Abstract
Gliomas are highly malignant tumors of the central nervous system, and their complex molecular heterogeneity poses major therapeutic challenges. Integrins are important members of the class of cell adhesion molecules (CAMs), consisting of α-subunits and β-subunits that form 24 different heterodimers. To elucidate the complex role of integrins in glioma pathogenesis, we analyzed integrin family genes. We used a scoring system based on gene set enrichment analysis (GSEA) to identify prognostic biomarkers and nonnegative matrix factorization (NMF) to establish a new integrin-based molecular classification of gliomas. Subsequent analyses of the clinical relevance of the molecular subtypes and the underlying mechanisms demonstrated a strong correlation between integrin-based molecular subtypes and glioma malignancy. We further characterized the different clinical features and tumor microenvironments (TMEs) associated with these subtypes. We identified subtype-specific driver genes using the limma R package and weighted gene coexpression network analysis (WGCNA). We subsequently identified key integrin-mediated genes that significantly contribute to poor prognosis through a combined approach of machine learning (ML) and protein‒protein interaction (PPI) network analysis. Finally, we performed in vitro cellular experiments on the integrin family gene ITGA7 and demonstrated that ITGA7 can serve as a biomarker for gliomas. Our findings provide important insights into the multifaceted roles of integrins in glioma biology, provide an opportunity for the discovery of novel targeted therapies on the basis of the subtype-specific vulnerability of integrins, and provide a basis for the study of the role of ITGA7 in gliomas.
Similar content being viewed by others
Introduction
Gliomas, a heterogeneous group of primary malignant brain tumors, pose a significant clinical challenge. Glioblastoma (GBM), the most aggressive subtype, accounts for 70–75% of all diffuse glioma diagnoses. Despite advancements in understanding the genetic basis of gliomagenesis, which involves mutations in genes such as epidermal growth factor receptor (EGFR), platelet-derived growth factor receptor alpha (PDGFRA), and isocitrate dehydrogenase (IDH), overall survival (OS) for glioma patients remains dismal, with a median OS of 14 to 17 months for GBM patients1. Integrins, transmembrane receptors that mediate cell adhesion and communication, play critical roles in tumorigenesis. Their involvement in cell‒cell and cell‒matrix interactions, as well as their ability to transmit mechanochemical signals into the cell, have been extensively studied2. Integrins are key regulators of diverse physiological and pathological processes, including development, immune responses, leukocyte trafficking, and hemostasis, and they are involved in the regulation of signaling in cancer, allowing tumor cells to proliferate unrestrictedly and invade through tissue boundaries2. Their interaction with the extracellular matrix (ECM) and cytoskeleton significantly impacts cell proliferation, differentiation, adhesion, and migration; ECM remodeling; immune system function; and the tumor microenvironment (TME)3. On the basis of their ligand-binding properties, integrins are classified into four types: leukocyte integrins, Arg-Gly-Asp (RGD)-binding integrins, collagen-binding integrins, and laminin-binding integrins4.
ITGA7 has emerged as a potential target in several cancers, including GBM, colorectal cancer, breast cancer, esophageal squamous cell carcinoma, and lung cancer1,5. ITGA7 has been implicated in tumorigenesis, metastasis, activation, and resistance to both oncogene and immune-targeted therapies5. Preclinical studies have demonstrated the involvement of ITGA7 in promoting cancer cell invasion and migration. Notably, ITGA7 knockdown in hepatocellular carcinoma (HCC) cells was found to suppress Hsp27-mediated cell invasion, whereas in lung cancer cells, ITGA7 knockdown was found to reduce S100P-mediated cell migration6. Additionally, ITGA7 knockdown has been shown to inhibit the proliferation of GBM cells by decreasing the phosphorylation of AKT and p38. These findings strongly suggest that ITGA7 plays a critical role in cancer development and progression.
To elucidate the mechanisms by which integrins contribute to glioma progression, we employed a multipronged approach. First, gene set enrichment analysis (GSEA) was used to assess the overall impact of integrin family genes on gliomas. We subsequently integrated prognostic gene sets using differential analysis and weighted gene coexpression network analysis (WGCNA) to identify the key drivers of integrin-mediated poor prognosis. To further explore the underlying molecular mechanisms, we established an integrin-based glioma subtype classification using nonnegative matrix factorization (NMF). This classification revealed two distinct subtypes: C1 and C2. The C2 subtype was found to have a significantly worse prognosis and was predominantly identified as World Health Organization (WHO) grade IV glioma.
Pathway analysis revealed that, compared with the C1 subtype, the C2 subtype exhibited aberrant activation of the mitogen-activated protein kinase (MAPK) signaling pathway and cell cycle, suggesting a link to increased tumor cell proliferation. Notably, the C2 subtype also showed significant enrichment of the epithelial‒mesenchymal transition (EMT) pathway, a process implicated in tumor proliferation and metastasis.
To identify potential therapeutic targets associated with the poor prognosis of integrin-based glioma subtypes, we integrated differential analysis, WGCNA, protein–protein interaction (PPI) network analysis, and machine learning (ML). This comprehensive approach yielded a core gene set that was used to construct a subtype-based risk model. Univariate and multivariate analyses validated the robustness of the model, and its stability was confirmed using an external dataset.
Our findings provide novel molecular targets for therapeutic interventions, leveraging the biological processes that integrin family genes promote in gliomas, ultimately leading to poor prognosis.
Methods and materials
Data sources
Transcriptomic and clinical data were obtained from three publicly available repositories: the Chinese Glioma Genome Atlas (CGGA) database (http://www.cgga.org.cn), The Cancer Genome Atlas (TCGA) database (https://portal.gdc.cancer.gov/)7,8,9,10,11,12,13. and Gene Expression Omnibus (https://www.ncbi.nlm.nih.gov/geo/). These databases provided a comprehensive dataset for our analysis.
Integrin-based glioma subtype classification
To explore the associations between integrin family genes and glioma, we performed single-sample gene set variation analysis (ssGSVA) on 26 integrin family genes (Table S1). The samples were subsequently divided into high- and low-enrichment clusters on the basis of the median value of the enrichment score. A list of differentially expressed genes (DEGs) was generated using the limma package and subsequently analyzed using Gene Ontology (GO) enrichment for cellular component (CC), molecular function (MF), and biological process (BP) terms. We further investigated the enrichment of Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways and conducted GSEA. NMF was used to identify distinct integrin-associated glioma subtypes7,8,9.
