Abstract
Handling the threats of intellectual property (IP) piracy and false IP ownership claim, are central from the perspective of IP vendor’ right as well as reliability of system-on-chip (SoC) designs. IP design piracy has become an emergent concern for the digital hardware design community, in the last few years. This paper presents a novel behavioral synthesis based hardware IP Protection methodology that exploits IP vendor’s proteogenomic Bio-marker as digital Watermark (BioW-IPP), for detective countermeasure against IP piracy and quashing false IP ownership claim. The proposed approach is capable of exploiting the IP vendor’s bio-watermark (proteogenomic signature) as secret digital evidence (BW-ID) to generate a robust hardware watermark for embedding into the IP design during behavioral synthesis/high level synthesis (HLS) process. The generated proteogenomic signature bio-watermark is embedded into the register allocation phase of behavioral synthesis (HLS) that ensures negligible design overhead post-embedding. The proposed approach on comparison with prior behavioral synthesis based watermarking techniques achieved greater security in terms of lower probability of coincidence (watermark collision) and higher tamper tolerance, at nominal design overhead.
Similar content being viewed by others
Introduction
In the field of electronics and multimedia systems, the efficient design of data and computation applications is critical for achieving enhanced performance. One of the most prominent computation intensive applications in the domain of image and video processing are joint photographic experts’ group compression decompression (JPEG-CODEC) application, image processing filters, etc. These applications play a pivotal role in handling data-intensive tasks (that involve identifying and isolating key attributes or characteristics of an image), that are fundamental to modern multimedia applications. Given the computational complexity and high throughput requirements of these operations, there is an increasing need for the development of dedicated hardware solutions tailored to address these challenges efficiently. Such specialized hardware ensures optimized performance, reduced latency, and lower energy consumption. The design process of these hardware systems involves various abstraction levels, including (a) algorithmic level, (b) system level, (c) register transfer level (RTL), (d) gate level, and (e) layout level. High level synthesis is a design process that automatically transforms a design representation (application) from the algorithmic level to the RTL counterpart. These data intensive applications can be efficiently designed as dedicated reusable intellectual property core using HLS framework1,2. Further, the electronic design industry relies heavily on the global integrated circuit supply chain of dedicated hardware IP core. As system-on-chip (SoC) design becomes more commoditized, the industry faces growing challenges, such as security vulnerabilities and escalating concerns about piracy, which pose significant risks to global supply chains. As the design and manufacturing processes often involve multiple parties across various geographical and organizational boundaries, they become susceptible to malicious activities by internal attackers within the SoC house. The potential hardware threats include IP piracy and unauthorized claim of IP ownership which may arise during the SoC integration or fabrication processes. Also, pirated/counterfeited IPs may contain malicious logic, resulting in risks such as functional malfunctions, overheating, data breaches, and reputational harm to original IP vendors. Additionally, IPs can be misused in various ways, including fraudulently claim by an adversary. From the perspective of original IP vendor, attacker might be present in the SoC house, which might be engaged into piracy and false claim of IP ownership rights. These situations can lead to significant financial losses, damage to the reputation of original IP vendor, and erosion of trust in the IP market design supply chain3,4,5,6,7,8,9,10,11,12. IP piracy involves the unauthorized replication or use of proprietary IP, while false claims of ownership can result in legal disputes and undermine the rightful owner’s ability to monetize their work.
In order to provide detective countermeasure/control against the aforementioned threats, security constraints (as IP vendor’s digital evidence) can be implemented across various design process abstraction levels, such as algorithmic, RTL, gate level, and physical level. Leveraging HLS for watermarking offers security at higher abstraction levels while reducing complexity and cost. This approach enhances IP protection by providing detective control against IP piracy, and false IP ownership claim by embedding security measures early in the design process1,2. HLS enables designers to integrate watermark constraints seamlessly during key stages of the design process, such as the register allocation and binding phase. This class of technique introduces minimal overhead to the overall design, ensuring that performance, area, and cost remain largely unaffected. This paper presents a novel hardware watermarking technique by employing proteogenomic bio-markers as robust digital watermark. The generated digital watermark is integrated as security/watermarking constraints into the hardware IP designs, which serve as robust and irrefutable evidence of original IP vendor. In cases of piracy or ownership disputes, these watermarks can be used as definitive proof of the rightful owner’s claims, providing a solid foundation for legal action and resolution. By adopting such advanced security techniques, IP vendors not only safeguard their assets but also contribute to the overall integrity and trustworthiness of the hardware IP design process.
Novel contribution of this paper
-
1.
This paper presents a novel hardware watermarking methodology that leverages the IP vendor’s proteogenomic biomarker to provide detective countermeasure against IP piracy and false IP ownership claim.
-
2.
The proposed approach is capable of exploiting the IP vendor’s bio-watermark (proteogenomic signature) as secret digital evidence (BW-ID) to generate a robust hardware watermark. The proposed approach has been demonstrated on JPEG-CODEC IP core.
