Abstract
RNA-based assays hold great potential for assessing viability molecularly in slow- or non-growing mycobacteria, and RNAseq has evolved into a powerful tool in infectious disease research. Such applications require efficient RNA preservation for optimal results. The performance of 70% ethanol as simpler alternative to commonly-used GTC-based storage of mycobacteria at − 80 °C was compared on cultured Mycobacterium tuberculosis H37Ra subjected to following setups: a 45 °C heat shock to test immediate stabilisation, five freeze-thaw scenarios to mimic different shipping conditions, and long-term storage at -80 °C, -20 °C, 4 °C or 30 °C for up to twelve months. Treatment with 70% ethanol yielded overall higher RNA quantities compared to GTC-TCEP and RNA integrity was maintained at -20 °C for twelve months. Both buffers are reportedly mycobactericidal and, in this study, prevented heat stress-induced transcriptomic changes, thereby conserving a transcriptomic snapshot. RNA yield and integrity remained unaltered after treatment with 70% ethanol, even with up to three freeze-thaw cycles. Based on these results, we recommend 70% ethanol over GTC-TCEP for RNA preservation of mycobacterial cultures. While freeze-thawing, short-term high-temperature and − 20 °C long-term storage results are promising, this inexpensive, widely available buffer needs further validation prior to applying it for RNA-based analysis in clinical samples.
Similar content being viewed by others
Introduction
Studying the RNA of infectious agents complements DNA analyses. For example, RNA-based testing can be an alternative to assess the viability of slow growing or non-culturable bacteria1. RNA sequencing (RNAseq) is increasingly affordable and can help decipher microbial transcriptomics in different in vitro conditions as well as host-pathogen interactions to understand disease mechanisms2.
Mycobacterial diseases like tuberculosis (TB) and leprosy remain public health problems. With 8.2 million newly detected patients in 2023 of whom approximately 400,000 infected with drug-resistant Mycobacterium tuberculosis, TB returned to being the main infectious killer worldwide after COVID-19 had been holding this position for three years3. For leprosy, more than 180,000 new patients were reported in 2023, a number known to be an underestimate4. Timely diagnosis and treatment follow-up are crucial to detect drug resistance and adjust treatment accordingly. Many mycobacteria are difficult to cultivate, taking weeks to months, while for M. leprae in vitro culture is impossible5.
Consequently, RNA-based analyses have been integrated into mycobacterial research to address this issue and assays have been developed, using RNA target quantity as a marker for mycobacterial burden, activity and viability. For M. tuberculosis the molecular bacterial load assay (MBLA)6 quantifies 16S rRNA transcripts while the RNA synthesis (RS) ratio calculates the ratio of pre-ribosomal ETS1 fragments over 23S rRNA as measure for metabolic activity7. The molecular viability assay (MVA) has been developed for M. leprae, assessing the quantity of two mRNA targets hsp18 and esxA normalized to the quantity of M. leprae DNA to characterise M. leprae viability8,9.
A major challenge for the success of RNA-based analyses is the transient nature of RNA molecules and omnipresent RNases that threaten RNA integrity. Therefore, to preserve RNA prior to extraction, storage conditions should stabilise the RNA molecules and inactivate the RNases.
Most commonly samples are flash-frozen and stored at -80 °C, commercially available RNA-preservation buffers are also frequently used. However, these treatments do not always achieve inactivation of mycobacterial cells, a critical step for biosafety and to ensure transcriptional arrest and a transcriptome snapshot. For mycobacteria, the chaotropic agent guanidinium thiocyanate (GTC)10 is frequently applied before storage at -80 °C because it inactivates RNases and kills the pathogen6,11,12,13. As such cold-chains are not always possible, more cost-effective and field-friendly alternative storage in acetic acid or 70% ethanol at -20 °C has been proposed for molecular viability testing of M. tuberculosis in sputum and M. leprae in skin biopsies8,9,14. When exposed to 70% ethanol, M. leprae and M. tuberculosis in sputum sediment are killed within 1 h at ambient temperature15,16. Therefore, in this study we systematically tested RNA stabilisation of mycobacterial cultures with widely available 70% ethanol in comparison to the reference buffer GTC-TCEP. We subjected these to a heat shock to assess transcriptomic arrest and analysed the quality and quantity of RNA during long term storage at -80 °C, -20 °C, 4 °C and 30 °C for up to 12 months as well as after up to three freeze-thaw cycles to mimic an interrupted cold chain.
