Abstract
The Philippines is a renowned marine biodiversity hotspot, home to several sea cucumber species with unusual biological traits. Among these, Stichopus cf. horrens is notable for its ability to undergo rapid body wall liquefaction when stressed, coupled with remarkable regenerative abilities. In contrast, Holothuria scabra has one of the most robust body walls in sea cucumbers and thrives in many regimes in the tropics. Despite their intriguing traits, the neurobiology and chemical diversity of these species remain underexplored. Neuropeptides are important components of an animal’s neurobiological toolkit that underlie various physiological and behavioral processes. Thus, the discovery of neuropeptides is a crucial step for understanding the molecular underpinnings of unique traits in sea cucumbers. Leveraging the throughput and sensitivity of tandem mass spectrometry, we obtained an unbiased view of the endogenous peptidomes of radial nerve cord tissues of non-model sea cucumber species, H. scabra and S. cf. horrens. In this work, we sequenced 60 mature peptides from S. cf. horrens that were derived from 22 precursor proteins, and 43 peptides originating from 25 precursor proteins in H. scabra nervous tissues. A total of seven previously unannotated and uncharacterized neuropeptide precursors were identified, thereby expanding the known animal neuropeptide repertoire. Furthermore, we discovered consistent structural features in mature neuropeptides based on the type of post-translational modifications while pushing forward potentially novel proteolytic processing sites during peptide maturation based on the enriched flanking amino acid residues. Collectively, our results provide preliminary data that expand our understanding of echinoderm neurobiology through neuropeptide discovery, potentially paving the way for innovative solutions to address the global demand for echinoderms.
Similar content being viewed by others
Introduction
Neuropeptides represent one of the most structurally and functionally diverse intercellular signaling molecules thought to be the earliest form of nervous control in animals1. Structurally, they range in size from three (3) to 91 amino acid residues, with distinct folds between families, and contain diverse post-translational modifications (PTMs). Single or multiple copies of mature neuropeptides originate from larger inactive precursor proteins. Neuropeptide precursors contain N-terminal signal peptides that target them to the regulated secretory pathway, where dedicated proteases cleave them at canonical endo-and exoprotease cleavage sites to produce mature neuropeptides. Subsequently, the peptides derived from precursor proteins also undergo further post-translational modifications until release through exocytosis, either via volume or synaptic transmission2,3,4,5. Functionally, neuropeptides can serve as neurotransmitters (short-range and fast), neuromodulators (short-range and slow), and neurohormones (systemic).
Sea cucumbers are deuterostomian marine invertebrates with pentaradial nervous systems that rely on neuropeptides to orchestrate numerous behavioral and physiological processes6. These neuropeptides are expressed in various anatomical structures, including tube feet, radial nerve cords, and visceral organs, suggesting that they can act both locally and systemically via coelomic fluid to modulate specific signaling circuits1,7,8,9,10,11,12. Regardless of their sites of synthesis, neuropeptides are involved in regulating many processes, such as osmoregulation, aestivation, tissue regeneration, and many developmental and reproductive processes5,6,7,8,9,10,11,12,13,14,15,16,17. Thus, neuropeptides have been used in Apostichopus japonicus, Holothuria glaberrima, and Holothuria leucospilota, to manipulate gamete and reproductive maturation, alter body wall stiffness, and regulate muscular activity related to feeding and movement to enhance sea cucumber production and fisheries13,14,15,16,17,18,19. This highlights the broad range of bioactivities associated with neuropeptides and their important role in echinoderm biology. However, the neuropeptide repertoire of other commercially important tropical sea cucumbers remains unexplored.
Holothuria scabra and Stichopus cf. horrens are high-value sea cucumber species that widely occur in the coastal areas of the Philippines and other tropical regimes in the Pacific20,21,22. H. scabra is most known for its broad distribution in the Indo-Pacific region and as a key species in sea cucumber aquaculture and fisheries. S.cf. horrens is less well-known and has recently been shown to belong to a cryptic species complex, thus the provisional qualifier cf. in the species assignment23,24.Previous studies have generated transcriptome resources for these species that investigated inter-individual growth rate variability and response to changing ocean conditions related to global warming25,26,27. To date, only H. scabra has been investigated for its neuropeptides which led to the discovery of 39 neuropeptide precursors being expressed in their nervous tissues. Some of these neuropeptides are homologs of myoactive neuropeptides that were previously shown to localize in the body wall and intestines10. However, due to the inherent variations in the generation of mature neuropeptides, transcriptomic and genomic approaches have been limited to the identification and comparative analysis of precursor molecules, with little to no structural information on mature peptides. Therefore, it is crucial to characterize the mature neuropeptides of echinoderm tissues in combination with publicly available genomic data in generating a more complete species neuropeptidome.
Mass spectrometry-based neuropeptide profiling has successfully described diverse classes of neuropeptides across a range of echinoderm species17,28,29,30,31,32. Mass spectrometry (MS) is a powerful analytical technique that enables high-throughput protein identification and characterization of post-translational modifications. In this study, we employed tandem mass spectrometry to provide a comprehensive characterization of the endogenous peptidomes of S. cf. horrens and H. scabra. Through our work, we significantly expanded the known sea cucumber neuropeptidome through the discovery of seven (7) novel putative neuropeptide precursors and sequenced 103 peptides, majority of which are novel peptides and peptide variants. By integrating mass spectrometry-based peptidomics and extensive bioinformatic analysis, we advance our understanding of the sea cucumber neuropeptidome through the discovery of potentially novel cleavage sites, peptide modifications, and sequence diversity of peptides between sea cucumber species. Ultimately, the peptides discovered in our study may offer important clues to some of their unique behaviors while providing new avenues for enhancing aquaculture practices using neuropeptides.
