Abstract
Lymphedema is a chronic condition characterized by lymphatic fluid accumulation, primarily affecting the limbs. Its diagnosis is challenging due to symptom overlap with conditions like chronic venous insufficiency (CVI), deep vein thrombosis (DVT), and systemic diseases, often leading to diagnostic delays that can extend up to ten years. These delays negatively impact patient outcomes and burden healthcare systems. Conventional diagnostic methods rely heavily on clinical expertise, which may fail to distinguish subtle variations between these conditions. This study investigates the application of artificial intelligence (AI), specifically deep learning, to improve diagnostic accuracy for lower limb edema. A dataset of 1622 clinical images was used to train sixteen convolutional neural networks (CNNs) and transformer-based models, including EfficientNetV2, which achieved the highest accuracy of 78.6%. Grad-CAM analyses enhanced model interpretability, highlighting clinically relevant features such as swelling and hyperpigmentation. The AI system consistently outperformed human evaluators, whose diagnostic accuracy plateaued at 62.7%. The findings underscore the transformative potential of AI as a diagnostic tool, particularly in distinguishing conditions with overlapping clinical presentations. By integrating AI with clinical workflows, healthcare systems can reduce diagnostic delays, enhance accuracy, and alleviate the burden on medical professionals. While promising, the study acknowledges limitations, such as dataset diversity and the controlled evaluation environment, which necessitate further validation in real-world settings. This research highlights the potential of AI-driven diagnostics to revolutionize lymphedema care, bridging gaps in conventional methods and supporting healthcare professionals in delivering more precise and timely interventions. Future work should focus on external validation and hybrid systems integrating AI and clinical expertise for comprehensive diagnostic solutions.
Similar content being viewed by others
Introduction
Lymphedema is a globally prevalent chronic pathological condition that exerts a significant burden on healthcare systems, with an estimated 140–250 million individuals affected worldwide1. This condition is characterized by the pathological accumulation of protein-rich lymphatic fluid within the interstitial spaces, leading to persistent tissue edema. In the absence of timely therapeutic intervention, this edematous state may progress to encompass both lymphatic fluid retention and fibrofatty tissue proliferation. Despite its high prevalence, lymphedema often remains under-recognized in clinical settings. Epidemiological studies conducted in European populations have reported prevalence rates ranging between 1.33 and 1.44 cases per 1,000 individuals2,3. However, these figures likely underestimate the global incidence due to multifaceted challenges in accurate case identification4.
One of the primary impediments to determining the true prevalence of lymphedema lies in the complexities inherent to its diagnosis. The condition presents considerable diagnostic challenges and is frequently misclassified as other pathologies. Standard diagnostic protocols typically involve a comprehensive review of the patient’s medical history and a detailed physical examination. However, confirmation often requires advanced imaging techniques such as lymphoscintigraphy. Despite these resources, conventional diagnostic approaches frequently fail to provide conclusive results, even when executed by experienced clinicians, necessitating reliance on sophisticated imaging modalities. These diagnostic limitations contribute to significant delays in disease identification and subsequent therapeutic intervention. Recent research suggests that the average diagnostic delay can extend to approximately ten years, during which patients remain undiagnosed and untreated, potentially resulting in disease progression and the deterioration of clinical outcomes5,6.
Moreover, the diagnostic complexity of lymphedema is exacerbated by its propensity to mimic the clinical manifestations of other conditions. The condition is often misdiagnosed as systemic diseases for example heart failure or renal insufficiency, venous disorders such as chronic venous insufficiency (CVI) or chronic deep vein thrombosis (DVT), or even as obesity in cases where no other significant comorbidities are apparent. The overlap in clinical presentations among these conditions frequently leads to misclassification. While these disorders may exhibit similar external symptoms, their underlying pathophysiological mechanisms differ significantly, necessitating precise and targeted therapeutic approaches. Lymphedema arises from impaired lymphatic drainage, resulting in the accumulation of protein-rich lymphatic fluid and subsequent chronic swelling7. Conversely, CVI is driven by venous valve incompetence, leading to elevated venous pressure and extravasation of fluid into surrounding tissues8,9. Systemic diseases such as hepatic, cardiac, or renal pathologies are typically characterized by generalized fluid retention resulting from imbalances in hydrostatic and oncotic pressures10.