Materials and methods
Bacterial cultures
Bacterial cultures were grown on solid, inclined Löwenstein-Jensen (LJ) medium at 37 °C. To simplify the handling, the avirulent M. tuberculosis H37Ra variant was used (BCCM/ITM CT1978-06710). After three weeks growing, log-phased bacilli were weighed and adjusted to 40 mg/ml in sterile physiological water.
Bacterial suspensions
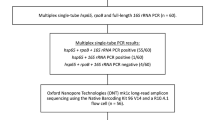
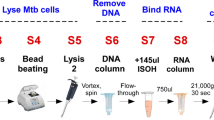
Bacterial suspensions were diluted 1:10 with two different preservation buffers: in ethanol to an end concentration of 70% and in GTC-TCEP (5 M guanidine-thiocyanate, 10 mM tris(2-carboxyethyl)phosphine) to an end concentration of 4.5 M, prepared at the Consortium for Applied Microbial Metrics, Aurora, CO, USA13. Samples with 4 mg bacilli were further processed for long-term storage, heat-shock and freeze-thaw challenges as described below and in Fig. 1. For all conditions technical triplicates were tested and a negative control containing sterile physiological water instead of bacterial suspension was included.
As control for the concentrations of bacterial suspensions, 10 µl of 1:2 × 104, 1:2 × 105 and 1:2 × 106 dilutions were grown on Middlebrook 7H10 agar plates for three weeks at 37 °C, 5–10% CO2 before colony counting.
Heat shock challenge
After treatment with the buffers for approximately 10 min samples were incubated for 20 min at 45 °C17 or at 37 °C as control before immediate RNA extraction.
Long-term storage
After a one-hour treatment with the buffers, baseline controls were extracted immediately and remaining samples were stored up to 12 months before RNA extraction at -80 °C, -20 °C (< -18 °C), 4 °C (2–8 °C) or 30 °C to approximate tropical ambient temperature .
Freeze-thaw challenge
After a one-hour treatment with the buffers, baseline controls were extracted immediately and remaining samples underwent different storage schedules over a course of six weeks, including one, two or three freeze-thaw cycles before RNA extraction: (i) two weeks − 20 °C, four weeks 4 °C, (ii) two weeks − 20 °C, one week 4 °C, one week − 20 °C, two weeks 4 °C, (iii) two weeks − 20 °C, three weeks 4 °C, one week − 20 °C, (iv) two weeks − 20 °C, one week 4 °C, one week − 20 °C, one week 4 °C, one week − 20 °C, (v) 30 °C overnight prior to scenario (iv). A baseline control extracted after a one-hour treatment with the buffers and negative controls for each condition were included.
RNA extraction and reverse transcription
For RNA extraction cells were pelleted for 20 min at 3,350 x g and resuspended in 800 µl RNeasy Lysis Buffer (RLT) buffer (Qiagen, Germany) with 1% β-mercaptoethanol. Cells were lysed in Lysing Matrix E tubes with a FastPrep-24 Classic instrument (both MP Biomedicals, Germany) four rounds at 6.0 m/s for 40 s each. Purification with the Maxwell RSC miRNA Tissue Kit (Promega, USA) resulted in ± 57.5 µl extract, of which 30 µl was DNase-treated with the TURBO DNA-free Kit (Invitrogen, USA).
Quality control of extracts
The quantity of DNase-treated RNA was assessed with the Qubit RNA High Sensitivity (HS) Kit (Invitrogen, USA). To estimate the quantity in the initial sample, results were corrected by a 1:100 dilution factor and multiplied by the Maxwell elution volume. The 4200 TapeStation system (Agilent, USA) with RNA Screentape reagents and Analysis Software version 5.1 were used to determine the RNA integrity number (RIN). A RIN of 7.0 has previously been reported as the cutoff for successful RNAseq for M. tuberculosis.18,19 All assays were run according to manufacturer instructions.
Reverse transcription and qPCR
DNase-treated RNA was reverse transcribed (RT) with the SuperScript IV First-Strand Synthesis System (Invitrogen, USA) according to manufacturer instructions. Each sample had a corresponding RT-negative control without the enzyme to control for DNA contamination. Digestion with 1 µl of RNase H (Invitrogen, USA) at 37 °C for 20 min eliminated remnant RNA.
