Abstract
Papaya (Carica papaya L.) is a commercially important fruit crop cultivated worldwide due to its nutritional and medicinal values. Papaya ringspot disease (PRSD), caused by the papaya ringspot virus (PRSV), poses a significant threat to papaya cultivation, resulting in substantial yield losses. In this study, two independent field experiments were conducted at Bagalkote located in the Northern Dry Zone of Karnataka state of India. The first experiment aimed to identify the optimal planting month for papaya to effectively manage PRSV disease. The second experiment was conducted to determine the susceptible papaya growth stage for PRSV infection. The results revealed that early planting of papaya in June or late planting in March were identified as the most optimal planting times across the majority of growth stages, as they exhibited the lowest disease incidence along with superior growth and yield characteristics compared to other planting months. In contrast, planting during the winter season (September to January) resulted in high disease severity due to an increased aphid population. Conversely, planting during periods of low aphid activity (spring season) delayed disease onset until the monsoon. By the time the aphid population increased, the plants had already progressed beyond the flowering and fruit-bearing stages. In the second experiment, the severity and frequency of foliar symptoms on PRSV-inoculated papaya plants were significantly higher in those inoculated at the early growth stage compared to those inoculated at a later growth stage. This indicates that the early growth stage (up to 180 days after transplanting) is a critical period for PRSV infection, necessitating the implementation of effective disease management measures during this time to minimize disease spread and enhance growth and yield. Furthermore, plants inoculated at the early stage exhibited a higher viral titer, more severe symptoms, and a higher percent transmission rate compared to those inoculated at a later stage. These findings were supported by qRT-PCR analysis, which demonstrated a highly significant and positive correlation between early inoculation and disease severity.
Similar content being viewed by others
Introduction
Carica papaya L. belongs to the family Caricaceae, commonly known as “papaya.” India is the largest and leading producer of papaya in the world, accounting for 44.04% of global production (6.05 Mt) and grown in a total area of 1.49 lakh ha with a productivity of 40.6 t/ha1. Papaya is susceptible to a range of pathogens, including viruses, bacteria, and fungi, which can lead to significant yield losses2,3,4,5,6,7,8,9,10. More than 29 different viruses have been reported to infect papaya worldwide. Of these, the papaya ringspot virus (PRSV) and the papaya leaf curl virus (PaLCuV) have gained global importance in all papaya-growing countries11,12,13,14,15,16,17,18,19,20,21,22. PRSV is a major constraint to papaya production not only in India but also in other tropical and subtropical regions of the world, including Southeast Asia (e.g., Thailand, Vietnam), Africa, the Central and South Americas (e.g., Brazil, Mexico), and the Pacific. The virus causes significant yield losses and reduces fruit quality, leading to economic losses for farmers. Understanding the epidemiology of PRSV and developing effective management strategies are critical for sustaining papaya production globally. PRSV is a major concern in India as it has the potential to cause yield loss up to 100%23. Papaya ringspot disease (PRSD) was first reported on the island of Oahu, Hawaii, which was well-known for papaya production24, where it was known as the “Wailu” disease25. Later, it was found to be of a viral nature and termed as papaya ringspot (PRS) by Jensen26. This disease was first reported in India from Pune in 194827and has since been reported from various parts of the country28,29,30,31,32,33.
PRSV belongs to the genus Potyvirus within the family Potyviridae and has two predominant pathotypes: PRSV-P, which infects both papaya and cucurbits, and PRSV-W, which infects only members of the Cucurbitaceae family11,34,35,36,37. Its genome comprises a monopartite, linear, single-stranded, positive-sense RNA of approximately 10,341 nucleotides, with a genome-linked viral protein (VPg) at the 5’ terminus38,39. Symptoms induced by PRSV infection on papaya typically appear two to three weeks after infection, which include an intense yellow mosaic on the leaves, shoestring-like deformities in new emerging leaves, dark green and slightly sunken rings on the fruit, numerous oily-looking streaks on the stem, and overall plant stunting. Fruits formed after infection are usually small and exhibit lichen-like lesions, ring spots with uneven bumps, and an unpleasant taste11. When plants are infected at an early age (pre-vegetative stage), they remain stunted and fail to produce fruit23,40. The severity of symptoms correlates with the titer of PRSV infection, with higher viral loads leading to more severe symptoms41. PRSV is primarily transmitted by aphid vectors in a non-persistent manner. It can also be mechanically transmitted from papaya to papaya under natural conditions42,43,44,45.
Epidemiological studies are more relevant as they reveal the dynamics of disease incidence in relation to climate46. The rate of virus spread has been shown to depend on various factors, including the season, age of the plant, variety, weather, vector population, size, and shape of the cropped area, and surrounding host crops33,46,47,48,49. Effective management of PRSV requires a comprehensive understanding of vector dynamics, including monitoring vector population, diversity, and incidence rates to implement timely and effective disease management practices49,50. Epidemiological strategies, provide a foundation for developing alternative, sustainable methods to mitigate the disease under field conditions51,52. While earlier research has investigated the influence of planting dates on PRSV management, present study provides a unique perspective by correlating PRSV incidence with weather parameters and analyzing viral titers across different crop growth stages. This approach offers deeper insights into how climatic factors interact with disease dynamics, facilitating the development of more tailored management strategies for PRSV.
The concept of planting date manipulation has been extensively studied and proven to be a highly effective strategy for managing viral diseases like soybean yellow mosaic virus, maize dwarf mosaic virus, zucchini yellow mosaic virus, and potato virus Y in several crops, including soybean, maize, cucumber, and potato53,54,55,56,57,58. By adjusting the planting schedule, crops can potentially escape peak disease periods and vector activity, leading to lower disease incidence and severity. For instance, delayed planting of cucumber has been shown to reduce exposure to peak aphid activity55, thereby minimizing the risk of non-persistent virus transmission. In the case of papaya ringspot virus (PRSV) in papaya, optimizing planting schedules to avoid periods of high aphid populations and favorable transmission conditions can significantly reduce disease pressure. Integrating planting date strategies with integrated pest management (IPM) practicessuch as the use of resistant varieties, vector control measures, and cultural practices—provides a more comprehensive and sustainable approach to disease management. Additionally, correlating climatic data with disease incidence and viral titers allows for better prediction of outbreaks, enabling timely and proactive interventions. Despite the potential of planting date manipulation in mitigating PRSV incidence in papaya, only a few studies have been conducted in India to evaluate the influence of planting dates, aphid population dynamics, and their interactions with disease occurrence49,59,60,61,62,63.
Not much studies have been conducted to determine the susceptible growth stage of papaya for PRSV infection or to correlate viral titer with the severity of papaya ringspot disease (PRSD) and transmission rates in papaya. However, well-designed studies on susceptibility, viral load, and transmission dynamics have been reported for other crops and viruses, providing valuable insights into virus-host interactions. In rice, the early growth phases (seedling and booting) have been found to be the most susceptible stages for rice yellow mottle virus (RYMV) infection64. Similarly, cucumber mosaic virus (CMV) in peanut and sweet pepper is more devastating and significantly reduces yield when infects the plants at early stages rather than at later stages65,66. The current epidemiological studies primarily focus on quantifying the virus titer in infected plants using qRT-PCR and its correlation with the appearance of symptoms67. Higher viral titers with severe symptom expression have been observed during the early infection stages in crops such as tomato68, cassava69, wheat70, and watermelon71. Also, the quantification of virus titers for viruses such as CMV, cucumber green mottle mosaic virus (CGMMV), and watermelon mosaic virus (WMV) infecting chili, cucurbits, and cucumber, respectively, has been extensively studied using qRT-PCR67,71,72.
Considering the research conducted so far and addressing the existing knowledge gap, the present study aimed to assess the impact of different planting months on PRSV incidence, evaluate growth and yield parameters of papaya, and identify the susceptible growth stage for PRSV infection by correlating it with viral titer.
Methods
Epidemiological studies were conducted at the Haveli experimental station, University of Horticultural Sciences (UHS), Bagalkote, Karnataka, India (16°12′07.0″N, 75°41′08.9″E), situated within the Northern Dry Zone of Karnataka.
Determining the effect of different months of planting on the incidence of PRSD, growth, and yield parameters of Papaya
The field experiments were conducted from 2019 to 2020 under natural disease pressure of PRSD using a Randomized Block Design (RBD) with three replications and ten treatments. Planting was carried out at monthly intervals, starting from the 1st week of June 2019 and continued until the 1st week of March 2020. Details of the treatments are provided in Supplementary Table 1. The susceptible non-genetically modified (non-GM) papaya cultivar ‘Red Lady’ was transplanted with a spacing of 1.5 m × 1.5 m, and all recommended agronomic practices outlined in the package of practices from UHS, Bagalkot73, were followed. Observations on percent disease incidence were assessed based on visual observations of symptoms typical to PRSV infection and confirmed through RT-PCR, using a set of CP gene-specific primers (MB 11A/MB 11B)36. Additionally, plant height (cm), plant girth (cm), number of leaves per plant, internodal length (cm), days taken for first flowering, number of flowers per plant, number of days from flowering to first fruit set, number of days to fruit harvest after fruit set, fruit length (cm), fruit breadth (cm), fruit diameter (cm), cavity diameter (cm), yield per plant (kg), and yield per hectare (t) were recorded at 30 day intervals, from 30 days after transplanting (DAT) to 270 DAT. Additionally, Area Under the Disease Progress Curve (AUDPC) analysis for percent disease incidence of PRSD was also carried out to assess the intensity and progression of the disease over a crop’s growth period using the formula74.
where, yi and yi+1 are disease incidence percentages at times ti and ti+1, respectively, ti and ti+1 are the respective time points in days and n is the total number of observations.