The pathophysiological differences between these conditions underscore the critical importance of accurate and timely diagnosis. Diagnostic errors or delays not only hinder appropriate treatment but may also exacerbate the primary condition, leading to unfavorable patient outcomes and increased healthcare expenditures. Given the significant diagnostic challenges associated with lymphedema, there is a compelling need for more sophisticated diagnostic tools capable of reliably distinguishing this condition from other etiologies.
Differentiating lymphedema from related conditions such as CVI & DVT, and systemic diseases necessitates advanced diagnostic expertise and innovative technological solutions. Recent advancements in artificial intelligence (AI) have shown great promising results in addressing these challenges. AI systems, particularly Convolutional Neural Networks (CNNs), have demonstrated remarkable capabilities in analyzing medical images and extracting complex patterns indicative of specific pathological conditions. For instance, Fu et al. (2018) reported an impressive 93.7% diagnostic accuracy in detecting lymphedema among breast cancer survivors using an artificial neural network trained on real-time symptom reports11.
Similarly, Wei et al.12 developed a symptom-warning model employing six machine learning algorithms, with logistic regression achieving the highest sensitivity (0.771) and specificity (0.883) for early detection. This model has been implemented as an open-access web-based application, allowing for real-time patient monitoring. Ozmen et al.13 further advanced AI applications in this field by proposing a deep learning model utilizing synthetic datasets generated through General Adversarial Networks (GANs). Their Feedforward Neural Network demonstrated a predictive accuracy of 95.25% for identifying lymphedema risk following breast cancer surgery. These findings underscore the transformative potential of AI models trained on extensive datasets to facilitate early diagnosis and enable personalized preventive measures.
Despite significant progress in AI-based lymphedema diagnostics, existing efforts predominantly focus on upper limb cases, particularly in breast cancer survivors, where differential diagnoses are less complex. In contrast, there remains a substantial research gap in applying AI to the diagnosis and management of lower limb edema, where differential diagnoses are more challenging. Additionally, the potential utility of software tools such as ImageJ in simulating expert clinical assessments has yet to be fully explored. Addressing these gaps is essential for developing comprehensive diagnostic tools for lower limb edema.
The application of AI in diagnosing lower limb edema offers significant promise for improving diagnostic precision, optimizing treatment planning, and enhancing patient care. Moreover, AI-driven diagnostic solutions could reduce healthcare costs by mitigating misdiagnoses and minimizing unnecessary diagnostic procedures.
This research aims to advance the field by developing an AI-based diagnostic framework specifically tailored to lower limb edema. By employing deep learning to analyze clinical images, the study seeks to differentiate between lymphedema, CVI, systemic diseases such as heart failure, and normal cases. This work represents a critical step toward improving diagnostic accuracy and timeliness, thereby addressing the substantial unmet needs in the clinical management of lower limb edema and enhancing patient outcomes.
Materials and methods
This study is a multicenter collaboration, having received ethical approval from the Institutional Review Board (IRB) under Certificate of Approval Si 090/2022 (Faculty of Medicine, Siriraj Hospital) and COA 063/2022 (Faculty of Medicine, Maharat Nakhon Ratchasima Hospital). Written informed consent was obtained from all participants prior to enrollment. All protocols and methodologies strictly adhered to the ethical guidelines outlined in the Declaration of Helsinki. The protocols were approved by the respective IRBs, and informed consent was obtained from all participants or their legal guardians.
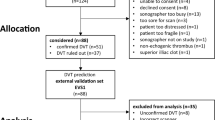
The study utilized a multi-step approach to develop and validate an AI-based diagnostic tool for classifying lower limb edema diseases as illustrated in Fig. 1. The process began with data collection, gathering a total of 1,622 clinical images of lower limbs. These images then underwent preprocessing, which involved cropping, padding, and resizing to ensure uniformity and suitability for model input.
The preprocessed images were used to train sixteen deep learning models: AlexNet14, VGG1915, DenseNet-16916, GoogLeNet17, ResNet-5018, SqueezeNet19, MobileNetV320, EfficientNetV221, ViT22, TnT23, Swin24, CvT25, PiT26, CCT27, MaxViT28, and DaViT29. These models were trained to classify the images into four categories: lymphedema, venous disease : chronic venous insufficiency and deep vein thrombosis (CVI & DVT), systemic disease, and normal. Following model training, a validation phase was conducted to assess the performance of each model.
To compare the AI models’ performance with human expertise, a group of healthcare workers participated in an evaluation. This group included medical students, residents from the surgery department, and fellows and staff from the surgery department. The evaluation involved a 28-question spot-diagnosis quiz, where participants were asked to classify images of lower limb conditions.
