Abstract
Sudden sensorineural hearing loss (SSNHL) is an acquired idiopathic hearing loss in which recent studies have suggested that microRNAs (miRNAs) have a pathogenetic role. We assessed six specific miRNAs linked to SSNHL in serum samples drawn from 11 SSNHL patients and 11 age and sex matched controls. Quantitative real-time PCR Cycle threshold (Ct) values were normalized to an internal control to determine relative miRNA levels. Inter-group mean age, sex, mean or median normalized Ct values were compared using Independent Sample t, Chi-squared or Mann-Whitney U tests. The relative expression (ΔΔCt) of miR-210-3p, miR-23a-2p, miR18b-5p -2.75, -3.33, and − 1.29, corresponded to statistically different ΔCt values (Students T-test, p = 0.006; Mann-Whitney U, p = 0.001 and 0.02) and fold changes of 6.73, 10.87, and 2.45, respectively, in SSNHL patients compared with controls. No significant difference in the expression levels of miR-15a-5p, miR-143-3p, or miR-183-5p was found. This study supports consistent dysregulation of miR-210-3p/-23a-3p and − 18b-5p across different populations of SSNHL patients, suggesting that they play a key role in the pathogenesis of SSNHL.
Similar content being viewed by others
Introduction
Sudden sensorineural hearing loss (SSNHL) refers to the development of an acquired idiopathic hearing loss within 72 h1. SSNHL is typically unilateral and often accompanied by symptoms of tinnitus, aural pressure, and, in just under 25% of cases, vertigo2. Epidemiological studies suggest different incidence rates in different study populations, ranging from 27 to 60 cases per 100,000 individuals in the United States and Japan, respectively3,4. The true incidence may be higher as individuals who recover spontaneously soon after onset may not seek treatment. Indeed, spontaneous recovery to within 10dB of the unaffected ear occurs in 45–65% of cases5,6. A slight male predominance was observed in the US study3, while the Japanese data revealed a threefold female predominance4. The pathophysiology of SSNHL is unknown, however viral and vascular mechanisms are widely proposed7,8.
MicroRNAs (miRNAs) are short RNA molecules, typically around 23 nucleotides in length. miRNAs exert control over target gene expression at the post-transcriptional level, primarily through the specific targeting of mRNA sequences by the miRNA’s 7-nucleotide seed region9. Over 2500 human miRNAs have been identified to date, with research suggesting their involvement in the regulation of approximately 30% of genes10. MiRNAs play pivotal roles in various signaling pathways governing processes such as organogenesis, hematopoiesis, differentiation, cell proliferation, apoptosis, and fat metabolism11. MiRNAs have been identified in body fluids including serum, plasma, saliva, and urine and are easily detectable reliable disease markers12.
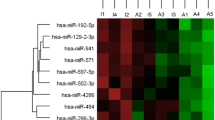
MiRNAs play a fundamental role in inner ear development, with evidence that miRNA regulation of gene expression is associated with the development of acquired hearing loss13,14. Dysregulation of some circulating miRNAs has been identified in SSNHL patients15,16,17. A better understanding of the cellular pathways influenced by these miRNAs changes may suggest novel treatments and point to the underlying cause of SSNHL. Li et al.’s15 study of miRNAs in plasma samples of 9 SSNHL patients and 3 controls recruited in Guandong China found miR-34a/15a/23a/210/18b/548n/143 to be differentially expressed, suggesting a role for these miRNAs in SSNHL pathogenesis. The age and sex of the participants was not presented15. Ha et al.17 searched for differential expression of the miRNAs reported by Li et al.15 and 2 ototoxicity related miRNAs in plasma samples of SSNHL patients and control participants recruited in Wonju, Korea. They did not provide participant demographic data and corroborated Li et al.’s findings with respect to differential expression of hsa-miR-23a/-18b, but not hsa-miR-15a in SSNHL patients. Furthermore, Ha et al. found hsa-miR-143/-183/-210 all implicated by Li et al. in SSNHL to be differentially expressed in SSNHL patients. Nunez et al.16 found a different set of differentially expressed miRNAs namely hsa-miR-590-5p/ -186-5p/ -195-5p/ -140-3p/ -128-3p/ -132-3p/ -375-3p, and − 30a-3p based on serum samples drawn from participants recruited in Vancouver, Canada.