Related works
In the literature, there have been some prior works that have been attempted to secure IP cores against false claim of IP ownership and potential threats of IP piracy. This includes hardware watermarking13,14,15,16 facial biometric17, steganography18, DNA biometric19, handwritten signature20 and digital signature21. The authors in13 proposed a hardware watermarking approach based on pragma insertion during the functional unit allocation phase of the HLS process. Additionally, the pragma insertion-based approach restricts the generation of large-size constraints, and the work lacks a comprehensive security analysis, including evaluations of watermark collision, resistance to brute-force attacks, and susceptibility to standard attack methodologies. Authors of17 proposed watermarking approach using facial biometric features of an IP vendor to generate watermark security constraints. Approach17 is incompetent of generating large watermark signature due to limited facial features. The watermarking approach described in14 employs a binary encoding technique to encode vendor’s signature into watermarking constraints, that are incorporated into the design process after embedding. This approach is capable of producing good watermark strength, however, doesn’t produce as strong watermark as the proposed approach in term of size and security. In15 authors proposed the method to embed digital signature bits within the hardware description language (HDL) design, utilizing message digest 5 (MD5) and secure hashing algorithm (SHA-1), but it leads to design area overhead. The approach in16 also uses SHA-1 and RSA at the HDL level, but it becomes vulnerable if the RSA key is revealed. This method relies entirely on the RSA key, as it is the sole security layer in the watermarking technique. Moreover, the approach in19 leverages the DNA biometric of IP vendors to generate security constraints. This approach is promising however, it doesn’t generate watermark signature based on proteogenomic bio marker of an IP vendor. This results in lesser security in terms of tamper tolerance as compared to the proposed approach. The approach in20 exploits the handwritten signature image of IP vendors to generate digital template, while the authors of21 proposed multi-level encoding and encrypted-hash-based digital signature for hardware protection. Both approaches20,21 generate lower watermark strength as digital evidence for IP protection. Similarly, the steganography-based approach in18 embeds secret marks through steganography constraints. However, its effectiveness diminishes if the design data or steganography encoding process is compromised, and the generation of steganography constraints adds significant implementation complexity. On the contrary, the proposed proteogenomic based watermarking employs the following several security layers, which an attacker needs to completely decode to regenerate the original watermark constraints to falsely claim IP ownership: (a) type of proteogenomic signatures (such as protein sequence signature and DNA sequence signature) provided by IP vendor, (b) proteogenomic signature fusion order to generate final watermark signature, (c) IP vendor’s signature generation encoding rule, and (d) watermark constraints mapping/embedding rule.
The proposed methodology: BioW-IPP
Motivation and overview of proposed approach
Proteogenomic is a hybrid combination of protein molecular biometric and DNA biometric information of an individual. Proteogenomic can serve as an important bio-marker during forensic analysis and identification. There are only about 20 different types of amino acids in the human body, but these can combine to make approximately 20,000 unique proteins. The order of amino acids in the polypeptide chain is determined by the order of nucleotides (the DNA sequence) of the gene that encodes it. Even a tiny change in the amino acid sequence of the polypeptide chain can alter the overall structure and function of the protein. This diversity of amino acids and the sheer number of possible combinations in their linear order allow for tremendous dissimilarity in the properties of proteins. The choice of amino acid sample (protein sample) augmented with the peptide chain (fragment) strength dictates the uniqueness generated from an IP vendor’s perspective22. Exploiting the formation of a robust integrated proteogenomic signature post-extraction of protein biometric signature and DNA signature from its respective samples, creates an exclusive, inimitable and imperative piece of digital bio-evidence that can be leveraged to generate a sturdy hardware watermark. Note: The respective sample of the IP vendor is taken after due consent of the respective entity and the obtained DNA/protein sequence from the lab is safely stored in a secure database. There will be a non-disclosure agreement (NDA) with the respective lab to preserve the privacy of the entity. Thereafter, obtaining/extracting the DNA/protein signature (to form proteogenomic signature) after receiving the respective sequences, is a straight-forward process using the flow in (Fig. 1). As established in several works earlier17,19,23,24, usage of biometric identifier is not extracted from a regular employee, but rather from an entity of the life board of trustees/owner/founder (who is also a trustworthy insider in the IP vendor’s house). In the proposed hardware watermarking approach, we bio-mimic the generation of proteogenomic signature from an IP vendor’s body sample for forming its respective bio-watermark as secret digital evidence (BW-ID). This BW-ID is embedded covertly in a hardware IP for providing detective countermeasure against IP piracy and false IP ownership. Our BW-ID enables us to create a powerful hardware watermarking framework. Even though the proposed approach combines the knowledge of extracting and encoding proteogenomic signature, however, it only requires the involvement of sequencing labs for providing the IP vendor with his/her DNA sequence and protein sequence respectively. Once the DNA sequence and protein sequence are received, then an IP vendor can algorithmically generate the alpha/beta chain protein signatures and DNA signature respectively for watermark generation, using the proposed flow in (Fig. 1). There is no additional knowledge of bioinformatics required for hardware watermark constraint generation. Therefore, the proposed proteogenomic signature-based watermark constraint generation process can be applied to any hardware IP designs with the knowledge of high-level synthesis design process and proposed watermark generation algorithm (Fig. 2).