The results from both the AI models and the healthcare worker evaluations were then compared to assess the relative performance of the AI-based diagnostic tool against human experts. This comprehensive approach aimed to validate the effectiveness of the AI models in accurately diagnosing lower limb edema conditions and to determine their potential utility in clinical practice.
Data collection and image acquisition
This study was conducted as a multi-center collaboration, with data collected from two independent medical institutions: Siriraj Hospital and Maharat Nakhon Ratchasima Hospital. These centers serve distinct patient populations, incorporating urban tertiary-care and provincial hospital settings, which contributes to a heterogeneous dataset. The dataset consists of 1622 clinical images of lower limbs, obtained from patients diagnosed with lymphedema (457), chronic venous insufficiency (CVI) & deep vein thrombosis (DVT) (362), systemic diseases associated with lower limb edema (360), and healthy controls (443).
Comprehensive demographic data, medical histories, and clinical characteristics of the participants are summarized in Table 1. All diagnoses were confirmed using gold-standard diagnostic techniques, including lymphoscintigraphy lymphoscintigraphy (sensitivity 92%, specificity 100%)30, duplex ultrasound (sensitivity 90%, specificity 84%)31, and disease-specific diagnostic tests. Standardized imaging protocols were followed at both centers to ensure consistency, with patients positioned uniformly and images acquired under controlled lighting conditions.
To ensure consistency and minimize variability in image quality, all photographs were captured in a dedicated imaging room with a controlled setup. An overhead 5000K LED light source was used to maintain uniform brightness, preventing variations in illumination that could affect model performance. The camera was positioned at a fixed 100 cm distance for full-length images and 60 cm for close-up images, ensuring a standardized field of view across all subjects. Fig. 2 shows examples of lymphedema, chronic venous insufficiency, systemic edema, and healthy controls, respectively
A DSLR camera (Canon EOS 5D Mark IV) was used for all image acquisitions, with manual exposure settings including f/8 aperture, 1/125 sec shutter speed, ISO 200, and a 60s exposure time to maintain consistent image clarity and depth of field. The camera was placed at a fixed distance of 1.0 meters from the patient’s legs to ensure a reproducible perspective for each photograph.
To further improve image quality and eliminate background distractions, we used a matte black, non-reflective backdrop to reduce shadows and reflections (Fig. 3). By controlling these parameters including lighting intensity, camera settings, and patient positioning. we can minimize variability in the collected images, thereby enhancing the reliability of our deep-learning-based analysis.
Image preprocessing
Each image included in the dataset contains both legs set against a black background. The collected images were saved in two file formats: PNG and JPEG. The image sizes range from 2048\(\times\)1536 pixels to 3888\(\times\)5184 pixels, all in RGB color space. The dataset consists of 1,622 images in total. To support deep-learning training and maintain the proportion of the given images, a preprocessing step is necessary. This preprocessing includes cropping, padding, and resizing, as illustrated in Fig. 4. Firstly, we split the lower leg image into two separate images, one for each leg. This was done by cropping the original image to isolate each leg. Subsequently, we added zero padding around each leg’s image to adjust its shape, resulting in a square image. This step ensures that the aspect ratio of the images is consistent, which is crucial for effective deep-learning model training. Finally, the padded images were resized to a standard size, which differed for each model since CNN and transformer-based models have varying input size requirements. This resizing step standardizes all images in the dataset, ensuring uniformity across the dataset and making it suitable for efficient training of the deep learning models. By performing these preprocessing steps, we ensure that the images maintain their proportions and are standardized. This is essential for achieving reliable and consistent results in the subsequent deep-learning analysis. As a result, no data augmentation was applied in this study.
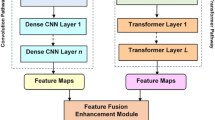
Illustration of the Architecture of EfficientNetV2, highlighting the use of Fused-MBConv layers for early-stage efficiency and MBConv layers for complex feature extraction. The comparison structure shows how Fused-MBConv replaces depthwise convolutions with standard convolutions, enhancing computational efficiency, while MBConv retains depthwise separable convolutions for capturing more detailed features.