Results
Discovery and structural elucidation of mature peptides from known neuropeptide precursors in the sea cucumber radial nerve cords
Identification of neuropeptides in the nervous tissues of sea cucumbers is key to understanding the neurobiological basis of their many distinct behaviors. Thus, we aimed to obtain a comprehensive view of the neuropeptidomes of two Philippine sea cucumbers, Stichopus cf. horrens, and Holothuria scabra, through mass spectrometry-based neuropeptidomics. Our neuropeptide profiling detected 38 mature peptides derived from 10 known neuropeptide precursors (Figs. 1 and 2). We sequenced peptides derived from precursors of L-type SALMFamides, GN19, kisspeptin-type 2, a calcitonin peptide, cholecystokinin-type peptides, TRH-type peptides, and pigment-dispersing factor-like peptides. Homologs of putative neuropeptide precursors, Neuropeptide 37 (HleNp37) and Neuropeptide 40 (HleNp40), first discovered in H. leucospilota were also detected by mass spectrometry, with HleNp40 peptides being found in the two study species. From a total of 38 peptides, 20 are hitherto new and unreported peptides processed from known neuropeptide precursors (full list found in Tables 1 and 2). More than two (2) mature peptides were sequenced from each precursor, with pedal-peptide-type precursor producing the greatest number of peptides and peptide variants (12 in S. cf. horrens and two in H. scabra tissues) that arise from different regions in the precursor protein. Despite the presence of 18 predicted neuropeptides with the motif Q—G-NH3 in the H. scabra TRH-type precursor, we did not detect any of these and only detected the much larger pyro-glutamylated N-terminal peptide, QLPAGWAFWEGKRLQGDALNDALRPAVVYGGYH. Our results show that we can robustly detect multiple peptides and their variants from different regions of the precursor proteins.
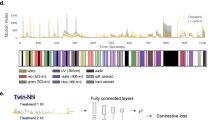
Sequences of neuropeptide precursors in Holothuria scabra that have been previously identified in sea cucumbers. Signal peptides predicted by SignalP5.0 are shown in blue while predicted neuropeptides are shown in red. Sequenced peptides are highlighted in yellow. C-terminal Glycine (G) used as substrate for C-terminal amidation is shown in dark green. Putative cleavage sites are shown in light green. aNeuropeptide precursors identified by Suwansa-ard et al., 2018, Peptides; bPredicted gene sequences from H. scabra genome assembly (GCA_026123075.1); cTranscript sequences generated by Ordoñez et al., 2021, Comp Biochem Physiol Part D. The annotations of known neuropeptide precursors were presented as they are reported in literature or appear in NCBI Databases for consistency.
Given the highly context-dependent manner of neuropeptide biosynthesis, transcriptomic datasets may not accurately reflect the presence of mature peptides in vivo. In fact, we observe that only a subset of reported precursor transcripts give rise to mature peptides (Supplemental Fig. 1). In S. cf. horrens, we detected 27 neuropeptide precursors, of which eight (8) were supported by peptide-level evidence. Conversely, in a separate analysis of H. scabra transcriptome, 39 neuropeptide precursors were identified, only eight (8) of which were found to produce mature peptides as detected in our peptidomics analysis. A peptide derived from the kisspeptin-type 2 precursor protein was detected in S. cf. horrens, but not in H. scabra. TRH-type and Calcitonin-type peptides were only detected in H. scabra. In both sea cucumber species, we detect the production of mature peptides from six (6) neuropeptide families (Supplemental Fig. 2). While several of these neuropeptides were predicted to be expressed in both sea cucumber species, as reflected in their transcriptomes, it is possible that we did not detect them because they were not expressed at detectable levels when we harvested and processed the samples.
Sequences of neuropeptide precursors in Stichopus cf. horrens that have been previously identified in sea cucumbers. Signal peptides predicted by SignalP5.0 are shown in blue while predicted neuropeptides are shown in red. Sequenced peptides are highlighted in yellow. C-terminal Glycine (G) used as substrate for C-terminal amidation is shown in dark green. Putative cleavage sites are shown in light green. aTSA sequence records of Stichopus horrens (ID 512756; PRJEB 29236); bPredicted gene sequences from S. horrens genome assembly through blast-all-v-all with S. monotuberculatus genome (SRX8106250); cPredicted genes from the S. horrens genome assembly (SRX8106250). The putative annotations of known neuropeptide precursors were presented as they are reported in literature or appear in NCBI Databases for consistency.
Our MS-based identification of peptides from both species provided structural information such as the average and range of peptide lengths, amino acid sequence, PTMs, and peptide cleavage site. Our MS-based identification of peptides from both species provided structural information such as the average and range of peptide lengths, amino acid sequence, PTMs, and peptide cleavage sites. The sequenced peptides ranged from nine (9) to 35 amino acid residues, which, on average, is longer than the most studied peptides like sulfated cholecystokinin-type peptides, pedal peptide/orcokinin-type, SALMFamides which vary in length, although ranging from eight (8) to 15 amino acids long (Fig. 3A)9,11,53. Our results have also enabled us to examine the possible mechanisms of neuropeptide processing. Based on the enriched residues at the N- and C-terminal ends of the mature peptides, we confirmed the presence of dibasic and monobasic residues which serve as substrates for prohormone convertases and other alkaline exo- and endopeptidases during mature peptide maturation34,35,36. Consistent with recent reports of acidic residues at the terminal ends of the mature peptides, we detected a preponderance of D and E residues that flank the N- and C- terminus of the mature peptide (Fig. 3B)37. C-terminal amidation was observed in 63% of all sequenced peptides derived from known neuropeptide precursors, demonstrating that C-terminal amidation is an evolutionarily conserved feature in neuropeptides among Bilaterians35,37,38,47 (Fig. 3C). All three enzymes necessary for C-terminal amidation were detected in the genomic dataset we used, including a multidomain PAM-A/B enzyme in the genome and the transcriptomes of study organisms. When comparing the masses and lengths of peptides sequenced from the two species, we found that their masses and lengths significantly differed indicating the presence of diverse peptide structures in their radial nerve cords despite the overlap in neuropeptide precursor families. On average, peptides from S. cf. horrens were larger and longer (One sample Wilcoxon test, p < 0.001). Post-translational modifications detected in the peptides in the two species also differed, which may be due to variations in the maturation process or extraction time, in the case of labile PTMs. Many C-terminally amidated peptides are observed in the peptidomes of both species while pyro-glutamylation (denoted as lowercase p) is only observed in H. scabra, found in the N-terminal Q of the pQLPAGWAFWEGKRLQGDALNDALRPAVVYGGYH peptide derived from a TRH-type precursor. Peptides that have these modifications are also referred to as capped peptides that may act over long distances primarily due to their recalcitrance to proteolytic degradation38. Sulfation of tyrosine in the cholecystokinin-like peptide, DYNDLGFFFG-NH3, was detected in the RNC of S. cf. horrens, but not in the cholecystokinin-like peptides of H. scabra. Our examination of their RNC endogenous peptidomes supports the adoption of untargeted peptidomics workflow that enables the detection of a diverse set of peptide structures that can be generated within and between species.