Primers and probes for esxA, icl, sigA, 16S rRNA, and hsp65 qPCR assays were designed with the Geneious Prime software (version 2020.0.3, Biomatters, New Zealand) or adapted from previous publications (Supplementary Table S1). To 18 µl of master mix (Sensimix II) and internal positive control (IPC) Yakima Yellow-BHQ-1 (both Eurogentec, Belgium), 2 µl of a 1:10 dilution of cDNA or RT-negative control were added. The qPCRs were run at 95 °C for 10 min followed by 45 cycles of 15 s at 95 °C and at 60 °C for 1 min on a StepOnePlus instrument (Applied Biosystems, USA). Each run included negative controls (nuclease-free water) and a 1:10 dilution series of plasmid DNA standard per target fragment (3 × 105 to 3 × 101 copies, TIB MOLBIOL, Germany), as standard curve for absolute quantification.
Data analysis
For all outcome measures, means of technical triplicates were calculated. For the statistical analysis of qPCR data, this mean was extrapolated to the original sample volume and log10-transformed. Qubit means were extrapolated to the original sample volume, RIN values were analysed unaltered. For the heat shock experiment and baseline comparisons of the long-term storage experiment, differences in the outcome measures were calculated with the non-parametric Wilcoxon Rank-Sum test. To further analyse the influence of the different storage factors and freeze-thaw scenarios on the RIN and Qubit results, the non-parametric Kruskal-Wallis Rank-Sum test was performed for each factor. Post hoc pairwise comparisons on each factor level were performed with the Wilcoxon Rank-Sum test. The Benjamini-Hochberg procedure corrected for multiple testing. The alternative hypothesis was accepted at a significance level α = 0.05. Analyses were performed with R version 4.4.0 (The R Foundation, Austria). Further, for each transcript quantified by qPCR, a ratio was calculated with the mean quantities per sample: \(\:{Ratio}_{t}=\:\frac{transcript\:quantity\:at-20,\:\:\:4\:or\:30\:^\circ\:C\:at\:timepoint\:t}{transcript\:quantity\:at-80\:^\circ\:C\:at\:timepoint\:t}\). The ratios for the different storage conditions were analysed descriptively.
Setup of three experiments to analyse RNA preservation with GTC-TCEP and 70% ethanol. The three experiments were started independently from different LJ cultures of the same M. tuberculosis H37Ra strain, aiming for 4 mg of bacilli per aliquot. For (b) and (c) samples were extracted at baseline and after different storage conditions. Per set of technical triplicates, a negative control with sterile physiological water was taken along. LJ: Löwenstein-Jensen medium, AT: ambient temperature (ca. 20 °C), i – v: freeze-thaw scenarios, described in Materials and Methods (“Freeze-thaw challenge”) and Fig. 5.
Results
Colony forming units counting
Plating the lowest dilution of the bacterial suspension resulted in confluent growth. On plates with the 1:2 × 105 and 1:2 × 106 dilutions 69 and 12 colony forming units (CFU) were counted, respectively. Considering the dilution factors, the concentration in the original suspension (40 mg/ml) was between 1.38 × 109 CFU/ml and 2.40 × 109 CFU/ml. Therefore, each sample with 4 mg bacilli contained approximately 2.0 × 108 CFU.
Assessment of effect of heat shock on M. tuberculosis gene expression in 70% ethanol and GTC-TCEP
The effect of a 20-minute heat shock at 45 °C on transcript quantities of hsp65, sigA and 16S rRNA (Supplementary Figures S1, S2 and S3) was determined. In both 70% ethanol and GTC-TCEP treated samples, the copy numbers of all targets were not significantly different after a heat shock compared to the 37 °C control. However, overall copy numbers were significantly lower in GTC-TCEP treated samples for hsp65 (p = 0.0022) and 16S rRNA (p = 0.0017).
Effects on RNA yield at baseline and during long-term storage in 70% ethanol and GTC-TCEP
At the baseline, when samples were treated for approximately 1 h at ambient temperature, it was possible to recover 2.4 x more total RNA (Qubit) after 70% ethanol treatment compared to GTC-TCEP (p = 0.05, Fig. 2). Similarly, log10-transformed copy numbers were higher in ethanol-preserved samples at baseline, around 1.7 x more esxA (p = 0.05), 2.9 x more icl (p = 0.05), 1.4 x sigA (p = ns), and 2.5 x more 16 S rRNA copies (p = 0.05) compared to GTC-TCEP preserved samples (Supplementary Figure S4).