WGCNA
WGCNA was conducted to identify gene modules associated with the two integrin-enriched clusters10. We employed a scale-free topology criterion to determine the soft threshold power beta. The minimum module size was set to 50 genes, and the dynamic tree cut method was utilized to identify modules with a MEDissThres parameter of 0.25.
Identification of key genes associated with subtype prognosis
ML algorithms were employed to identify key genes associated with the distinct prognostic profiles of the integrin-based glioma subtypes. Differential analysis was performed on the gene dataset of DEGs. These DEGs were then intersected with genes associated with modules identified through WGCNA, yielding genes related to different phenotypes. Furthermore, the intersection of these genes with those identified using ML algorithms, including random forest (RF) and least absolute shrinkage and selection operator (LASSO) regression, was obtained.
Construction of a subtype-based risk model
On the basis of the intersection of genes identified through the above analyses, we constructed a subtype-based risk model. To validate the model’s accuracy, we utilized the dataset as a training group and the TCGA301 dataset as a testing group. The model’s performance was evaluated through univariate and multivariate analyses.
Cell culture
The glioma cell lines (U87, U251, LN229, A172, and U118) and normal human astrocytes (HAs) were obtained from the Chinese Academy of Science. These cells were grown in a carefully sustained, clean, and nontoxic environment in which a stable temperature of 37 °C was maintained. The surrounding atmosphere was composed of 95% air and 5% CO2. The culture medium utilized was DMEM supplemented with 10% FBS (fetal bovine serum) + 1% dual antibiotic (penicillin and streptomycin, P/S) for the most effective sustenance and promotion of growth. All materials, including the culture medium and supplements, were procured from Sanofi (Shanghai, China). The culture medium was changed every two days to create the best possible conditions for cell expansion and proliferation. Every two months, the cells were refreshed to maintain their viability.
CCK-8 and colony formation assays
We used a cell counting kit-8 (CCK-8) and cell clones to detect cell proliferation. For the cell counting assays, 2500 U87/U251 cells/well were seeded in 96-well plates and cultured. Cell viability was assessed with a CCK-8 kit (YiSen, Shanghai). CCK-8 reagent (10 μL) was mixed with 100 μL of culture medium and added to each well, after which the cells were incubated for 2 h. The CCK-8 assay was then performed by measuring the absorbance using a Bio-Tek microplate reader (USA). The colony formation assay consisted of seeding 1000 cells into each well (2 ml) of a 6-well plate followed by 2 weeks of culture in a cell incubator. Following incubation, the plates were washed with PBS, fixed with 4% paraformaldehyde, and finally stained with crystal violet.
Flow cytometry analysis
We used an Annexin V-FITC/PI apoptosis detection kit (Yeasen Shanghai, 0302ES20) to evaluate the percentage of apoptotic cells. Apoptosis was detected according to the manufacturer’s instructions. The cells were seeded in 6-well plates, digested with 0.25% trypsin, washed twice with PBS and transferred to Eppendorf tubes. Annexin V-FITC/PI was added. The mixture was incubated for 15 min at room temperature in the dark. The degree of apoptosis in the samples was detected with a CytoFLEX S (Beckman, USA).
Western blotting
Proteins were extracted from cells and glioma tissues using RIPA lysis buffer (RIPA, Beyotime, P0013K) containing protease and phosphatase inhibitors. The proteins were fully centrifuged to separate them (12,000 r, 15 min, 4 °C) and boiled for denaturation (98 °C, 10 min). Then, the proteins were separated by 10% sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE, P0052A, Beyotime) (90 V, 120 min) and transferred to a polyvinylidene fluoride membrane (Millipore) (300 mA, 120 min), followed by blocking with 5% skimmed milk at room temperature (60 r, 1 h). Then, the membrane was incubated with primary antibodies ITGA7 (1:1000, ab182941, 129 kDa, Abcam) and GAPDH (1: 40,000, 60,004-1-Ig, 37 kDa, Proteintech) at 4 °C overnight. The membrane was washed on a shaker with TBST containing Tween-20 (120 r, 5 min × 3 times). Then, the membrane was incubated with goat anti-mouse IgG (H + L) secondary antibody (1:4000, AS003, ABclonal) and goat anti-rabbit IgG (H + L) secondary antibody (1:4000, AS014, ABclonal) at room temperature for 2 h, and then washed with PBST (120 r, 5 min × 3 times). Finally, the protein bands were visualized by enhanced chemiluminescence reagent and Image Quant LAS 500 system (GE).
Immunofluorescence
Tumor samples were fixed with formaldehyde for at least 1 week, and then dehydrated using a fully enclosed tissue processor (Leica, ASP300), and the tumor samples were embedded in paraffin. The embedded tissues were sectioned into 4um slices on an automatic microtome, and then the 4um slices were baked in an oven at 65 °C for 6 h. The slices were then placed successively in xylene I, xylene II (10 min), 100% absolute ethanol I, 100% absolute ethanol II, 95% ethanol, 90% ethanol, 80% ethanol, and 70% ethanol (8 min) for dehydration and dewaxing. Then, the slices were immersed in citrate repair solution and heated in a microwave oven at high power for 15 min for antigen repair. After cooling to room temperature, they were washed with PBS (120 r, 5 min × 3 times). A histological pen was used to draw a circle around the tissue, and then 5% goat serum was dropped for blocking at room temperature for 1 h. Subsequently, the primary antibody ITGA7 (1:500, ab203254, Abcam) was dropped on the tumor tissue and incubated overnight at 4 °C. The primary antibody was washed with PBS (120 r, 5 min × 3 times). Then, the tumor tissue was incubated with the corresponding fluorescent secondary antibody (1:200, Cat NO 550043, Zbnbio) in the dark for 2 h, and then washed with PBST (120 r, 5 min × 3 times). DAPI (AB104139, Abcam) was used for sealing the slices, and fluorescence images were captured in the dark using a Leica DM4B microscope.