Mature peptides from the two species exhibit extensive structural variations. (A) Length (number of amino acid residues) distribution of sequenced peptides originating from neuropeptide precursors (B) Flanking amino acid residues at the N-term and C-term of the mature peptides. (C) Post translational modifications of sequenced peptides.
Discovery of novel neuropeptide precursors
Our study has successfully expanded the neuropeptidome of Echinodermata by discovering mature peptides from seven (7) proteins annotated in the genome as “hypothetical” proteins (Figs. 4 and 5). Protein-level evidence in the form of peptide fragmentation spectra points to the fact that these are not hypothetical proteins anymore, but are, in fact, expressed genes. In addition, by matching the protein sequence against existing genome, transcriptome and proteome databases, we find that these hypothetical proteins have been previously detected in other echinoderms, albeit have remained functionally unassigned.
Leveraging machine learning-based functional annotation, we searched for enriched keywords related to neuropeptide function, such as positive and negative regulation, signaling, cell-cell communication, and receptor binding, among many others, to classify them as putative neuropeptides. We also searched for typical cleavage sites, PTMs, and signal peptide sequences in the precursor as additional evidence for their assignment as precursor neuropeptides. Based on these criteria, we report and confirm the identity of these seven (7) hypotheticals proteins as putatively novel neuropeptide precursors. On average, the novel neuropeptide precursors contained at least three (3) peptide variants structurally similar to peptides derived from known neuropeptide classes, in terms of flanking amino acid residues and their PTMs. SHNP3, one of the novel precursors that gave rise to mature peptides in vivo, matched to precursors from other sea cucumbers but not all echinoderm groups (Supplemental Fig. 3A). It is predicted to play a role in several biological and signaling activities such as receptor binding, chemotaxis, immune response, and cytotoxic activity. Compiling all the functional GO annotations for all putatively novel neuropeptides, we detect enrichment of keywords like positive and negative regulation, receptor binding, and cell-cell signaling comparable with the functional GO annotations of eight (8) known non-neuropeptide precursors (p < 0.00001). While the SHNP3 peptide tends to exhibit a higher number of acidic residues atypical in neuropeptides, we find from our dataset that peptides from known neuropeptide precursors have a broad range of isoelectric points, as well. Additionally, when inspecting for sites of cleavage by signal peptidases, we also found that there can be disagreements between the cleavage site assignment between the two earlier versions of SignalP version 6.043. But given the high confidence of our peptide sequencing results, we accurately delineate the signal peptide and other proteolytic cleavage sites regardless of bioinformatic predictions (Supplemental Fig. 3B). While all this evidence points to the possibility of novel neuropeptide precursors, functional studies need to be performed to validate their activities common in neuropeptides.
Discovery of other endogenous peptides in the nervous tissues of sea cucumbers
While the focus of our study is on the identification of neuropeptides, we detected peptide sequences derived from non-neuropeptide precursors (see Tables 1 and 2). Peptides that arise from non-hormone or non-neuropeptide precursors are typically categorized as cryptides, some of which have recently been shown to be bioactive39. Notable among these peptides is holokinin, which is derived from a collagen precursor, shown to influence body wall viscosity in many species of the genus Holothuria7. Endogenous peptides like holokinin originating from non-neuropeptide precursors further support the important roles of cryptides in biological systems. The other peptides detected in our study are peptides derived from proteins involved in muscle movement such as twitchin, actin, myosin and myophilin, many of which also exhibit variations on their status of modification. Specifically, acetylation of the N-terminal methionine was observed in these proteins.
Discussion
Recent discoveries in neurobiology have emphasized the important roles of non-classical neurotransmitters and long-range neuronal signaling molecules like neuropeptides in the physiology and behavior of many animals6. While genomic approaches have facilitated the discovery of neuropeptide precursors in animals through sequence-based predictions and homology searching, they often fall short in unraveling the diversity of neuropeptide structures in vivo. In contrast, mass spectrometry-based peptidomics is capable of providing structural information on the mature peptides, agnostic of key diagnostic features required for neuropeptide precursor prediction, thus enabling the discovery of previously unreported peptides and neuropeptide precursors2,35,38. This study leverages the throughput and sensitivity of mass spectrometry to explore the endogenous peptidomes of commercially important sea cucumbers, the S. cf. horrens and H. scabra, to discover neuropeptides and other endogenous peptides that may underpin their exceptional regenerative, wound-healing capabilities and other unique biological traits. Through this approach, we identified and sequenced 103 peptides derived from known and novel putative neuropeptide precursors, and other proteins found in the sea cucumber radial nerve cords.
In this study, we detected peptides from known neuropeptide precursors that exhibited variations in their structures based on the terminal amino acids and their PTMs. This demonstrates the analytical power of peptidomics that does not heavily rely on diagnostic features for identification. By comparing enriched flanking amino acid residues, we also support the prevailing hypothesis for neuropeptide processing which involves a battery of endo-and exo-peptidases that act on basic amino acids36. Historically, neuropeptide maturation is primarily thought to only involve prohormone convertases and a few alkaline proteases34. Our findings, on the other hand, revealed the presence of acidic residues that flank the mature peptides suggesting the involvement of as-yet-uncharacterized proteases in neuropeptide maturation. This was similarly observed in peptides sequenced from cnidarians, sponges and ctenophores35,37. Moreover, our analysis of post-translational modifications on the peptides highlighted the crucial role of peptide capping through C-terminal amidation for peptide stability35,38. Aside from increasing the half-lives of peptides in vivo, C-terminal amidation has also been demonstrated to influence peptide potency35. In our study, C-terminal amidation was the most common PTM consistent with peptidomic reports across diverse animal taxa.