The paper presents a novel HLS methodology to secure hardware IP cores using BW-ID of authentic IP vendor. This gives a distinctive molecular level identification of the authentic IP provider. These bio-markers are taken from her/his body sample (insulin and DNA). The primary inputs of the proposed secure HLS framework (in Fig. 1) consist of the module library, data flow graph (DFG) or control data flow graph (CDFG) of the hardware application, resource constraints, scheduling algorithm, protein sequence of IP vendor (alpha chain and beta chain), DNA sequence of IP vendor, watermark encoding mechanism (watermark constraint generator), and fusion order to generate proteogenomic based watermark. This proposed secure HLS flow consists of four main sections, viz. (i) protein signature generation from alpha chain (1st polypeptide chain), (ii) DNA signature generation from DNA sequence, (iii) protein signature generation from beta chain (2nd polypeptide chain), and (iv) HLS design process. After applying IP vendor’s provided fusion order, an integrated proteogenomic digital watermark template (BW-ID) is generated. Further, by using the watermark encoding rule of the IP vendor, the generated watermark template is converted into its respective hidden security constraints. Based on the primary inputs, different HLS steps of scheduling, allocation, binding, and register allocation are performed, followed by construction of the initial register allocation table (RAT), which describes the assignment of initial storage variable into distinct registers for the given hardware application (DFG). Subsequently, the previously generated watermark security constraints are integrated into the register allocation phase of HLS process for the target hardware application. Finally, secured register transfer level (RTL) hardware design is generated carrying proteogenomic watermark impression. This design contains the IP vendor’s robust watermark signature that can provide detective countermeasure against IP piracy and resolution of false IP ownership. Detailed explanations of the proposed approach are given in the next section.
Details of the proposed methodology
The proposed methodology to generate proteogenomic digital watermark for securing hardware IP cores against IP piracy/false claim of authentic IP ownership are demonstrated in detail through the following modules (section III.B.1 – III.B.5). The secured HLS design flow of the proposed approach is explained in (Fig. 2).
Protein watermark signature generation from alpha chain—1st polypeptide chain
The structure of human body protein (extracted from insulin) consists of a sequence chain of amino acids. The connection between these amino acids is formed by bonding the carboxylic acid group of one amino acid with the amine group of another amino acid, which is known as a peptide bond. Peptide bonds link amino acids together to create a series of polypeptide chains or a protein sequence. As shown in (Fig. 3), there are two different protein sequences (polypeptide chain) of IP vendor’s human body insulin. 1st polypeptide chain is named/termed as ‘Alpha chain’, and second polypeptide chain is named/termed as ‘Beta chain’. There are nearly 500 amino acids in the nature, but in human protein sequence, there are 20 unique amino acids25,26 (in Fig. 3) which link together to create the amino acid chain. Figure 2 explains the process of generation of digital watermark (AC) from its respective alpha chain. Based on the IP vendor’s protein sequence (as input), each amino acid is encoded into unique encoding bit using the proposed encoding rule, which subsequently after concatenation of the binarized bits generates a unique protein watermark signature (AC). Further, Fig. 4 shows the encoding of the 1st polypeptide chain, which consists of 21 amino acids (Note: The shaded protein elements in the table are not part of the alpha chain). Each amino acid is encoded uniquely according to their positions in the alphabet (encoding rule), like, the amino acid Glycine (G) has an alphabet position of 7 and is encoded with binary bits 111. Similarly, for Isoleucine (I), the alphabet position is 9 and corresponding binary representation is 1001. Finally, ‘AC’ is generated by combining the binarized bits that represent the amino acids in the alpha chain (as shown in Fig. 4). Finally, the total size of watermark strength from IP vendor’s 1st polypeptide chain is 81 bits.
DNA signature generation
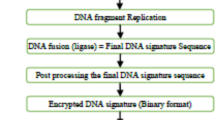
In the proposed approach, chromosomal DNA sequence contains two base pairs (BP). First BP is generated with two chemical elements (T-Thymine and A-Adenine), and second BP is generated with other two chemical elements (G-Guanine and C-Cytosine). Subsequently, DNA sequence can be generated in two ways, either taking alternative base pairs of similar type (like, TA-AT-GC-CG) or taking alternative base pairs of distinct type (like, TA-GC-AT-CG). In our proposed approach, combinations of base pairs of similar type are assumed in the DNA sequence. The polynucleotide (sugar phosphate backbone—represented as ‘S’) has been included as both the leading and lagging strands in the DNA sequence to create the final DNA sequence. Figure 2 explains the process of generation of digital watermark (D) from the respective IP vendor’s DNA sequence. Based on the IP vendor’s DNA sequence (as input), each chemical elements are encoded into unique encoding bit using the proposed encoding rule (according to their positions in the alphabet), which subsequently generates a unique DNA watermark signature (D). IP vendor’s DNA sequence (STAS-SATS-SGCS-SCGS-STAS-SATS-SGCS-SCGS-STAS-SATS-SGCS-SCGS-STAS-SATS-SGCS-SCGS) is depicted in (Fig. 5) (adopted from19). Subsequently, it is converted into digital template using the encoding rule. For example, the chemical element Guanine (G) has an alphabet position of 7 which is encoded with binary bits 111. Similarly, for Polynucleotide (S), the alphabet position is 19 which is encoded with binary bits 10011. Finally, the corresponding encoded DNA watermark signature (D) is shown in Fig. 5 where the total size of watermark strength is 248 bits.
