Abstract
Ensemble learning aggregates several models’ outputs to improve the overall model’s performance. Ensemble feature selection separating the appropriate features from the extra and non-essential features. In this paper, the main focus will be to expand the scope of Ensemble Learning to include Feature Selection. We will propose an Ensemble Feature Selection Method called AdaptDiabfor Diabetes Prediction that is Model-Agnostic. Our approach combines diverse feature selection techniques, such as filter and wrapper methods, harnessing their complementary strengths. We have used an adaptive combiner function, which dynamically selects the most informative features based on the characteristics of the ensemble members. We demonstrate the effectiveness of our proposed AdaptDiab method through empirical studies using various classification models. Empirical Results of Our Proposed Ensemble Feature Selection Model outperforms traditional methods. This paper contributes to Ensemble Learning Methods and provides a Practical and Better Framework for Feature selection.
Similar content being viewed by others
Introduction
Diabetes is a primary global health concern with significant economic and societal burdens. Early detection and intervention are crucial for preventing severe complications and improving patient outcomes1. Accurate diabetes prediction is vital for early diagnosis and intervention, allowing for improved patient outcomes and potentially preventing complications. However, achieving precise prediction can be challenging due to the complex nature of diabetes itself. High-dimensional datasets used for prediction often involve numerous features, which can lead to overfitting and reduced model interpretability.
Additionally, irrelevant or redundant features can hinder the model’s identifying genuinely informative patterns. These challenges necessitate robust feature selection techniques to identify the most relevant features and improve the accuracy and generalizability of diabetes prediction models. Feature selection selects relevant and non-redundant features from an ample feature space. In many machine learning and pattern recognition applications, feature selection is used to select the optimal feature set from the feature space to train the learning model.
High-dimensional datasets with many features result in increased computational complexity, overfitting, and reduced generalization performance. Machine learning has emerged as a promising tool for predicting diabetes risk, allowing for preventative measures and better patient management. However, existing machine learning approaches encounter several challenges, including data quality issues, lack of interpretability, and limited generalizability2. Models trained on these datasets exhibit a decrease in interpretability. Optimal feature selection techniques address challenges associated with high dimensional datasets, computational overhead, reduced interpretability, and suboptimal model performance. The main objective of feature selection is to improve prediction accuracy, remove redundant features, and reduce time consumption during analysis. However, single feature-selection techniques are sensitive to specific data characteristics, don’t capture interaction and dependencies among features, and are vulnerable to noisy and redundant features. So, it leads to poor generalization and decreased model robustness. Many single feature-selection methods, such as mutual information or ANOVA, rely heavily on specific assumptions (e.g., linear relationships or independence among variables) about the dataset. Techniques like the Fisher score evaluate features individually, ignoring interactions or dependencies between features. Single-feature selection methods are not inherently optimized for multiple machine-learning models. So, we propose a model based on ensemble feature selection techniques by leveraging the strength of numerous feature selection techniques as a synergistic method. In the ensemble feature selection method, multiple feature selection techniques are combined to produce an optimal subset of features using a combination of feature ranking that improves accuracy. The ensemble feature selection method ensures robustness through diversity, comprehensive exploration, noise reduction, and enhanced stability. In the first step of the ensemble method, a set of different feature selectors is chosen, and each selector provides a sorted order of features. The second step aggregates the selected subsets of features using different aggregation techniques. This research investigates the effectiveness of the model-agnostic AdaptDiab architecture in improving the performance of diabetes prediction models compared to traditional feature selection techniques.
Related works
The quality of data used for training machine learning models significantly impacts their accuracy. Issues like missing values, imbalanced datasets, and noisy data can hinder model performance. The approach proposed3 demonstrates how dynamic optimization techniques can effectively address imbalanced data challenges. While their focus is on synthetic sample generation, our work, AdaptDiab, addresses a complementary challenge by emphasizing the importance of selecting relevant and non-redundant features, which helps mitigate the bias caused by imbalanced class distributions. Models trained on specific datasets might not perform well on unseen data. Ensuring generalizability across diverse patient populations and healthcare institutions is crucial for real-world application. The study4 presents a novel resampling strategy that effectively leverages labeled and unlabeled data to address class imbalance. This approach highlights the importance of balancing data distributions to improve the predictive performance of machine learning models. Our proposed method complements such strategies by selecting the most relevant and non-redundant features, which inherently mitigates the effects of class imbalance. By ensuring that features contributing to the minority class are preserved and amplified through an ensemble feature selection process, Our technique enhances the model’s ability to generalize across imbalanced datasets. Moreover, the semi-supervised perspective of the cited work inspires potential future directions for our work, where semi-supervised or unsupervised methods could be integrated further to refine feature selection in low-label or imbalanced scenarios.