Our peptidome profiling also revealed species-specific variations in peptide masses and lengths, demonstrating the unique ways by which different species process neuropeptide precursors into mature peptides. Despite arising from the same class of neuropeptide precursors, a species may process it differently, thereby resulting in the production of unique structural variants. Differences in response to stimuli may also have driven this divergence in peptide profiles despite our best efforts at standardizing the conditions during analysis. Despite these variations in their peptidomes, there was a notable overlap in the neuropeptide precursor families from which most peptides were derived. This suggests that some of these peptides may be constitutively expressed or induced in the conditions they were subjected to during our experiments. CCK-type peptides from the cholecystokinin-type precursor 1 were detected in both species, with S. cf. horrens producing both sulfated and non-sulfated forms (Supplemental Fig. 4).In mammalian systems, the sulfated form has been shown to be more potent and tends to exhibit a distinct bioactivity profile than the non-sulfated form49,50. This is similarly observed in the sea star, Asterias rubens, wherein the sulfated form is five to six orders of magnitude more potent than the non-sulfated form in influencing feeding behavior and muscular contraction11.GN19 neuropeptides are echinoderm-specific neuropeptides that were first observed to influence contraction and relaxation of sea cucumber intestines7. GN19 peptides are characterized by being 19 residues long and containing an N-terminal G residue and a C-terminal N residue7,18. In our peptidomic dataset, the N-terminal residues of GN19 peptides from S. cf. horrens and H. scabra were D and N, and were 24 and 22 amino acids long, respectively (Supplemental Fig. 7). All sequenced pedal peptide-type peptides have an F residue at the N-terminus and a C-terminal amidation which is consistent with other MS-based structural characterization in other sea cucumber species14,17. Pedal peptides in sea cucumbers have been shown to influence muscular activity by encouraging movement while reducing velocity51. We provide the first peptide-level evidence to a C-terminally amidated peptide from the pigment-dispersing factor-like precursor also detected in the transcriptomes of other sea cucumber species like H. leucospilota, A. japonicus, and H. scabra10,14,18. To date, there is nothing that is known about the role of PDF-like peptides in echinoderms, but in flies, they have been shown to tightly regulate circadian rhythmicity45,46. At least two peptides originating from the L-type SALMFamide precursors were detected in our mass spectrometric analysis of RNC extracts, all of which bearing a C-terminal amidation. In the sea star, A. rubens, and sea cucumber, A. japonicus, L-type SALMFamides have muscle relaxant activities52. Neuropeptide 40 or HleNp40 is a sea cucumber-specific neuropeptide precursor that were first discovered in H. leucospilota14. Interestingly, we sequenced a much larger neuropeptide 33–35 AA long N-terminal peptide in both species compared to the seven AA long N-terminal peptide from the RNCs of H. leucospilota. It is worth noting that the same 35 AA long peptide is contained in the HleNp40 precursor but is interrupted by dibasic residues at the 8th -9th position from the N-terminal cut site. Peptides derived from Neuropeptide 37 and kisspeptin-type precursor 2 were detected in S. cf. horrens, but not in H. scabra. Peptides from these precursors that were detected in this study remains to be functionally characterized, although several C-terminally amidated kisspeptin-type peptides have already been shown to modulate feeding behavior in Apostichopus japonicus8. TRH-like and Calcitonin-like mature peptides were exclusively detected in H. scabra. TRH-type peptides are known to modulate developmental processes in in H. scabra, where it has been demonstrated to induce oocyte maturation11,54. Calcitonin-like peptides have been demonstrated to be potent muscle relaxants in the sea star species, Asterias rubens and Patiria pectinifera44. This was similarly observed in the sea cucumber, A. japonicus, whereby peptides have been shown to have dose-dependent effect on intestine and body wall relaxation, and influence growth and feeding48.
Transcriptomic mining revealed more neuropeptide precursors than our peptidomic analysis, which is no surprise considering that many of these peptides may not undergo full maturation and degrade when not required by the animal. Alternatively, the peptides may exhibit very localized expression that may then evade detection and high-quality sequencing through mass spectrometry8,29,35,42,44.Remarkably, our peptidomic profiling discovered more unreported neuropeptide precursors than transcriptomic analysis, highlighting the advantage of mass spectrometry-based peptidomics for the discovery of novel precursors independent of the presence of homologs in other taxa. Additionally, we uncovered six (6) constitutively expressed neuropeptide families in the two sea cucumber species, evidenced by their expression in both peptidomic and transcriptomic datasets. These constitutively expressed peptides may be important for maintaining homeostasis at unperturbed/ non-stressful conditions. Some of these neuropeptides may be leveraged to expedite some processes in their aquaculture. Peptides that modulate reproductive processes, metabolism, sociality, and other important ecologically relevant behaviors may be leveraged for more efficient aquaculture and their conservation in the wild. Our dataset contained spectra that did not result in significant peptide sequence matches. This is primarily due to the lack of a dedicated database from genomic data of our study organism belonging to the Philippine population for peptide-to-spectrum matching that is highly sensitive to the type of PTMs and variations in the peptide sequence. These spectra can be subjected to automated de novo analysis, which can then be matched against a larger database. Despite pooling samples from at least 12 animals, there were still low-abundance peptides whose fragmentation spectra were insufficient to be significantly matched to any peptide sequence in the transcriptome. In these cases, further sample enrichment may yield a more comprehensive profile.
Taken together, our results present the most comprehensive peptidome profile of these species to date. Peptidomics, coupled with extensive bioinformatic analysis and mining of publicly available sequences from related species, is a powerful tool for describing neuropeptidomes in non-model invertebrate systems.