Li et al.’s bioinformatic analysis of SSNHL differentially expressed miRNAs identified the highest enrichment of their target genes in the MAPK signaling, Arachidonic acid metabolism and complement and coagulation cascade cellular pathways15. Nunez et al. ‘s similar analysis identified the highest target gene enrichment in PI3K/Akt and MAPK signaling pathways and HTLV-1 infection pathway14. PI3K/Akt and MAPK cellular signaling pathways are involved in ischemia/reperfusion and hypoxia response mediation mechanisms through regulatory effects on oxidative stress, inflammation, apoptosis and other cellular processes18,19. Arachidonic acid, complement/coagulation cascade, and HTLV-1 pathways are inflammation related pathways. Therefore, Li et al.’s and Nunez et al.’s findings together suggest that oxidative stress and inflammation are part of the underlying cellular disease processes in idiopathic SSNHL.
Identification of a common dysregulated group of miRNAs will help in to determine which of the above pathways and processes play the greatest role in SSNHL pathogenesis. A follow-on study investigated if the blood fraction studied accounted for the difference in Nunez et al.’s14 miRNA findings and that of Ha15 & Li et al.13. No difference in the expression levels of the differentially expressed miRNAs in paired serum and plasma samples drawn from another cohort of 17 SSNHL patients recruited in Vancouver, Canada was found20. Therefore, the blood fraction studied does not explain the discrepancy between Nunez et al.’s findings and the studies by Ha15 and Li et al.17. The incongruence in SSNHL associated miRNA findings in Asian and North-American studies maybe due to population specific miRNA variation which has been demonstrated especially in oncogenic related miRNAs across the globe21. Hence, we specifically evaluate here the expression levels of hsa-miR-15a/-23a/-210/-18b/-143/-183 previously found to be differentially expressed in SSNHL patients recruited in China15 and Korea17 in a cohort of participants recruited in Vancouver Canada. The null hypothesis is that none of these miRNAs will be dysregulated in participants in Canada.
Results
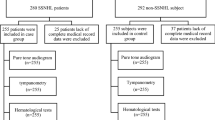
A total of 11 SSNHL patients and 11 controls matched by age and sex, were included in the study. Each group consisted of 4 males and 7 females (Fishers’ exact, p value = 1.00). There was also no statistically significant difference in mean age between SSNHL patients (mean age = 44.27 years, s.dev. = 9.6) and controls (mean age = 42.64 years, s.dev. = 11.3) (p value = 0.4, independent samples t-test) confirming that the groups were effectively matched by age (Table 1).
Candidate reference miRNAs (miR-103a-3p, miR-16-5p, miR-191-5p) were evaluated for stability across triplicate PCR runs. RefFinder confirmed miR-191-5p as the most stable reference across both SSNHL and control groups, supporting its use for normalization.
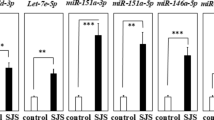
In this case–control study, serum miRNA normalized mean or median ΔCt distributions in SSNHL patients and controls were compared using Student’s t or Mann–Whitney U tests depending on data normality and homoscedasticity. The mean (s. dev) and median (interquartile range, IQR) ΔCt values of miR-210-3p, miR-23a-3p and miR-18b-5p 4.61 (2.36), -3.04 (2.10) and 2.99 (2.77) in the SSNHL group were statistically different (Student t-test, p = 0.006; Mann-Whitney U, p = 0.001; Mann WhitneyU, p = 0.028) to those in the controls 7.37(1.78), 1.75 (0.7) and 4.40 (1.23) respectively. The corresponding relative expression (ΔΔCt) levels (SSNHL – Control) were − 2.75, − 3.44, and − 1.29, consistent with fold changes of 6.73, 10.87, and 2.45, respectively, in SSNHL patients. However, no significant case-control differences were observed in the ΔCt values of miR-15a-5p, miR-143-3p, or miR-183-5p corresponding to fold changes of 1.53, 2.55, and 2.83, respectively (Fig. 1; Table 2).