The population dynamics of aphids were regularly monitored using yellow sticky traps installed at two different locations, 50 m apart. The number of aphids collected was recorded weekly, and the cumulative aphid population was calculated monthly. Additionally, daily weather data were obtained from the automatic weather station located at UHS, Bagalkote, Karnataka, India (16°12′07.0″N, 75°41′08.9″E). The recorded weather parameters included daily mean, high, and low temperatures (°C), mean relative humidity (%), solar radiation (W/m²), wind speed (m/s), and rainfall (mm). Further, correlation coefficients between PRSD incidence, aphid population, and various growth and yield parameters were analyzed at different growth stages of papaya, specifically at 30, 60, 90, 120, 150, 180, 210, and 240 DAT.
Identification of the susceptible stage of papaya for PRSV infection
The study was conducted during 2019–2020 under a screen house to determine the susceptible growth stage of papaya (cv. Red Lady) for PRSV infection (Suppl. Table 2) using viruliferous aphids, which transmits the virus in a naturally, efficiently, and rapidly in a non-persistent manner43,44,45. Aphis craccivora Koch. major vector of PRSD which was identified to be associated with papaya orchards in Karnataka was individually collected from papaya orchards and maintained under an insect-proof nylon mesh cage on cotton (Gossypium hirsutum cv. Varalakshmi) (Suppl. Figure 1). Since, the average lifespan of an aphid is approximately one month75, aphids from the maintained colony were used transmission after one month of colony establishment to rule out the possibility of PRSV contamination from field collected aphihids. Before virus acquisition, these non-viruliferous aphids were starved for approximately one hour and then allowed to feed on PRSV-infected papaya plants for a brief acquisition access period (AAP) of 30 min. Given the short retention time of PRSV on aphids, which ranging from a few seconds to minutes, viruliferous aphids 15 per plant and 60 per each replication in a treatment which contain 4 plants were subsequently transferred and kept in the insect proof net cages and given an inoculation access period (IAP) for 30 min. The experiment was conducted under insect-proof polyhouse conditions to prevent external contamination and ensure controlled virus transmission76.
To ensure uniformity, all plants were grown under uniform environmental and cultural conditions, allowing for comparability across treatments and replicates. Within each treatment group, all plants were transplanted on the same day, ensuring synchronization of growth stages across replicates. This approach maintained consistency in observations and minimized variability due to external factors, ensuring reliable and reproducible results. For each replication of the treatments, the plants were covered with an insect-proof nylon mesh of 40X gauge to prevent the spread of viruliferous aphids. Observations on percent transmission, days taken for symptom expression after post-inoculation (dpi), typical symptoms induced, plant height (cm), plant girth (cm), number of leaves per plant (cm), internodal length (cm), days taken for first flowering, number of flowers per plant, number of days from flowering to first fruit set, number of days from fruit set to harvest after fruit set, fruit length (cm), fruit breadth (cm), fruit diameter (cm), cavity diameter (cm), yield per plant (kg), and yield per hectare (t) were recorded.
Data from all experiments were analyzed using analysis of variance (ANOVA) to determine the significance of treatment effects at a 5% probability level. Post-hoc comparisons were conducted using Tukey’s Honestly Significant Difference (HSD) test to identify significant differences among treatment means. Additionally, correlation analysis was performed to assess the relationships between disease incidence, growth parameters, and aphid population dynamics. All statistical analyses were carried out using OPSTAT online software (http://14.139.232.166/opstat/)77, and significant differences are indicated in both figures and tables. Results are presented with standard errors (S.Em ±) and critical differences (CD @ 5%) for each parameter.
Quantification of PRSV titer at different growth stages of the crop using qRT-PCR
The qRT-PCR analysis was conducted using the Applied StepOnePlus Real-Time PCR System to evaluate the PRSV titer in papaya plants. Sampling was performed 15 days after inoculation (DAI) for each treatment, as this timeframe corresponds to the onset of classical symptoms in PRSV-infected papaya plants10. Newly emerging young, symptomatic leaves from PRSV-infected plants and asymptomatic uninoculated virus-free plants maintained under insect-proof net house conditions were collected, placed in labeled polythene bags, snap-frozen in liquid nitrogen, and stored at −80 °C for subsequent RNA quantification.
Total RNA was extracted from 100 mg of leaf tissue using the Spectrum™ Plant Total RNA Kit (Sigma-Aldrich, Catalog No. STRN50) following the manufacturer’s protocol. The extracted RNA was treated with DNase I (Thermo Fisher Scientific) to remove genomic DNA contamination and then quantified using a NanoDrop spectrophotometer. Total RNA was extracted from the collected leaf samples and quantified, followed by cDNA synthesis for qRT-PCR analysis. To ensure reliable quantification, uninoculated, virus-free plants were used as negative controls, while fully symptomized plants from early stages served as positive controls. Synthesized cDNA was subjected to qRT-PCR using TB Green Premix Ex Taq II (Tli RNaseH Plus, Cat. #RR820A) and specific primers for PRSV (PRSV2F and PRSV2R), which were designed based on sequence data obtained from amplification of the HC-Pro region of the PRSV-BGK isolate (GenBank accession: OL677454) using NCBI Primer Blast (https://www.ncbi.nlm.nih.gov/tools/primer-blast/index.cgi) online tool. Additionally, the TATA-binding protein 2 (TBP2F and TBP2R) was used as a candidate reference gene, as per Zhu et al.78. Details of the primers and PCR conditions are provided in Table 1. This methodological approach ensured precise quantification of PRSV titers, enabling accurate assessment of the virus’s impact at various growth stages.
Results
Determining the effect of different months of planting on the incidence of PRSD, growth and yield parameters of Papaya
PRSD incidence
Planting papaya (cv Red Lady) in different months significantly influenced the incidence PRSD (p < 0.05). Plants established in June showed no incidence of PRSD up to 90 DAT, with a minimal incidence of 8.33% at 120 DAT, eventually reaching 100% at 240 DAT (Suppl. Table 3, Fig. 1). Similarly, July planting recorded the first incidence of PRSD (11.11%) at 90 DAT and reached 100% at 210 DAT. The progression to 100% incidence varied across planting months: it took 180 days for August and February plantings, 120 days for September planting, 90 days for October, November, and January plantings, and only 60 days for December planting. In contrast, March planting exhibited a significantly lower incidence of PRSD at all stages. The first incidence (5.56%) in March planting was recorded at 60 DAT, and it required 270 DAT to reach 100% incidence, demonstrating a slower disease progression compared to all other planting months.
AUDPC analysis
Planting papaya in June recorded the lowest AUDPC value (8499.90), indicating the least incidence and progression of PRSD, followed by March planting, which also showed relatively lower disease pressure (9166.80) (Suppl. Table 4, Fig. 2). In contrast, plantings during the October to January months exhibited the highest AUDPC values, with December having the highest peak value (23666.70). This indicates that planting during this period resulted in the most severe PRSD. The AUDPC reveals a seasonal trend where plantings during the winter months (October to January) in the study region experienced higher disease pressure compared to those planted in the early monsoon (June) and early spring (March) months. Thus, adjusting the planting schedule to avoid peak disease months can significantly reduce the incidence and severity of PRSD, enhancing crop health and yield potential.
Growth parameters
The plant height, internodal length, plant girth, and leaves per plant at various growth stages showed significant differences (p ≤ 0.05) among the treatments for all stages. March planting showed significantly superior plant height compared to all other treatments at every growth stage. At 270 DAT, March planting (227.19 cm) had the highest plant height, while the least plant height was observed with January planting (132.13 cm) (Suppl. Table 5). March planting recorded significantly superior internodal length compared to all other treatments, while January and December plantings showed the least internodal length at all growth stages (Suppl. Table 6). At 270 DAT, March planting (41.47 cm) recorded the highest plant girth, while January planting (36.19 cm) recorded the least (Suppl. Table 7). Similarly, March planting also had significantly more leaves per plant, ranging from 6.80 to 29.72 at all growth stages (Suppl. Table 8).
Yield and yield parameters
The number of days taken for the first flowering was significantly lower in March planting (89.33 days), followed by June (91.11 days) and July (91.56 days) plantings, while it was maximum in the January planting (101.89 days) (Suppl. Table 9). Similarly, a significantly higher number of flowers per plant, the least number of days taken from flowering to the first fruit set (91.97 days), the highest number of days taken for the first fruit set to harvest (122.41 days) were recorded at all growth stages of March planting, followed by June planting compared to all other planting dates (Suppl. Tables 9 & 10).
