (2023).
You, J., Li, Y. & Chong, W. The role and therapeutic potential of SIRTs in sepsis. Front. Immunol. 15, 1394925. https://doi.org/10.3389/fimmu.2024.1394925 (2024).
Zhang, J. et al. YAP1 alleviates sepsis-induced acute lung injury via inhibiting ferritinophagy-mediated ferroptosis. Front. Immunol. 13, 884362. https://doi.org/10.3389/fimmu.2022.884362 (2022).
Li, Y. & Kan, X. Mendelian randomization analysis to analyze the genetic causality between different levels of obesity and different allergic diseases. BMC Pulm. Med. 23, 352. https://doi.org/10.1186/s12890-023-02636-9 (2023).
Wang, G. et al. Identification of hepatocellular carcinoma-related subtypes and development of a prognostic model: a study based on ferritinophagy-related genes. Discov. Oncol. 14, 147. https://doi.org/10.1007/s12672-023-00756-6 (2023).
Liu, S., Xie, X., Lei, H., Zou, B. & Xie, L. Identification of key circRNAs/lncRNAs/miRNAs/mRNAs and pathways in preeclampsia using bioinformatics analysis. Med. Sci. Monitor Int. Med. J. Exper. Clin. Res. 25, 1679–1693. https://doi.org/10.12659/MSM.912801 (2019).
Dai, W. et al. LPIN1 is a regulatory factor associated with immune response and inflammation in sepsis. Front. Immunol. 13, 820164. https://doi.org/10.3389/fimmu.2022.820164 (2022).
Cheng, Q., Chen, X., Wu, H. & Du, Y. Three hematologic/immune system-specific expressed genes are considered as the potential biomarkers for the diagnosis of early rheumatoid arthritis through bioinformatics analysis. J. Transl. Med. 19, 18. https://doi.org/10.1186/s12967-020-02689-y (2021).
Hanzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinf. 14, 7. https://doi.org/10.1186/1471-2105-14-7 (2013).
Wang, S. et al. An Eight-CircRNA assessment model for predicting biochemical recurrence in prostate cancer. Front. Cell Develop. Biol. 8, 599494. https://doi.org/10.3389/fcell.2020.599494 (2020).
Langfelder, P. & Horvath, S. WGCNA: An R package for weighted correlation network analysis. BMC Bioinf. 9, 559. https://doi.org/10.1186/1471-2105-9-559 (2008).
Chen, J., Huang, Z., Qin, C., Pang, Z. & Chen, Y. Identification of mitophagy-related biomarkers in human rheumatoid arthritis using machine learning models. Artificial Cells Nanomed. Biotechnol. 53, 287–303. https://doi.org/10.1080/21691401.2025.2533361 (2025).
Kanehisa, M., Furumichi, M., Sato, Y., Matsuura, Y. & Ishiguro-Watanabe, M. KEGG: Biological systems database as a model of the real world. Nucleic Acids Res. 53, D672–D677. https://doi.org/10.1093/nar/gkae909 (2025).
Wu, T. et al. clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innovation 2, 100141. https://doi.org/10.1016/j.xinn.2021.100141 (2021).
Zhou, H. et al. Education and lung cancer: a Mendelian randomization study. Int. J. Epidemiol. 48, 743–750. https://doi.org/10.1093/ije/dyz121 (2019).
Zhao, M. et al. Identification of growth-related snps and genes in the genome of the pearl oyster (Pinctada fucata) using GWAS. Fishes 8, 296. https://doi.org/10.3390/fishes8060296 (2023).
Bowden, J., Davey Smith, G. & Burgess, S. Mendelian randomization with invalid instruments: Effect estimation and bias detection through Egger regression. Int. J. Epidemiol. 44, 512–525. https://doi.org/10.1093/ije/dyv080 (2015).
Burgess, S., Zuber, V., Gkatzionis, A. & Foley, C. N. Modal-based estimation via heterogeneity-penalized weighting: model averaging for consistent and efficient estimation in Mendelian randomization when a plurality of candidate instruments are valid. Int. J. Epidemiol. 47, 1242–1254. https://doi.org/10.1093/ije/dyy080 (2018).