Box-and-whisker plots of ΔΔCt values (SSNHL – Control) for six miRNAs. Each box represents the interquartile range (IQR, 25th–75th percentile), with the horizontal line inside the box showing the median and the “X” indicating the mean. Whiskers extend to 1.5×IQR, and circles represent outliers. Negative ΔΔCt values indicate higher relative expression (up-regulation) in SSNHL patients, while positive ΔΔCt values indicate lower relative expression (down-regulation). Statistically significant differences were observed for miR-210-3p (t-test, p = 0.006), miR-23a-3p (Mann–Whitney U test, p = 0.001), and miR-18b-5p (Mann–Whitney U test, p = 0.02). No significant differences were detected for miR-143-3p, miR-15a-5p, or miR-183-5p. Significance levels are indicated as p < 0.05 (*), p < 0.01 (**), and p < 0.001 (***).
Discussion
We found increased expression of miR-210 in our patients with SSNHL compared to controls, which agrees with Ha et al.’s17 and Li et al.’s15 findings. We also found significantly increased expression of miR-23a in our SSNHL patients compared to controls, while Ha et al.17 and Li et al.15 found decreased expression in SSNHL patients. We also found a significant increase in miR-18b expression in SSNHL patients, which agrees with Li et al.15 but not Ha et al.17, who reported a significant decrease in miR-18b expression. Inter-group miR-15a, miR-143, and miR-183 expression levels were not significantly different in our study. Ha et al.17 similarly found no significant inter-group difference in miR-15a expression levels, but they reported significant differences in miR-143 and miR-183 expression between SSNHL patients and controls. In contrast, Li et al.15 found that miR-15a, and miR-143 were differentially expressed in SSNHL patients versus controls.
One factor that may contribute to the different findings reported here compared with Ha et al.17 and Li et al.15 is the choice of reference miRNAs. Our values are reported relative to miR-191, whereas Ha et al.‘s. values were reported relative to either miR-103 or miR-16 and Li et al.’s15 to a data distribution derived model. We conducted a thorough evaluation of potential reference miRNAs to guide our choice and thus achieve accurate normalization in this study’s qPCR analysis. Specifically, we undertook triplicate PCR runs with previously described candidate reference miRNAs miR-103a-3p /-16-5p17 and /-191-5p22,23 with all SSNHL and control samples. Among these candidates, miR-191-5p was identified with RefFinder30 as the most stable reference miRNA. Therefore, miR-191-5p was selected as the reference miRNA for this study. The decision to use miR-191-5p is further supported by its frequent use as a stable reference miRNA in similar studies20,22.This stability is critical for normalization, as it reduces potential variability and ensures the reliability of comparative expression analysis.
Interestingly, there was a significant difference in the variance of most of the tested miRNA expression values in the control and SSNHL patient groups. The SSNHL patient group had a much greater variance in delta Ct values in all tested miRNAs except miR-210 and − 143-3p. This difference may be due to variable dysregulation of miRNAs caused by SSNHL. Ha et al.17 demonstrated similar, albeit less stark, differences in variance of values between their SSNHL patients and controls, supporting a SSNHL disease effect. Strikingly, three different groups of investigators in three widely disparate geographic locations studying participants from differing populations have consistently identified dysregulation of miR − 23a/-210 and − 18b-5p in SSNHL patients. One of miR-23a’s target genes, NR3C1, may have an effect on the vulnerability of the cochlea to noise injury, and may also have neuroprotective effects24,25. MiR-210 is speculated to regulate the genes YWHAG and SIN3A, which have been linked to age-related hearing loss and inner-ear hair cell proliferation, respectively15. MiR-18b-5p appears to be associated with the MDM2 gene, a negative regulator of p53 that has been linked to developmental auditory disturbances17. These miRNAs are particularly worthy of further study.
The discrepancies between our findings and those of Ha et al.17 and Li et al.15 may reflect that while patients with SSNHL share the same symptom phenotype there are differences in the underlying disease processes in individual patients. Additionally, or alternatively miRNA genetic variation across different populations may be responsible for the difference21. The current study investigated 22 participants, while Ha et al.17 and Li et al.15 investigated 45 and 12 participants respectively. Further studies with greater sample sizes may identify other SSNHL associated miRNAs and be better powered to undertake sub-group analysis by age, sex, hearing loss severity, treatment and hearing loss recovery. Such work is required to corroborate whether miRNAs can be used as predictive biomarkers of SSNHL pathogenesis, severity and recovery.