Feature selection plays a crucial role in enhancing the performance of classification tasks. By selecting a subset of relevant features, researchers have observed significant improvements in model accuracy5 and interpretability6. The paper7 highlights the importance of reducing noise and redundancy in high-dimensional medical datasets. Similarly, our method tackles these issues in tabular datasets by employing an ensemble feature selection technique. It ensures the selection of the most relevant features, enhancing model performance and demonstrating the importance of robust feature selection approaches in healthcare-related machine learning tasks.
Several feature selection techniques have been established to achieve this goal, each with strengths and limitations. Filter methods like chi-square8 and information gain9 analyze the intrinsic properties of features to identify those most correlated with the target variable. Wrapper methods like sequential forward selection10 and recursive feature elimination11 iteratively evaluate feature subsets using a classification algorithm, leading to potentially superior results. Embedded methods like LASSO regression12 and Elastic Net13 integrate feature selection within the model training process, balancing efficiency and effectiveness.
To overcome the limitations of individual techniques, researchers have explored ensemble feature selection methods that combine the strengths of multiple approaches. These methods have been shown to improve robustness and reduce bias compared to a single technique14. However, designing effective ensemble methods can be challenging, and their computational complexity can be significant15.
In conclusion, feature selection offers a powerful tool for improving classification performance. While various techniques exist, each with advantages and disadvantages, the field actively explores ensemble methods for even more robust and effective feature selection.
Preprocessing
Data preprocessing is an essential step in the data analysis pipeline that plays a vital role in enhancing the quality and reliability of the dataset for subsequent modeling tasks. It involves a series of operations designed to clean, transform, and organize raw data to make it suitable for analysis by machine learning algorithms. The real-world datasets are often noisy, incomplete, or contain irrelevant information, which can adversely impact the performance of predictive models. The primary objectives of preprocessing are to address these challenges, improve data quality, and create a more conducive environment for accurate and meaningful model training.
These are the following methods that we used for preprocessing:
(1)) Replacing null values with mean: Missing values in a dataset can cause issues during model training, leading to biased results and reduced accuracy. Filling null values with the mean ensures that missing data does not disrupt the overall statistical characteristics of the feature. This approach maintains data integrity and prevents the loss of valuable information.
(2) Handling Outliers: Outliers can distort statistical measures, leading to biased model training and compromised generalization. Outliers can skew data distribution and adversely affect specific models’ performance. Removing outliers based on the IQR ensures a more robust and representative dataset.
The lower (Q1) and upper (Q3) quartiles are calculated for a feature Xi. The IQR is then determined as IQR = Q3 − Q1. Outliers below Q1 − 1.5×IQR or above Q3 + 1.5×IQR are removed.
(3) Normalization: Features with varying scales can lead to biased model training, as certain features may dominate the learning process. Normalization ensures that features with different scales contribute equally to the model training process. It improves convergence and stability in algorithms sensitive to input features’ scale.
For a feature Xi, normalization is expressed as:
where\(\:{X}_{i}^{{\prime\:}}\)′ is the normalized feature, mean (Xi) is the mean of Xi, and std (Xi) is the standard deviation of Xi.
(4) Oversampling with SMOTE16: Class imbalance can result in models with high accuracy but poor performance in minority class instances. SMOTE is the Synthetic Minority Over-sampling Technique. Class imbalance can lead to biased models that favor the majority class. Oversampling the minority class using SMOTE helps mitigate this imbalance, improving the model’s ability to classify minority instances correctly.
SMOTE generates synthetic instances for the minority class by interpolating between existing cases. Given a minority instance X(k), a synthetic instance is created by selecting X(m) from the k-nearest neighbors and generating a synthetic instance X′ as:
where λ is a random value between 0 and 1.
Replacing missing values with the remaining data’s mean ensures a complete ensemble-based feature selection analysis dataset. It allows the ensemble feature selection methods to evaluate features based on their distribution within the diabetic and non-diabetic populations. Also, it leads to a more accurate selection of features that discriminate between the two groups. IQR-based outlier removal helps identify and eliminate outliers that might disproportionately influence the choice of features by ensemble feature selection algorithms. This approach ensures ensemble feature selection focuses on the core trends within the diabetic and non-diabetic populations, leading to a more robust selection of features representing most patients. For example, an unusually high number of pregnancies for a single data point might be an outlier. It shouldn’t unduly influence the selection of features related to pregnancy as a risk factor. The Pima Indian Medical Association dataset17 might be imbalanced, with fewer cases of diabetic patients compared to non-diabetic patients. If the Pima Indian Medical Association dataset exhibits class imbalance, SMOTE oversampling can be beneficial. By creating synthetic data points for the minority class (diabetic patients), SMOTE helps balance the dataset.