Immunohistochemistry
The tumor samples were fixed with formaldehyde for at least 1 week. Then, after dehydration using a fully enclosed tissue processor (Leica, ASP300), the tumor samples were embedded in paraffin. The embedded tissues were cut into 4um slices on an automatic microtome. Then, the 4um slices were baked in an oven at 65 °C for 6 h. The slices were then placed successively in xylene I, xylene II (10 min), 100% absolute ethanol I, 100% absolute ethanol II, 95% ethanol, 90% ethanol, 80% ethanol, and 70% ethanol (8 min) for dehydration and dewaxing. Then, the slices were immersed in citrate repair solution and heated in a microwave oven at high power for 15 min for antigen repair. After cooling to room temperature, they were washed with PBS (120 r, 5 min × 3 times). A histochemical pen was used to draw a circle around the tissue, and then 5% goat serum was dropped for blocking at room temperature for 1 h. Subsequently, the primary antibody ITGA7 (1:500, ab203254, Abcam) was dropped on the tumor tissue and incubated overnight at 4 °C. The primary antibody was washed with PBS (120 r, 5 min × 3 times). HRP-labeled rabbit secondary antibody polymer was added and incubated for 30 min (keep away from light) (abs996, Absin), and washed with PBST (120 r, 5 min × 3 times). Freshly prepared DAB chromogenic solution (G1212, Servicebio) was used and incubated at room temperature for 3 min (observed at any time under the microscope). The glass slides were rinsed with tap water, and 100 ul of hematoxylin was added and incubated for 3 min (the time can be adjusted appropriately according to the chromogenic situation). The slides were rinsed with tap water for bluing. Subsequently, dehydration was performed successively with 70% ethanol, 80% ethanol, 90% ethanol, 95% gradient ethanol, 100% absolute ethanol II, and 100% absolute ethanol I (2 min), and transparency was achieved with xylene II and xylene I (2 min). Neutral gum was used for sealing the slices. Bright-field images were captured using a Leica DM4B microscope.
ITGA7 small interfering RNA (siRNA) construction and transfection
Transfection was executed using the RNAFit Transfection Reagent (Biological lndustries) according to the manufacturer’s instructions. Approximately 200,000 cells were seeded one day before transfection. ITGA7 was silenced by facilitating transfection with 8 µl of siRNA for 72 h. The specific siRNA sequences used in this investigation were ITGA7siRNAs with the following sequences: siRNA-1, positive sense strand: GCUUGGAUAUGGAUGGGAATT; complementary strand: UUCCCAUCCAUAUCCAAGCTT; siRNA-2, sense strand: GCUCCAGGAAAAUGUCAAATT; complementary strand: UUUGACAUUUUCCUGGAGCTT; siRNA-3, sense strand: CAGUGAAGUCCUCCAUAAATT; and complementary strand: UUUAUGGAGGACUUCACUGTT. siRNA NC, positive strand: UUCUCCGAACGUGUCACGUTT and the complementary strand: ACGUGACACGUUCGGAGAATT. The previously mentioned siRNAs were procured from HanHen Biotechnology (Shanghai) Co.
Statistical analysis
Statistical analyses were performed using R version 4.3 (http://www.r-project.org). For continuous data, the Wilcoxon test was used to measure differences between the two subtypes. The Pearson correlation coefficient was used to assess the relationship between two variables. Kaplan‒Meier survival analysis was used to compare survival rates associated with different subtypes. Statistical significance was defined as p < 0.05.
All the statistical analyses were performed using GraphPad Prism version 9.5. The experiments were repeated independently at least three times. Differences between two groups were compared using unpaired Student’s t tests, and differences among multiple groups were compared using one-way ANOVA with Dunnett’s post hoc test. Significance was accepted at P values < 0.05 for all the statistical tests.
Results
Initial exploration of integrin family gene expression profiles
A dataset comprising 1,018 samples was acquired from the CGGA database. Integrin family genes were extracted (Table S1), and GSEA scores were calculated for these genes. On the basis of the median GSEA score, the samples were categorized into high-integrin and low-integrin groups.
Differential analysis revealed a greater prevalence of oncogene expression in the high-integrin group (Fig. 1A), which was associated with a worse prognosis (Fig. 1B). Pathway analysis revealed aberrant activation of several signaling pathways in the high-integrin group, including the Janus kinase/signal transducers and activators of transcription (JAK/STAT) pathway, the p53 signaling pathway, and the tumor necrosis factor (TNF) signaling pathway (Fig. 1C–D, Table S2). These findings suggest potential molecular mechanisms underlying the association between integrin family gene expression and poor prognosis in gliomas.
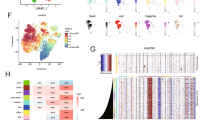
provides an overview of the integrin gene family scores. (A) Shows the top 40 most significantly up-regulated and down-regulated genes in the high-score group versus the low-score group. (B) Illustrates the difference in survival between the high-low and low-rated groups. (C) and (D) present the results of GSEA analysis and GO analysis, respectively, between the high and low scoring groups. (E) Transcription factor activity and cellular trafficking were significantly enriched in the high integrin group. (F) and (G) show that WCGNA was utilized to identify score-related module genes. (H) Highlights the intersection of differential genes and WCGNA module genes.
GO analysis of CC, MF, and BP revealed significant enrichment of transcription factor activity and cell transport in the high-integrin group (Fig. 1E), potentially contributing to the aggressive glioma phenotype.
WGCNA of the high- and low-integrin groups revealed a strong correlation between the MEdarkgreen module and the high-integrin group (R = 0.61, Fig. 1F–G). Intersecting DEGs (n = 988) with module genes (n = 990) yielded 385 genes associated with distinct phenotypes (Fig. 1H).