During storage, the overall RNA quantity determined by Qubit was significantly influenced by the buffer (p < 0.0001). Pairwise comparisons showed a significant detrimental effect of 30 °C compared to -80 °C (p = 0.0039), -20 °C and 4 °C (both p = 0.0364) for ethanol-preserved samples while for GTC-TCEP 4 °C had a positive influence on RNA quantity compared to -80 °C (p = 0.0323) and − 20 °C (p = 0.0067) (Fig. 2). For 70% ethanol-treated samples the storage time did not have a significant influence on the total RNA yield.
Quantities of all transcripts were significantly higher during long-term storage in ethanol and increased over time at -80 °C, -20 °C and 4 °C, except for timepoint month 6 where yields in both buffers were lower (Supplementary Figure S4 and Supplementary Table S2). Storing ethanol-preserved samples at 4 °C and 30 °C led to lower yields than in frozen samples for almost all targets (p-values in Supplementary Table S2). For GTC-TCEP preserved samples the same effect was observed for 30 °C storage compared to the other temperatures.
Total RNA quantity from samples stored in 70% ethanol or GTC-TCEP at different temperatures. Aliquots of a suspension of M. tuberculosis H37Ra colonies (~ 2.0 × 108 CFU) were treated in triplicate with either 70% ethanol or 4.5 M GTC-TCEP and extracted directly after a 1-hour incubation at ambient temperature (fresh) and after additional storage overnight, for two weeks, one, three, six and twelve months. During storage samples were kept at -80 °C, -20 °C, 4 °C or 30 °C. RNA extracts, were treated with TURBO DNase and RNA quantity was determined by Qubit fluorometry. The graphs show the median quantity of the replicates of all time points per temperature with the interquartile range.
Effect of long-term preservation in 70% ethanol and GTC-TCEP on different RNA targets
For each RNA target a ratio at -20 °C, 4 °C and 30 °C and each time point was determined (Fig. 3). In ethanol-preserved samples, a similar pattern was observed for all four targets: At -20 °C the ratio remained close to 1.0 with a drop at 12 months of storage. This drop was most pronounced for sigA, the target with the largest amplicon size (160 bp). At 4 °C the ratio dropped already after three months but not further thereafter, and at 30 °C was only close to the 1.0 mark at the overnight timepoint.
For GTC-TCEP preserved samples, no such pattern was observable across all targets and standard deviations were considerably higher than in ethanol. The ratios were comparably around 1.0 at -20 °C and 4 °C for the smallest transcript esxA (288 bp). For the other targets values were more scattered.
Effect of long-term storage in 70% ethanol and GTC-TCEP on the ratio of different transcript quantities. A suspension of M. tuberculosis H37Ra colonies (~ 2.0 × 108 CFU) was treated in triplicate with 70% ethanol (left panels) or 4.5 M GTC-TCEP (right panels) and extracted after storage for up to 12 months (12 M). The quantity of four targets esxA (a, b), icl (c, d), sigA (e, f) and 16S rRNA (g, h) was determined by qPCR after TURBO DNase treatment and reverse transcription. Graphs show the ratio of the mean transcript quantity at -20 °C (blue), 4 °C (green) or 30 °C (yellow) relative to the respective mean quantity at -80 °C at each tested time point with the standard deviation of the ratio.
Influence of up to twelve-month storage on RNA integrity
To analyse the influence of buffer, time and temperature on RNA integrity, RIN values were calculated and compared between storage conditions. RNA of the same quality was recovered from ethanol and GTC-TCEP preserved samples at baseline (RIN 8.07 ± 0.1 s.d.) and after overnight storage at -80 °C, -20 °C and 4 °C (RIN 7.7 ± 0.2 s.d.) (Fig. 4). At 30 °C a drop in RIN was observed after overnight storage in GTC-TCEP, while in 70% ethanol the RNA remained highly intact (RIN 7.6 ± 0.1 s.d.) compared to -80 °C. Further, in 70% ethanol, RIN values never dropped below 7.0 during frozen storage (-80 °C and − 20 °C), but after six months at 4 °C. For GTC-TCEP, RNA with a RIN above the 7.0 cutoff was still extracted after 12 months at -80 °C. A pairwise comparison of samples stored in GTC-TCEP showed that 4 °C and 30 °C have a significantly detrimental effect on the RIN values compared to the baseline (p = 0.0176 and p = 0.0207), after 6 months at 30 °C the RIN remained undetermined. In the same comparison for samples stored in 70% ethanol, the detrimental influence of a 30 °C storage was significant compared to the baseline (p = 0.021), the remaining p-values are summarised in Supplementary Table S2. Overall, increasing storage time and temperature had a significant detrimental influence on the RIN values (both p < 0.0001), as well as treating with GTC-TCEP compared to 70% ethanol (p = 0.0371). A pattern was observable where samples stored in GTC-TCEP showed a reduction in RIN earlier and at lower temperatures than samples stored in 70% ethanol.