By incorporating these preprocessing steps, the research aims to ensure a clean, balanced, and appropriately scaled dataset, thereby mitigating potential issues that could impact the performance of machine learning models during training and evaluation.
Materials and methods
Dataset
We use various materials and methods to analyze diabetes prediction thoroughly. The dataset we use in this study is the Pima Indian Medical Association Indians diabetes dataset17. Pima Indian Medical Association Indians are among the most popular machine-learning data sets for predicting and evaluating diabetes. It is frequently used as a reference for performing various algorithms to indicate diabetes. The NIDDK originally developed the Pima Indian Medical Association Indian data set in partnership with Native American communities. It consists of 768 data points, each representing one person. Each data point contains eight health-related characteristics and binary outcome variables to determine whether or not the individual has diabetes. The health-related characteristics are Pregnancy (number of times pregnant), Glucose (level of glucose concentration in an oral glucose test), Diastolic blood pressure, Triceps skinfold thickness, Insulin (level of Insulin in an insulin test), Body mass index, DiabetesPedigreeFunction (a measure of heritable risk) and Age.
The glucose value ranges from 0 to 199 (the non-diabetic range is lower). The pregnancies have no missing values. The values of skin thickness and Insulin require imputation. The values of DBF range from 0.078 to 2.42 and need imputation. The dataset contains no categorical variables. Most features have skewed distributions, indicating the need for scaling or transformation for better model performance.
Feature selection
Feature selection is an essential step in machine learning that involves choosing relevant subsets of features from all the features to improve model performance, interpretability, and efficiency. Feature selection techniques are broadly classified into filter, wrapper, and embedded. In this experiment, we will use four feature selection filter types: ANOVA F-Score18, Fisher Score19, Variance Threshold20, and Point Biserial21. These Filter type methods evaluate the relevance of features independently of the chosen learning algorithm.
ANOVA F-score
ANOVA stands for anomaly detection of variance. It is also referred to as F-statistics.ANOVA is a statistical technique used to compare the mean values of two groups of participants. An ANOVA F-score determines the significance of specific features associated with a target variable. ANOVA (Anomaly detection of variance) is beneficial for classifying tasks that search for features that show significant deviation across classes.
The ANOVA F-Score for a feature i in a dataset with k classes is calculated as follows:
Where, MSR = Mean Square due to Regression, MSE = Mean Square due to Error, and F is ANOVA F-Score.
In the context of feature selection, higher ANOVA F- scores demonstrate significant differences in mean across classes. A higher ANOVA F-score indicates more significant variability between class means than within-class variability, showing that a particular feature is essential for classification.
Fisher score
Fisher’s score or discriminant analysis is a statistical tool mainly used for binary classification tasks. It assesses the discriminative power of individual features by evaluating the variance ratio between classes and within classes. The goal is to identify features that maximize the separation between class means while minimizing the spread within each class.
The Fisher Score for feature i is calculated as follows:
Where, Fi is the Fisher Score for feature i, Xˉ0i and Xˉ1i are the means of feature i for class 0 and 1, respectively, S0i2 and S1i2 are the variances of feature i for Class 0 and Class 1.
The numerator represents the squared difference between the means of the two classes, while the denominator is the sum of the variances within each class. The larger the Fisher Score, the better the feature discriminates between the two classes. Features with higher Fisher Scores are considered more relevant for classification tasks and are often selected for inclusion in predictive models.
Variance threshold
Variance Threshold involves removing features with low variance. Eliminating these low-variance features aims to improve model performance and computational efficiency.
For a feature i, the variance (Vari) is computed as follows:
Vari is the variance of feature i, n is the total number of instances, xij is the jth instance of feature i, ˉi is the mean of feature i.
The variance thresholding method involves setting a predefined threshold (T) and removing features with variance below this threshold. The mathematical representation of this process is:
Remove feature i if Vari<T.
The variance threshold is beneficial in scenarios where features with slight variation do not contribute significantly to the model’s predictive power and may introduce noise.
Point biserial
The Point Biserial Correlation Coefficient is a statistical measure that assesses the relationship between binary and continuous variables. The Point Biserial Correlation Coefficient ranges19 from − 1 to 1, where − 1 indicates a perfect negative correlation, 1 indicates a perfect positive correlation, and 0 indicates no correlation.