Methods
Animal and tissue collection
Adult sea cucumbers (Holothuria scabra and Stichopus cf. horrens) from aquaculture grow out and wild stocks were obtained from Bolinao Marine Laboratory Hatchery and the coasts of Bolinao and Anda, Pangasinan. Live animals were transported in filtered seawater to The Marine Science Institute at the University of the Philippines Diliman for dissection. Before tissue collection, the animals were acclimatized in a 10 L tank containing filtered seawater with sufficient aeration for approximately one (1) hour until reduced activity. To induce anesthesia, the animals were injected with 50mM MgCl2 solution in isotonic saline (5mM) until they showed reduced response to tactile stimuli on their tube feet. A perfusion solution containing 50 mM EDTA and 2 mM PMSF (phenylmethylsulphonyl fluoride, ThermoScientific) in isotonic saline was injected into the coelomic cavity of the echinoderms through the aboral end using a syringe at a flow rate of approximately 0.5 ml/s. The sea cucumbers were longitudinally cut from the ventral side, avoiding damage to the five longitudinal muscle bands (LMBs). The LMBs were separated from the body wall, and the radial nerve cords (RNCs) were carefully isolated from the LMBs and radial canals using a scalpel. The RNCs were either flash-frozen in liquid nitrogen or heated before resuspension in lysis buffer composed of 6 M Urea, 5mM EDTA, 5mM MgCl2, and 1mM PMSF before storage at -80 °C until further analysis. All experiments were done in three biological replicates.
Sequence of putatively novel neuropeptide precursor in Holothuria scabra. Signal peptides predicted by SignalP5.0 are shown in blue while predicted neuropeptides are shown in red. Sequenced peptides are highlighted in yellow. C-terminal Glycine (G) used as substrate for C-terminal amidation is shown in dark green. Putative cleavage sites are shown in light green. aQuery sequences generated from the work of Suwansa-ard et al., 2018, Peptides; bPredicted gene sequences from H. scabra genome assembly (GCA_026123075.1); cTranscript sequences generated by Ordoñez et al., 2021, Comp Biochem Physiol Part D.
Sequences of putatively novel neuropeptide precursors in Stichopus cf. horrens. Signal peptides predicted by SignalP5.0 are shown in blue while predicted neuropeptides are shown in red. Sequenced peptides are highlighted in yellow. C-terminal Glycine (G) used as substrate for C-terminal amidation is shown in dark green. Putative cleavage sites are shown in light green. aTSA sequence records of Stichopus horrens (ID 512756; PRJEB 29236); bPredicted gene sequences from S. horrens genome assembly through blast-all-v-all with S. monotuberculatus genome (SRX8106250); c Predicted genes from the S. horrens genome assembly (SRX8106250).
Tissue lysis and protein extraction
To identify the peptides found in the radial nerve cords, tissue lysis, and protein extraction methods based on acidified methanol precipitation and ultrafiltration were done, as described elsewhere 14,29,33. The heat-treated or flash-frozen RNCs from 12 animals were pooled and weighed. These were homogenized and resuspended in acidified methanol (60:40:1, Methanol, LC/MS Grade H2O, Formic acid). The tissues were further homogenized on ice using a QSonica Ultrasonicator (QSonica, United States of America) at five (5) cycles of 20s:30s sonication: rest at 50% amplitude. Lysates were centrifuged at 16,000 x g for 20 min at 4 °C to separate the protein pellet from the supernatant.
The supernatant was collected and then dried using a SpeedVac concentrator (BIOBASE, China) at 50 °C for three (3) hours or until little to no visible solvent remained. The drying process was monitored regularly to avoid over-drying of samples. The samples were resuspended in 100 µl of 0.1% FA in water by vortexing for three (3) minutes. The samples were pooled and mixed by pipetting up and down several times to ensure homogeneity and maximum yield. Conversely, ultrafiltration was performed using Amicon Ultra-0.5 mL centrifugal filters (Merck Millipore, Germany) with a 3 kDa molecular weight cutoff to separate endogenous peptides < 3 kDa or more than > 3 kDa MW. The filtrate (< 3 kDa) and retentate (> 3 kDa) were stored at -80 °C until mass spectrometry analysis. Total peptide concentration was determined using Bicinchoninic Acid (BCA) Assay and NanoDrop (ThermoScientific, United States) using a standard 1 mg/ml BSA (Bovine Serum Albumin, Sigma-Aldrich, United States of America). Before LC-MS analysis, peptides were desalted using C18 ZipTip (Merck Millipore, Germany) to remove salts that may interfere with data collection. A 1–2 µg/µL sample was transferred into an LC-MS 96-well plate.
LC/MS/MS analysis
Endogenous peptide fractions from the RNCs were analyzed using a Waters Acquity UHPLC System coupled with a Xevo-G2-XS-QTOF mass spectrometer. A total of 1.5µg of peptide per sample were loaded using an autosampler into an analytical column (Phenomenex Kinetex, 1.7 μm C18 100 A 50 × 2.1 mm). Separation of peptides was performed with a segmented 12-minute stepwise run of 5- 40%, 45-95%B (mobile phase A, 0.1%FA in LC-MS grade water; mobile phase B, 0.1% FA in Acetonitrile) with a flow rate of 300 uL/min. The instrument was set to positive mode, and peptides from 400 to 2000 Da were scanned during MS1 and 50 to 2000 Da during MS2. MS/MS data was obtained through fast data-dependent acquisition (DDA) for ions with total ion chromatogram intensities above 5e^4. Each sample was injected twice to obtain technical replicates. All experiments were also done in three biological replicates. Peptides that appeared consistently across runs were considered true matches.
MS data analysis and protein identification
The generated raw MS/MS data were visualized and processed (peak processing and deconvolution) using MassLynx (Waters Corp., United States of America), MASCOT Distiller (Matrix Science Ltd.), and MZmine3 (http://mzmine.github.io/). Peptide-to-spectrum matching was done using MASCOT Daemon and Distiller Search engine against an neuropeptide precursor sequence database generated from genome and transcriptome mining concatenated with a reverse decoy database. We also downloaded sequences of neuropeptide precursors from UniProt using the following search keywords Taxa: Animalia, Keywords: Neuropeptides, Neuropeptide Precursors.