Materials and methods
Study populations and sampling
Adult patients aged 18 years and older presenting with sudden sensorineural hearing loss (SSNHL) as defined by the criteria outlined by the American Academy of Otolaryngology-Head and Neck Surgery (AAOHNS)1 were recruited for this study. Inclusion criteria comprised individuals willing to provide signed informed consent, aged 18 years or older, and experiencing pure tone audiometrically documented sensorineural hearing loss of at least 30 dB across three contiguous frequencies within 72 h of onset.
Controls were selected to match the demographic characteristics of the SSNHL patients and were recruited from individuals without any history of hearing loss or otologic disorders.
Exclusion criteria for both SSNHL patients and controls included the inability to provide blood samples, complete diagnostic physical and hearing tests, or the presence of known causes for hearing loss, major intercurrent medical illnesses, or coexisting ear pathologies. All methods were performed in accordance with the guidelines and regulations approved by the Ethics Committee of the University of British Columbia.
Clinical Examination, patient Recruitment, and pure tone audiometry
Clinical evaluation and participant recruitment involved comprehensive examination conducted by Otolaryngologist Head and Neck Surgeons registered with the College of Physicians and Surgeons of British Columbia. Otoscopic examination of the ear canals and tympanic membranes was performed to identify potential causes of conductive hearing loss, such as middle ear fluid, otitis media, foreign bodies, cerumen impaction, otitis externa, or trauma. Additionally, Rhinne’s tests utilizing 256–512 Hz tuning forks were administered to identify sensorineural hearing losses26. Ethical approval for this study was obtained from the University of British Columbia’s Clinical Research Ethics Board, and enrollment of SSNHL patients occurred at the Division of Otolaryngology Head and Neck Surgery, Vancouver General Hospital, between 2017 and 2022. Individuals without hearing impairments, whose hearing thresholds were confirmed to be 25 dB or better across all pure tone audiometric (PTA) test frequencies, were recruited from hospital staff or their acquaintances as controls.
All participants underwent comprehensive pure tone audiometric assessment conducted by Audiologists or Hearing Instrument Practitioners registered with the College of Speech and Hearing Health Professionals of British Columbia. These assessments were conducted in soundproof environments using either insert or supra-aural earphones and bone vibrators, adhering to established practice guidelines27. The recorded results from these assessments were collected by the investigators to ascertain the suitability of participants for inclusion as either controls or SSNHL patients.
For the purposes of this study, sensorineural hearing loss was defined as a loss of > 25dB at any frequency with an air-bone gap of < 10dB28,29, while conductive hearing loss was defined as an air-bone gap of > 25 dB at any frequency. Mixed hearing loss was characterized by both air conduction (AC) and bone conduction (BC) losses of > 25 dB, with an air-bone gap > 10dB. During audiometric assessment, if a patient did not respond to the maximum sound capacity of the audiometer, 5 dB was added to the maximum level to represent the threshold level.
Pure tone audiometry was employed to validate the diagnosis of SSNHL, classify the severity of hearing impairment upon presentation, and assess the patients’ hearing recovery. Air conduction pure-tone average thresholds were determined for affected ears at either four low frequencies (0.5, 1, 2, and 3–4 kHz) or three high frequencies (3 or 4, 6, and 8 kHz)30. The degree of SSNHL was categorized based on the hearing threshold in the affected ear31. The selection of frequencies to be averaged for each patient (high or low) was contingent upon the frequencies demonstrating the greatest degree of hearing loss in the initial pure tone audiogram.
Only SSNHL patients with a sensorineural hearing loss ≥ 30 dB across three contiguous frequencies were eligible for enrollment in the study. Control participants with any hearing loss > 25dB at any test frequency were excluded. Additionally, SSNHL patients with associated conductive or mixed hearing loss were also excluded from the study.