Hemani, G. et al. The MR-Base platform supports systematic causal inference across the human phenome. Elife https://doi.org/10.7554/eLife.34408 (2018).
Burgess, S. et al. Using published data in Mendelian randomization: a blueprint for efficient identification of causal risk factors. Eur. J. Epidemiol. 30, 543–552. https://doi.org/10.1007/s10654-015-0011-z (2015).
Hartwig, F. P., Davey Smith, G. & Bowden, J. Robust inference in summary data Mendelian randomization via the zero modal pleiotropy assumption. Int. J. Epidemiol. 46, 1985–1998. https://doi.org/10.1093/ije/dyx102 (2017).
Friedman, J., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33, 1–22 (2010).
Yang, H. et al. The prognostic role of blood inflammatory biomarkers and EGFR mutation status in stage IIIA/N2 non-small cell lung cancer patients treated with trimodality therapy. Front. Oncol. 11, 707041. https://doi.org/10.3389/fonc.2021.707041 (2021).
Craven, K. E., Gokmen-Polar, Y. & Badve, S. S. CIBERSORT analysis of TCGA and METABRIC identifies subgroups with better outcomes in triple negative breast cancer. Sci. Rep. 11, 4691. https://doi.org/10.1038/s41598-021-83913-7 (2021).
Goncharova, I. A. et al. Genetic predisposition to early myocardial infarction. Mol. Biol. 54, 224–232. https://doi.org/10.31857/S0026898420020044 (2020).
Chin, C. H. et al. cytoHubba: identifying hub objects and sub-networks from complex interactome. BMC Syst. Biol. 8(Suppl 4), S11. https://doi.org/10.1186/1752-0509-8-S4-S11 (2014).
Yu, L. et al. Characterization of cancer-related fibroblasts (CAF) in hepatocellular carcinoma and construction of CAF-based risk signature based on single-cell RNA-seq and bulk RNA-seq data. Front. Immunol. 13, 1009789. https://doi.org/10.3389/fimmu.2022.1009789 (2022).
Qiu, X. et al. Dynamic changes in human single-cell transcriptional signatures during fatal sepsis. J. Leukoc. Biol. 110, 1253–1268. https://doi.org/10.1002/JLB.5MA0721-825R (2021).
Fang, Z., Li, J., Cao, F. & Li, F. Integration of scRNA-Seq and Bulk RNA-Seq reveals molecular characterization of the immune microenvironment in acute pancreatitis. Biomolecules https://doi.org/10.3390/biom13010078 (2022).
Rao, Y. et al. Single-cell transcriptome profiling identifies phagocytosis-related dual-feature cells in a model of acute otitis media in rats. Front. Immunol. 12, 760954. https://doi.org/10.3389/fimmu.2021.760954 (2021).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30. https://doi.org/10.1093/nar/28.1.27 (2000).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. Publ. Protein Soc. 28, 1947–1951. https://doi.org/10.1002/pro.3715 (2019).
Scicluna, B. P. et al. Classification of patients with sepsis according to blood genomic endotype: A prospective cohort study. Lancet Respir. Med. 5, 816–826. https://doi.org/10.1016/S2213-2600(17)30294-1 (2017).
Li, M. et al. Experimental study on ratio optimization and application of improved bonded dust suppressant based on wetting effect. J. Air Waste Manag. Assoc. 73, 394–402. https://doi.org/10.1080/10962247.2023.2189173 (2023).
Villanueva, J. et al. alpha3beta4 acetylcholine nicotinic receptors are components of the secretory machinery clusters in chromaffin cells. Int. J. Mol. Sci. https://doi.org/10.3390/ijms23169101 (2022).
Li, J. Y. et al. Ferritinophagy: A novel insight into the double-edged sword in ferritinophagy-ferroptosis axis and human diseases. Cell Prolif. 57, e13621. https://doi.org/10.1111/cpr.13621 (2024).