Integrin-based glioma subtype classification
To identify and integrate different glioma subtypes associated with integrins, we utilized the CGGA325 and CGGA693 datasets as validation sets and performed NMF (Non-negative Matrix Factorization) analysis. During the analysis, we selected the Brunet algorithm and set the range of K values from 2 to 10 (Fig. 2A). For each K value, we conducted NMF decomposition and subsequently calculated statistical indicators, including the Cophenetic coefficient, silhouette score, and incorporated clinical information features of the samples for biological significance validation. After comprehensive evaluation, we determined the optimal K value to be 2. To ensure the stability and reliability of this selection, we further confirmed it through cross-validation and validation with independent datasets. The optimal clustering results revealed two distinct transcriptional subtypes, which we named C1 and C2 (Fig. 2B). Through subtype-specific survival analysis, we found that the prognosis of the C1 subtype was significantly better than that of the C2 subtype (Fig. 2C). To verify that this classification result was not overfitted and had biological significance in different patient cohorts, we obtained multiple independent external datasets as validation sets, including GEO16011 (Fig. 2D), CGGA301 (Fig. 2E), TCGA (Fig. 2F), and GEO18494 and GEO4412 (Fig. 2G, Table S4). The results showed that in these external independent datasets, the samples could also be divided into two subgroups with clear prognostic significance. Further clinical feature analysis revealed that the prevalence of high-grade gliomas was significantly higher in the C2 subtype (Fig. 2H). Additionally, through KEGG pathway analysis7, we found abnormal activation of the MAPK signaling pathway and cell cycle in the C2 subtype, suggesting a potential association between pathway enrichment and the poor prognosis observed in this subtype (Fig. 2I,J).
illustrates the process of constructing a glioma model. (A) Presents the performance of Non-negative Matrix Factorization (NMF) under different factorization ranks. It includes six subplots, each displaying the variation of six indicators: cophenetic, dispersion, evar, residuals (RSS), silhouette, and sparseness, with respect to the factorization rank. Each indicator showcases the performance of different decomposition methods, including Basis, Best fit, Coefficients, and Consensus. (B) Presents the consensus matrix when the fractalization is set to 2. (C) Depicts the survival differences observed in the typing. Independent external datasets as validation sets, including GEO16011 (D), CGGA301 (E), TCGA (see F), and GEO18494 and GEO4412 (G). (H) Shows the proportion of samples between the classification and the WHO grade. (I) features a differential volcano diagram of the classification. Finally, (J) displays the KEGG results for the typing.
Molecular and immune landscape of glioma subtypes
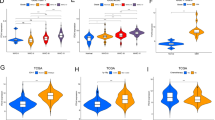
To further investigate the distinct molecular and immune characteristics of these subtypes, we assessed the enrichment of 50 tumor-associated gene sets. The C2 subtype exhibited significant enrichment of gene sets associated with malignant phenotypes (Fig. 3A). Pathway analysis revealed upregulated and increased enrichment scores for EMT in the C2 subtype (Fig. 3B–C), suggesting a correlation between EMT and the poor prognosis of this subtype. EMT is known to promote tumor proliferation and metastasis, highlighting a potential mechanism contributing to the aggressive nature of the C2 subtype.
presents the specimen molecular landscape. (A) Displays a heatmap showing the distribution of 50 hotspot gene sets. (B) Highlights the up-regulated hallmark pathway, while (C) shows the down-regulation of the hallmark pathway. (D) Presents the landscape of gene set scores related to immune genes and immune pathways, (E) shows the differences in tumor microenvironment scores between the two subtypes, and (F) displays the expression differences of the HLA gene family between the two subgroups.
Analysis of the molecular landscape of integrin-related genes across subtypes revealed significant enrichment of poor prognosis-associated genes in the C2 subtype (Fig. 3D). Furthermore, analysis of the TME revealed a greater proportion of stromal cells and lower tumor purity in the C2 subtype group than in the C1 subtype group (Fig. 3E). Intriguingly, a greater proportion of human leukocyte antigen (HLA)-related genes was observed in the C1 subtype group (Fig. 3F). Previous research has demonstrated that an increased number of human leukocytes is associated with worse survival in glioma patients. This observation may provide further insight into the biological processes mediated by integrin family genes, potentially contributing to the progression from low-grade to high-grade gliomas and ultimately leading to poor prognosis.
Identification of key gene modules associated with glioma subtypes
We employed WGCNA to identify gene modules associated with the distinct subtypes. A soft threshold power (β) of 19 was determined on the basis of the scale-free topology criterion and the dendrogram (Fig. S1A). This analysis revealed eight coexpression modules (cut height ≥ 0.25) representing clusters of genes with similar expression patterns. These modules were designated MEsienna3, MEred, MEbrown, MEdarkgreen, MEviolet, MEgreen, MEpurple, MEdarkgrey, MEcyan, MElightgreen, MEgrey, and MEgrey (Fig. S1E–F). Hierarchical clustering of the modules Fig. S1B–D) show that modules belonging to the same branch presented similar gene expression patterns. Further analysis of the correlations between modules highlighted a strong association between the MElightgreen and MEpurple modules, particularly the MElightgreen module, and the C2 subtype (cor = 0.53, p = 2e−71; Table S3). This strong correlation identifies the MElightgreen module as a key module for further investigation.
Identification of key genes and construction of a prognostic risk model
To further pinpoint key genes associated with the C2 subtype, we performed dimensionality reduction and ML analyses. We extracted 898 genes from the MElightgreen module identified through WGCNA and 1291 DEGs from the differential analysis. The intersection of these gene sets revealed 286 DEGs strongly correlated with the C2 subtype (Fig. 4A). PPI analysis was then performed on these 286 genes to identify core genes (Fig. 4B). We subsequently employed two ML algorithms, LASSO regression and RF, to further refine the list of key genes. LASSO regression identified 63 genes (Fig. 4C), whereas RF identified 126 genes (Fig. 4D,E). The intersection of these two gene sets yielded 39 genes, suggesting potential interactions among these genes in promoting integrin-associated poor prognosis in gliomas.
illustrates the process of obtaining core genes related to typing. (A) Displays the differential genes and module genes. (B) Presents the PPI network analysis of the intersection genes. (C) Showcases the LASSO regression analysis. Lastly, (D) and (E) provide the results of the random forest analysis.