Protein watermark signature generation from beta chain—2nd polypeptide chain
Figure 2 explains the process of generation of digital watermark (BC) from its respective beta chain. Similar to the 1st Polypeptide chain, based on the IP vendor’s protein sequence, each amino acid is encoded into unique encoding bit using the proposed encoding rule, which subsequently after concatenation of the binarized bits generates a unique protein watermark signature (BC). Further, Fig. 6 shows the encoding of the 2nd polypeptide chain, which consists of 30 amino acids (Note: The shaded protein elements in the table are not part of the beta chain). Each amino acid is encoded uniquely according to their positions in the alphabet (encoding rule). The amino acid Phenylalanine (F) has an alphabet position of 6 which is encoded with binary bits 110. Similarly, for Valine (V), the alphabet position is 22 which is encoded with binary bits 10,110. Finally, ‘BC’ is generated by combining the binarized bits that represent the amino acids in the beta chain (as shown in Fig. 6). Finally, the total size of watermark strength of IP vendor’s 2nd polypeptide chain is 116 bits.
Details of JPEG-CODEC DFG
In the proposed approach, JPEG-CODEC benchmark (DFG) (in Fig. 7, adopted from27) is used as an example to demonstrate the watermark (security) constraints generation process and its mapping/embedding process (Note: DFG of any other benchmark could also have been used for demonstration purpose). It comprises of 2-dimensional discrete cosine transformation (2D-DCT) in the core of its operation process. Post-DCT process, it includes the quantization procedure as well. To perform DCT on a given image input block, the 2D-DCT coefficient matrix (T) is applied to the input block (M) using the following formula27:
where, ‘D’ is calculated through (2)
Elements of ‘D’ matrix (d11,d12,…,d88) indicates column wise transformed elements of input image block. Further, elements of ‘X’ matrix (X11, X12,…,X88) indicates both row and column wise transformed elements of input image block. Thus, Ttrans is the transpose of 2D-DCT coefficient matrix ‘T’, and ‘X’ is the corresponding discrete cosine transformed block of input image block M.
For a dedicated IP core design, the matrix relationship must be transformed into a hardware function, for that reason—the pixels (mij) of the input image block (M) are transformed into compressed image pixel (Xij) (using (1)). For example, X11 which is the first pixel of the compressed image is modeled as follows27:
where, d11, d12,…,d18 is calculated as follows:
Similar transformations are made to the other pixels of the block (M), where the input pixels stay the same, but the 2D-DCT coefficients change. Therefore, the equation’s structure stays the same, but the inputs used to calculate the various modified image pixel intensities alters. Each macro-IP is created using eight structurally similar micro-IPs (named IP1-IP8), and each micro-IP uses (4)-(5) to perform its dij. Every micro-IP operation, multiplication, and addition operation are represented by light brown, green, and blue color nodes respectively. As illustrated in (Fig. 7), the result of operation 135 produces the pixel intensity of the transformed image (X). All the scheduling, binding and allocation operations of the JPEG-CODEC are shown in (Table 1).
After the DCT transformation, the pixel intensities of the 8 × 8 image block (X) are subsequently compressed using quantization technique. To obtain various quality levels (from 1 to 100), a distinct quantization matrix is utilized. Quality level 100 signifies reduced compression resulting in a higher quality image, while quality level 0 signifies increased compression resulting in a lower quality image. This is accomplished by operation 136, which is executing the quantization on the transformed image pixel intensities (X) using the matrix element ‘q’ of the quantization matrix (Q). This generates the final output as compressed/quantized image pixel values (X’). The next sub-section of this paper utilizes JPEG-CODEC SDFG to derive the register allocation information for watermark constraints generation and its subsequent embedding.
Final digital template and conversion into security constraints, and its embedding
The final proteogenomic based digital watermark signature is generated after following the IP vendor’s provided fusion order i.e., the concatenation of ‘AC’, ‘D’ and ‘BC’ (TWS = AC ╫D╫BC) accordingly. Consequently, this watermark signature is converted into the watermark (security) constraints using IP vendor’s chosen mapping/embedding rule.