RNA integrity for long-term stored samples in ethanol (a) or GTC-TCEP (b) at different temperatures. Aliquots of a suspension of M. tuberculosis H37Ra colonies (~ 2.0 × 108 CFU) were treated in triplicate with either 70% ethanol (a) or 4.5 M GTC-TCEP (b) and extracted immediately (fresh) or after storage overnight, for two weeks (2 W), one, three, six and twelve months (1 M, 3 M, 6 M, 12 M). The stored samples were kept at -80 °C (purple), -20 °C (blue), 4 °C (green) or 30 °C (yellow), fresh samples were processed at ambient temperature. After RNA extraction, samples were treated with TURBO DNase and RIN values were determined by TapeStation. The graphs show the mean RIN values of triplicates with the standard deviation, the dashed line indicates the RIN-cutoff of 7.0.
Effect of freeze-thawing on RNA preservation in 70% ethanol and GTC-TCEP
To assess the influence of freeze-thawing, cultures preserved with 70% ethanol and GTC-TCEP underwent five different storage schedules with one, two or three freeze-thaw cycles before extraction (Fig. 5). Again, using 70% ethanol instead of GTC-TCEP yielded overall around 2.5 x more RNA (p < 0.0001, data not shown) and RIN values between 6.0 and 7.0 for all scenarios (Fig. 5 and Supplementary Figure S5). Pairwise comparisons that were performed for 70% ethanol and GTC-TCEP separately showed no significant detrimental effect of the different freeze-thaw scenarios on the RNA quantity. The same was the case for the RIN in ethanol-preserved samples (Fig. 5). For GTC-TCEP preserved samples the RIN values could not be determined for all but two replicates and making statistical analysis impossible.
Influence of freeze-thawing on mycobacterial mRNA from cultures stored in 70% ethanol. Aliquots of a suspension of M. tuberculosis H37Ra colonies (~ 2.0 × 108 CFU) were in triplicate treated with 70% ethanol and extracted fresh or after undergoing five different freeze-thaw scenarios (a). After RNA extraction, samples were treated with TURBO DNase and RNA quantity was determined by Qubit fluorometry (b) and RNA integrity by TapeStation (c). The graphs show the mean quantity of triplicates with the standard deviation.
Discussion
In this study, we compared RNA preservation of cultured M. tuberculosis complex in two different buffers, GTC-TCEP and 70% ethanol. Our results show that the success of RNA preservation is highly dependent on the storage conditions and that 70% ethanol performed equally or better than GTC-TCEP in the assays used.
Firstly, both buffers stabilise the expression levels of hsp65, sigA and 16S rRNA during a heat shock assay used to analyse stress-induced transcriptomic changes20. The mycobactericidal effect of GTC-TCEP11,12 and 70% ethanol16,21 is probably part of the reason for this unaltered expression as in viable mycobacterial cells changes in hsp65 expression have been reported21. Although the full transcriptome was not analysed, the results indicate that de novo transcription induced by metabolic changes in the bacteria is unlikely after treatment with GTC-TCEP and 70% ethanol.
A baseline comparison between the two buffers in the independently conducted long-term storage and freeze-thaw experiments revealed that ± 2.5 x more total RNA was recovered from ethanol compared to GTC-TCEP preserved samples. In line with this, not only 16S rRNA but also mRNA targets (esxA, icl, sigA) were more abundant in ethanol-preserved samples, a finding of special importance when RNA is recovered from low bacillary burden samples. For GTC-TCEP the unexpected peaks in the Qubit data at 4 °C and 30 °C can possibly be explained by contamination. The higher RNA yield after ethanol storage could be attributable to disrupted peptidoglycan crosslinks, leading to more efficient lysis and increased RNA availability for extraction22. The time-dependent increase in copy numbers for most targets suggests that this cell-wall weakening effect builds over longer storage times.