Blood collection and MicroRNA extraction
Blood samples were obtained for serum separation, and aliquots were frozen at -80˚C for subsequent RNA extraction. Total RNA was isolated from 200 µL of serum using the miRNeasy Mini Kit (Qiagen, Toronto, ON, Canada) following the manufacturer’s protocol. Briefly, 1 mL of Qiazol Lysis reagent was added to 200 µL of serum and incubated at room temperature for 5 min. Subsequently, 200 µL of Chloroform (Sigma-Aldrich, St. Louis, MO, USA) was added, and the samples were vortexed and incubated for an additional 5 min at room temperature before centrifugation at 12,000×g for 15 min at 4 °C. The resulting aqueous phase containing RNA was transferred to a new tube, and isopropanol (Fisher-Scientific, Thermo Fisher Scientific, Waltham, MA, USA) was added. After a 10 min incubation at room temperature, the samples were centrifuged at 12,000xg for 15 min at 4 °C. Next, 1.5 volumes of 100% ethanol were added and thoroughly mixed by pipetting. Up to 700 µL of sample, including any precipitate, was pipetted into an RNeasy MinElute spin column in a 2 ml collection tube and centrifuged at ≥ 8000 x g for 15 s at room temperature. The spin column was washed with 700 µL Buffer RWT and then 500 µL Buffer RPE before centrifuging for 15 s at ≥ 8000 x g. Subsequently, 500 µL of 80% ethanol was added to the spin column and centrifuged for 2 min at ≥ 8000 x g. Afterward, the spin column was placed in a new 2 ml collection tube with an open lid and centrifuged at full speed for 5 min to dry the membrane. Following the removal of the flow-through, 14 µL RNase-free water was added directly to the center of the spin column membrane.
MicroRNA reverse transcription
For microRNA reverse transcription, the TaqMan Advanced cDNA synthesis kit (Applied Biosystem) was utilized, incorporating a preamplification step followed by TaqMan™ advanced miRNA Assay as previously described16. In brief, a total of 30 µL and 2 µL of microRNAs, 3 µL of poly(A) reaction mix, 10 µl of ligation reaction mix, and 15 µL of reverse transcriptase reagents were incubated at 42˚C for 15 min and deactivated at 85˚C for 5 min. Subsequently, 5 µL of the RT reaction product was combined with 45 µL of the miR-Amp Reaction Mix before extracting and diluting cDNA 1:10 with nuclease-free water for microRNA real-time PCR analysis.
MicroRNA real-time PCR
For microRNA real-time PCR, Taqman® Advanced miRNA Assays and Taqman Fast Advanced master mix were employed on a QuantStudio Real-time PCR System (Applied Biosystems, Thermo Fisher Scientific, Waltham, USA). Individual qPCR assays were conducted in triplicate with a total reaction volume of 20 µL using hsa-miR-15a-5p/-18b-5p/-143-3p/-23a-3p/-210-3p/-183-5p primers (Table 3) as previously described22. The PCR thermocycler was programmed for denaturation at 95 °C for 20 s, followed by 40 annealing cycles of 95 °C for 1 s and 60 °C for 20 s. Normalization of microRNA values was carried out based on the reference microRNA, hsa-miR-191-5p, using the delta Ct method32. Reference miRNAs known to be stable in serum and not associated with SSNHL were identified from published studies17,20,22. To select the reference miRNA, we evaluated candidate miRNAs (miR-103a-3p17, miR-16-5p17,33, and miR-191-5p20,22,23 by performing triplicate PCR runs with all test samples and miRNA primers. Stability was evaluated using RefFinder34 which integrates geNorm, NormFinder, BestKeeper, and the comparative ΔCt method to generate a comprehensive stability ranking. miR-191-5p emerged as the most stable.
Statistical analysis
SPSS version 26 (IBM, Armonk, USA) was used for analyses. A Shapiro–Wilk test was used to determine if the ΔCt values obtained with each test miRNA in the SSNHL patient and control groups were normally distributed. Homoscedasticity was determined by Levene’s test of variance with a cut off of p < 0.05. The mean ΔCt values of the test miRNAs in the serum of SSNHL patients and controls were then compared with an Independent Sample Students’ t-test when Ct values were normally distributed and the data in both groups was homoscedastic. A Mann-Whitney U test was used to compare median expression levels when normality or equal variance assumptions were not satisfied. ΔΔCt values (SSNHL – Control) were calculated for each miRNA, and fold changes were derived using the 2^-ΔΔCt method, with controls as the comparative group for interpretability. Both ΔΔCt and fold-change values are reported in the Results. Box and Whisker plots showing median, mean, interquartile ΔΔCt values were generated using Excel, with statistical significance indicated as p < 0.05 (*), p < 0.01 (**), and p < 0.001 (***). Pearson’s chi-square test or Fisher’s exact test was used to assess the distribution of participants by sex in both groups. An independent t-test was used for inter-group comparison of the distribution of participants by age.