Database search parameters implemented were as follows: 10 ppm for precursor tolerance, 0.3 Da for fragment tolerance, no enzyme used, variable modifications such as methionine oxidation, cyclization of N-terminal glutamine to pyroglutamic acid, deamidation of asparagine and glutamine, acetylation at peptide N-termini, sulfation of STY and amidation at peptide C-termini were searched. Briefly, the peaks in the fragmentation spectra were manually annotated and corroborated with the MASCOT assignment. We also used MSProspector and an in-house code to predict fragmentation based on the provided peptide sequence.
Identification of inactive precursor neuropeptides from transcriptome and genome data
Publicly available high-quality transcriptome and whole genome assemblies of S. cf. horrens (SRX8106250; PRJEB29236) and H. scabra (GIRH01000000; GCA_026123075.1) were downloaded from NCBI Transcriptome Shotgun Assembly database (TSA) and the Sequence Read Archive (SRA) database using the SRAToolkit. Gene calling was performed using Augustus trained with Apostichopus japonicus and Stichopus monotuberculatus genomic data. Previously identified neuropeptide precursors from other echinoderms were used as tBLASTn and BLASTp queries to identify precursors in our in-house database of predicted ORFs generated using the software TransDecoder (https://github.com/TransDecoder/TransDecoder) for ORF prediction. Sequence hits were translated using Expasy (https://www.expasy.org/translate) before domain analysis using InterPro, HMMER, and SignalP versions 4.0, 5.0, and 6.0.
Protein sequences that contained (1) signal peptides, (2) mono/dibasic cleavage sites, and (3) in some cases, C-terminal glycinewere considered putative neuropeptide precursors. No nucleotide sequencing was done in this study.
Bioinformatics and statistical analysis
Mining of neuropeptide precursors was performed using tBlastN, tBlastX, and BlastP (https://blast.ncbi.nlm.nih.gov/Blast.cgi) using default settings for searching for homologs from closely related species, but were adjusted to an e-value of 10 to detect orthologs in the Animalia database. Sequence matches were then downloaded and submitted to CLANS (https://toolkit.tuebingen.mpg.de/tools/clans) for cluster analysis of sequences with an e-value threshold of 1. Clusters with p-value < 1 × 10^-10 were considered as true clusters. We then utilized the find clusters function of CLANS java applet to obtain further support for network clustering. Here, we used the Linkage clustering algorithm with two (2) as the minimum number of links and jackknifing with 1000 replicates. Query sequences that formed clusters with published precursor neuropeptide sequences were further verified using Blast and domain analysis. Protein alignments were done in Jalview 2.11.1 using the MUSCLE algorithm. Sequences were manually trimmed before alignment construction. Web-based machine learning-based protein function prediction tools such as PROST and ProteinFer were used using default parameters40,41. We used the NeuroPred webserver to predict cleavage sites in neuropeptide precursors (http://stagbeetle.animal.uiuc.edu/cgi-bin/neuropred.py).
Sequence analysis of the peptides and their precursors was done using a homemade Python script that integrated the functionalities of Pyteomics (https://github.com/levitsky/pyteomics) for predicting fragmentation, EXPASY for determining peptide physicochemical characteristics, and text-mining. The .csv outputs were manually inspected before statistical analysis. All statistical analyses were performed using GraphPad Prism 9 (Prism, United States of America).
Data availability
The datasets used and analyzed in this study are available as partial submissions in the ProteomeXchange with identifiers PXD057178/MSV000096207 and PXD057188/MSV000096212.
References
Elphick, M. R., Mirabeau, O. & Larhammar, D. Evolution of neuropeptide signalling systems. J. Exp. Biol. 221, jeb151092 (2018).
Fälth, M. et al. Neuropeptidomics strategies for specific and sensitive identification of endogenous peptides. Mol. Cell. Proteom. 6, 1188–1197 (2007).
Grimmelikhuijzen, C. J. P. & Hauser, F. Mini review: the evolution of neuropeptide signaling. Regul. Pept. 177, S6–S9 (2012).
Koch, T. L. & Grimmelikhuijzen, C. J. P. Global neuropeptide annotations from the genomes and transcriptomes of Cubozoa, Scyphozoa, Staurozoa (Cnidaria: Medusozoa), and Octocorallia (Cnidaria: Anthozoa). Front. Endocrinol. 10, (2019).
Wood, N. J. et al. Neuropeptidergic Systems in Pluteus Larvae of the Sea Urchin Strongylocentrotus purpuratus: neurochemical complexity in a simple nervous system. Front. Endocrinol. 9, 628 (2018).
Semmens, D. C. & Elphick, M. R. The evolution of neuropeptide signalling: insights from echinoderms. Brief. Funct. Genomics. 16, 288–298 (2017).
Elphick, M. R. The protein precursors of peptides that affect the mechanics of connective tissue and/or muscle in the Echinoderm Apostichopus japonicus. PLOS ONE. 7, e44492 (2012).
Escudero Castelán, N. et al. Receptor deorphanization in an echinoderm reveals kisspeptin evolution and relationship with SALMFamide neuropeptides. BMC Biol. 20, 187 (2022).
Lin, M., Egertová, M., Zampronio, C. G., Jones, A. M. & Elphick, M. R. Pedal peptide/orcokinin-type neuropeptide signaling in a deuterostome: the anatomy and pharmacology of starfish myorelaxant peptide in Asterias rubens. J. Comp. Neurol. 525, 3890–3917 (2017).
Suwansa-ard, S. et al. Transcriptomic discovery and comparative analysis of neuropeptide precursors in sea cucumbers (Holothuroidea). Peptides 99, 231–240 (2018).
Tinoco, A. B. et al. Ancient role of sulfakinin/cholecystokinin-type signaling in inhibitory regulation of feeding processes revealed in an echinoderm. eLife 10, e65667.