Furthermore, March planting recorded a significantly higher number of fruits per plant (25.92), the highest diameter of the fruits (12.31 cm), the highest length of the fruits (29.92 cm), the highest breadth of the fruits (14.63 cm), and a larger cavity diameter (6.97 cm) compared to other treatments. Correspondingly, January planting recorded the lowest number of fruits (7.86), the smallest diameter of the fruits (8.14 cm), the smallest fruit length (19.75 cm), and the smallest cavity diameter (5.91 cm) (Suppl. Table 9).
Yield per plant and yield per hectare were significantly highest in March (60.12 kg/plant and 185.54 t/ha) planting compared to all the treatments. Planting in June (55.46 kg/plant and 171.15 t/ha) and July (51.23 kg/plant and 158.09 t/ha) months was the next best, while it was the least in January (13.84 kg/plant and 42.70 t/ha) planting, followed by December (15.40 kg/plant and 47.53 t/ha) planting (Suppl. Table 9, Suppl. Figure 2, and Fig. 3).
Aphid population dynamics during papaya crop growth stage at monthly intervals
The papayas planted in June, July, and August showed no infestation of aphids at 30 DAT. However, plantings in September (30 No.), October (94 No.), November (111 No.), December (192 No.), January (344 No.), February (329 No.), and March (31 No.) showed a moderate aphid population at 30 DAT (Suppl. Table 11). The plantings in June, July, and August had the lowest aphid population (124, 235, and 427, respectively) up to 150 DAT, after which a sudden increase was observed (235, 427, and 771, respectively, at 180 DAT). On the other hand, the plantings from September to February had initially high aphid activity, but after 150 DAT, aphid activity was recorded as low. The March planting had an initial aphid load, but the cumulative aphid population at each growth stage was lower than that of all the other planting months.
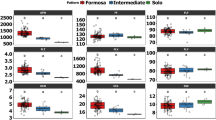
The environmental conditions, monitored daily during the trial period (Suppl. Table 12 and Fig. 4) showed significant seasonal variability, which has likely influenced aphid activity and PRSV incidence. From June to August, mean daily temperatures ranged between 27.4 °C and 31.8 °C, accompanied by high relative humidity (65 to 90%), moderate solar radiation (~ 200 W/m²), and wind speeds varying from 3.0 to 8.6 m/s, occasionally peaking at 9.0 m/s. Frequent rainfall, with several days recording over 20 mm of precipitation, was noticed during this period. These cool, humid, and windy conditions likely suppressed aphid activity and reduced viruliferous rates, correlating with the low PRSV incidence observed during this time.
In contrast, the months from October to January experienced mean daily temperatures stabilizing between 24 °C and 30 °C, with lower relative humidity (40 to 70%), increased solar radiation (exceeding 250 W/m²), reduced wind speeds, often below 3 m/s and as low as 0.2 to 1.6 m/s on many days. Minimal rainfall during these dry and warm months likely created favorable conditions for aphid activity and the efficient spread of PRSV. This shift in environmental factors corresponded with the higher disease incidence recorded during this period. These findings highlight the strong influence of environmental conditions on vector behavior and PRSV transmission, emphasizing the importance of aligning planting schedules with periods unfavorable to aphid activity to reduce disease incidence.
Correlation studies between disease incidence with aphid population, growth, yield, and yield parameters at different crop growth stages
The correlation between the incidence of PRSD, aphid population, growth, and yield parameters were analyzed at different growth stages (i.e., 30 DAT to 240 DAT) (Fig. 5). The incidence of PRSD showed a significant and strong positive correlation with the aphid population in all growth stages except at 30 DAT, where it was non-significant (0.501). Additionally, the days to first flowering (up to 120 DAT) and first fruit set (up to 210 DAT) showed a strong positive correlation with yield.
The remaining growth and yield attributes showed a negative correlation. Fruit length, fruit breadth, and fruit cavity diameter exhibited significant correlations up to 210 DAT. Yield per plant, fruits per plant, and fruit diameter displayed a strong negative correlation up to 180 DAT. Days from fruit set to harvest showed strong negative significance, limited to 150 DAT. The number of leaves per plant also exhibited a negative correlation with the incidence of PRSD up to 120 DAT. Plant height, internodal length, and plant girth showed no significant correlation with the incidence of PRSD at a few growth stages. Simultaneously, the number of flowers was recorded as non-significant from the beginning (at 150 DAT) to the end of flowering (at 240 DAT).
Identification of susceptible stage of papaya for PRSV infection
Percent transmission and days taken for symptom expression
PRSV inoculation at 30 and 60 DAT has shown early symptomatic expression, with symptoms starting to appear on the 15th DAI with 100% transmission. The plants inoculated at 60 DAT also showed 100% transmission, but it took 17 days for symptom expression to occur. Subsequent inoculations at 120, 150, and 180 DAT recorded a gradual decrease in the transmission rate (75.0%, 58.0%, and 25.0%, respectively) and a delay in symptom expression (20th, 24th, and 25th DAI, respectively). Furthermore, plants inoculated at 210, 240, and 270 DAT had the lowest transmission rate (8.33%) and a further delay in symptom expression (25th, 26th, and 28th DAI, respectively) (Table 2; Fig. 6).
Regarding the type of symptoms expressed, plants inoculated at 30, 60, and 90 DAT showed the most severe symptoms, such as vein clearing, mosaic, leaf reduction, leaf distortion, shoestring, stunted growth, mottling, chlorotic ring, and chlorotic spots on fruits. The plants inoculated at 120 and 150 DAT showed vein clearing, mosaic, leaf reduction, leaf distortion, shoestring, stunted growth, mottling, and chlorotic spots on fruits. Only vein clearing, mosaic, leaf reduction, and leaf distortion were recorded in plants inoculated at 180 DAT. Delayed inoculation at 210 and 240 DAT showed a reduced severity of symptoms, with the expression of vein clearing and mosaic symptoms on leaves. Finally, only vein clearing was recorded in plants inoculated at 270 DAT (Suppl. Figure 3).
Quantification of viral titer in inoculated papaya plants by qRT-PCR
Data regarding the inoculation of PRSV at different growth stages and the corresponding viral titers in plants are summarized in Table 3; Fig. 7. The melting curve and amplification plots for the TATA-binding protein-2 (a housekeeping gene) and PRSV, obtained through qRT-PCR, are shown in Suppl. Figures 4, 5, and 6. The highest viral titer, 615.97, was observed in plants inoculated at 30 DAT, representing the early growth stage. The viral titer recorded for plants inoculated at 60 DAT was 414.78. As the stage of inoculation was delayed, there was a corresponding decrease in the viral titer. Specifically, plants inoculated at 90, 120, 150, and 180 DAT had viral titers of 302.36, 235.09, 210.42, and 105.65, respectively. Further delays in inoculation resulted in even lower viral titers, with 72.58 at 210 DAT, 44.89 at 240 DAT, and the lowest titer of 9.86 recorded in plants inoculated at 270 DAT. The overall result indicates that the early growth stages of papaya are more susceptible to PRSV infection, with higher viral titers observed when inoculated earlier in the plant’s development. Delaying inoculation reduces the viral load, indicating a lower susceptibility as the plant matures.
Effect of PRSV inoculation at different growth stages of Papaya on growth and yield
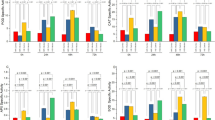
The results of PRSV inoculation at different growth stages demonstrated significant differences in the growth and yield parameters of papaya (Suppl. Table 13). As inoculation was delayed, there was a linear increase in growth and yield parameters, including the number of flowers per plant, the number of days to first fruit set and harvest, the number of fruits per plant, fruit diameter, length, breadth, cavity diameter, and overall yield. In contrast, a linear decrease was observed in the number of days from first flowering to the first fruit set and from first flowering to harvest. The highest plant height was recorded in the uninoculated control plants (242.07 cm), which was comparable to the height of plants inoculated at 270 DAT (241.72 cm). The lowest plant height was observed in plants inoculated at the early stages, specifically at 30 DAT (145.69 cm), followed by plants inoculated at 60 DAT (168.58 cm) (Fig. 8). These findings indicate that delaying PRSV inoculation allows papaya plants to maintain better growth and higher yield parameters. Early inoculation, particularly at 30 and 60 DAT, resulted in significantly stunted plant growth and development.
The plants inoculated at 30 DAT recorded shorter internodal length (4.03 cm), the lowest plant girth (33.74 cm), the lowest number of leaves (28.70), the highest number of days taken for the first flowering (105.04), the lowest number of flowers per plant (9.21) (Fig. 9), and the highest number of days taken from flowering to the first fruit set (91.56). Conversely, control plants recorded the highest internodal length (4.26 cm), the highest plant girth (43.64 cm), the highest number of days taken for the first flowering (93.68), the highest number of flowers per plant (23.49), and the least number of days taken from flowering to the first fruit set (85.81).