For further examination of RNA stability, a ratio was calculated per timepoint (see Materials and Methods) with a value of 1.0 reflecting “perfect” preservation without loss in yield compared to -80 °C. A stable ratio of 1.0 over the whole 12-month study period was not achieved for any of the targets, indicating RNA degradation. In GTC-TCEP, degradation was inconsistent for all targets but esxA where the ratio remained relatively stable at -20 and 4 °C for twelve months, possibly because this small 288 bp transcript is less affected by degradation processes. In contrast, for ethanol-preserved samples the ratio followed a temperature- and time-dependent pattern. Within the first six months, results were comparable between − 20 °C and − 80 °C storage across all targets. Higher temperatures reduced this window to one month at 4 °C and to only overnight when stored at 30 °C. Therefore, samples stored at different temperatures remain difficult to compare. However, when stored at the same temperature, the ratio remains comparable within these time limits for different targets, while comparing absolute transcript numbers requires more caution. The role of transcript-specific degradation rates is not fully understood as previously published studies arrived at contradicting conclusions23. It is assumed that different transcripts do not degrade uniformly in dying tissues, however, in this study this effect was not observed for ethanol storage, favouring this buffer for long-term sample storage. Although both mRNA and rRNA targets were included, the limited data of only four targets cannot exclude that also during ethanol preservation, the transcriptome-wide degradation pattern may differ and should therefore be subject of future research.
Further, changes in RNA integrity during the long-term storage of mycobacterial culture samples were investigated. Although low RIN values are not an exclusion criterium for RNAseq23,24 because they are rRNA-based and imperfectly mirror mRNA integrity, a cutoff value of RIN > 7.0 is commonly used to predict M. tuberculosis RNAseq success18,19. In this study, samples stored in 70% ethanol at -80 and − 20 °C maintained high RIN values (> 7.3) for twelve months, while storage at 4 °C caused a slight drop from RIN 7.2 (3 months) to RIN 6.1 (6 months). This aligns with studies demonstrating ethanol’s effectiveness in preserving good-quality RNA in spiders and freshwater insects when stored in the fridge or freezer25,26. Conversely, in GTC-TCEP the RIN declined earlier, after two weeks at 4 °C and after six months at -20 °C, indicating a less efficient RNA stabilization. Interestingly, storing samples overnight at 30 °C had a highly detrimental effect on the RNA in GTC-TCEP but not in 70% ethanol.
To our knowledge, this study is the first to evaluate the impact of repeated freeze-thaw cycles on RNA preservation in mycobacteria. Five different freeze-thaw scenarios were applied to simulate handling of specimens, e.g. as part of different experimental procedures or when samples require transport to a centralized lab for resource-intensive RNA analyses. The finding that up to three freeze-thaw cycles do not affect the RNA yield from cultured M. tuberculosis H37Ra is promising. The same is the case for RNA integrity that also remained stable in ethanol-preserved samples; however, no conclusion could be drawn for GTC-TCEP due to RIN calculation failure. Therefore, our findings suggest a potential cryoprotective effect of ethanol27 that reduces crystal formation during freezing of mycobacterial cultures. For human samples, freeze-thaw effects on RNA quality vary from minor to significant, depending on the study setup and tested tissue28. For example, in a study with human leukocytes in RNAlater researchers found a 3’ bias in coverage after freeze-thawing29. As we did not perform RNAseq, our data cannot draw similar conclusions for mycobacterial cells preserved in 70% ethanol or GTC-TCEP.
A future direction will be testing 70% ethanol for use in clinical samples containing virulent mycobacterial strains. Using 4–5 volumes of GTC-based buffer to preserve a 0.5 ml sputum sample6, costs approximately € 0.27, while diluting to 70% ethanol would account for only € 0.027. Additionally, (molecular grade) ethanol can be purchased worldwide and can be shipped under less strict IATA guidelines when the volume is below 1 ml30, while other RNA-preserving buffers may be subject to varying availability. Although, 70% ethanol has been reported as insufficient to disinfect sputum spiked with M. tuberculosis H37Rv within 1 min31, a longer contact time with this buffer inactivates cultures (15 min) and spiked sputum sediment (1 h)16,32. This suggests that sputa may have to be homogenised prior to ethanol treatment to yield efficient inactivation. Based on this, 70% ethanol shows potential field-friendliness, with no impact on RNA yield and integrity after one day at tropical temperature and three freeze-thaw cycles, possibly simplifying shipment workflows, though confirmation with clinical samples is needed.