For a binary target variable Y and a continuous feature X, the Point Biserial Correlation Coefficient (rpb) is calculated as follows:
Where, rpb is the Point Biserial Correlation Coefficient, n is the number of instances, Xi is the value of the continuous feature, for example,i, Xˉ is the mean of the constant feature, Yi is the binary target variable, for instance,i, Yˉ is the mean of the binary target variable.
The magnitude of rpb indicates the strength of the relationship, with values closer to -1 or 1 indicating a stronger association. A higher absolute value of rpb in feature selection suggests that the feature is more correlated with the binary target variable, making it a potentially valuable predictor in classification tasks.
Classification models
This section provides an insightful overview of 5 diverse classification algorithms we used in our research, each with unique strengths and applications. These algorithms have been chosen to explore our dataset and proposed ensemble feature selection technique and compare it with traditional feature selection techniques. The five chosen classification algorithms are Gaussian Naive Bayes, Logistic Regression, Random Forest, Gradient Boosting, and MultiLayer Perceptron. Logistic Regression’s interpretability makes it a valuable benchmark for evaluation. SVMs are robust to high-dimensional datasets and effectively handle overlapping data points using kernel methods. It Highlights how feature selection techniques improve model performance by reducing irrelevant features, which is critical for SVM to avoid overfitting. Gradient Boosting Demonstrates robustness by enhancing the performance of a highly complex and adaptive model. The chosen models represent a mix of linear (LR) and non-linear (SVM, RF, GBM), ensuring the generalizability of ensemble feature selection across varied learning paradigms. This classification algorithm array ensures a thorough evaluation of our proposed ensemble feature selection technique. The subsequent sections delve into each classification model’s specific methodologies and results, providing an understanding of their contributions to the overall predictive framework.
Gaussian Naive Bayes
Gaussian Naive Bayes is a probabilistic classification algorithm based on Bayes’ theorem, which calculates the probability of a hypothesis given the observed evidence. It is designed explicitly for datasets with continuous features, assuming that the values of each feature follow a Gaussian (normal) distribution within each class.
The probability of an instance x belonging to class Ck in GNB is calculated using Bayes’ theorem as follows:
Where, P(Ck∣x) is the probability of class Ck given the observed instance x, P(x∣Ck) is the likelihood of observing x given class Ck, P(Ck) is the prior probability of class Ck, P(x) is the probability of observing x across all classes, acting as a normalization factor, mean µk and variance 2σk2.
Gaussian Naive Bayes is chosen for the following reasons: (1) Assumption of feature independence, (2) Simplicity and Interpretability, and (3) Robustness to noisy data.
By incorporating Gaussian Naive Bayes in the ensemble of classification models, the research aims to provide insights into the adaptability and effectiveness of the proposed feature selection methodology.
Logistic regression
Logistic Regression is a widely used linear model for binary classification tasks. The model estimates the probability that an instance belongs to a particular class and makes predictions based on a logistic function. The sigmoid function is used to transform the output of a linear combination of features into a probability between 0 and 1:
P(Y = 1∣X) is the probability of belonging to class 1 given the feature values X, β0, β1,…, βn are the model parameters (coefficients), and X1, X2,…, Xn are the feature values.
The logistic regression model is trained by optimizing these coefficients to maximize the likelihood of the observed outcomes.
Logistic Regression was selected as one of the classification models for evaluating the proposed ensemble feature selection technique for several reasons, such as linear decision boundary, interpretability, sensitivity to feature changes, and benchmark for simplicity. The analysis with Logistic Regression contributes to understanding how the ensemble approach influences both linear and non-linear models, emphasizing the versatility of the feature selection technique.
Random forest
Random Forest is an ensemble learning algorithm that builds many decision trees during training and merges their predictions to improve accuracy and robustness. It is known for its versatility, ability to handle complex relationships, and resistance to overfitting.
The Random Forestalgorithm combines the predictions of multiple decision trees. For a binary classification task, the probability of an instance belonging to class 1 can be expressed as an average over all the individual trees:
P(Y = 1∣X) is the probability of belonging to class 1 given the feature values X, N is the total number of trees in the forest, and Pi(Y = 1∣X) is the probability predicted by the ith tree.
Random Forest was included as one of the classification models for evaluating the proposed ensemble feature selection technique due to the following considerations: Ensemble nature, robustness and generalization, feature importance, non-linearity, and interaction. Incorporating Random Forest into the ensemble of classification models enhances the research’s comprehensiveness, enabling a thorough examination of the proposed feature selection methodology’s performance across different learning paradigms and model complexities.