Similarly, plants inoculated at 30 DAT had the fewest days from the first fruit set to harvest, the lowest number of fruits per plant (1.75) (Fig. 10), the smallest fruit diameter and cavity diameter (8.95 cm and 6.11 cm), and the shortest length of the fruit (21.21 cm), and the smallest fruit breadth (9.73 cm), while control plants recorded the longest number of days (119.72), the largest number of fruits per plant (38.50), the largest fruit diameter and cavity diameter were recorded in the control plants (13.07 cm and 7.30 cm), and the longest length of the fruit (31.78 cm), the largest fruit breadth (15.28 cm).
The highest fruit yield per plant and yield per hectare were recorded in the uninoculated control (59.39 kg/plant and 183.28 t/ha), which was comparable to that in plants inoculated at 270 DAT (58.64 kg/plant and 180.95 t/ha) and 240 DAT (57.52 kg/plant and 177.51 t/ha). On the other hand, the lowest yield per plant was recorded in plants inoculated at 30 DAT (1.92 kg/plant and 5.94 t/ha), which was similar to that in plants inoculated at 60 DAT (2.22 kg/plant and 6.84 t/ha) (Fig. 11).
Correlation studies between papaya ringspot viral titer in Papaya plant with percent transmission, growth, and yield parameters
The correlation between the papaya ringspot viral titer in the host and percent transmission, growth, yield, and yield parameters was analyzed and presented in Fig. 12. The quantity of papaya ringspot viral titer had a strong positive correlation with percent transmission (0.917), which is highly significant. All growth attributes, such as the number of leaves per plant (−0.947 and −0.962), plant girth at 270 days (−0.943 and −0.900), internode length (−0.882 and −0.881), and height (−0.985 and −0.928), showed highly significant negative correlations with both the viral titer and percent transmission. Similarly, all flower and fruit parameters, including the number of flowers per plant (−0.994 and −0.934), days to first fruit set to harvest (−0.957 and −0.963), number of fruits per plant (−0.917 and −0.986), fruit diameter (−0.973 and −0.918), length (−0.948 and −0.973), breadth (−0.936 and −0.978), and cavity diameter (−0.953 and −0.949), also recorded highly significant negative correlations with the viral titer and percent transmission. On the other hand, the number of days to first flowering (0.988 and 0.866) and the number of days from flowering to first fruit set (0.866 and 0.949) had positive and highly significant correlations. Finally, the fruit yield showed a highly significant negative correlation between both the viral titer (−0.942) and percent transmission (−0.995).
Discussion
Papaya ringspot virus (PRSV) is a serious pathogen affecting papaya cultivation, particularly in tropical and subtropical regions, leading to substantial yield losses, reduced fruit quality, and increased management costs. Effective management requires a multifaceted approach that includes understanding environmental factors, optimizing planting timings, and potentially utilizing resistant cultivars. In the present study, two independent field experiments were conducted to identify the optimal planting month for papaya and determine the papaya growth stage, that is most susceptible to PRSV infection. The results indicated that no disease incidence was observed up to 90 DAT in June-planted papaya, and up to 60 DAT in July-planted papaya. Additionally, June planting recorded the lowest AUDPC value (8499.90). This can be attributed to environmental conditions, from June to August, which was not conducive to the proliferation of both the virus and its aphid vectors. The reduced aphid population during this period likely contributed to the low disease incidence. The June planting reached peak disease incidence at 240 DAT, while subsequent plantings took progressively fewer days to reach peak incidence. The December planting, reached peak incidence within just 60 DAT.
The increased PRSV incidence observed from October to January, with December planting showing the highest cumulative disease pressure (23666.7), may be linked to the availability of various crops in surrounding areas, including mungbean, urdbean, cowpea, horse gram, red gram, cucurbits, and chili, which served as hosts for aphid vectors. Additionally, the collected weather data underscore the significant role of environmental conditions in influencing PRSV incidence and vector dynamics, highlighting the importance of managing these factors in disease management strategies.
From June to August, daily mean temperatures ranged from 27.4 °C to 31.8 °C, accompanied by high relative humidity (65 to 90%), moderate solar radiation (approximately 200 W/m²), and wind speeds ranging from 3 to 8.6 m/s. These conditions likely reduced aphid activity and viruliferous rates, correlating with the low PRSV incidence observed in June-planted papaya. Additionally, frequent rainfall during the early phase of development, particularly with daily totals occasionally exceeding 20 mm, further suppressed aphid populations, contributing to the absence of disease incidence. Higher wind speeds also disrupted aphid movement and feeding behavior, which reduced virus transmission rates and contributed to the lower disease incidence observed during this period49.
In contrast October to January, coinciding with winter, aphid population peaked, reaching maximum activity in January, which resulted in increased disease incidence in the later stages of growth. During this period, environmental conditions shifted, with mean temperatures stabilizing between 24 °C and 30 °C, relative humidity decreasing to 40 from 70%, solar radiation increasing to above 250 W/m², and wind speeds dropping significantly (often below 3.0 m/s, with some days as low as 0.2 m/s). These dry, warm conditions likely created an optimal environment for aphid proliferation. Aphids thrive in this temperature range, and the lower relative humidity diminished the effectiveness of natural predators and pathogens that typically control aphid populations49,79. Increased solar radiation promoted plant growth, providing abundant food for aphids, while reduced wind speeds facilitated aphid settlement, activity, and reproduction, enhancing virus transmission efficiency, leading to higher PRSV incidence in later-planted crops49.
Although the March planting exhibited mild disease incidence during early phase, it took 210 days to reach peak incidence of 100%, with relatively lower disease pressure (9166.80). Aphid activity was recorded during the March planting but gradually decreased, leading to reduced disease spread, due to extreme weather conditions. These findings underscore the importance of environmental factors in influencing vector dynamics and disease progression, emphasizing the need for region-specific strategies to mitigate PRSV incidence and improve yield. Similarly, several studies also demonstrate that temperature significantly influences aphid population dynamics. Moderate cool temperatures promote population growth, while extremes in temperature negatively affect survival and reproduction80,81,82. Hence, early planting (June) or late planting (March) of papaya was identified as the most optimal planting months across the majority of growth stages, as it exhibited the lowest disease incidence along with superior growth and yield characteristics compared to other planting months, highlighting the role of environmental factors in shaping PRSV dynamics and vector population. Future studies incorporating single-aphid qPCR could also provide a clearer understanding of the relationship between aphid populations, virus transmission efficiency, and environmental conditions. Even though, the data generated provides a brief insight into dynamics of PRSV incidence, a long-term studies will enhance by validating the robustness and consistency of findings across the different geographical regions of the world. Which can be utilized for designing the date of sowing based management strategies for different regions.
This study focused on the impact of planting dates and environmental factors on PRSV incidence under controlled field conditions. However, farmers’ agricultural practices, such as fertilization, irrigation and pest management, significantly influence disease dynamics in real-world conditions. Standardized practices were used to minimize variability and isolate the effects of planting dates and environmental factors.
The incidence of PRSV exhibited a significant positive correlation with the aphid population across growth stages, while other growth and yield attributes showed a negative correlation. Since this study was conducted under natural field conditions with viruliferous aphids, the findings reflect the combined influence of aphid population dynamics and virus transmission.
Similarly, Chandrashekar et al.62 recorded the peak aphid population in January, which sharply declined by March and remained low from March onwards until September. It then progressively reached its peak again by January in papaya. This resulted in the maximum (100%) incidence of PRSV observed in papaya planted from September to January, and the incidence of PRSV showed a positive correlation with the aphid population under Pune conditions. Correspondingly, no deviation in the month-wise pattern in the dynamics of the aphid population was observed during the present experiment in Bagalkote.
The findings of Chavan et al.83, which documented a maximum aphid population (10.47/plant) during December on papaya plants in Pune, Maharashtra, India, are in line with the present investigations. Many researchers also noticed the maximum population of aphids from November, reaching its peak during December and January84,85. Additionally, planting during the spring season, characterized by a lower aphid population, leads to a delayed incidence until the monsoon period. By the time the aphid population begins to build up, the plants have already passed the flowering and fruit-bearing stages. These results are consistent with the findings of Sharma et al.61. The optimal time for papaya planting was determined to be in March, followed by June. To minimize PRSV damage, it is recommended to avoid planting during the months of September to January under the Northern Dry Zone of Karnataka, India.
This conclusion is supported by the research of Mora-Aguilera et al.59, who showed that the aphid population had a bimodal distribution, with the highest population peak occurring from December to February and a secondary peak from August to September. They demonstrated that the incidence of PRSV disease in papaya was influenced by the aphid population of the previous month. However, Thiribhuvanamala et al.63reported that papaya planted between August and December had the lowest incidence of PRSV (10 to 20%) compared to crops planted in February and March. This is attributed to the influence of weather parameters on the population dynamics of different aphid species in that location48.
The decreased plant height and internode length are responses to alterations in genomic regions related to virus replication, movement, and processes regulating virus accumulation in infected plants86,87,88,89,90,91,92,93,94. Similarly, Sheikh et al.95 documented a decrease in plant height, number of fruits, fruit length, stem girth, root length, and fruit malformation in papaya plants infected with PRSV.