Abdul Rehman, S. A. et al. Discovery and characterization of noncanonical E2-conjugating enzymes. Sci. Adv. 10, 0123. https://doi.org/10.1126/sciadv.adh0123 (2024).
Fahmidehkar, M. A., Shafiee, S. M., Eftekhar, E., Mahbudi, L. & Seghatoleslam, A. Induction of cell proliferation, clonogenicity and cell accumulation in S phase as a consequence of human UBE2Q1 overexpression. Oncol. Lett. 12, 2169–2174. https://doi.org/10.3892/ol.2016.4860 (2016).
Grzmil, P. et al. Embryo implantation failure and other reproductive defects in Ube2q1-deficient female mice. Reproduction 145, 45–56. https://doi.org/10.1530/REP-12-0054 (2013).
Zhang, B. et al. Upregulation of UBE2Q1 via gene copy number gain in hepatocellular carcinoma promotes cancer progression through beta-catenin-EGFR-PI3K-Akt-mTOR signaling pathway. Mol. Carcinog. 57, 201–215. https://doi.org/10.1002/mc.22747 (2018).
Chang, R. et al. Upregulated expression of ubiquitin-conjugating enzyme E2Q1 (UBE2Q1) is associated with enhanced cell proliferation and poor prognosis in human hapatocellular carcinoma. J. Mol. Histol. 46, 45–56. https://doi.org/10.1007/s10735-014-9596-x (2015).
Shafiee, S. M. et al. UBE2Q1 expression in human colorectal tumors and cell lines. Mol. Biol. Rep. 40, 7045–7051. https://doi.org/10.1007/s11033-013-2824-8 (2013).
Seghatoleslam, A. et al. Expression of the novel human gene, UBE2Q1, in breast tumors. Mol. Biol. Rep. 39, 5135–5141. https://doi.org/10.1007/s11033-011-1309-x (2012).
Huang, M., Cai, S. & Su, J. The pathogenesis of sepsis and potential therapeutic targets. Int. J. Mol. Sci. https://doi.org/10.3390/ijms20215376 (2019).
Hosseini, S. M. et al. E2 ubiquitin-conjugating enzymes in cancer: Implications for immunotherapeutic interventions. Clinica Chimica Acta International Journal of Clinical Chemistry 498, 126–134. https://doi.org/10.1016/j.cca.2019.08.020 (2019).
Saito, Y. et al. Cardiac-specific ITCH overexpression ameliorates septic cardiomyopathy via inhibition of the NF-kappaB signaling pathway. J. Mol. Cell. Cardiol. Plus 2, 100018. https://doi.org/10.1016/j.jmccpl.2022.100018 (2022).
Chen, X. S. et al. Losartan attenuates sepsis-induced cardiomyopathy by regulating macrophage polarization via TLR4-mediated NF-kappaB and MAPK signaling. Pharmacol. Res. 185, 106473. https://doi.org/10.1016/j.phrs.2022.106473 (2022).
Qin, F., Tan, H., Yang, Y., Xu, L. & Yang, X. Upregulation of Cullin1 neddylation promotes glycolysis and M1 polarization of macrophage via NF-kappaB p65 pathway in sepsis. Funct. Integr. Genomics 24, 204. https://doi.org/10.1007/s10142-024-01483-z (2024).
Manning, J. A. & Kumar, S. Physiological functions of Nedd4-2: lessons from knockout mouse models. Trends Biochem. Sci. 43, 635–647. https://doi.org/10.1016/j.tibs.2018.06.004 (2018).
Goel, P., Manning, J. A. & Kumar, S. NEDD4-2 (NEDD4L): the ubiquitin ligase for multiple membrane proteins. Gene 557, 1–10. https://doi.org/10.1016/j.gene.2014.11.051 (2015).
Duerr, J. et al. Conditional deletion of Nedd4-2 in lung epithelial cells causes progressive pulmonary fibrosis in adult mice. Nat. Commun. 11, 2012. https://doi.org/10.1038/s41467-020-15743-6 (2020).