Gradient boosting
Gradient Boosting is an ensemble learning algorithm that builds a series of weak learners (typically decision trees) sequentially, with each tree compensating for the errors of its predecessors. It is a powerful and flexible algorithm known for its high predictive accuracy and ability to handle complex relationships in the data.
For a binary classification task, the prediction of an instance belonging to class 1 in a Gradient boosting model can be expressed as:
P(Y = 1∣X) is the probability of belonging to class 1 given the feature values X. F (X) is the additive function representing the sum of predictions from all the weak learners in the ensemble.
Gradient Boosting was chosen as one of the classification models to evaluate the proposed ensemble feature selection technique for the following reasons: sequential nature, ensemble characteristics, and handling non-linearity. The model minimizes a loss function during training, and each subsequent tree is trained to correct the errors made by the previous trees. This analysis contributes to the broader assessment of the feature selection technique’s applicability across different ensemble learning paradigms.
Multilayer perceptron
A Multilayer Perceptron is an artificial neural network consisting of multiple layers of nodes, including input, hidden, and output layers. MLPscan learns complex non-linear relationships in data, making them suitable for various applications, including classification tasks.
For a binary classification task, the output of an MLP can be expressed as follows:
Where, P(Y = 1∣X) is the probability of belonging to class 1 given the feature values X, W(i), and b(i) represent the weights and biases of layer i, ϕ is the activation function applied element-wise to the layer’s input, σ is the sigmoid activation function applied to the final layer.
Multilayer Perceptron (MLP) was included in the ensemble of classification models for evaluating the proposed ensemble feature selection technique due to several key factors: non-linearity and complexity, representation learning, adaptability to high dimensional data, and benchmark for deep-learning models. By incorporating Multilayer Perceptron into the ensemble of classification models, the research aims to comprehensively evaluate the proposed feature selection methodology across diverse neural network architectures, offering insights into its adaptability and efficacy in deep learning.
Proposed architecture
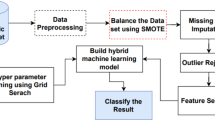
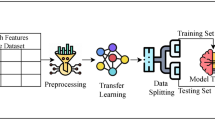
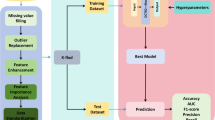
AdaptDiab – Adaptive Diabetes Prediction Feature Selection Tool In this research, we introduce anAdaptDiab algorithm to enhance the effectiveness of feature selection algorithms. Many traditional feature selection approaches rely on a single algorithm, which may overlook important information in the data. In contrast, the AdaptDiabarchitecture presented here combines the outputs of different feature selection techniques to create a collaborative decision-making process that leverages the strengths of various methods, as shown iFig. . 1.
The ensemble approach is versatile, adaptive, and robust. Ensemble learning combines predictions from several models to enhance forecasting accuracy and reliability. The goal is to reduce model errors or biases using an ensemble’s collective intelligence. Ensemble buildings can be classified as homogeneous and heterogeneous ensemble techniques22. A homogenous ensemble is a collection of feature selection algorithms of the same type, built upon a different subset of data. A homogeneous ensemble distributes the dataset on other nodes. A heterogeneous ensemble is a set of feature selection algorithms of various types built upon the same data. A heterogeneous ensemble combines the result of different feature selection methods. The heterogeneous approach takes account of the strengths and weaknesses of the individual techniques.
Our proposed approach combines the power of both homogeneous and heterogeneous ensemble techniques by splitting the processed data into subsets, applying feature selection algorithms on them similar to the homogenous ensemble feature selection technique, and using different types of feature selection algorithms on the same data identical to the heterogenous ensemble feature selection technique.
The AdaptDiab splits processed data into three subsets using a random shuffle. Four feature selection techniques, ANOVA F-score, fisher score, variance threshold, and Point biserial, select features from 3 data subsets. The combiner function in the AdaptDiab architecture is used to aggregate the results from all individual feature selection algorithms applied to different subsets of data. This combiner function produces the final selected features using the AdaptDiab technique.
The combiner function in AdaptDiab’s architecture works based on the principle of assigning weights to each feature selection algorithm, and these weights determine the contribution of each feature selection technique to the results of the combined ensemble feature selection algorithm. The weights for each feature selection algorithm are assigned using mutual information.
p(x, y): The joint probability distribution of X and Y.
p(x): The marginal probability distribution of X.
p(y): The marginal probability distribution of Y.
Mutual information between two random variables is a non-negative value, which measures the dependency between the variables. It equals zero if two random variables are independent; higher values mean higher dependency. This mutual information is used to find the dependency between each feature selected using the individual feature selection algorithm and the target variable.