Disease incidence at the early to middle stages of crop growth significantly and negatively impacted yield. Severely infected plants experience delayed flowering due to viral multiplication96,97. Prakash et al.98 observed that the papaya cultivar ‘PusaNanha’ infected with PRSV exhibited delayed flowering time of 81.50 days compared to healthy plants, which flowered normally in 64.67 days.
Fruiting traits are linked to growth, flowering, and photosynthetic activity, all of which can be affected by viral infections. Viral infections are known to increase the content of malondialdehyde and alter the activities of peroxidase enzymes, resulting in quantitative and qualitative changes in their molecular isoforms99,100.
Altering traditional planting times can help crops avoid viruliferous vectors. The best planting time depends on the timing of vector migration. Mora-Aguilera et al.60reported the usefulness of transplanting dates in controlling PRSV in Central Veracruz, Mexico, through multivariate comparison of its epidemics. However, the effectiveness of changing planting dates to minimize virus infection for a particular crop should be considered in relation to other economic factors101,102.
Plants inoculated at early stages exhibited severe symptoms, including vein clearing, mosaic symptoms, leaf reduction, leaf distortion, shoestring, stunted growth, mottling, chlorotic rings, and chlorotic spots on fruits, and displayed symptoms more quickly. Additionally, a strong and significant positive correlation (0.917) was observed between the viral titer and the percent transmission rate. Quantifying changes in the viral titer to measure the accumulation of viral RNA, replication of viral RNA, and levels of viral and host translation in PRSV-infected papaya plants has been extensively studied41,103,104. Moreover, there have been studies on the antagonistic effects of different viruses on PRSV that employed absolute quantification105. However, research exploring the correlation between viral titer, percent transmission, and disease severity is limited. Hence, this study provides valuable insights into the correlation between viral titer, percent transmission, and disease severity.
Early viral infections exhibit a high replication rate and increased cell-to-cell movement of viral particles, which are crucial steps in initiating and spreading virus infections in plants. This process occurs through intercellular connections known as plasmodesmata and is typically mediated by one or more virus-encoded movement proteins that interact with multiple cellular factors106. Tran et al.107reported that BAM1 is one such cellular factor required for efficient cell-to-cell movement of the TMV movement protein, suggesting that BAM1 interacts with the TMV movement protein to facilitate early virus movement, thereby increasing the number and spread of viral particles. Furthermore, these virus particles manipulate auxin signaling pathways, leading to morphogenic alterations in cells and exacerbating viral symptoms80.
Host plants may acquire or increase their resistance as they grow and develop108,109,110. Wang et al.111 found that wheat lines infected by the barley yellow dwarf virus (BYDV) at the adult stage upregulated certain defense responses, such as ROS scavenging, GA and JA biosynthesis genes. This suppressed BYDV replication, movement, and accumulation. In Arabidopsis and turnip, long-distance transport of the cauliflower mosaic virus (CaMV) occurs in the phloem. During the host’s development, the sink-source relationships change, and the region that CaMV can invade progressively decreases, leading to resistance, restricted migration, and reduced viral titer112.
As the viral inoculation was delayed, the plants reported a lower viral titer and minimum yield loss, as evidenced by the negative and highly significant correlation between yield and viral titer (−0.924). Zhu et al.113 showed that sugarcane plants infected with the sugarcane yellow leaf virus (SCYLV) recorded high virus titers and severe symptoms, such as yellow leaf symptoms, yielding only 54.0 to 60.0% of cane and sugar tonnage compared to plants with low virus titers. Agrios et al.114 noticed that early inoculations had severe symptoms, whereas later inoculations had mild or no symptoms regarding CMV on pepper plants. Furthermore, CMV titers were greater in early-inoculated plants. Levy and Lapidot115, while working with TYLCV, recorded maximum yield loss of tomato in plants inoculated at an early stage, i.e., 14 DAS, compared to plants when inoculated at 45 DAS, confirming the occurrence of age-related (or mature plant) resistance in tomato plants against TYLCV.
In conclusion, the selection of the ‘Red Lady’ cultivar for the present study was based on its prominence as the most commonly cultivated papaya variety in Karnataka, known for its high market value and widespread adoption by local farmers. Given the significant impact of environmental factors, planting dates, and susceptible growth stages on this cultivar, understanding how these variables influence PRSV incidence is crucial for improving disease management practices in the region. While resistant papaya varieties could offer effective control of viral diseases, their adoption has been limited due to factors such as cost, availability and market demand.
The findings of this study have wide-ranging implications for papaya-growing regions worldwide, especially in areas with comparable climatic conditions. Identifying the optimal planting time and susceptible growth stages for PRSV infection can contribute to more effective disease management strategies in regions facing similar challenges. Furthermore, the methodological approach employed in this study, particularly the use of qRT-PCR for viral titer quantification, can be extended to the study of other viral pathogens across different crops. This enhances the study’s scientific relevance, offering valuable insights for global plant virology research and disease management programs.
This study provides valuable insights into disease management strategies specifically tailored to the ‘Red Lady’ cultivar, offering practical recommendations for farmers in Karnataka. Moving forward, research that incorporates susceptible and resistant varieties, alongside the environmental and agronomic factors explored in this study, will be essential for developing integrated disease management strategies. These strategies should consider the diverse farming practices and crop types across Karnataka but also provide a framework for sustainable papaya production in other tropical and subtropical regions, ensuring relevance and accessibility to the global agricultural community.
Conclusions
The results of the current study contribute to our understanding of the influence of planting month and papaya susceptibility to PRSV infection. Early June planting or late March planting of papaya was emerged as a non-chemical, eco-friendly, and effective management strategy against PRSD under field conditions. Farmers should avoid planting from September to January to mitigate disease and yield losses, as this aligns with the high aphid population during winter, particularly in the Northern dry zone of Karnataka. The severity and frequency of foliar symptoms in PRSV-infected papaya plants were notably higher in those infected at early growth stages compared to later stages. Consequently, early viral infections generally result in severe local and systemic symptoms, while later inoculation leads to milder effects. The viral titer of PRSV positively correlated with symptom severity and percent transmission, with higher titers recorded in plants inoculated early in crop growth, showcasing severe symptoms and higher transmission rates than later stages. Thus, the early crop growth stage (up to 180 DAT) emerges as a critical period for PRSV infection, urging effective disease management measures up to this period to optimize growth and yield attributes. This study provides valuable insights into the epidemiology of PRSV and offers practical recommendations for disease management that can be adapted to other papaya-growing regions worldwide. The findings also contribute to the broader scientific understanding of virus-host interactions and the role of environmental factors in disease dynamics, which are critical for sustainable agriculture.
Data availability
All data generated or analysed during this study are included in this published article [and its supplementary information files].
References
Anonymous. FAOSTAT Data Base (2023); accessed 1 September 2023, http://nhb.gov.in/database.
Din, B. N. B. M. Isolation, characterization, and histopathological study of pathogenic bacteria associated with crown rot of papaya (Carica papaya L.) in Peninsular Malaysia. M.Sc. Thesis (2014).
Fullerton, R. A. et al. First record of bacterial crown rot of Papaya (Carica papaya L.) caused by an Erwinia papayae-like bacterium in the Kingdom of Tonga. Plant. Dis. 95, 70. https://doi.org/10.1094/PDIS-06-10-0455 (2011).
Gómez-Godínez, L. J. et al. Isolation and characterization of fungal pathogens associated with Carica papaya L. and their biocontrol with Trichoderma Sp. Agro Product. 17, 185–197. https://doi.org/10.32854/agrop.v17i17.2802 (2024).
Guo, T. et al. Comparison of transmission of papaya leaf curl china virus among four cryptic species of the whitefly Bemisia tabaci complex. Sci. Rep. 5, 15432. https://doi.org/10.1038/srep15432 (2015).
He, Y. et al. First report of anthracnose fruit rot of Papaya caused by Colletotrichum gigasporum in China. Plant. Dis. 108, 810. https://doi.org/10.1094/PDIS-10-23-2162-PDN (2024).
Mustofa, S., Ahad, M. T., Emon, Y. R. & Sarker, A. Papaya leaf: A dataset of Papaya leaf for disease detection, classification, and analysis. Data Brief. 57, 110910. https://doi.org/10.1016/j.dib.2024.110910 (2024).
Rafiq, R. & Fatima, H. Morphological characterization of Sclerotinia sclerotiorum causing fruit rot of Papaya and its management using biopesticides. J. Agric. Biol. 1, 47–52. https://doi.org/10.55627/agribiol.001.02.0652 (2023).
Sneha, K. B., Indra, N., Murugavel, K. & Thangeswari, S. Occurrence of Gilbertella persicaria causing soft rot of Papaya in India. Physiol. Mol. Plant. Pathol. 102481. https://doi.org/10.1016/j.pmpp.2024.102481 (2024).
Wu, Z., Mo, C., Zhang, S. & Li, H. Characterization of papaya ringspot virus isolates infecting transgenic Papaya ‘huanong 1’ in South China. Sci. Rep. 8, 8206. https://doi.org/10.1038/s41598-018-26596-x (2018).
Tripathi, S., Suzuki, J. Y., Ferreira, S. A. & Gonsalves, D. Papaya ringspot virus-P: Characteristics, pathogenicity, sequence variability and control. Mol. Plant. Pathol. 9, 269–280. https://doi.org/10.1111/j.1364-3703.2008.00467.x (2008).