Zhang, Y. et al. Somatostatin-type and allatostatin-C–type neuropeptides are paralogous and have opposing myoregulatory roles in an echinoderm. Proc. Natl. Acad. Sci. U.S.A. 119, e2113589119 (2022).
Chieu, H. D., Suwansa-ard, S., Abramov, T., Elizur, A. & Cummins, S. F. In vitro oocyte maturation by radial nerve extract and early development of the black sea cucumber (Holothuria leucospilota). Aquaculture 495, 247–254 (2018).
Chieu, H. D., Suwansa-ard, S., Wang, T., Elizur, A. & Cummins, S. F. Identification of neuropeptides in the sea cucumber Holothuria leucospilota. Gen. Comp. Endocrinol. 283, 113229 (2019).
Chieu, H. D. et al. Aquaculture breeding enhancement: maturation and spawning in Sea cucumbers using a recombinant relaxin-like gonad-stimulating peptide. Front. Genet. 10, (2019).
Hoang Dinh, C. Genetic investigation and molecular identification of reproduction neuropeptides in sea cucumber. (2019). https://doi.org/10.25907/00515
Lee, E. Y. et al. PACAP is a pathogen-inducible resident antimicrobial neuropeptide affording rapid and contextual molecular host defense of the brain. Proceedings of the National Academy of Sciences. 2021 [accessed 2024 Apr 23];118(1):e1917623117. https://www.pnas.org/doi/https://doi.org/10.1073/pnas.1917623117. https://doi.org/10.1073/pnas.1917623117.
Chen, M., Talarovicova, A., Zheng, Y., Storey, K. B. & Elphick, M. R. Neuropeptide precursors and neuropeptides in the sea cucumber Apostichopus japonicus: a genomic, transcriptomic and proteomic analysis. Sci. Rep. 9, 8829 (2019).
Zhang, X. et al. The sea cucumber genome provides insights into morphological evolution and visceral regeneration. PLoS Biol. 15, e2003790 (2017).
Nocillado, J. et al. Spawning induction of the high-value white teatfish sea cucumber, Holothuria fuscogilva, using recombinant relaxin-like gonad stimulating peptide (RGP). Aquaculture 547, 737422 (2022).
Ajik, J., Ajik, M-N. & Tahiluddin, A. Population Density, distribution, and Species Composition of Sea cucumbers in Pagasinan tidal flat, Bongao, Tawi-Tawi, Philippines. TPJF 143-149 https://doi.org/10.31398/tpjf/28.2.2021-0008 (2022).
Jr, E. C. C. Growth and survival of sea urchin (Tripneustes gratilla) fed different brown algae in aquaria.
Palomar-Abesamis, N., Abesamis, R. A. & Juinio-Meñez, M. A. Distribution and microhabitat associations of the juveniles of a high-value sea cucumber, Stichopus cf. horrens, in northern Philippines. Aquat. Ecol. 51, 17–31 (2017).
Lizano, A. M. D., Kim, K. M., Juinio-Meñez, M. A. & Ravago-Gotanco, R. Pseudocryptic diversity and species boundaries in the sea cucumber Stichopus cf. horrens (Echinodermata: Stichopodidae) revealed by mitochondrial and microsatellite markers. Sci. Rep. 14, 4886 (2024).
Liu, R., Liu, J. & Zhang, H. Positive selection analysis reveals the deep-sea adaptation of a hadal sea cucumber (Paelopatides sp.) to the Mariana Trench. J. Ocean. Limnol. 39, 266–281 (2021).
Mongiardino Koch, N. et al. A phylogenomic resolution of the sea urchin tree of life. BMC Evol. Biol. 18, 189 (2018).
Transcriptomic data of sea. cucumber Stichopus horr. (ID 512756) - BioProject - NCBI. https://www.ncbi.nlm.nih.gov/bioproject/PRJEB29236
Ordoñez, J. F. F., Galindez, G. G. S. T., Gulay, K. T. & Ravago-Gotanco, R. Transcriptome analysis of growth variation in early juvenile stage sandfish Holothuria scabra. Comp. Biochem. Physiol. Part. D Genomics Proteom. 40, 100904 (2021).
Monroe, E. B. et al. Exploring the sea urchin neuropeptide landscape by mass spectrometry. J. Am. Soc. Mass. Spectrom. 29, 923–934 (2018).
Zheng, Y. et al. Nervous System Development and neuropeptides characterization in embryo and larva: insights from a non-chordate deuterostome, the Sea Cucumber Apostichopus japonicus. Biology 11, 1538 (2022).
Smith, M. K. et al. Structure and proteomic analysis of the crown-of-thorns starfish (Acanthaster sp.) radial nerve cord. Sci. Rep. 13, 3349 (2023).
Smith, M. K. et al. The neuropeptidome of the Crown-of-Thorns Starfish, Acanthaster planci. J. Proteom. 165, 61–68 (2017).
Van Bael, S. et al. A Caenorhabditis elegans Mass Spectrometric Resource for Neuropeptidomics. J. Am. Soc. Mass. Spectrom. 29 (5), 879–889. https://doi.org/10.1007/s13361-017-1856-z (2018). https://doi.org/10.1007/s13361-017-1856-z [accessed 2024 Apr 23].
Bhat, U. S., Shahi, N., Surendran, S. & Babu, K. Neuropeptides and behaviors: how small peptides regulate nervous system function and behavioral outputs. Front. Mol. Neurosci. 14, (2021).
Chew, Y. L., Grundy, L. J., Brown, A. E. X., Beets, I. & Schafer, W. R. Neuropeptides encoded by nlp-49 modulate locomotion, arousal and egg-laying behaviours in Caenorhabditis elegans via the receptor SEB-3. Philosophical Trans. Royal Soc. B: Biol. Sci. 373, 20170368 (2018).
Zhang, X. et al. Neuropeptidomic analysis establishes a major role for Prohormone Convertase-2 in Neuropeptide Biosynthesis. J. Neurochem. 112, 1168–1179 (2010).
Hayakawa, E. et al. Mass spectrometry of short peptides reveals common features of metazoan peptidergic neurons. Nat. Ecol. Evol. 6, 1438–1448 (2022).