EfficientNet and its extension for image classification
EfficientNet is a group of convolutional neural networks (CNNs) designed to be both powerful and efficient21. The key idea behind EfficientNet is to carefully increase the network’s width, depth, and resolution together in a balanced way. This approach helps achieve better accuracy with fewer parameters and lower computational costs. EfficientNet was introduced by Mingxing Tan and his colleagues in 2019.
EfficientNet includes several models, each designed at different scales to balance better accuracy with fewer parameters and lower computational costs. It starts with EfficientNet-B0, a small model (the base architecture) developed using neural architecture search (NAS), and extends to larger models like EfficientNet-B7 using the compound scaling method. The compound scaling method is a core concept of EfficientNet and solves the challenge of effectively scaling up convolutional neural networks. Instead of scaling the network’s depth, width, and resolution separately, which can result in poor performance, EfficientNet scales all three dimensions together in a systematic and balanced way. This approach uses fixed scaling coefficients to uniformly adjust the network’s depth, width, and resolution, ensuring that each contributes appropriately to the model’s capacity and efficiency. Traditionally, networks are scaled in one of three ways: (i) Depth Scaling (d) by increasing the number of layers (e.g., ResNet), (ii) Width Scaling (w) by increasing the number of channels in each layer, or (iii) Resolution Scaling (r) by increasing the input image resolution. However, scaling only one of these dimensions can create issues. For example, increasing depth alone might lead to overfitting, while increasing resolution alone can significantly raise computational demands without a proportional improvement in accuracy. To address this, EfficientNet introduced a compound scaling method that scales the network’s depth, width, and resolution simultaneously using a compound coefficient. The compound scaling approach uses the following equations to produce the compound coefficient:
where \(\alpha\), \(\beta\), and \(\gamma\) are constants that determine how much to scale each dimension (depth, width, and resolution, respectively). \(\phi\) is the compound coefficient that controls the overall scaling of the network. By increasing \(\phi\), you can scale up the model to create larger versions. Each scaling parameter plays a specific role in the network’s performance. Depth scaling increases the number of layers, allowing the network to learn more complex features. Width scaling increases the number of channels in each layer, enabling the network to capture more features per layer. Resolution scaling increases the input image resolution, helping the network capture finer details in images. The constants \(\alpha\), \(\beta\), and \(\gamma\) must follow specific rules to ensure that the network scales evenly across all dimensions-depth, width, and resolution.
This constraint ensures that when \(\phi\) is increased, the model’s computational cost (in terms of FLOPs) roughly doubles. The constants \(\alpha\), \(\beta\), and \(\gamma\) are typically determined using grid search to find the best balance between model accuracy and computational efficiency.
After that, EfficientNetV2 was introduced by Mingxing Tan and his colleagues at Google AI in 202121. EfficientNetV2 builds on the foundation of EfficientNetV1 by introducing several key improvements aimed at enhancing training speed and model efficiency. Three major innovations in EfficientNetV2 are Fused-MBConv layers, Progressive Learning with Adaptive Regularization, and Fine-Tuned Scaling Factors. These advancements collectively contribute to making EfficientNetV2 faster, more accurate, and more efficient in various deep-learning tasks
Fused-MBConv layers are an improvement over the MBConv layers used in EfficientNetV1. The MBConv layers utilized depthwise separable convolutions, which were efficient in terms of parameters and FLOPs but often didn’t fully take advantage of modern hardware like GPUs and TPUs. Fused-MBConv layers address this by combining the depthwise convolution and expansion convolution operations into a single regular convolution with size \(3\times 3\) followed by a pointwise convolution with size \(1\times 1\) as shown in Fig. 5 . This combination reduces computational overhead and speeds up training, especially in the earlier stages of the network where depthwise convolutions are less effective. The result is a more hardware-friendly architecture that still delivers strong performance across various tasks. Progressive learning with adaptive regularization in EfficientNetV2 is a training strategy where both the input image resolution and regularization intensity are gradually increased during the training process. Initially, the model is trained on lower-resolution images, allowing for faster training with larger batch sizes. As the training progresses, the image resolution is incrementally increased, which enables the model to capture finer details without significantly increasing the computational cost. This approach not only speeds up the early stages of training but also improves the model’s ability to generalize. By exposing the model to a wider variety of features at different resolutions throughout training, it learns to adapt better to new data. Regularization techniques like dropout, weight decay, and data augmentation play a crucial role in preventing overfitting. In progressive learning with adaptive regularization, these regularization techniques are adjusted based on the current training phase and image resolution. During the early phase with lower resolution images, lighter regularization is applied, as the model is focusing on learning broader, less detailed features. Applying too much regularization at this stage could hinder learning or cause the model to underfit. As the resolution increases and the model begins to focus on finer details, it becomes more prone to overfitting. To counter this, stronger regularization methods are introduced, ensuring that the model continues to generalize well and does not merely memorize the training data. The key to this strategy is finding the right balance between underfitting and overfitting throughout the training process. By dynamically adjusting regularization as training progresses, the model remains robust across different training stages and resolutions, leading to improved performance on unseen data.