Watermark mapping/embedding rule
If the proteogenomic digital watermark signature bit is 0, then even-even storage variable pairs < Vi, Vj > from the SDFG are generated as watermarking constraints, where ‘i’ and ‘j’ are the storage variable numbers. In contrast, for an encrypted watermark signature bit 1, the generated constraints are odd-odd storage variable pairs in the SDFG.
This indicates that the mapping of the watermark constraints happens during the register allocation phase of the HLS process. In the JPEG-CODEC DFG (shown in Fig. 7), different registers (used to store variables) are represented with different colors, and storage variables are represented as V0-V272. In SDFG, storage variables are used for storing the primary and intermediate inputs/outputs of the operations temporarily. The SDFG of JPEG-CODEC is used to generate the corresponding RA table, which shows the optimal allocation of storage variables to registers (using register sharing). The following section provides a detailed explanation of how to create watermark security constraints using the proteogenomic digital watermark and its embedding (explained with the same JPEG-CODEC example). In the proposed method, final proteogenomic based digital watermark (TWS) obtained is shown in (Fig. 8), where total size of watermark strength is 445 bits (which consists of 168 zeros and 277 ones). These 445 bits are mapped and embedded as watermark constraints in the RA table of the IP design (using the above mapping rule). Therefore, the watermark constraints obtained for JPEG-CODEC IP design are as follows: < V0, V2 > , < V0, V4 > ,…, < V0, V270 > , < V0, V272 > , < V1, P3 > , < V1, V5 > ,…, < V1, V269 > , < V1, V271 > , < V2, V4 > ,…, < V2, V64 > , < V2, V66 > , < V3, V5 > , …, < V3, V271 > , < V5, V7 > , …, < V5, V19 > , and < V5, V21 > .
In the HLS process, the register allocation stage integrates these constraints into the RAT. To ensure storage variable pairs corresponding to watermarking constraints are not assigned to the same register, registers are either locally swapped, or a new register is allocated. This approach leads to unique register allocations corresponding to storage variable pairs of watermarking constraints. Table 2 shows the RAT for the JPEG-CODEC prior and after the integration of the proposed watermark constraints, with modified register allocation highlighted in red color. In this final RA table, registers are defined as (R1—R137), control steps are defined as (C0—C30), and storage variables are defined as (V0—V272). This RAT, after embedding watermark constraints, is subjected to classical HLS process to generate a secure JPEG-CODEC RTL IP design. Therefore, the existence of proteogenomic based watermark constraints, based on the BW-ID of authentic IP vendor, serves as detective countermeasure against potential threats of IP piracy and fraudulent claims of IP ownership.
Nullification of false IP ownership claim and detection of IP piracy
An attacker or third-party IP vendor may attempt to participate in false claiming IP ownership or IP piracy using tampering attack, brute-force attack and watermark collision attack. To converse these threats, the proposed proteogenomic based digital watermark must ensure robust protection/security against potential threats of IP piracy and fraudulent claims of IP ownership. In instances of disputes regarding IP ownership, the conflict is settled using the integrated IP vendor’s proteogenomic based digital watermark constraints which is embedded in the IP design as digital proof. Ownership validation involves extracting watermark constraints from the IP’s RTL file and matching them with the original watermark constraints (which can be recreated using the proposed proteogenomic based digital watermark signature generation algorithm, fusion order, and mapping/embedding rule). Only the authentic IP owner can effectively meet these requirements, prohibiting the opposition from successfully claiming the ownership in IP court. Moreover, to identify IP piracy, the original watermarking constraints can be compared with those constraints which are generated from the suspected chip; a match would verify/confirm the presence of IP piracy.
Security properties of the proposed approach
The proposed approach is resilient against potential forgery attack or misuse after forgery. It is also resilient against potential false claim of IP ownership threat. The proposed approach includes several security layers of an IP vendor for watermark constraints generation which an attacker needs to decode such as: (a) determination of polypeptide alpha chain length, (b) encoding of the amino acids of the alpha chain, (c) determination of polypeptide beta chain length, (d) encoding of the amino acids of the beta chain, (e) DNA sequence length, (f) encoded DNA chemical element, (g) proteogenomic signature fusion order, and (h) watermark embedding rule. An attacker needs to decode all the parameters (a)–(h) in succession, in order to regenerate, establish and prove the original watermark in front of the third-party authenticator. Therefore, reverse engineering only the encoding rule or fusion order is not sufficient to regenerate, establish and prove the original watermark in front of the third-party authenticator. Additionally, even if the alpha chain or beta chain or DNA sequence of the IP vendor is forged/compromised, still an attacker needs to decode security layers (a)–(h) to regenerate and establish the original embedded watermark for falsely claiming IP ownership.
Results and analysis
This section analyses the proposed methodology in terms of the impact of proposed BW-ID watermark feature on achieved watermark strength (used as digital proof/evidence), followed by its security potency (robustness against IP piracy), and design overhead. The experimental evaluation of the proposed methodology is carried out on a 1.80 GHz processor and 8 GB memory. For area and latency estimation, the proposed methodology employs a 15 nm technology based on the NanGate library28.