Davis, R. I. & Tsatsia H. A survey for plant diseases caused by viruses and virus-like pathogens in the Solomon Islands. Australas Plant. Pathol. 38, 193–201. https://doi.org/10.1071/AP08114 (2009).
Cabrera Mederos, D., Giolitti, F., Torres, C. & Portal, O. Distribution and phylodynamics of papaya ringspot virus on Carica papaya L. in Cuba. Plant. Pathol. 68, 239–250. https://doi.org/10.1111/ppa.12950 (2019).
Hasiów-Jaroszewska, B., Borodynko, N., Rymelska, N. & Pospieszny, H. First report of papaya ringspot virus infecting zucchini plants in Poland. Plant. Dis. 94, 633. https://doi.org/10.1094/PDIS-94-5-0633A (2010).
Kim, O. K., Mizutani, T., Natsuaki, K. T., Lee, K. W. & Soe, K. First report and the genetic variability of cucumber green mottle mosaic virus occurring on bottle gourd in Myanmar. J. Phytopathol. 158, 572–575. https://doi.org/10.1111/j.1439-0434.2010.01634.x (2010).
Noshad, Q. Q. et al. First record of papaya ring spot virus (PRSV) strain in Malir district Sindh and in Islamabad Pakistan. Int. J. Agric. Biol. 17, 399–402. https://doi.org/10.17957/IJAB/17.3.15 (2015).
Khanal, V. & Ali, A. First report of cucurbit aphid-borne yellows virus infecting Cucurbita pepo in Oklahoma. Plant. Dis. 102, 1046–1047. https://doi.org/10.1094/PDIS-11-17-1684-PDN (2018).
Begum, F. et al. Surveillance of the disease incidence and severity of papaya ringspot virus at four selected districts of Bangladesh. Int. J. Environ. Agric. Biotechnol. 3, 2083–2090. https://doi.org/10.22161/ijeab/3.6.16 (2018).
Listihani, Damayanti, T. A., Hidayat, S. H. & Wiyono, S. Karakterisasi Molekuler papaya ringspot virus Tipe P Pada Tanaman mentimun Di Jawa. J. Fitopatol. Indones. 14, 75–82. https://doi.org/10.14692/jfi.14.3.75 (2018).
Esquivel-Fariña, A., Kraide, H. D., Camelo-García, V., Rezende, J. A. M. & Kitajima, E. W. Detection of the papaya strain of papaya ringspot virus (PRSV-P) in Paraguay. J. Plant. Pathol. 103, 1. https://doi.org/10.1007/s42161-021-00805-5 (2021).
Medina-Salguero, A. X. et al. Genetic characterization of a mild isolate of papaya ringspot virus type-P (PRSV-P) and assessment of its cross-protection potential under greenhouse and field conditions. PLoS ONE 16, e0241652. https://doi.org/10.1371/journal.pone.0241652 (2021).
Premchand, U. et al. Survey, detection, and characterization of papaya ringspot virus from Southern India and management of Papaya ringspot disease. Pathogens 12, 824. https://doi.org/10.3390/pathogens12060824 (2023).
Basavaraj, Y. B., Parameshwari, B., Kumar, A., Jain, R. K. & Tripathi, S. Papaya ring spot virus: Status of 80 years of global research. In Plant RNA Viruses, 135–172 (Academic Press, 2023). https://doi.org/10.1016/B978-0-323-95339-9.00024-7.
Parris, G. K. A new disease of Papaya in Hawaii. Proc. Am. Soc. Hortic. Sci. 36, 263–265 (1938).
Linder, R. C., Jensen, D. D. & Ikeda, W. Ring spot: New Papaya plunderer. Hawaii. Farm. Home. 8, 10–14 (1945).
Jensen, D. D. Papaya virus diseases with special reference to Papaya ringspot. Phytopathology 39, 191–211 (1949).
Capoor, S. P. & Verma, P. M. A mosaic disease on Carica papaya L. in the Bombay Province. Curr. Sci. 17, 265–266 (1948).
Reddy, L. R., Nagaraju, C.N., Kumar, M.K.P. & Venkataravanappa, V. Incidence of Papaya ringspot virus disease in Bangalore district. J. Plant Dis. Sci. 2, 104–106 (2007).
Talukdar, A. et al. Genetics of yellow mosaic virus (YMV) resistance in cultivated soybean [Glycine max (L) Merr]. Legume Res. 36, 263–267 (2013).
Hemavati, R., Sharma, S. K., Pinki, P. & Ngachan, S. V. Molecular evidence for association of papaya ringspot virus with Papaya from North East hill region of India. Indian Phytopathol. 70, 246–251. https://doi.org/10.24838/ip.2017.v70.i2.70755 (2017).
Babu, K. S. & Banerjee, A. Biological and molecular evidence of papaya ringspot virus pathotype P from mid-hills of Meghalaya, India. Indian Phytopathol. 71, 611–620. https://doi.org/10.1007/s42360-018-0097-9 (2018).
Harish, A. Characterization, host range, and management of papaya ringspot virus. MSc thesis, Kerala Agric. Univ. 204 (2018).
Basavaraj, Y. B. et al. Molecular diversity of Papaya ringspot virus in India: Genetic recombination and mutations between the isolates from different hosts and geo-climatic locations are role players in virus evolution. Indian Phytopathol. 72, 497–511. https://doi.org/10.1007/s42360-019-00157-2 (2019).
Quiot-Douine, L., Lecoq, H., Quiot, J. B., Pitrat, M. & Labonne, G. Serological and biological variability of virus isolates related to strains of papaya ringspot virus. Phytopathology 80, 256–263. https://doi.org/10.1094/Phyto-80-256 (1990).
Baker, C. A., Lecoq, H. & Purcifull, D. E. Serological and biological variability among papaya ringspot virus type-W isolates in Florida. Phytopathology 81, 722–728. https://doi.org/10.1094/Phyto-81-722 (1991).
Bateson, M. F., Henderson, J., Chaleeprom, W., Gibbs, A. J. & Dale, J. L. Papaya ringspot potyvirus: Isolate variability and the origin of PRSV type P (Australia). J. Gen. Virol. 75, 3547–3553. https://doi.org/10.1099/0022-1317-75-12-3547 (1994).
Premchand, U. et al. Comparative host range and molecular studies of papaya ringspot virus. BFAIJ 13, 370–373 (2021).
Yeh, S. D. & Gonsalves, D. Translation of Papaya ringspot virus RNA in vitro: Detection of a possible polyprotein that is processed for capsid protein, cylindrical-inclusion protein, and amorphous-inclusion protein. Virology 143, 260–271. https://doi.org/10.1016/0042-6822(85)90306-3 (1985).
Yeh, S. D. et al. Complete nucleotide sequence and genetic organization of papaya ringspot virus RNA. J. Gen. Virol. 73, 2531–2541. https://doi.org/10.1099/0022-1317-73-10-2531 (1992).
Premchand, U. et al. Identification of novel begomoviruses associated with leaf curl disease of Papaya (Carica papaya L.) in India. Agronomy 13, 3. https://doi.org/10.3390/agronomy13010003 (2023).
Chávez-Calvillo, G. et al. Antagonism or synergism between papaya ringspot virus and papaya mosaic virus in Carica papaya L. is determined by their order of infection. Virology 489, 179–191. https://doi.org/10.1016/j.virol.2015.11.026 (2016).
Purcifull, D. E., Edwardson, J., Hiebert, E. & Gonsalves, D. Papaya ringspot virus. CMI/AAB. Description of Plant Viruses, No. 292, Wellesbourne, UK: Association of Applied Biologists, 209 (1984).
Kalleshwaraswamy, C. M. & Kumar, N. K. Transmission efficiency of papaya ringspot virus by three aphid species. Phytopathology 98, 541–546. https://doi.org/10.1094/PHYTO-98-5-0541 (2008).
Singh, V. & Singh, D. Studies on natural transmission of papaya ringspot virus disease in Eastern Uttar Pradesh. Ann. Plant. Prot. Sci. 18, 188–192 (2010).
Gadhave, K. R., Dutta, B., Coolong, T. & Srinivasan, R. A non-persistent aphid-transmitted Potyvirus differentially alters the vector and non-vector biology through host plant quality manipulation. Sci. Rep. 9, 39256. https://doi.org/10.1038/s41598-019-39256-5 (2019).
Jain, R. K. et al. Variability in the coat protein gene of Papaya ringspot virus isolates from multiple locations in India. Arch. Virol. 149, 2435–2442. https://doi.org/10.1007/s00705-004-0392-x (2004).
Pushpa, R.N., Shantamma, P., Pappachan, A., Manjunath, B., Sumit, B., Kumar, S., Rangaswamy, K.T., Girish, T.R. & Nagaraju, N. Molecular characterization, epidemiology and management of the Papaya ringspot virus (PRSV) in papaya under southern Indian conditions. Int. J. Agric. Sci. 10, 5029–5038.