Development and validation of a subtype-based risk model
Following the prognostic analysis of the 39 identified genes (Fig. 5A), we constructed a subtype-based risk score (Fig. 5B), which was validated using an external dataset (Fig. 5C). Univariate and multivariate analyses demonstrated that the risk score had strong prognostic value (Fig. 5D,E), serving as a potential prognostic risk factor for gliomas. Next, we performed time-dependent AUC and ROC curve analyses and found that the risk score exhibited good predictive performance in both the validation and test sets. Specifically, in the validation set (Fig. 6A), the AUC values for 1, 2, and 3 years were 0.78, 0.829, and 0.831, respectively; in the test set (Fig. 6B), the corresponding AUC values were 0.694, 0.791, and 0.816, respectively. In addition, we incorporated clinical information into the ROC curve analysis (Fig. 6C). The results showed that the AUC value for the risk score was the highest (0.772), followed by Grade in the WHO classification (AUC = 0.745), gender (AUC = 0.512), age (AUC = 0.565), IDH mutation (AUC = 0.322), 1p19q codeletion (AUC = 0.390), and MGMT promoter methylation (AUC = 0.489). Considering the important influence of IDH status, MGMT methylation, and 1p19q codeletion classification systems on the prognosis of glioma, we incorporated these systems into our analysis. Both the calibration plot and decision curve analysis showed that our risk score and risk level had excellent predictive effects on patient prognosis (Fig. 6D–F). The decision curve (Fig. 6F) clearly indicated that the risk score and risk level were superior to other classification indicators. Furthermore, the calibration curve showed that the predictive effect of the risk level was comparable to that of the WHO grade, but the risk level was slightly better. At the same time, the 5-year survival rate prediction based on the risk level was also slightly better than that based on the WHO grade. The analysis conclusions in the test set were consistent with those in the validation set (Fig. 6G–I). It is particularly noteworthy that the predictive ability of this model is comparable to that of the WHO glioma classification system, providing a new theoretical basis for the staging of glioma and potentially enhancing clinical decision-making and patient management.
details the construction of a prognostic model. (A) Displays the model gene risk score. (B) Presents the survival analysis for the experimental group based on the risk model. (C) Shows the survival analysis for the test group using the risk model. (D) provides the results of the one-factor analysis for the risk model. Finally, (E) displays the multi-factor analysis of the risk model.
Model ROC Curves, Calibration Plots, and Decision Curve Analysis. (A) Presents the time-dependent ROC curve for the validation set, while (B) shows the time-dependent ROC curve for the test set. (C) Displays the ROC curve for clinical traits and risk scores. (D) Illustrates the calibration curve for WHO grades in the validation set, and (E) presents the calibration curve for risk levels in the validation set. (F) Depicts the decision curve for the validation set. (G) Shows the calibration curve for WHO grades in the test set, (H) presents the calibration curve for risk levels in the test set, and (I) depicts the decision curve for the test set.
Impact of ITGA7 expression levels on glioma prognosis
We conducted an in-depth assessment of the distribution characteristics of integrin gene family members and clinical traits across two subgroups and risk scores (Fig. 7A–C). The results revealed that ITGA7 stands out with the highest baseline expression level among the integrin gene family. Additionally, ITGA7 exhibits highly significant differential expression in subtype classification and prognostic scoring. Notably, ITGA7 expression is markedly elevated in the C2 subtype and High group, which are associated with poorer prognosis. Subsequently, we utilized Cytoscape software to perform a visual analysis of the protein–protein interaction (PPI) network for the integrin gene family. The results clearly demonstrate strong interactions and correlations among the proteins of this gene family (Fig. 7D). ITGA7 has recently been reported to promote a poor prognosis in a wide range of tumors, including hepatocellular carcinomas11, breast carcinomas12 and endometrial carcinomas13, Therefore, we further validated the expression of ITGA7 in multiple tumors using the TCGA database. The findings indicate that ITGA7 expression is generally high in most tumors, with particularly prominent expression in gliomas, especially glioblastoma. Furthermore, we observed a gradual increase in ITGA7 expression with advancing tumor grade (Fig. 7E,F). Concurrently, high ITGA7 expression is closely associated with poorer patient prognosis, aligning with our observations in the C2 subtype and High group, and further confirming the tight correlation between high ITGA7 expression and adverse tumor outcomes (Fig. 7G).
explores the impact of ITGA7 expression on glioma prognosis. (A–C) show the expression abundance of the integrin gene family and its clinical correlation between the two subgroups and the risk score. (D) Displays the integrin gene families used for PPI network analysis. (E and F) showcase ITGA7 expression in pan-cancer. Finally, (G) assesses the relationship between ITGA7 expression and glioma prognosis using Kaplan–Meier analysis.
ITGA7 expression in GBM
To validate the findings from the TGGA database, we assessed the expression levels of ITGA7 in normal brain tissues, glioma cells, and GBM tissues. The results revealed that ITGA7 levels were significantly elevated in GBM (Fig. 8A). ITGA7 expression was detected in several established glioma cell lines and normal human astrocytes, and the expression level of ITGA7 was elevated in the human glioma cell lines U251, U87, A172 and LN229 compared with the level in normal HA cells (Fig. 8B). Furthermore, the results of immunohistochemical analysis (Fig. S2A,B) and immunofluorescence experiments (Fig. S2C–H) both indicate that the expression level of ITGA7 is significantly higher in tumor tissues compared to normal tissues, with a more pronounced difference. Compared with HA cells, GBM cell lines(including U251, U87, A172 and LN229) had greater expression levels of ITGA7; therefore, the U251 and U87 cell lines were selected for further knockdown experiments.
illustrates the expression of ITGA7 in tumour tissues and tumour cells, as well as its effect on tumour cell proliferation and apoptosis. (A) Compares the expression of ITGA7 in normal brain tissues and GBM specimens. (B) Shows the expression of ITGA7 in various cell lines, including HA, LN229, U251, U118, U87, and A172. (C and D) verify the efficiency of Si1-ITGA7, Si2-ITGA7, and Si3-ITGA7 in U251 and U87 cells, respectively, compared to the NC group. (E and F) utilize CCK8 to detect the proliferation ability of U251 and U87 cells in the Si group and NC group. (G and H) Display the results of a clone formation assay for U251 and U87 cells in the Si group and NC group. Lastly, (I and J) use flow cytometry to detect the degree of apoptosis in U251 and U87 cells in the Si group and NC group.