Analysis on final digital watermark signature generation for BW-ID feature
In the proposed approach, final digital watermark signature is generated from BW-ID features (DNA and protein). DNA sequence contains 248 bits watermark strength and protein sequence contains 197 bits watermark strength. These 445 bits watermark strength are mapped and embedded as watermark constraints in the RA table of the IP design (using the above mapping/embedding rule). The storage/register comparison before embedding and after embedding (for JPEG-CODEC benchmark) is shown in (Table 3). It is observed that there is no register overhead in the JPEG-CODEC benchmark for embedding the proteogenomic based watermark.
Security analysis
The results of the proposed approach have been evaluated on different designs such as JPEG-CODEC, Blur filter, Sharpening filter, LED filter, and Cardiac-Pacemaker, adopted from29,30. The security analysis of the proposed methodology has been assessed using two important security metrics: probability of coincidence (PC) and tamper tolerance (TT)18,21,31,32,33. The PC (also indicates false positive) is an important metric for evaluating the security robustness of a system, particularly in terms of how likely an IP vendor’s watermark may be present in an unsecured IP design. It is also a measure of IP owners’ digital proof that can be used to nullify the false IP ownership during litigation in IP court. A lower PC value indicates stronger security, which signifies a lesser likelihood of coincidentally matching the watermark constraints in an unsecured design (watermark collision). The PC is evaluated through the following equation18,21,32,33:
where, ‘r’ stands for the total number of utilized unique registers in the IP design before embedding watermark constraints, while ‘c’ represents the total watermark constraints embedded. Table 4 reports the probability of coincidence for the proposed approach corresponding to JPEG-CODEC benchmark for different values of digital watermark constraints. It shows that the value of constraints size, which is desirable for yielding robust security. Further, the values of PC for different benchmarks corresponding to the proposed approach are depicted in (Table 5).
Next, TT metric is the critical measure of secure design’s robustness against tampering attack which is launched by an adversary. An increment in TT value signifies the generation of more signature combinations (search space), diminishing an attacker’s chances of decoding the precise signature combination and identifying the original secret watermarking constraints. This higher TT value enhances the system’s resilience against tampering. It is evaluated through the following equation21,31,32,33:
where, ‘v’ is the number of distinct embedding variables (like 0 and 1), and ‘c’ represents the total generated watermark constraints. Furthermore, Table 6 depicts the comparison of PC and TT among the proposed method and prior works (13,14,15,16,17,18,19,20,21,34). The proposed method shows more robust security with a lesser value of PC and higher value of TT than all similar state-of-the-art methods. This is because the proposed proteogenomic based digital watermark generation method can generate and embed larger watermark constraints compared to prior methods. As the proposed bio-hardware watermark method involves several security layers such as protein sequence signature generation and DNA sequence signature generation, proteogenomic signature fusion order, and watermark constraints mapping/embedding rule etc., therefore, the proposed methodology makes it harder for an attacker to exactly regenerate the embedded digital watermark security constraints or tamper with the proposed embedded digital watermark by exactly decoding the watermark security constraints. Overall, these aforementioned reasons increase the robustness of the proposed approach against IP piracy and fraudulent claims of IP ownership.
Design cost analysis
The overhead/design cost analysis of the proposed method is evaluated using the following metric21,31,32,33:
where, ‘W1’ and ‘W2’ represent the weightage factors for normalizing the watermarked design latency and area (in the proposed approach, W1 = W2 = 0.5). ‘L’ and ‘A’ represent the watermarked design latency and area corresponding to the hardware. Further, ‘Lmax’ and ‘Amax’ represent the corresponding maximum latency and maximum area. Note: The area and latency design metrics of the watermarked IP designs indicate the design area and latency after embedding the digital watermark signature of the IP vendor. Tables 7 and Table 8 depict the registers needed in the IP design before and after the embedding watermark constraints and simultaneous design cost for the proteogenomic based proposed approach corresponding to the LED filter and JPEG-CODEC benchmarks. The proposed proteogenomic based watermark incurs very less (0.037% only) design cost overhead (in terms of register count) for the LED filter IP design because of its smaller size, but it does not incur any design overhead for larger size JPEG-CODEC IP design after embedding. The register count indicates the quantity of storage hardware utilized during the HLS register binding phase (illustrated in Table 2 earlier, which shows the requirement of 137 registers, i.e., R1-R137) of the proposed approach, which subsequently reveals the number of storage variables in the HLS-generated RTL data path of the watermarked IP core. The details of the security/watermark strength generated corresponding to different proteogenomic feature types have been already discussed earlier in section III.B.5 and V.A. These registers generally hold the primary inputs, intermediate outputs of the data processing computation, and final output.