The combiner function assigns a mutual information score for each feature. The average of these mutual information scores obtained from each feature selected is the mutual information score of that particular feature selection algorithm. The average of all these mutual information scores from each feature selection algorithm processed that particular data subset is the mutual information score of that specific data subset. Based on this average mutual information score, weightage is given to results obtained from each data subset, and based on this weightage, the combiner function in the AdaptDiab architecture selects features from the features proposed by different feature selection algorithms from different data subsets.
Results and discussion
The evaluation was divided into two stages. The first focused on assessing the feature relevance; thus, we compared the performance of several classification algorithms using all features selected by existing feature selection algorithms with the performance achieved when we used subsets of relevant features chosen by theAdaptDiab approach.
In the second stage, the stability of the subset of relevant features was calculated. We calculate stability to measure the consistent performance of the tool across different data subsets, robustness, and interpretation. Stability metrics like the Jaccard Index help ensure that your model is not overfitting or excessively sensitive to the particularities of a specific dataset, which is especially important in real-world applications.
Performance
Our proposed AdaptDiab technique selects a feature subset from the whole feature set. As mentioned in section "Preprocessing", we train our models in these features. First, we train it on the entire feature set, then on individual feature subsets selected by individual feature selection methods, and then on the feature subset selected by our proposed feature selection tool. We calculate the models’ performance on each feature subset.
Table 1 shows the accuracy score of all the models on each feature subset. It indicates that models that use features selected by our AdaptDiab technique outperform features selected by other individual feature selection techniques in model performance.
Figure 2 shows the plot for comparing accuracy scores of different feature selection techniques in the Gaussian Naive Bayes model. The plot shows that the proposed AdaptDiab approach performs better than other feature selection techniques.
Figure 3 shows the plot for comparing the accuracy scores of different feature selection techniques in the Logistic Regression model. The plot shows that the proposed AdaptDiab approach performs better than other feature selection techniques.
Figure 4 shows the plot for comparing the accuracy scores of different feature selection techniques in the Random Forest model. The plot shows that the proposed AdaptDiab approach performs better than other feature selection techniques.
Figure 5 shows the plot for comparing the accuracy scores of different feature selection techniques in the Gradient Boosting model. The plot shows that the proposed AdaptDiab approach performs better than other feature selection techniques.
Figure 6 shows the plot for comparing the accuracy scores of different feature selection techniques in the Multilayer Perceptron model. The plot shows that the proposed AdaptDiab approach performs better than other feature selection techniques.
ANOVA F-score technique might favor features with high variance, which could be informative for some models like Gaussian naive Bayes but lead to potentially irrelevant features for other models like a random forest. By combining diverse feature techniques, AdaptDiab can capture various informative features, including those with high and low variance but with interaction effects with other features. It shows our proposed technique’s consistent performance across multiple models.
The logistic regression model thrives on features that effectively separate classes. Point Biserial’s strength lies in identifying features that distinguish between classes. By combining different techniques, AdaptDiab selects a mixture of features that capture class separability and potentially informative features with a more complex relationship to diabetes. This broader selection could contribute to the consistent performance of our ensemble feature selection across various models, including Logistic Regression.
The random forest model is known for handling complex interactions between features and feature importance scores. Variance threshold techniques might remove features with subtle variations that could hold valuable information for capturing complex interactions, potentially explaining its lower performance in Random Forest than other models. Our approach might better identify these subtle variations and interaction effects.
The Gradient boosting model performs well with diverse features, including those with lower individual importance. Some techniques, like Fisher Score or ANOVA F-Score, might prioritize features with high individual variance, potentially missing features that contribute within the boosting framework. By combining methods, our approach might more likely capture this broader range of features, including those that are less prominent individually but contribute to the boosting process. It could explain why our AdaptDiabperforms well in Gradient Boosting compared to some individual techniques.
The tool is handy in real-world diabetes prediction applications, where datasets are often noisy and high-dimensional. By consistently selecting the most critical features across different models, AdaptDiab ensures better generalization and reduces overfitting, making it highly applicable in healthcare settings. This ability to outperform existing methods across multiple models makes AdaptDiab a powerful tool for developing reliable and interpretable diabetes prediction systems.
In conclusion, our proposed AdaptDiab approach demonstrates consistent and superior performance in feature selection across various machine learning models for diabetes prediction compared to individual feature selection techniques. This improvement can be attributed to several factors. The ensemble approach mitigates the limitations of individual techniques by leveraging their diverse strengths. It avoids biases towards specific feature types, explores complex feature interactions, and potentially reduces the impact of noisy data. The consistently high accuracy scores achieved byAdaptDiab across models like Gaussian Naive Bayes, Logistic Regression, Random Forest, Gradient Boosting, and Multilayer Perceptron strongly support its effectiveness.