ITGA7 knockdown efficiency
To investigate the effects of ITGA7 on GBM cell proliferation and apoptosis, siRNA transfection was subsequently performed. Compared with si-NC, Si3-ITGA7 significantly inhibited ITGA7 protein levels in both the U251 and U87 cell lines (Fig. 8C,D). Si3-ITGA7 had the most effective knockdown efficiency. Therefore, it was used for further experiments.
Effects of Si-ITGA7 on the proliferation and spoptosis of U87 and U251 cells
The results of the CCK-8 assay revealed that the cell proliferation ability was lower in the Si-ITGA7 group than in the NC group (Fig. 8E,F), and the results of the cell colony formation experiments were consistent with those of the CCK8 assay (Fig. 8G,H). Immediately afterward, we used flow cytometry to detect the effects of Si-ITGA7 and NC on tumor cell apoptosis, and the experiments revealed that Si-ITGA7 promoted cell apoptosis (Fig. 8I,J).
Discussion
Gliomas, which are highly malignant and heterogeneous neuroepithelial tumors, pose a significant clinical challenge because of their high intracranial recurrence rate and debilitating nature. The 2021 WHO update for glioma classification, which incorporates both neuropathological and molecular pathological features, was created with the aim of improving diagnostic accuracy and guiding the implementation of more tailored treatment strategies.
Recent advancements in next-generation sequencing technologies and the expansion of molecular databases have revolutionized research in the field of molecular biology. These advancements not only increase the precision and efficiency of gene sequencing but also provide a wealth of molecular data for investigation. This has spurred a surge of research exploring the intricate molecular mechanisms underlying cancer, including the crucial role of programmed cell death in tumor development and progression. This research not only deepens our understanding of cancer biology but also has the potential to facilitate the identification of novel therapeutic targets.
Integrins, key members of the CAM family, are composed of two subunits, α and β, forming 24 distinct combinations. These receptors play critical roles in cell adhesion and communication, significantly influencing tumorigenesis by relaying signals into the cell. Integrins regulate cell‒cell adhesion, activate various intracellular signaling pathways, and are essential for numerous biological processes, including development, immune responses, and leukocyte trafficking.
In the context of cancer, integrins have emerged as key players in tumor progression, enabling uncontrolled proliferation, the invasion of surrounding tissues, and interactions with the ECM and cytoskeleton. This multifaceted role of integrins profoundly influences numerous biological processes in cancer, including cell proliferation, differentiation, adhesion, migration, and interactions with the immune system and TME.
To elucidate the role of integrin family genes in glioma, we conducted GSEA. On the basis of the median GSEA score, glioma samples were categorized into low- and high-integrin groups, designated the C1 and C2 subtypes, respectively. A comparison of these subtypes revealed significant differences in their molecular and clinical characteristics. The C2 subtype was associated with a greater prevalence of oncogene expression, a worse prognosis, and aberrant activation of the JAK-STAT signaling pathway, p53 signaling pathway, and TNF signaling pathway. These pathways are commonly activated in various malignancies, contributing to unfavorable outcomes. GO analysis revealed significant enrichment of transcription factor activity and cellular transport processes in the C2 subtype.
We subsequently used DEGs to develop a molecular classification model for gliomas on the basis of integrin family genes. Analysis revealed a significant enrichment of malignant phenotype-associated gene sets in the C2 subtype, accompanied by upregulation and increased scores for the EMT pathway. EMT is a crucial process during normal tissue development and wound healing. However, its dysregulation contributes to the pathogenesis of cancer and fibrosis14,15. During EMT, epithelial cells undergo a transformation in which they lose polarity, detach from each other, gain deformability and motility, and ultimately migrate to distant sites. EMT has been implicated as a crucial factor in the development of malignant phenotypes, such as invasion and metastasis, in various epithelial-derived cancers, including breast cancer16, liver cancer17, bladder cancer18, and lung cancer19,20.
Our findings suggest a strong association between the unfavorable prognosis of the C2 subtype and EMT, indicating that this subtype is associated with increased tumor proliferation and metastasis.
Analysis of the molecular landscape of integrin-related genes across the C1 and C2 subtypes revealed a significant enrichment of poor prognosis-associated genes in the C2 subtype. ITGA7 is particularly noteworthy in the C2 subtype.
ITGA7, a member of the laminin-binding integrin family, is located on chromosome 12p13 and comprises at least 27 exons spanning approximately 22.5 × 103 kb21. ITGA7 serves as a major laminin receptor in skeletal and cardiac muscle, plays a crucial role in muscle development and is described as a key regulator in skeletal muscle diseases22. ITGA7 is also highly expressed in the heart, and mutations in ITGA7 have been reported in children with congenital cardiomyopathy23.
Emerging evidence suggests that aberrant ITGA7 expression is associated with clinicopathological characteristics in various cancers24. As a cell surface receptor, ITGA7 mediates interactions between cancer cells and the TME, promoting tumor growth and driving cancer progression. For example, in breast cancer, high ITGA7 expression has been linked to increased invasiveness and metastasis25. In lung cancer, ITGA7 is considered a crucial gene associated with invasion and metastasis26. In colorectal cancer, ITGA7 modulates downstream targets that regulate colorectal cancer metastasis, promoting tumor cell invasion and metastasis24. Moreover, ITGA7 overexpression has been correlated with higher pathological grade and larger tumor size27. Yuan et al.28 reported that ITGA7 is associated with CD133 and aldehyde dehydrogenase 1 (ALDH1) and is linked to poor differentiation, lymph node metastasis, and reduced survival in patients with cervical cancer. Bhandari et al.21 identified ITGA7 as a tumor suppressor that regulates breast cancer migration and invasion, and low ITGA7 expression potentially affects the migration of breast cancer cells and is associated with a poor prognosis. Other studies have revealed that reduced ITGA7 expression significantly inhibits tumor cell proliferation and invasion. Therefore, ITGA7 has emerged as a crucial marker for grading and assessing the prognosis of various cancers22.