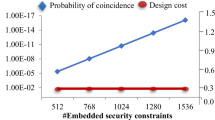
Further, Fig. 9 demonstrates the probability of coincidence vs. design cost trade-off for different watermark constraints corresponding to JPEG-CODEC application for the proposed approach. As shown in this figure, with the increment in the watermark constraints, the PC value decreases without occurring any design cost overhead, because the JPEG-CODEC is a very large size IP design which includes several registers in the datapath.
Table 9 reports the total computational time corresponding to each benchmark, when executed on a system with 1.6 GHz and 8 GB RAM. Computational time contains the process of protein signature and DNA sequence extraction, encoding, and integration; followed by watermark constraints generation and embedding time of the benchmark. As evident from Table 9, the computational time overhead is very low despite the steps involved for proposed watermark signature generation and embedding.
Limitations of the proposed work
There are some limitations of this proposed work:
-
(i)
In the process of converting watermark constraints from the proteogenomic based watermark signature (during HLS), it is essential to carefully generate the storage variable pairs (sorted in ascending order) that correspond to each bit of the watermark signature. If storage variable pairs are not generated carefully as watermark constraints, there is a risk of improperly mapping of signature bits to storage variable pairs.
-
(ii)
The embedding process of the proposed proteogenomic based watermark constraints requires careful local alterations/swapping of storage variables to the corresponding registers during behavioral synthesis process. Without proper precautionary swapping, there is a risk of register conflicts during watermark embedding.
Conclusion and future work
This paper presented a novel hardware IP protection methodology by exploiting IP vendor’s proteogenomic bio-marker to generate a robust digital watermark based digital evidence (BW-ID), for detective countermeasure against IP piracy. The proposed methodology makes it harder for an attacker to exactly regenerate the embedded digital watermark security constraints or tamper with the proposed embedded digital watermark by exactly decoding the watermark security constraints due to several security layers involved. This proposed method has performed better compared to similar prior works in terms of robust security, at negligible design overhead.
In our future work, we intend to explore low-cost proteogenomic bio-watermark for hardware IP protection of loop based applications based on simultaneous exploration of loop unrolling factor and resource configuration (with respect to design area-delay tradeoff).
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Rostami, M., Koushanfar, F. & Karri, R. A Primer on hardware security: Models, methods, and metrics. Proc. IEEE 102 (8), 1283–1295 (2014).
Sengupta, A., Bhadauria, S. & Mohanty, S. P. TL-HLS: Methodology for low cost hardware trojan security aware scheduling with optimal loop unrolling factor during high level synthesis. IEEE Trans. Comput. Aided Des. Integr. Circ. Syst. 36 (4), 655–668 (2017).
Sengupta, A., Roy, D., Mohanty, S. P. & Corcoran, P. DSP design protection in CE through algorithmic transformation based structural obfuscation. IEEE Trans. Consum. Electron. 63 (4), 467–476 (2017).
Sengupta, A., Roy, D. & Mohanty, S. P. Triple-phase watermarking for reusable IP core protection during architecture synthesis. IEEE Trans. Comput. Aided Des. Integr. Circ. Syst. 37 (4), 742–755 (2018).
Zhao, Y. et al. Side channel security oriented evaluation and protection on hardware implementations of Kyber. IEEE Trans. Circuits Syst. I Regul. Pap. 70 (12), 5025–5035 (2023).
Kim, D., Hong, J. P., Lee, J. & Nam, J. W. High-speed light detection sensor for hardware security in standard CMOS technology. IEEE Trans. Circ. Syst. II: Express Br. 70 (10), 3917–3921 (2023).
Yasin, M., Rajendran, J. J., Sinanoglu, O. & Karri, R. On improving the security of logic locking. IEEE Trans. Comput. Aided Design Integr. Circ. Syst. 35 (9), 1411–1424 (2016).
Roy, D. B., Bhasin, S., Nikolić, I. & Mukhopadhyay, D. Combining PUF with RLUTs: A two-party pay-per-device IP licensing scheme on FPGAs. ACM Trans. Embed. Comput. Syst. 18 (2), 22 (2019).
Yilmaz, Y., Aniello, L. & Halak, B. ASSURE: A hardware-based security protocol for resource-constrained IoT systems. J. Hardw. Syst. Secur. 5, 1–18 (2021).
Anandakumar, N. N. et al. Tehranipoor, rethinking watermark: Providing Proof of IP Ownership in Modern SoCs, Cryptology ePrint Archive. https://eprint.iacr.org/2022/092 (2022)
Das, U., et al. PSC-watermark: Power side channel based IP watermarking using clock gates. 2023 IEEE European Test Symposium (ETS), Venezia, Italy, 1–6 (2023)
Slpsk, P., Nair, A. A., Rebeiro, C. & Bhunia, S. SIGNED: A challenge-response scheme for electronic hardware watermarking. IEEE Trans. Comput. 72 (6), 1763–1777 (2023).
Chen, J & Schafer, B. C. Watermarking of Behavioral IPs: A Practical Approach,” 2021 Design, Automation & Test in Europe Conference & Exhibition (DATE), 1266–1271 https://doi.org/10.23919/DATE51398.2021.9474071. (2021)
Koushanfar, F., Hong, I. & Potkonjak, M. Behavioral synthesis techniques for intellectual property protection. ACM Trans. Des. Autom. Electron. Syst. 10 (3), 523–545 (2005).