Concluding from the results of this study, 70% ethanol is superior to GTC-TCEP in terms of RNA-preservation of pure cultures of M. tuberculosis H37Ra and can be used without freezing to -80 °C. For maximal RNA preservation we recommend storing mycobacterial cultures in 70% ethanol at 4 °C for up to one month, while for long-term storage − 20 °C are preferred. However, as at 4 °C RNA integrity remains high and qPCR results of different targets remain comparable after the same storage time, storage at 4 °C for three months or longer may be considered dependent on the downstream application.
Data availability
Data is provided within the manuscript and supplementary information files. Additional raw data are available from the corresponding author on reasonable request.
References
Kumar, S. S. & Ghosh, A. R. Assessment of bacterial viability: A comprehensive review on recent advances and challenges. Microbiol. (United Kingdom). 165, 593–610 (2019).
Westermann, A. J. & Vogel, J. Cross-species RNA-seq for Deciphering host–microbe interactions. Nat. Rev. Genet. 22, 361–378 (2021).
World Health Organization (WHO). Global tuberculosis report 2024. (2024).
World Health Organization. Global leprosy (Hansen disease)update, 2023: elimination of leprosydisease is possible – Time to act! Wkly. Epidemiol. Rec Relev Épidémiologique Hebd. 99, 501–521 (2024).
Wallace, E. et al. Culturing Mycobacteria. in 1–58 (2021). https://doi.org/10.1007/978-1-0716-1460-0_1
Honeyborne, I. et al. Molecular bacterial load assay, a culture-free biomarker for rapid and accurate quantification of sputum Mycobacterium tuberculosis bacillary load during treatment. J. Clin. Microbiol. 49, 3905–3911 (2011).
Walter, N. D. et al. Mycobacterium tuberculosis precursor rRNA as a measure of treatment-shortening activity of drugs and regimens. Nat. Commun. 12, 2899 (2021).
Davis, G. L. et al. Molecular assays for determining Mycobacterium leprae viability in tissues of experimentally infected mice. PLoS Negl. Trop. Dis. 7, (2013). https://doi.org/10.1371/journal.pntd.0002404
Lenz, S. M. et al. Utility of a Mycobacterium leprae molecular viability assay for clinical leprosy: an analysis of cases from the Philippines, Ethiopia, and Nepal. Front. Trop. Dis. 3, 1–17 (2022).
Hanke, M., Hansen, N., Tomm, E., Grundmeier, G. & Keller, A. Time-Dependent DNA Origami denaturation by guanidinium chloride, guanidinium sulfate, and guanidinium thiocyanate. Int. J. Mol. Sci. 23, 8547 (2022).
Krausser, L. et al. Mycobacterium tuberculosis retains viability in RNAlater buffer but not in GTC-TCEP and DNA/RNA shield. Diagn. Microbiol. Infect. Dis. 106, 115905 (2023).
Mtafya, B. et al. A practical approach to render tuberculosis samples safe for application of tuberculosis molecular bacterial load assay in clinical settings without a biosafety level 3 laboratory. Tuberculosis (Edinb). 138, 102275 (2023).
Musisi, E. et al. Reproducibility of the ribosomal RNA synthesis ratio in sputum and association with markers of Mycobacterium tuberculosis burden. Microbiol. Spectr. 9, 1–7 (2021).
Palekyte, A. et al. Acetic acid enables molecular enumeration of Mycobacterium tuberculosis from sputum and eliminates the need for a biosafety level 3 laboratory. Clin. Chem. 70, 642–652 (2024).
Lahiri, R., Randhawa, B. & Krahenbuhl, J. Application of a viability-staining method for Mycobacterium leprae derived from the athymic (nu/nu) mouse foot pad. J. Med. Microbiol. 54, 235–242 (2005).
Williams, D. L., Gillis, T. P. & Dupree, W. G. Ethanol fixation of sputum sediments for DNA-based detection of Mycobacterium tuberculosis. J. Clin. Microbiol. 33, 1558–1561 (1995).
Manganelli, R., Dubnau, E., Tyagi, S., Kramer, F. R. & Smith, I. Differential expression of 10 Sigma factor genes in Mycobacterium tuberculosis. Mol. Microbiol. 31, 715–724 (1999).