Data availability
The data presented in this study are available on request from the corresponding author.
References
Chandrasekhar, S. S. et al. Clinical practice guideline: sudden hearing loss (Update). Otolaryngology–Head Neck Surg. 161, S1–S45 (2019).
Xu, R. Z. et al. A retrospective comparative chart review of hearing recovery in neural and sensory type sudden sensorineural hearing loss patients. The J. Laryngology & Otology. 138, 1090–1094 (2024).
Alexander, T. H. & Harris, J. P. Incidence of sudden sensorineural hearing loss. Otology Neurotology. 34, 1586–1589 (2013).
Nakashima, T. et al. Idiopathic sudden sensorineural hearing loss in Japan. Acta Otolaryngol. 134, 1158–1163 (2014).
Mattox, D. E. & Simmons, F. B. Natural history of sudden sensorineural hearing loss. Annals Otology Rhinology Laryngology. 86, 463–480 (1977).
Byl, F. M. Jr Sudden hearing loss: eight years’ experience and suggested prognostic table. Laryngoscope 94, 647–661 (1984).
Linthicum, F. H. Jr, Doherty, J. & Berliner, K. I. Idiopathic sudden sensorineural hearing loss: vascular or viral? Otolaryngology–Head Neck Surg. 149, 914–917 (2013).
Tsuzuki, N. & Wasano, K. Idiopathic sudden sensorineural hearing loss: A review focused on the contribution of vascular pathologies. Auris Nasus Larynx. 51, 747–754 (2024).
Mitchell, P. S. et al. Circulating MicroRNAs as stable Blood-Based markers for cancer detection. Proc. Natl. Acad. Sci. 105, 10513–10518 (2008).
Rothenberg, M. & Lu, T. M. R. N. A. J. Allergy Clin. Immunol. 141, 1202–1207. (2018).
Ferrante, M. & Conti, G. O. Environment and neurodegenerative diseases: an update on MiRNA role. Microrna 6, 157–165 (2017).
Mohr, A. M. & Mott, J. L. Overview of microRNA Biology. In Proceedings of the Seminars in liver disease; Thieme Medical Publishers, 35, 003–011. (2015).
Chen, H., Wijesinghe, P. & Nunez, D. MicroRNAs in acquired sensorineural hearing loss. J. Laryngology Otology. 133, 650–657 (2019).
Pang, J. et al. Circulating miR-34a levels correlate with Age-Related hearing loss in mice and humans. Exp. Gerontol. 76, 58–67 (2016).
Li, Q. et al. RNA Sequencing Uncovers the Key microRNAs Potentially Contributing to Sudden Sensorineural Hearing Loss. Medicine 96. (2017).
Nunez, D. A., Wijesinghe, P., Nabi, S., Yeh, D. & Garnis, C. MicroRNAs in sudden hearing loss. Laryngoscope 130, E416–E422 (2020).
Ha, S. M. et al. Circulating MicroRNAs as potentially new diagnostic biomarkers of idiopathic sudden sensorineural hearing loss. Acta Otolaryngol. 140, 1013–1020 (2020).
Nunez, D. A. & Guo, R. C. Acquired sensorineural hearing Loss, oxidative Stress, and MicroRNAs. Neural Regeneration Res. 20, 2513–2519 (2025).
Han, Y. et al. PI3K/AKT pathway: A potential therapeutic target in cerebral Ischemia-Reperfusion injury. European J. Pharmacology. 998, 177505 (2025).
Nunez, D. A., Abgoon, R., Wijesinghe, P. & Garnis, C. Serum and plasma MiRNA expression levels in sudden sensorineural hearing loss. Int. J. Mol. Sci. 26, 1245 (2025).
Rawlings-Goss, R. A., Campbell, M. C. & Tishkoff, S. A. Global Population-Specific variation in MiRNA associated with cancer risk and clinical biomarkers. BMC Med. Genom. 7, 53. https://doi.org/10.1186/1755-8794-7-53 (2014).