In-depth analysis of the TME revealed a significantly greater proportion of stromal cells in the C2 subtype group than in the C1 subtype group, accompanied by lower tumor purity. Conversely, the C1 subtype group presented abnormally elevated expression of HLA-related genes. Previous studies have indicated that a greater number of leukocytes in glioma patients is often associated with shorter survival. This observation suggests a potential link between the biological processes associated with integrin family genes and the progression of GBM from low-grade to high-grade malignancy, ultimately contributing to a worse prognosis.
To elucidate the underlying mechanisms driving these changes, we first employed WGCNA to identify key modules. We subsequently utilized the limma R package to identify modular genes and DEGs between the two clusters. Building on these analyses, we employed ML algorithms, specifically LASSO regression and RF, to further refine the dataset. Through intersection analysis with survival-related genes, we successfully identified 39 key genes.
On the basis of these 39 genes, we constructed survival models for the two subtypes and derived a subtype-based risk model. Univariate and multivariate analyses demonstrated that these risk model genes effectively predicted survival outcomes in both subtypes. Importantly, the risk score system we developed serves as a significant prognostic risk factor for gliomas, with predictive power comparable to that of the WHO grade. As mentioned earlier, the specific role of integrin family genes in tumors has been discussed in a variety of cancers, and given the specificity of ITGA7 studies in a variety of tumors, we performed in vitro experiments on ITGA7 and reported that ITGA7 expression was elevated in GBM compared with normal brain tissue and was correlated with a worse prognosis. In cellular experiments, Si-ITGA7 decreased the proliferation of tumor cells compared with that in the NC group and increased the apoptosis of tumor cells in the experimental group, as detected by flow cytometry. This research provides not only a new theoretical basis for the typing of gliomas but also further validates the effectiveness of the WGCNA and limma R packages for identifying modular genes and DEGs and demonstrates the powerful application of advanced algorithms, such as ML, LASSO regression and RF, to bioinformatics analysis.
In summary, we developed a molecular typing method based on the integrin gene family that more accurately predicts the survival of glioma patients. We delved into the similarities and differences between the two subtypes in terms of clinical features, molecular characteristics, immune function, pathways of genesis, and the tumor microenvironment. This study reveals new research pathways for the molecular typing of gliomas. However, we have not yet fully clarified how these related genes and subtypes specifically affect the prognosis of gliomas. Therefore, we need to further explore the intrinsic mechanisms in these subtypes in gliomas, which is an important challenge at present and a major direction for future research.
Data availability
This study utilized publicly available datasets accessible from the following sources: The CGGA database (http://www.cgga.org.cn) and TCGA database (https://portal.gdc.cancer.gov/ ). Raw files for Western Blot analysis used in the manuscript are available in the Supplementary Materials.
Abbreviations
- CAM:
-
Cell adhesion molecule
- GSEA:
-
Gene set enrichment analysis
- NMF:
-
Negative matrix factorisation
- TME:
-
Tumour mi-croenvironment
- WGCNA:
-
Weighted gene co-expression network analysis
- ML:
-
Machine learning
- PPI:
-
Protein–protein interaction
- GBM:
-
Glioblastoma
- EGFR :
-
Epidermal growth factor receptor
- PDGFRA :
-
Platelet-derived growth factorreceptor alpha
- IDH :
-
Isocitrate dehydrogenase
- OS:
-
Overall survival
- ECM:
-
Extracellular matrix
- TME:
-
Tumor microenvironment
- RGD:
-
Arg-Gly-Asp
- ITGA7:
-
Integrin α7
- HCC:
-
Hepatocellular carcinoma
- GSEA:
-
Gene set enrichment analysis
- WHO:
-
World Health Organization
- MAPK:
-
Mitogen-activated protein kinase
- EMT:
-
Epithelial-mesenchymal transition
- CGGA:
-
Chinese Glioma Genome Atlas
- TCGA:
-
The Cancer Genome Atlas
- ssGSVA:
-
Single-sample gene set variation analysis
- DEGs:
-
Differentially expressed genes
- GO:
-
Gene Ontology
- CC:
-
Cellular component
- MF:
-
Molecular function
- BP:
-
Biological process
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- RF:
-
Random forest
- LASSO:
-
Least absolute shrinkage and selection operator
- TNF:
-
Tumor necrosis factor
References
Carrasco-Garcia, E., Auzmendi-Iriarte, J. & Matheu, A. Integrin α7: A novel promising target in glioblastoma stem cells. Stem Cell Investig. 5, 2. https://doi.org/10.21037/sci.2017.12.05 (2018).
Hynes, R. O. Integrins: Bidirectional, allosteric signaling machines. Cell 110(6), 673–687. https://doi.org/10.1016/s0092-8674(02)00971-6 (2002).
Hirtz, A., Rech, F., Dubois-Pot-Schneider, H. & Dumond, H. Astrocytoma: A hormone-sensitive tumor?. Int. J. Mol. Sci. 21(23), 9114. https://doi.org/10.3390/ijms21239114 (2020).
Calderwood, D. A. Integrin activation. J. Cell Sci. 117(Pt 5), 657–666. https://doi.org/10.1242/jcs.01014 (2004).
Network, T. C. Corrigendum: Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 494(7438), 506. https://doi.org/10.1038/nature11903 (2013).
Jia, Z. et al. Identification of new hub genes associated with bladder carcinoma via bioinformatics analysis. Tumori 101(1), 117–122. https://doi.org/10.5301/tj.5000196 (2015).
Kanehisa, M., Furumichi, M., Sato, Y., Matsuura, Y. & Ishiguro-Watanabe, M. KEGG: Biological systems database as a model of the real world. Nucleic Acids Res. 53(D1), D672–D677. https://doi.org/10.1093/nar/gkae909 (2025).
Ogata, H. et al. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 27(1), 29–34. https://doi.org/10.1093/nar/27.1.29 (1999).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 28(11), 1947–1951. https://doi.org/10.1002/pro.3715 (2019).
Langfelder, P. & Horvath, S. WGCNA: An R package for weighted correlation network analysis. BMC Bioinform. 9, 559. https://doi.org/10.1186/1471-2105-9-559 (2008).
Ge, J. C., Wang, Y. X., Chen, Z. B. & Chen, D. F. Integrin alpha 7 correlates with poor clinical outcomes, and it regulates cell proliferation, apoptosis and stemness via PTK2-PI3K-Akt signaling pathway in hepatocellular carcinoma. Cell Signal. 66, 109465. https://doi.org/10.1016/j.cellsig.2019.109465 (2020).