EfficientNetV2 also improves upon the compound scaling strategy introduced in EfficientNetV1. Instead of scaling the depth, width, and resolution uniformly, EfficientNetV2 uses a non-uniform scaling strategy. The scaling factors for depth, width, and resolution are fine-tuned through extensive experimentation and the NAS to optimize performance on modern hardware. This refined scaling ensures that the model scales more effectively, focusing computational resources where they have the most impact on performance.
Training and testing an efficientnet for lower limb edema
The process for classifying and analyzing lower limb edema using EfficientNetV2 is divided into two main phases. The first is the training phase, and the second is the testing phase.
In the training phase, the first step involves gathering a dataset of clinical images depicting various types of lower limb edema. These images, collected using cameras, are carefully annotated by medical professionals to accurately label the specific conditions. The images then undergo preprocessing steps, including cropping, padding, and resizing, to prepare them for training. EfficientNetV2 employs progressive learning, where the images start at a lower resolution and gradually increase in resolution as training progresses. This method allows the model to learn general patterns before focusing on finer details. The model is initialized with pre-trained weights for example from ImageNet to leverage transfer learning, which helps accelerate convergence and improve accuracy. During training, the model is trained using the prepared dataset, often with techniques like k-fold cross-validation. The model’s performance is evaluated using metrics such as accuracy, F1 score, precision, sensitivity, confusion matrices, and ROC-AUC. Misclassified cases are closely analyzed to understand the model’s limitations, which assists in fine-tuning the model or making necessary adjustments to the training process.
In the testing phase, the best-performing model from the training phase is tested with a separate testing dataset. The results are analyzed, with particular attention to any misclassified cases, and evaluated in collaboration with healthcare workers to ensure the model’s effectiveness and reliability in a real-world clinical setting.
This structured approach ensures that the EfficientNetV2 model is completely trained, evaluated, and tested, resulting in a robust tool for classifying and analyzing lower limb edema.
Comparison of AI and human performance
In addition to training and validating the AI model, we conducted a comparative analysis between the AI and human performance using a 28-question spot-diagnosis quiz. This quiz was designed to assess diagnostic accuracy and included a diverse range of cases. We recruited 67 healthcare professionals consisting of 22 medical students, 23 residents, and 22 fellows and attending physicians to complete the quiz (Table 2).
Each participant was asked to complete the quiz independently, and their performance was compared to the AI model’s predictions. Metrics such as accuracy, precision, sensitivity and F1 were calculated for both the AI and human participants. This comparison aimed to evaluate the AI’s diagnostic capabilities relative to human expertise and assess its potential utility in clinical settings.
Results
Comparative performance of AI in lymphedema and other lower limb edema diagnosis
This experiment aims to evaluate and compare the performance of various convolutional neural network (CNN) and transformer-based models for lower limb edema disease classifications based on clinical images. There are four specific condition classes: Chronic Venous Insufficiency & Deep Vein Thrombosis (CVI&DVT), Lymphedema, Systemic Disease, and Normal (images without any of the specified conditions). The performance of these models is assessed using key metrics to determine their effectiveness and efficiency in medical image classification tasks. To ensure a robust and unbiased evaluation, k-fold cross-validation was applied. The dataset was randomly divided into five equal parts, with each fold preserving the same class distribution as the entire dataset. The filenames and corresponding labels for each fold were saved and consistently used across all models to maintain comparability. This k-fold cross-validation approach, with k set to 5, was implemented to ensure a reliable and consistent performance evaluation.