Pushpa, R. N., Nagaraju, N., Joshi, S. & Jagadish, K. S. Epidemiology of papaya ringspot virus-P (PRSVP) infecting Papaya (Carica papaya Linn.) and influence of weather parameters on population dynamics of predominant aphid species. J. Entomol. Zool. Stud. 7, 434–439 (2019).
Anu, O. P., Sheoran, M. S., Tonk, K. & Devi, K. Quantitative analysis of environmental influences on mustard aphid population dynamics using statistical modelling. Int. Res. J. Eng. Technol. 10, 555–562 (2023).
Azad, M. A. K., Amin, L. & Sidik, N. M. Gene technology for Papaya ringspot virus disease management. Sci. World J. 768038. https://doi.org/10.1155/2014/768038 (2014).
Hull, R. Plant Virology (Academic Press, 2013).
Premchand, U. et al. Management of papaya ringspot virus (PRSV) using insecticides and bio-rationals under field conditions. BFAIJ 13, 743–748 (2021).
Thiara, S. K., Cheema, S. S. & Kang, S. S. Effect of date of sowing and plant density on incidence of soybean yellow mosaic virus (SoYMV) in soybean. J. Res. 40, 207–215 (2003).
Clemente-Orta, G., Albajes, R. & Achon, M. A. Early planting, management of edges and non-crop habitats reduce potyvirus infection in maize. Agron. Sustain. Dev. 40, 21. https://doi.org/10.1007/s13593-020-00625-4 (2020).
Kone, N. et al. Influence of planting date on incidence and severity of viral disease on cucurbits under field condition. Ann. Agric. Sci. 62, 99–104. https://doi.org/10.1016/j.aoas.2017.05.005 (2017).
Broadbent, L., Heathcote, G., McDermott, N. & Taylor, C. The effect of date of planting and of harvesting potatoes on virus infection and on yield. Ann. Appl. Biol. 45, 603–622. https://doi.org/10.1111/j.1744-7348.1957.tb00407.x (1957).
MacKenzie, T. D., Fageria, M. S., Nie, X. & Singh, M. Effects of crop management practices on current-season spread of potato virus Y. Plant. Dis. 98, 213–222. https://doi.org/10.1094/PDIS-04-13-0403-RE (2014).
Abbas, G. et al. Effect of planting dates on agronomic crop production. In Agronomic Crops: Volume 1: Production Technologies 131–147 (2019). https://doi.org/10.1007/978-981-32-9151-5_8.
Mora-Aguilera, G., Téliz, D., Campbell, C. L. & Avila, C. Temporal and spatial development of Papaya ringspot in Veracruz, Mexico. J. Phytopathol. 136, 27–36. https://doi.org/10.1111/j.1439-0434.1992.tb01278.x (1992).
Mora-Aguilera, G., Nieto-Angel, D., Téliz, D. & Campbell, C. L. Development of a prediction model for Papaya ringspot in Veracruz, Mexico. Plant. Dis. 77, 1205–1211. https://doi.org/10.1094/PD-77-1205 (1993).
Sharma, S. K. et al. Integrated management of papaya ringspot virus. Acta Hortic. 851, 473–480. https://doi.org/10.17660/ActaHortic.2010.851.73 (2010).
Chandrashekar, K., Chavan, V. M., Sharma, S. K. & Bhosle, A. B. Management of PRSV-P in Papaya through time of planting and border cropping. Indian J. Hortic. 72, 423–425. https://doi.org/10.5958/0974-0112.2015.00083.3 (2015).
Thiribhuvanamala, G., Sridharan, S., Soorianathasundaram, K. & Reddy, M. K. Influence of weather factors on aphid population and the incidence of papaya ringspot virus disease. Pest Manag. Hortic. Ecosyst. 22, 91–93 (2016).
Onwughalu, J. T., Abo, M. E., Okoro, J. K., Onasanya, A. & Sere, Y. Rice yellow mottle virus infection and reproductive losses in rice (Oryza sativa Linn). Trends Appl. Sci. Res. 6, 182–189. https://doi.org/10.3923/tasr.2011.182.189 (2011).
De Breuil, S., Giolitti, F. J., Bejerman, N. & Lenardon, S. L. Effects of cucumber mosaic virus on the yield and yield components of peanut. J. Plant. Pathol. 669, 669–673 (2012).
Olobashola, N., Salaudeen, M. T. & Achikwu, M. Growth and yield responses of sweet pepper (Capsicum annuum L.) cultivars to infections with cucumber mosaic virus disease. NJAFE 13, 201–205 (2017).
Kumar, J. V. et al. Rapid real-time viral load estimation technique for Chilli leaf curl virus and its validation in different Chilli genotypes from Eastern Himalayan plains (2022). https://doi.org/10.21203/rs.3.rs-1906135/v1.
Elvira González, L., Peiró, R., Rubio, L. & Galipienso, L. Persistent Southern tomato virus (STV) interacts with cucumber mosaic and/or pepino mosaic virus in mixed-infections modifying plant symptoms, viral titer, and small RNA accumulation. Microorganisms 9, 689. https://doi.org/10.3390/microorganisms9040689 (2021).
Shirima, R. R. et al. Absolute quantification of cassava brown streak virus mRNA by real-time qPCR. J. Virol. Methods 245, 5–13. https://doi.org/10.1016/j.jviromet.2017.03.003 (2017).
Ranabhat, N. B., Bruce, M. A., Fellers, J. P. & Shoup Rupp, J. L. A reproducible methodology for absolute viral quantification and viability determination in mechanical inoculations of wheat streak mosaic virus. Trop. Plant. Pathol. 47, 553–561. https://doi.org/10.1007/s40858-022-00507-y (2022).
Rubio, L. et al. Detection and absolute quantitation of watermelon mosaic virus by real-time RT-PCR with a TaqMan probe. J. Virol. Methods. 300, 114416. https://doi.org/10.1016/j.jviromet.2021.114416 (2022).
Chai, A. et al. Rapid quantification of infectious cucumber green mottle mosaic virus in watermelon tissues by PMA coupled with RT-qPCR. Viruses 14, 2046. https://doi.org/10.3390/v14092046 (2022).
Anonymous. Package of Practice: Papaya. University of Horticultural Sciences, Bagalkot, India (2023), accessed on 10 May 2023; http://uhsbagalkot.edu.in/downloads/Horticulture_POP.pdf.
Shaner, G. & Finney, R. E. The effect of nitrogen fertilization on the expression of slow-mildewing resistance in Knox wheat. Phytopathology 67, 1051–1056 (1977).
Zhaozhi, L. et al. Differences in the high-temperature tolerance of Aphis craccivora Koch (Hemiptera: Aphididae) on cotton and soybean: Implications for ecological niche switching among hosts. Appl. Entomol. Zool. 52, 9–18. https://doi.org/10.1007/s13355-016-0446-z (2017).
Dijkstra, J. & de Jager, C. P. Virus transmission by aphids. In Practical Plant Virology: Protocols and Exercises 148–158 (Springer, 1998). https://doi.org/10.1007/978-3-642-72030-7_28.
Sheoran, O. P., Tonk, D. S., Kaushik, L. S., Hasija, R. C. & Pannu, R. S. Statistical software package for agricultural research workers. Recent Advances in Information Theory, Statistics & Computer Applications by D.S. Hooda & R.C. Hasija, Department of Mathematics Statistics, CCS HAU, Hisar 8, 139–143 (1998).
Zhu, X. et al. Evaluation of new reference genes in Papaya for accurate transcript normalization under different experimental conditions. PLoS ONE 7, e44405. https://doi.org/10.1371/journal.pone.0044405 (2012).
Wang, J. J. & Tsai, J. H. Effect of temperature on the biology of Aphis spiraecola patch (Homoptera: Aphididae). Ann. Entomol. Soc. Am. 93, 874–883. https://doi.org/10.1603/0013-8746(2000)093[0874:EOTOTB]2.0.CO;2 (2000).
Alyokhin, A., Drummond, F. A., Sewell, G. & Storch, R. H. Differential effects of weather and natural enemies on coexisting aphid populations. Environ. Entomol. 40, 570–580 (2011).
Sun, J., Tan, X., Li, Q., Francis, F. & Chen, J. Effects of different temperatures on the development and reproduction of Sitobion miscanthi from six different regions in China. Front. Ecol. Evol. 10, 794495 (2022).
Brabec, M., Honěk, A., Pekár, S. & Martinková, Z. Population dynamics of aphids on cereals: Digging in the time-series data to reveal population regulation caused by temperature. PLoS ONE 9, e106228 (2014).
Chavan, V. M., Tomar, S. P. S. & Dhale, M. G. Management of Papaya ringspot virus (PRSV-P) of Papaya under Pune conditions. Acta Hortic. 851, 447–452. https://doi.org/10.17660/ActaHortic.2008.851.63 (2008).
Rashmi, P., Mayank, R., Kuldeep, S. & Deepti, C. Studies on population dynamics of Myzus persicae on potato crop with special reference to its relation with various weather parameters. Veg. Sci. 34, 167–169 (2007).