Bai, X., Gao, C., Zhang, L. & Yang, S. Integrin α7 high expression correlates with deteriorative tumor features and worse overall survival, and its knockdown inhibits cell proliferation and invasion but increases apoptosis in breast cancer. J. Clin. Lab. Anal. 33(8), e22979. https://doi.org/10.1002/jcla.22979 (2019).
Liang, M., Liu, C., Lei, T., Guo, S. & Min, J. Effect of integrin α7 on cell proliferation, invasion, apoptosis and the PI3K/AKT pathway, and its association with clinicopathological features in endometrial cancer. Oncol. Lett. 25(1), 26. https://doi.org/10.3892/ol.2022.13612 (2023).
Peinado, H., Olmeda, D. & Cano, A. Snail, Zeb and bHLH factors in tumour progression: An alliance against the epithelial phenotype?. Nat. Rev. Cancer 7(6), 415–428. https://doi.org/10.1038/nrc2131 (2007).
Yang, J. et al. Guidelines and definitions for research on epithelial-mesenchymal transition. Nat. Rev. Mol. Cell Biol. 21(6), 341–352. https://doi.org/10.1038/s41580-020-0237-9 (2020).
Jiang, G., Fang, H., Shang, X., Chen, X. & Cao, F. CHFR-mediated epithelial-to-mesenchymal transition promotes metastasis in human breast cancer cells. Mol. Med. Rep. https://doi.org/10.3892/mmr.2021.12090 (2021).
Lei, Y. et al. Comprehensive analysis of partial epithelial mesenchymal transition-related genes in hepatocellular carcinoma. J. Cell Mol. Med. 25(1), 448–462. https://doi.org/10.1111/jcmm.16099 (2021).
Monteiro-Reis, S., Lobo, J., Henrique, R. & Jerónimo, C. Epigenetic mechanisms influencing epithelial to mesenchymal transition in bladder cancer. Int. J. Mol. Sci. 20(2), 297. https://doi.org/10.3390/ijms20020297 (2019).
Franco-Chuaire, M. L., Magda Carolina, S. C. & Chuaire-Noack, L. Epithelial-mesenchymal transition (EMT): Principles and clinical impact in cancer therapy. Invest. Clin. 54(2), 186–205 (2013).
Iwatsuki, M. et al. Epithelial-mesenchymal transition in cancer development and its clinical significance. Cancer Sci. 101(2), 293–299. https://doi.org/10.1111/j.1349-7006.2009.01419.x (2010).
Bhandari, A. et al. ITGA7 functions as a tumor suppressor and regulates migration and invasion in breast cancer. Cancer Manag. Res. 10, 969–976. https://doi.org/10.2147/cmar.S160379 (2018).
Mizumoto, M. et al. Reirradiation for recurrent malignant brain tumor with radiotherapy or proton beam therapy. Technical considerations based on experience at a single institution. Strahlenther. Onkol. 189(8), 656–663. https://doi.org/10.1007/s00066-013-0390-6 (2013).
Bugiardini, E. et al. Integrin α7 mutations are associated with adult-onset cardiac dysfunction in humans and mice. J. Am. Heart Assoc. 11(23), e026494. https://doi.org/10.1161/jaha.122.026494 (2022).
Chen, J. R., Zhao, J. T. & Xie, Z. Z. Integrin-mediated cancer progression as a specific target in clinical therapy. Biomed. Pharmacother. 155, 113745. https://doi.org/10.1016/j.biopha.2022.113745 (2022).
Chen, J. et al. A restricted cell population propagates glioblastoma growth after chemotherapy. Nature 488(7412), 522–526. https://doi.org/10.1038/nature11287 (2012).
Chen, Y., Wang, Y., Cai, Z., Fan, X. & Zhang, Y. Integrin α7 is overexpressed and correlates with higher pathological grade, increased T stage, advanced TNM stage as well as worse survival in clear cell renal cell carcinoma patients: A retrospective study. J. Clin. Lab. Anal. 34(1), e23034. https://doi.org/10.1002/jcla.23034 (2020).
Huang, J., Tian, Y. & Chen, J. Overexpression of integrin α7 correlates with advanced disease condition and poor prognosis in rectal cancer patients. Int. J. Clin. Exp. Pathol. 11(9), 4630–4636 (2018).
Yuan, N. et al. ITGA7, CD133, ALDH1 are inter-correlated, and linked with poor differentiation, lymph node metastasis as well as worse survival in surgical cervical cancer. J. Obstet. Gynaecol. Res. 48(4), 1011–1018. https://doi.org/10.1111/jog.15163 (2022).
Acknowledgements
In the preparation of this work, we used chatgtp3.5 as well as ERNIE Bot 4.0 to linguistically edit the already written manuscript in order to make our manuscript more rigorous and standardised in this one aspect of language. After using this tool, we reviewed and edited the content as needed and take full responsibility for the content of the publication.
Funding
This study was supported by the Natural Science Foundation of Gansu Province (22JR5RA959).
Author information
Authors and Affiliations
Contributions
In completing this thesis, each author made the following contributions. Hongxi Han was responsible for writing the main manuscript and completed all experimental work. Peng Feng was responsible for the bioinformatics work. Finally, Guoqiang Yuan, as the leader of our team, participated in the overall design of the experiments and reviewed all the manuscripts.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
We enrolled nine human glioma samples and nine derived from with Ventricular decompression fistula organization, all collected from the Lanzhou University Second Hospital. According to the Declaration of Helsinki specifications, this experiment was given written informed consent and approval for human research (2022A-401) by the Ethics Committee of the Second Hospital of Lanzhou University. Written informed consent was obtained from all subjects.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Han, H., Feng, P. & Yuan, G. Molecular typing of gliomas on the basis of integrin family genes and a functional study of ITGA7. Sci Rep 15, 12306 (2025). https://doi.org/10.1038/s41598-025-97342-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-97342-3
Keywords
This article is cited by
-
The role of genes related to excitotoxicity in glioma prognosis
Discover Oncology (2025)