The CNN models being evaluated include AlexNet14, VGG1915, DenseNet16916, GoogLeNet17, ResNet-5018, SqueezeNet19, MobileNetV320, and EfficientNetV221, while the transformer-based models consist of ViT22, TnT23, Swin24, CvT25, PiT26, CCT27, MaxViT28, and DaViT29. All models were trained for 50 epochs with a batch size of 16, using the cross-entropy loss function. The Adam optimizer was employed across all models. The learning rates varied among the architectures, with CNN and transformer models utilizing values of \(1 \times 10^{-5}\) or \(1 \times 10^{-4}\) depending on the specific architecture. The models are evaluated based on their ability to classify images into the following classes which are CVI & DVT, Lymphedema, Systemic Disease, and Normal. All hyperparameters were set according to the recommended configurations for each model or as specified in the provided implementation code. This ensures that each architecture operates under optimal conditions, following best practices for training and evaluation. Tables 3 and 4 show the performance metrics such as Precision, Sensitivity (Recall), F1 Score, and Accuracy to evaluate the models. Precision measures the accuracy of the model in predicting positive cases, Sensitivity measures the model’s ability to identify true positives, the F1 Score provides a balance between Precision and Recall, and Accuracy reflects the overall correctness of the model.
From Table 3, model performance varies across different metrics and conditions during the training phase with our training and validation datasets (1,622 clinical images). To provide a clear understanding, the CNN and transformer-based models are divided into three groups based on their overall accuracy: (i) models with an overall accuracy of less than 0.80, (ii) models with an overall accuracy between 0.80 and 0.90, and (iii) models with an overall accuracy of more than 0.90. This categorization helps in highlighting the strengths and weaknesses of each model more distinctly.
First, CvT is the only model falling into this category, with an overall accuracy of \(0.748\pm 0.022\). The lower accuracy suggests that CvT struggles in correctly distinguishing between different classes. This is also reflected in its relatively lower precision and sensitivity across all conditions, particularly in CVI&DVT (\(0.701\pm 0.053\) precision, \(0.569\pm 0.086\) sensitivity) and normal (\(0.639\pm 0.045\) precision, \(0.713\pm 0.073\) sensitivity). These values suggest that the CvT model performs relatively worse than the other models in this study. Second, several CNN-based models, including AlexNet (\(0.870 \pm 0.023\)), MobileNetV3 (\(0.809 \pm 0.013\)), and SqueezeNet (\(0.870 \pm 0.022\)), as well as transformer-based models such as ViT (\(0.874 \pm 0.013\)) and TnT (\(0.873 \pm 0.010\)), fall within the overall accuracy range of 0.80 to 0.90. These models show moderate performance with relatively balanced precision and sensitivity across different classes. However, MobileNetV3 has a lower F1 score in CVI&DVT (\(0.747 \pm 0.033\)) and normal classification (\(0.719 \pm 0.027\)), suggesting difficulty in correctly identifying affected regions. Similarly, while AlexNet and SqueezeNet achieve high F1 scores in systemic disease classification (\(0.964 \pm 0.024\) and \(0.953 \pm 0.011\), respectively), their F1 scores for CVI&DVT, lymphedema, and normal classification remain below 0.900, indicating challenges in distinguishing these conditions accurately. Among transformer models in this group, ViT and TnT perform well in systemic disease classification, with ViT achieving an F1 score of \(0.943 \pm 0.021\) and TnT reaching \(0.952 \pm 0.014\). However, their performance in CVI&DVT, lymphedema, and normal classification remains moderate, with F1 scores below 0.900. Third, the top-performing models, with accuracy exceeding 0.90, include both CNN-based architectures (DenseNet169, GoogLeNet, ResNet50, VGG16, and EfficientNetV2) and transformer-based models (Swin, PiT, CCT, MaxViT, and DaViT). Among the CNN models, DenseNet169 (\(0.945 \pm 0.009\)) and EfficientNetV2 (\(0.941 \pm 0.009\)) achieve the highest accuracy, with precision and sensitivity scores above 0.90 across all classes. GoogLeNet (\(0.923 \pm 0.017\)), ResNet50 (\(0.930 \pm 0.013\)), and VGG16 (\(0.925 \pm 0.014\)) also perform well, though their F1 scores for normal cases fall below 0.900. Among the transformer-based models, DaViT (\(0.939 \pm 0.014\)) and MaxViT (\(0.944 \pm 0.011\)) show the best classification performance, with consistently high precision and sensitivity across all conditions. Swin (\(0.919 \pm 0.007\)) and PiT (\(0.927 \pm 0.011\)) also achieve strong results, demonstrating that transformer models in this group can match the classification accuracy of top-performing CNNs.