Stability
Jaccard coefficient23, also known as the Jaccard similarity index, is a statistic used to measure the similarity between two sets. It calculates the proportion of shared elements between two sets relative to the total number of distinct elements. The Jaccard index helps measure stability in feature selection.
Mathematically, the Jaccard index J(A, B) between two sets A and B is defined as:
Where, |A∩B∣ represents the cardinality (number of elements) of the intersection of sets A and B. ∣A∪B∣means the cardinality of the union of sets A and B.
The Jaccard index ranges from 0 to 1, where J = 0 indicates no overlap between the sets (complete dissimilarity). J = 1 suggests that the sets are identical (complete similarity).
In summary, the Jaccard index quantitatively measures the similarity between sets. It is a valuable tool for assessing stability in feature selection algorithms, clustering methods, or any scenario where comparing the overlap between different subsets of elements is relevant.
The subset of features generated to support the classification of epileptiform events was evaluated to determine AdaptDiab’s reliability. First, the AdaptDiab method was used ten times to create relevant features with ten random samples from the dataset. Then, the ten subsets generated were compared according to the Jaccard index to determine their difference. For ten executions, the AdaptDiab method obtained the same subset of relevant features, which means that the stability measured by the Jaccard index is perfect in 100% of the cases evaluated. Considering the previous, it is concluded that at least for datasets with complete and correctly balanced data, such as the one used in this test, the AdaptDiab method achieved 100% stability.
This work demonstrates that the ensemble feature selection method selects the same relevant features (100% stability) across ten executions with different random data samples. This stability ensures that the identified features are genuinely informative and not random selections that might vary significantly with small data fluctuations. This reliability is essential for building robust and generalizable machine-learning models for epileptiform event classification.
Unstable feature selection methods can lead to highly variable sets of features depending on the specific data sample used. This variability can hinder the model’s generalizability, meaning it might perform well in the particular training data but not on unseen data. The stability of our approach helps mitigate this issue by consistently selecting a core set of informative features, leading to models that are likely to perform well on new data. When the AdaptDiab consistently identifies the same features, it strengthens our confidence in their importance for classifying epileptiform events.
In summary, the high stability of our AdaptDiabtechnique ensures reliable feature selection, reduces variability in model performance, and increases our confidence in the identified features for accurate classification of epileptiform events. This stability is a significant advantage of our approach compared to methods that might exhibit high variability in feature selection.
Unlike traditional feature selection methods that may favor specific models or metrics, AdaptDiab consistently enhances the performance of five distinct machine learning models across all evaluation metrics.
Conclusion and future work
In this paper, we have introduced AdaptDiab, an ensemble of feature selection, which combines subsets of features selected by different filters, ANOVA F-score, Fisher score, variance threshold, and Point biserial that yields an optimal subset of features. To evaluate the performance ofAdaptDiab, we have used different classifiers, Gaussian naive Bayes, Logistic Regression, Random forest, Gradient boosting, and Multilayer perception on the Pima Indian Medical Association Indians diabetic dataset. From our average classification accuracy (ACA) analysis, it can be observed that the proposed AdaptDiab technique mostly overcomes the local optimal problem of the individual filters, especially for high-dimensional datasets. The excellent stability achieved by the AdaptDiabtechnique shows consistency in feature selection, robustness to variation, generalizability, and reduced overfitting.
AdaptDiab can be used for Improved Diagnosis and Screening, Personalized Treatment Plans, Integration into Clinical Decision Support Systems, and Educational and Training Use. Its model-agnostic nature makes AdaptDiab applicable to diverse datasets, ensuring its utility in large-scale healthcare systems or other complex data domains. AdaptDiab reduces false positives/negatives by improving predictive accuracy, avoiding unnecessary medical tests or treatments, and saving resources. While designed for diabetes classification, AdaptDiab has potential applications in other fields like agriculture (predicting crop diseases), material science (selecting critical parameters in experimental studies), and finance (identifying key factors influencing credit risk).
Future work will explore methods for feature interpretation, visualization, and domain-specific validation to understand the underlying factors driving predictive performance. Investigate the role of ensemble diversity in the ensemble feature selection technique and its impact on performance and stability.
Data availability
The dataset used in this study was already available in the Internet resources.https://www.kaggle.com/datasets/uciml/pima-indians-diabetes-database.