Sarvendra, S., Akhilesh, K. & Awasthi, B. K. Study of sucking and leaf-feeding insects in relation to weather parameters on the Brinjal crops. Veg. Sci. 32, 210–212 (2005).
Lazarowitz, S. G. Geminiviruses: Genome structure and gene function. CRC Crit. Rev. Plant. Sci. 11, 327–349. https://doi.org/10.1080/07352689209382350 (1992).
Timmermans, M. C., Das, O. P. & Messing, J. Geminiviruses and their uses as extrachromosomal replicons. Annu. Rev. Plant. Physiol. Plant. Mol. Biol. 45, 79–112. https://doi.org/10.1146/annurev.pp.45.060194.000455 (1994).
Brown, C. R., Corsini, D., Pavek, J. & Thomas, P. E. Heritability of field resistance to potato leafroll virus in cultivated potato. Plant Breed. 116, 585–588. https://doi.org/10.1111/j.1439-0523.1997.tb02194.x (1997).
Gan, S. Mitotic and postmitotic senescence in plants. Sci. Aging Knowl. Environ. 38, re7. https://doi.org/10.1126/sageke.2003.38.re7 (2003).
Ndunguru, J. & Rajabu, A. C. Effect of Okra mosaic virus disease on the above-ground morphological yield components of Okra in Tanzania. Sci. Hortic. 99, 225–235. https://doi.org/10.1016/S0304-4238(03)00108-0 (2004).
Zhu, S. et al. The rice Dwarf virus P2 protein interacts with ent-kaurene oxidases in vivo, leading to reduced biosynthesis of gibberellins and rice Dwarf symptoms. Plant. Physiol. 139, 1935–1945. https://doi.org/10.1104/pp.105.072306 (2005).
Espinoza, C., Medina, C., Somerville, S. & Arce-Johnson, P. Senescence-associated genes induced during compatible viral interactions with grapevine and Arabidopsis. J. Exp. Bot. 58, 3197–3212. https://doi.org/10.1093/jxb/erm165 (2007).
Benjamins, R. & Scheres, B. Auxin: The looping star in plant development. Annu. Rev. Plant. Biol. 59, 443–465. https://doi.org/10.1146/annurev.arplant.58.032806.103805 (2008).
Jin, L. et al. Rice Dwarf virus P2 protein hijacks auxin signaling by directly targeting the rice OsIAA10 protein, enhancing viral infection and disease development. PLoS Pathog. 12, e1005847. https://doi.org/10.1371/journal.ppat.1005847 (2016).
Sheikh, M. A., Safiuddin, Khan, Z. & Mahmood, I. Effect of bhendi yellow vein mosaic virus on yield components of Okra plants. J. Plant. Pathol. 95, 391–393 (2013).
Li, W., Han, Y., Tao, F. & Chong, K. Knockdown of SAMS genes encoding S-adenosyl-l-methionine synthetases causes methylation alterations of DNAs and histones and leads to late flowering in rice. J. Plant Physiol. 168, 1837–1843. https://doi.org/10.1016/j.jplph.2011.05.020 (2011).
Shalit-Kaneh, A. et al. The flowering hormone florigen accelerates secondary cell wall biogenesis to harmonize vascular maturation with reproductive development. Proc. Natl. Acad. Sci. USA 116, 16127–16136. https://doi.org/10.1073/pnas.1906405116 (2019).
Prakash, J., Singh, K., Goswami, A. K. & Singh, A. K. Comparison of plant growth, yield, fruit quality, and biotic stress incidence in Papaya Var. Pusa Nanha under polyhouse and open field conditions. Indian J. Hortic. 72, 183–186. https://doi.org/10.5958/0974-0112.2015.00036.5 (2015).
Huseynova, I. M. et al. Virus-induced changes in photosynthetic parameters and peroxidase isoenzyme contents in tomato (Solanum lycopersicum L.) plants. Photosynthetica 56, 841–850. https://doi.org/10.1007/s11099-017-0737-9 (2018).
Lal, M. K. et al. Effect of potato apical leaf curl disease on glycemic index and resistant starch of potato (Solanum tuberosum L.) tubers. Food Chem. 359, 129939. https://doi.org/10.1016/j.foodchem.2021.129939 (2021).
Broadbent, D. E. Listening to one of two synchronous messages. J. Exp. Psychol. 44, 51–55. https://doi.org/10.1037/h0056491 (1952).
Cadman, C. H. & Chambers, J. Factors affecting the spread of aphid-borne viruses in potato in Eastern Scotland: III. Effects of planting date, rouging and age of crop on the spread of potato leaf roll and Y viruses. Ann. Appl. Biol. 48, 729–738. https://doi.org/10.1111/j.1744-7348.1960.tb03572.x (1960).
Mangrauthia, S. K., Singh, P. & Praveen, S. Genomics of helper component proteinase reveals effective strategy for papaya ringspot virus resistance. Mol. Biotechnol. 44, 22–29. https://doi.org/10.1007/s12033-009-9205-5 (2010).
Sahana, N. et al. Inhibition of the host proteasome facilitates Papaya ringspot virus accumulation and proteasomal catalytic activity is modulated by viral factor HcPro. PLoS ONE 7, e52546. https://doi.org/10.1371/journal.pone.0052546 (2012).
Vargas-Mejía, P., Vega-Arreguín, J., Chávez-Calvillo, G., Ibarra-Laclette, E. & Silva-Rosales, L. Differential accumulation of innate and adaptive immune response derived transcripts during antagonism between papaya ringspot virus and Papaya mosaic virus. Viruses 12, 230. https://doi.org/10.3390/v12020230 (2020).
Hong, J. S. & Ju, H. J. The plant cellular systems for plant virus movement. Plant Pathol. J. 33, 213–228. https://doi.org/10.5423/PPJ.RW.09.2016.0198 (2017).
Tran, P. T., Zhang, C. F. & Citovsky, V. Rapid generation of inoculum of a plant RNA virus using overlap PCR. Virology 553, 46–50. https://doi.org/10.1016/j.virol.2020.11.001 (2021).
Develey-Riviere, M. P. & Galiana, E. Resistance to pathogens and host developmental stage: A multifaceted relationship within the plant Kingdom. New. Phytol. 175, 405–416. https://doi.org/10.1111/j.1469-8137.2007.02130.x (2007).
Hu, L. & Yang, L. Time to fight: Molecular mechanisms of age-related resistance. Phytopathology 109, 1500–1508. https://doi.org/10.1094/PHYTO-11-18-0443-RVW (2019).
Chikh-Ali, M., Tran, L. T., Price, W. J. & Karasev, A. V. Effects of the age-related resistance to potato virus Y in potato on the systemic spread of the virus, incidence of the potato tuber necrotic ringspot disease, tuber yield, and translocation rates into progeny tubers. Plant Dis. 104, 269–275. https://doi.org/10.1094/PDIS-06-19-1201-RE (2020).
Wang, X., Rong, W., Liu, Y., Wang, X. & Zhang, Z. Investigation of the mechanism of adult-stage resistance to barley yellow dwarf virus associated with a wheat Thinopyrum intermedium translocation. Crop J. 6, 394–405. https://doi.org/10.1016/j.cj.2018.02.002 (2018).
Leisner, S. M., Turgeon, R. & Howell, S. H. Long distance movement of cauliflower mosaic virus in infected turnip plants. Mol. Plant. Microbe Interact. 5, 41–47. https://doi.org/10.1094/MPMI-5-041 (1992).
Zhu, Y. J., Lim, S. T., Schenck, S., Arcinas, A. & Komor, E. RT-PCR and quantitative real-time RT-PCR detection of sugarcane yellow leaf virus (SCYLV) in symptomatic and asymptomatic plants of Hawaiian sugarcane cultivars and the correlation of SCYLV titre to yield. Eur. J. Plant. Pathol. 127, 263–273. https://doi.org/10.1007/s10658-010-9591-3 (2010).
Agrios, G. N., Walker, M. E. & Ferro, D. N. Effect of cucumber mosaic virus inoculation at successive weekly intervals on growth and yield of pepper (Capsicum annuum) plants. Plant. Dis. 69, 52–55. https://doi.org/10.1094/PD-69-52 (1985).
Levy, D. & Lapidot, M. Effect of plant age at inoculation on expression of genetic resistance to tomato yellow leaf curl virus. Arch. Virol. 153, 171–179. https://doi.org/10.1007/s00705-007-1086-y (2008).
Acknowledgements
We would like to thanks to Department of Horticulture, Government of Karnataka, India for approving Deputation to first author for pursuing Ph.D and University of Horticultural Sciences, Bagalkote, India for providing research facility.
Funding
This research received no external funding.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study’s conception and design. Material preparation, data collection, analysis, and interpretation were performed by P.U., R.K.M., and M.P.B. The initial draft of the manuscript was written by P.U., V.V., L.R.C.N.R., and K.S.S. All authors reviewed and provided critical feedback on the manuscript. All authors read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Premchand, U., Mesta, R.K., Basavarajappa, M.P. et al. Epidemiological studies on the incidence of papaya ringspot disease under Indian sub-continent conditions. Sci Rep 15, 6973 (2025). https://doi.org/10.1038/s41598-025-91612-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-91612-w