Figure 6 presents Grad-CAM visualizations for various deep-learning models, highlighting the regions of interest when classifying normal, systemic disease, CVI&DVT, and lymphedema conditions. The left half of the figure displays CNN-based models (AlexNet, GoogLeNet, ResNet50, VGG-16, MobileNetV3, DenseNet-169, SqueezeNet, and EfficientNetV2), while the right half presents transformer-based models (ViT, TnT, Swin, CvT, PiT, CCT, MaxViT, and DaViT). Each row corresponds to a specific model, and each column represents a disease category. The heatmaps illustrate where the models focus when making their predictions. Red regions indicate higher attention, suggesting the areas the model considers most important for classification. A checkmark indicates correct classification, while a cross represents incorrect classification. From the visualizations, EfficientNetV2 and DenseNet-169 demonstrate strong focus on the affected regions, particularly for systemic disease and CVI&DVT cases. In contrast, models such as GoogLeNet, ResNet50, and VGG-16 show inconsistent focus, often misplacing attention on irrelevant areas, leading to incorrect predictions. Transformer-based models exhibit mixed performance, with some models like DaViT effectively identifying key regions, while others struggle with certain classifications, as indicated by widespread activation beyond the lesion areas.
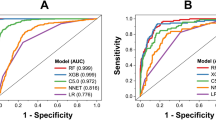
Moreover, Fig. 7 presents the Receiver Operating Characteristic (ROC) curves for deep learning models, evaluating their classification performance for lower limb edema diseases. The ROC curve plots the True Positive Rate against the False Positive Rate, providing insight into each model’s ability to distinguish between different conditions. The Area Under the Curve (AUC) values displayed in the legend quantify the overall performance of each model. Higher AUC values indicate better classification performance, with EfficientNetV2 (AUC=0.9955) and DenseNet-169 (AUC=0.9925) achieving the highest scores. Other models also demonstrate strong classification performance. The diagonal dashed line represents an AUC of 0.5, serving as a baseline. The fact that all models significantly outperform this line indicates that they effectively differentiate between the different disease conditions.
Finally, among all models, DenseNet169, EfficientNetV2, and MaxViT demonstrated the most consistent and accurate performance, achieving overall accuracies above 0.940 and F1 scores exceeding 0.900 across all classes, which is consistent with the patterns observed in the confusion matrices (Fig. 8). The ROC curve analysis further confirmed their classification capabilities, with EfficientNetV2 achieving the highest AUC score, demonstrating its strong ability to distinguish between different conditions. These results identify EfficientNetV2 as the most reliable model for classifying lower limb edema diseases using clinical images. With performance validated across multiple evaluation metrics, these models are well-suited for classifying lower limb edema diseases based on clinical images.
Comparative performance of AI on the 28-spot-diagnosis quiz
After training, the models were evaluated using the 28-spot-diagnosis quiz. Table 4 summarizes the classification performance of various deep learning models in identifying CVI&DVT, lymphedema, normal, and systemic disease conditions. To better understand these results, the models are categorized into three groups based on overall accuracy: (i) below 0.65, (ii) between 0.65 and 0.74, and (iii) above 0.74.
The first group consists of models with overall accuracy below 0.65, including AlexNet (0.643), ResNet50 (0.607), CvT (0.607), PiT (0.643), CCT (0.643), and MaxViT (0.643). These models exhibited weaker classification performance, particularly in distinguishing normal and CVI&DVT cases, with F1 scores consistently below 0.72. AlexNet and ResNet50 struggled the most in classifying normal cases, with F1 scores below 0.500, while their performance in lymphedema, CVI&DVT, and systemic disease classification was comparatively better, with F1 scores exceeding 0.700. Similarly, CvT, PiT, CCT, and MaxViT also had difficulty accurately classifying normal and CVI&DVT cases, as indicated by their lower F1 scores in these categories. The overall accuracy of models in this group remained below 0.65 primarily due to their poor prediction performance in these two classes. This suggests that these models struggle to extract relevant features for distinguishing normal legs and CVI&DVT conditions, leading to a high rate of misclassification. The second category includes GoogLeNet (0.679), VGG16 (0.679), MobileNetV3 (0.714), ViT (0.714), TnT (0.714), and DaViT (0.714). These models demonstrated moderate performance, with improved sensitivity and F1 scores across all classes. However, their classification accuracy for normal and CVI&DVT cases varied, as reflected in their lower F1 scores. ViT and TnT showed a slightly more balanced performance across categories, but models in this group still struggled with certain classifications. The third group consists of the best-performing models, with overall accuracies above 0.74, including DenseNet169 (0.750), Swin (0.750), SqueezeNet (0.750), and EfficientNetV2 (0.786). Among these, EfficientNetV2 achieved the highest accuracy, demonstrating strong and consistent performance across all categories except for normal classification. DenseNet169 and Swin also performed well, particularly in systemic disease and lymphedema classification, but showed some difficulty in identifying normal cases. The results from this group suggest that these models are more effective at extracting relevant features for distinguishing between lower limb edema conditions.