References
Centers for Disease Control and Prevention. National Diabetes Statistics Report (2023). https://www.cdc.gov/diabetes/php/dataresearch/?CDC_AAref_Val=https://www.cdc.gov/diabetes/data/statistics-report/index.html
AlZu’bi, S. Diabetes monitoring system in smart health cities based on Big Data intelligence. Future Internet (2023).
Hao, J., Wang, C. & Yang, G. Annealing genetic GAN for imbalanced web data learning. IEEE Trans. Multim. (2021). https://doi.org/10.1109/TMM.2021.3120642.
Jiang, Z., Zhao, L. & Lu, Y. A semi-supervised resampling method for class imbalanced learning. Elsevier Expert Syst. Appl. (2023). https://doi.org/10.1016/j.eswa.2023.119733.
Hongfang, Z. A diabetes prediction model based on Boruta feature selection and ensemble learning. BMC Bioinform. (2023).
Francesco & Prendin Machine Learning-Based Feature Selection Techniques for Diabetes Prediction (Current Medical Research and Opinion, 2021).
Chen, C., Zhou, K., Wang, Z., Zhang, Q. & Xiao, R. All answers are in the images: A review of deep learning for cerebrovascular segmentation. Comput. Med. Imaging Graph. 107, 102229. https://doi.org/10.1016/j.compmedimag.2023.102229 (2023).
Han & Kamber. Feature Selection for Knowledge Discovery and Data Mining (MIT Press, 1999) Chap. 6.
Salton & McGill. Introduction to Information Retrieval (McGraw-Hill, Chap. 11, 2005).
Aha, D.W. & Bankert, R.L. A Comparative Evaluation of Sequential Feature Selection Algorithms (1996).
Kuhn, M. & Johnson, K. Feature Engineering and Selection: A Practical Approach for Predictive Models, Chap. 11 (2021).
Härdle, W.K. & Prastyo, D.D. Embedded Predictor Selection for Default Risk Calculation: A Southeast Asian Industry Study, Chap. 7.
Zou, H. & Hastie, T. Regularization and variable selection via the elastic net (2004).
Saeys et al. Ensembles for Feature Selection: A Review and Future Trends (Springer, 2016).
Mohsen & Moshki. Scalable feature selection in high-dimensional data based on GRASP. Appl. Artif. Intell. (2015).
Chawla, N.V. SMOTE: Synthetic minority over-sampling technique. J. Artif. Intell. (2002).
UCI Machine Learning Repository. https://archive.ics.uci.edu/dataset/34/diabetes
Keselman, H.J., Kowalchuk, R.K., Laureen, L.L., Petoskey, M.D., Keselman, J.C. & Levin, J.R. An Analysis of Their ANOVA, MANOVA (2002).
Hastie, T., Tibshirani, R. & Friedman, J. The Elements of Statistical Learning (2009).
Guyon & Elisseeff. An introduction to variable and feature selection. J. Mach. Learn. Res. (2003).
Glass, V. & Hopkins, K.D. Statistical methods in education and psychology. Gene (1995).
Seijo-Pardo, B., Porto-Díaz, I., Bolón-Canedo, V. & Alonso-Betanzos, A. Ensemble feature selection: Homogeneous and heterogeneous approaches. Knowl.-Based Syst. 118 (2017).
Jaccard, P. ‘Etude comparative de la distribution florale dans une portion des alpes et des jura’ (1901).
Acknowledgements
Nil.
Funding
No funding was provided for this research work.
Author information
Authors and Affiliations
Contributions
Natarajan K, Dhanalakshmi Baskaran, and Selvakumar Kamalanathan Authors are done equal contribution on this research work.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
This work was approved by the “NIT Tiruchirappalli.” All procedures performed in this study followed the ethical standards of the institution and national research committee, as well as its later amendments or comparable ethical standards. Also, informed Consent was obtained from all participants before their inclusion in the study.
Consent for publication
The dataset used in this study was benchmark data. Hence, no issues cover identifiable information such as images, personal details, or any other information that could potentially identify a participant.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Natarajan, K., Baskaran, D. & Kamalanathan, S. An adaptive ensemble feature selection technique for model-agnostic diabetes prediction. Sci Rep 15, 6907 (2025). https://doi.org/10.1038/s41598-025-91282-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-91282-8
Keywords
This article is cited by
-
Probrank: a feature probability estimation-based framework for feature selection and ranking
Evolutionary Intelligence (2026)
-
A hybrid method for fusion cardiac biomarkers and echocardiography videos in the experimental classification of Trypanosoma cruzi infection
BioMedical Engineering OnLine (2025)