Abgoon, R., Wijesinghe, P., Garnis, C. & Nunez, D. A. The expression levels of MicroRNAs differentially expressed in sudden sensorineural hearing loss patients’ serum are unchanged for up to 12 months after hearing loss onset. Int. J. Mol. Sci. 24, 7307 (2023).
Shew, M., Wichova, H., St. Peter, M., Warnecke, A. & Staecker, H. Distinct MicroRNA profiles in the Perilymph and serum of patients with Menière’s disease. Front. Neurol. 12, 646928 (2021).
Ohlemiller, K. K., Rosen, A. D. & Gagnon, P. M. A major effect QTL on chromosome 18 for noise injury to the mouse cochlear lateral wall. Hear. Res. 260, 47–53 (2010).
Lee, Y. R. et al. Amelioration of Auditory Response by DA9801 in Diabetic Mouse. Evidence-Based Complementary and Alternative Medicine 2015, 230747. (2015).
Butskiy, O., Ng, D., Hodgson, M. & Nunez, D. A. Rinne test: does the tuning fork position affect the sound amplitude at the ear? J. Otolaryngology-Head Neck Surg. 45, 1–8 (2016).
Association, A. S. -L.-H.; others Guidelines for Manual Pure-Tone Threshold Audiometry. (2005).
Hearing, C. on; Equilibrium Committee on Hearing and Equilibrium Guidelines for the Diagnosis and Evaluation of Therapy in Meniere’s Disease. Otolaryngology–Head and Neck Surgery 113, 181–185. (1995).
Committee on Hearing and Equilibrium Guidelines for the Evaluation of Results of Treatment of Conductive Hearing Loss. AmericanAcademy of Otolaryngology-Head and neck surgery Ffoundation, inc. Otolaryngol. Head Neck Surg. 113, 186–187. https://doi.org/10.1016/S0194-5998(95)70103-6 (1995).
Joshua, T. G., Ayub, A., Wijesinghe, P. & Nunez, D. A. Hyperbaric oxygen therapy for patients with sudden sensorineural hearing loss: A systematic review and Meta-Analysis. JAMA Otolaryngology–Head & Neck Surgery. 148, 5–11 (2022).
G. Clark, J. Uses and abuses of hearing loss classification. Asha 23, 493–500 (1981).
Schmittgen, T. D. & Livak, K. J. Analyzing Real-Time PCR data by the comparative CT method. Nat. Protoc. 3, 1101–1108. https://doi.org/10.1038/nprot.2008.73 (2008).
Wang, X. et al. Evaluation of the performance of serum MiRNAs as normalizers in MicroRNA studies focused on cardiovascular disease. J. Thorac. Disease. 10, 2599 (2018).
Xie, F., Xiao, P., Chen, D., Xu, L. & Zhang, B. MiRDeepFinder: A MiRNA analysis tool for deep sequencing of plant small RNAs. Plant Mol. Biol. 80, 75–84 (2012).
Acknowledgements
We kindly thank the clinic administrative staff of the Vancouver General Hospital Division of Otolaryngology–Head and Neck Surgery and the Audiology Services. We extend our gratitude to Dr. Sedigheh Shams for her valuable input on the statistical analysis. Additionally, we thank the Rotary Club of Vancouver Hearing Foundation, the Pacific Otolaryngology Foundation, and individual donors for their financial support.
Funding
This research was funded by Vancouver Coastal Health Research Institute, grant number RD47, University of British Columbia grant number GR026281, and Rotary Club of Vancouver Hearing Research Foundation grant number 008074.
Author information
Authors and Affiliations
Contributions
DAN generated the idea; RA performed the experiments and analyzed the data; RA and DAN drafted the paper; RA, CN and DAN recruited participants; RA, RG, CN and DAN revised the paper; DAN examined participants, secured funding and supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Institutional review board statement
The study was conducted in accordance with the Declaration of Vancouver, and approved by Ethics Committee of University of British Columbia (protocol code H15-03024 initial date of approval 2/17/2016).
Informed consent statement
Informed consent was obtained from all subjects involved in the study.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Abgoon, R., Guo, R., Needham, C. et al. Differential expression of miR-23a-3p/-210-3p and /-18b-5p in sudden sensorineural hearing loss patients: A North American cohort study. Sci Rep 16, 2465 (2026). https://doi.org/10.1038/s41598-025-32213-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-32213-5