Sanjabi, S., Oh, S. A. & Li, M. O. Regulation of the immune response by TGF-beta: From conception to autoimmunity and infection. Cold Spring Harbor Perspect. Biol. https://doi.org/10.1101/cshperspect.a022236 (2017).
Li, S., Wang, Z., Tong, H., Li, S. & Yan, Y. TCP11L2 promotes bovine skeletal muscle-derived satellite cell migration and differentiation via FMNL2. J. Cell. Physiol. 235, 7183–7193. https://doi.org/10.1002/jcp.29617 (2020).
Bongers, K. S. et al. The gut microbiome modulates body temperature both in sepsis and health. Am. J. Respir. Crit. Care Med. 207, 1030–1041. https://doi.org/10.1164/rccm.202201-0161OC (2023).
Brown, G. C. The endotoxin hypothesis of neurodegeneration. J. Neuroinflammation 16, 180. https://doi.org/10.1186/s12974-019-1564-7 (2019).
Eyenga, P., Rey, B., Eyenga, L. & Sheu, S. S. Regulation of oxidative phosphorylation of liver mitochondria in sepsis. Cells https://doi.org/10.3390/cells11101598 (2022).
Yao, R. Q. et al. Single-cell transcriptome profiling of the immune space-time landscape reveals dendritic cell regulatory program in polymicrobial sepsis. Theranostics 12, 4606–4628. https://doi.org/10.7150/thno.72760 (2022).
Wan, J. et al. Infiltrated IL-17A-producing gamma delta T cells play a protective role in sepsis-induced liver injury and are regulated by CCR6 and gut commensal microbes. Front. Cell. Infect. Microbiol. 13, 1149506. https://doi.org/10.3389/fcimb.2023.1149506 (2023).
Bird, M. F. et al. In vitro sepsis up-regulates Nociceptin/Orphanin FQ receptor expression and function on human T- but not B-cells. Br. J. Pharmacol. 180, 2298–2314. https://doi.org/10.1111/bph.16088 (2023).
Yao, R. Q. et al. Single-cell transcriptome profiling of sepsis identifies HLA-DR(low)S100A(high) monocytes with immunosuppressive function. Mil. Med. Res. 10, 27. https://doi.org/10.1186/s40779-023-00462-y (2023).
Du, J. et al. Research progress of the chemokine/chemokine receptor axes in the oncobiology of multiple myeloma (MM). Cell Commun. Signal 22, 177. https://doi.org/10.1186/s12964-024-01544-7 (2024).
Lin, X. et al. Quercetin ameliorates ferroptosis of rat cardiomyocytes via activation of the SIRT1/p53/SLC7A11 signaling pathway to alleviate sepsis-induced cardiomyopathy. Int. J. Mol. Med. https://doi.org/10.3892/ijmm.2023.5319 (2023).
Yu, J. A. et al. mechanism linking ferroptosis and ferritinophagy in melatonin-related improvement of diabetic brain injury. iScience 27, 109511. https://doi.org/10.1016/j.isci.2024.109511 (2024).
Gallenstein, N., Tichy, L., Weigand, M. A. & Schenz, J. Notch signaling in acute inflammation and sepsis. Int. J. Mol. Sci. https://doi.org/10.3390/ijms24043458 (2023).
Author information
Authors and Affiliations
Contributions
Weichuan Xiong:Conceptualization, Data curation, Formal analysis, Writing – original draft,Writing–review & editing,Software;Fangpeng Liu:Supervision,Validation, Visua- lization, Methodology, Project admini- stration, Resources.
Corresponding author
Ethics declarations
Competing interests
The authors have declared that no conflict of interest exists.
Consent for publication
All listed authors consented to the submission, and all data were used with the consent of the person who generated them.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xiong, W., Liu, F. Ferritinophagy-related prognostic genes UBE2Q1, NEDD4L, and TCP11L2 for prognosis prediction in sepsis. Sci Rep 15, 43314 (2025). https://doi.org/10.1038/s41598-025-29680-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-29680-1