Wiggenhorn, A. L. et al. A class of secreted mammalian peptides with potential to expand cell-cell communication. Nat. Commun. 14, 8125 (2023).
Abd El-Aal, A. A. A., Jayakumar, F. A., Lahiri, C., Tan, K. O. & Reginald, K. Novel cationic cryptides in Penaeus vannamei demonstrate antimicrobial and anti-cancer activities. Sci. Rep. 13, 14673 (2023).
Kilinc, M., Jia, K. & Jernigan, R. L. Improved global protein homolog detection with major gains in function identification. Proc. Natl. Acad. Sci. U.S.A. 120 (9), e2211823120. https://doi.org/10.1073/pnas.2211823120 (2023).
Sanderson, T., Bileschi, M. L., Belanger, D. & Colwell, L. J. ProteInfer, deep neural networks for protein functional inference. eLife 12, e80942 (2023).
Tian, S. et al. Urbilaterian origin of paralogous GnRH and corazonin neuropeptide signalling pathways. Sci. Rep. 6, 28788 (2016).
Teufel, F. et al. SignalP 6.0 predicts all five types of signal peptides using protein language models. Nat. Biotechnol. 40 (7), 1023–1025. https://doi.org/10.1038/s41587-021-01156-3 (2022).
Cai, W. et al. Biochemical, anatomical, and pharmacological characterization of calcitonin-type neuropeptides in Starfish: Discovery of an ancient role as muscle relaxants. Front. NeuroSci. 12, 382. https://doi.org/10.3389/fnins.2018.00382 (2018).
Hofbauer, B. et al. The neuropeptide pigment-dispersing factor signals independently of bruchpilot-labelled active zones in daily remodelled terminals of Drosophila clock neurons. Eur. J. Neurosci. 59 (10), 2665–2685. https://doi.org/10.1111/ejn.16294 (2024).
Helfrich-Förster, C. & Rieger, D. A clock for all seasons. J. Comp. Physiol. A. https://doi.org/10.1007/s00359-024-01711-8 (2024). 210:4, (473–480), Online publication date: 1-Jul-2024.
Merkler, D. J. C-terminal amidated peptides: production by the in vitro enzymatic amidation of glycine-extended peptides and the importance of the amide to bioactivity. Enzym. Microb. Technol. 16 (6), 450–456. https://doi.org/10.1016/0141-0229(94)90014-0 (1994).
Cong, X. et al. Functional characterisation of neuropeptides that act as ligands for both calcitonin-type and pigment-dispersing factor-type receptors in a deuterostome. bioRxiv. Jan 1:2024.08.29.610391. http://biorxiv.org/content/early/2024/08/30/2024.08.29.610391.abstract. (2024). https://doi.org/10.1101/2024.08.29.610391
Penke, B. & New York Academy of Sciences. In Vivo Sulfation of Cholecystokinin Octapeptide. Annals of the. [accessed 2024 Dec 4];448(1):293–305. https://onlinelibrary.wiley.com/doi/abs/10.1111/j.https://doi.org/10.1111/j.1749-6632.1985.tb29925.x(1985).
Bonetto, V. et al. Isolation and characterization of sulphated and nonsulphated forms of cholecystokinin-58 and their action on gallbladder contraction. Eur. J. Biochem. 264 (2), 336–340. https://doi.org/10.1046/j.1432-1327.1999.00599.x (1999). https://onlinelibrary.wiley.com/doi/abs/10.1046/j.1432-1327.1999.00599.x [accessed 2024 Dec 4].
Ding, K. et al. The Effect of Pedal Peptide-Type Neuropeptide on Locomotor Behavior and Muscle Physiology in the Sea Cucumber Apostichopus japonicus. Frontiers in Physiology. 2020 [accessed 2024 Dec 4];11. https://www.frontiersin.org/journals/physiology/articles/10.3389/fphyshttps://doi.org/10.3389/fphys.2020.559348. .
Ding, K. et al. Influence of an L-type SALMFamide neuropeptide on locomotory performance and muscle physiology in the sea cucumber Apostichopus japonicus. J. Exp. Biol. 224 (19), jeb242566. https://doi.org/10.1242/jeb.242566 (2021). [accessed 2024 Dec 4];.
Elphick MR, Achhala, S. & Martynyuk, N. The evolution and diversity of SALMFamide neuropeptides. PLOS ONE. 8 (3), e59076. https://doi.org/10.1371/journal.pone.0059076 (2013). https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0059076 [accessed 2024 Dec 25].
Chaiyamoon, A. et al. Characterization of TRH/GnRH-like peptides in the sea cucumber, Holothuria scabra, and their effects on oocyte maturation. Aquaculture 518, 734814. https://doi.org/10.1016/j.aquaculture.2019.734814 (2020). https://linkinghub.elsevier.com/retrieve/pii/S0044848619321751
Acknowledgements
The authors would like to acknowledge Vicenzo Paolo Torreno for his technical assistance, Dr. Annette Juinio-Meñez and the UP MSI Marine Invertebrate Ecology Laboratory for the provision of samples, and the Mass Spectrometry Facility at the Institute of Chemistry, UP Diliman for instrument use. This research was supported by the Department of Science and Technology—Philippine Council for Agriculture, Aquatic and Natural Resources Research and Development (DOST PCAARRD), the PCAARRD-Graduate Research and Education Assistantship for Technology program, and the University of the Philippines Office of the Vice Chancellor for Research and Development Outright Research Grant (UP-OVCRD ORG Project #23218).
Author information
Authors and Affiliations
Contributions
All authors were involved in the conception of the study, interpretation of data, and preparation of the manuscript. J.A.R. performed the experiments, data analysis, and drafted the manuscript and prepared the figures. All authors contributed to the manuscript revision and approved the submitted version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Resoles, J.A., Yu, E.T. The neuropeptidomes of the sea cucumbers Stichopus cf. horrens and Holothuria scabra. Sci Rep 15, 7032 (2025). https://doi.org/10.1038/s41598-025-85696-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-85696-7