Castillo, E., Parrilla, L., Garcia, A., Meyer-Baese, U., Botella, G. & Lloris, A. Automated signature insertion in combinational logic patterns for HDL IP core protection. 2008 4th Southern Conference on Programmable Logic 183–186 (2008)
Yu, T. & Zhu, Y. A new watermarking method for soft IP protection. 2011 International Conference on Consumer Electronics, Communications and Networks (CECNet). 3839–3842 (2011).
Sengupta, A. & Rathor, M. Facial biometric for securing hardware accelerators. IEEE Trans. Very Large Scale Integr. (VLSI) Syst. 29 (1), 112–123 (2021).
Sengupta, A. & Rathor, M. IP core steganography for protecting DSP kernels used in CE systems. IEEE Trans. Consum. Electron. 65 (4), 506–515 (2019).
Sengupta, A. & Chaurasia, R. Securing IP cores for DSP applications using structural obfuscation and chromosomal DNA impression. IEEE Access 10, 50903–50913 (2022).
Rathor, M., Sengupta, A., Chaurasia, R. & Anshul, A. Exploring handwritten signature image features for hardware security. IEEE Trans. Depend. Secure Comput. 20 (5), 3687–3698 (2023).
Sengupta, A., Kumar, E. R. & Chandra, N. P. Embedding digital signature using encrypted-hashing for protection of DSP cores in CE. IEEE Trans. Consum. Electron. 65 (3), 398–407 (2019).
Proteins & Amino Acid, Chapter 5, Harvard University, USA. (accessed on December 2024); https://projects.iq.harvard.edu/files/lifesciences1abookv1/files/5_-_proteins_and_amino_acids_revised_9-24-2018.pdf.
Sengupta, A. & Anshul, A. Secure hardware IP of GLRT cascade using color interval graph based embedded fingerprint for ECG detector. Nat. Sci. Rep. 24, 13250 (2024).
Guo, Z., Karimian, N., Tehranipoor, M. M. & Forte, D. Hardware security meets biometrics for the age of IoT. IEEE International Symposium on Circuits and Systems (ISCAS) 1318–1321 (2016).
Duong, V. A., Park, J. M., Lim, H. J. & Lee, H. Proteomics in forensic analysis: Applications for human samples. Appl. Sci. 11, 3393 (2021).
Abdelaziz, G. Synthesis of human insulin gene in vitro through computational methodology. Life Sci. J. 11, 27–34 (2014).
Sengupta, A., Roy, D., Mohanty, S. P. & Corcoran, P. Low-cost obfuscated JPEG CODEC IP core for secure CE hardware. IEEE Trans. Consum. Electron. 64 (3), 365–374 (2018).
NanGate 15 nm Open Cell Library. (accessed on Dec 2024); http://www.nangate.com/?pageid=2328.
Sengupta, A. & Chaurasia, R. Secure implantable cardiac pacemaker for medical consumer electronics. NPJ Biomed. Innov. https://doi.org/10.1038/s44385-025-00008-y (2025).
Express benchmark suite, University of California Santa Barbara (UCSB), (accessed on February 2025); https://web.ece.ucsb.edu/EXPRESS/benchmark/
Hu, W. et al. An overview of hardware security and trust: Threats, countermeasures, and design tools. IEEE Trans. Comput. Aided Des. Integr. Circ. Syst. 40 (6), 1010–1038 (2021).
Potkonjak, M. Methods and Systems for the Identification of Circuits and Circuit Designs (Google Patents, 2006).
Rathor, M. & Rathor, G. P. Hard-sign: A hardware watermarking scheme using dated handwritten signature. IEEE Design Test 41 (2), 75–83 (2024).
Karmakar, R., Jana, S. S. & Chattopadhyay, S. A cellular automata guided finite-state-machine watermarking strategy for IP protection of sequential circuits. IEEE Trans. Emerg. Topics Comput. 10 (2), 806–823 (2022).
Acknowledgements
This work is technically and financially supported by CSIR Grant No. 22/0856/23/EMR-II.
Author information
Authors and Affiliations
Contributions
Anirban Sengupta—Idea generation, development, writing, project supervision Nabendu Bhui—experiment and writing Vishal Chourasia—experiment and writing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Approval for human experiments
We confirm that all methods were carried out in accordance with relevant guidelines and regulations. We confirm that all experimental protocols were approved by Indian Institute of Technology Indore. We confirm that informed consent was obtained from all subjects and/or their legal guardian(s).
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sengupta, A., Bhui, N. & Chourasia, V. Hardware IP protection by exploiting IP vendor’s proteogenomic BioMarker as digital watermark during behavioral synthesis. Sci Rep 15, 12718 (2025). https://doi.org/10.1038/s41598-025-96495-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-96495-5