Walker, J. E. et al. Measuring up: A comparison of tapestation 4200 and bioanalyzer 2100 as measurement tools for RNA quality in postmortem human brain samples. Int. J. Mol. Sci. 24, 13795 (2023).
Ma, S., Jones, R. M., Gleason, N. S., Farrow-Johnson, J. & Sherman, D. R. Experimental and computational workflow for RNA sequencing in Mycobacterium tuberculosis: from total RNA to differentially expressed genes. in 481–512 (2021). https://doi.org/10.1007/978-1-0716-1460-0_21
Stewart, G. R. et al. Dissection of the heat-shock response in Mycobacterium tuberculosis using mutants and microarrays. Microbiology 148, 3129–3138 (2002).
Stewart, G. R. et al. The heat shock response of Mycobacterium tuberculosis: linking gene expression, immunology and pathogenesis. Comp. Funct. Genomics. 3, 348–351 (2002).
Ingram, L. O. & Vreeland, N. S. Differential effects of ethanol and hexanol on the Escherichia coli cell envelope. J. Bacteriol. 144, 481–488 (1980).
Gallego Romero, I., Pai, A. A., Tung, J. & Gilad, Y. RNA-seq: impact of RNA degradation on transcript quantification. BMC Biol. 12, 42 (2014).
Puchta, M., Boczkowska, M. & Groszyk, J. Low RIN value for RNA-Seq library construction from Long-Term stored seeds: A case study of barley seeds. Genes (Basel). 11, 1190 (2020).
Kono, N., Nakamura, H., Ito, Y., Tomita, M. & Arakawa, K. Evaluation of the impact of RNA preservation methods of spiders for de Novo transcriptome assembly. Mol. Ecol. Resour. 16, 662–672 (2016).
Astrid, T., Margit, E., Leopold, F. Ethanol: A simple and effective RNA-preservation for freshwater insects living in remote habitats. Limnol. Oceanogr. Methods. 14, 186–195 (2016).
Lewis, J. G., Learmonth, R. P. & Watson, K. Cryoprotection of yeast by alcohols during rapid freezing. Cryobiology 31, 193–198 (1994).
Shabihkhani, M. et al. The procurement, storage, and quality assurance of frozen blood and tissue biospecimens in pathology, biorepository, and biobank settings. Clin. Biochem. 47, 258–266 (2014).
Kellman, B. P. et al. Multiple freeze-thaw cycles lead to a loss of consistency in poly(A)-enriched RNA sequencing. BMC Genom. 22, 69 (2021).
International Air Transport Association (IATA). Dangerous Goods Regulations (DGR). (2024).
Best, M., Sattar, S. A., Springthorpe, V. S. & Kennedy, M. E. Efficacies of selected disinfectants against Mycobacterium tuberculosis. J. Clin. Microbiol. 28, 2234–2239 (1990).
Dunne, W. M. et al. Rapid inactivation of Mycobacterium and nocardia species before identification using Matrix-Assisted laser desorption Ionization–Time of flight mass spectrometry. J. Clin. Microbiol. 52, 3654–3659 (2014).
Acknowledgements
L.K. was supported by the Research Foundation Flanders (FWO), grant numbers 1SE7522N + 1SE7524N. The study was kindly supported by the IntegrOmics project (FWO, grant number G0B0222N) and the TB/NTM Research Cluster (FWO, grant number W001822N).
Author information
Authors and Affiliations
Contributions
L.K.: Conceptualisation, Methodology, Investigation, Formal Analysis, Visualisation, Writing –Original Draft, Writing – Review and Editing; M.V.D.: Investigation, Validation, Writing – Review and Editing; R.B.: Investigation, Writing – Review and Editing, R.R.: Investigation, Writing – Review and Editing; R.A.: Investigation, Writing – Review and Editing; L.R.: Supervision, Writing – Review and Editing; N.D.W.: Resources, Writing – Review and Editing; M.I.V.: Resources, Writing – Review and Editing, A.V.R.: Supervision, Writing – Review and Editing; B.C.J.: Conceptualisation, Writing – Review and Editing, Project Administration, Supervision; S.M.B.: Conceptualisation, Methodology, Project Administration, Supervision, Writing – Original Draft, Writing – Review and Editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Krausser, L., Van Dyck-Lippens, M., Balde, R. et al. 70% ethanol preserves mycobacterial RNA from cultures more efficiently than GTC-TCEP. Sci Rep 15, 12322 (2025). https://doi.org/10.1038/s41598-025-93699-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-93699-7