This result highlights the effectiveness of different deep learning models in classifying lower limb edema diseases using clinical images. Among them, EfficientNetV2 stands out as the most reliable model, achieving the highest classification performance in the 28-question spot-diagnosis quiz. The results in Table 4 confirm its strong performance, which is also supported by the patterns observed in the confusion matrices (Fig. 8). Additionally, the ROC curve in Fig. 9 shows that EfficientNetV2 achieved the highest AUC score of 0.9546, indicating strong overall classification ability.
Comparative performance of AI and human participants on the 28-spot-diagnosis quiz
The comparison between human performance and AI models on the 28-spot-diagnosis quiz in classifying medical conditions reveals interesting insights into the strengths and weaknesses of both approaches. From the AI perspective according to Table 4, models like EfficientNetV2, DenseNet169, SqueezeNet, Swin emerged as the top performers. EfficientNetV2 stood out with the highest overall accuracy of 0.786, showcasing strong performance across multiple disease categories, particularly in terms of sensitivity and F1 scores. However, the AI models struggled with normal class detection, as reflected by their lower sensitivity and precision values in this category. This was also evident in the confusion matrices (Fig. 10) as shown in , where normal classes often had lower true positive rates.
The human results clearly demonstrate a trend of increasing diagnostic accuracy correlating with greater clinical experience, as shown in Table 5 and 6. Fellows and staff, the most experienced group, achieved the best overall classification results with an accuracy of 0.61, surpassing medical students and residents. This group displayed higher precision, sensitivity and F1 scores. Residents performed moderately well with an accuracy of 0.601, showing better precision in detecting lymphedema conditions but struggling with systemic disease conditions. Medical students, on the other hand, had the lowest sensitivity and accuracy at 0.55.
Diagnostic accuracy is significantly influenced by the number of cases handled. Individuals with more than 20 cases achieved the highest accuracy at 0.63 and sensitivity at 0.627, particularly excelling in detecting normal conditions. In contrast, those with 0 to 5 cases had the lowest accuracy at 0.56 and faced challenges in identifying systemic diseases, highlighting a significant performance gap. This trend indicates that classification abilities improve with experience but remain below the performance of AI models like EfficientNetV2 in both sensitivity and accuracy.
Human performance, especially among medical students, displayed much greater confusion between lymphedema and other conditions (Fig. 11. Medical students often misclassified lymphedema as CVI & DVT, reflecting a lower overall diagnostic accuracy. While experienced professionals, such as fellows and staff, performed better than students, they still fell short of achieving the high precision and consistency demonstrated by the AI models.
The trend of misclassification among human professionals, especially those with less experience, indicates a tendency to confuse lymphedema with CVI & DVT. This issue was more pronounced in less experienced individuals, such as medical students or practitioners with fewer than 20 cases of experience. Conversely, AI models showed a stronger ability to differentiate between these conditions, with far fewer errors, reinforcing the reliability of AI in diagnostic settings.
Overall, AI models generally outperform humans in diagnostic accuracy, particularly in detecting conditions such as lymphedema and CVI& DVT. EfficientNetV2 consistently achieved higher scores across most categories compared to human professionals, especially less experienced ones.
Discussions
This study highlights the potential of deep learning models in medical diagnostics, particularly in addressing the challenges associated with lower limb edema classification. The diagnostic performance of the AI models was assessed using both the validation set as well as a 28-question spot-diagnosis quiz designed to compare AI performance against human evaluators with varying levels of clinical experience. Among all tested architectures, EfficientNetV2 demonstrated superior performance, achieving an accuracy of 0.94 on the validation set and 0.786 on the quiz. It consistently outperformed other CNN and transformer-based models across key metrics, including accuracy, F1 scores, and the area under the ROC curve.
When evaluated EfficientNetV2’s performance against human evaluators to assess its clinical applicability, EfficientNetV2 demonstrated superior diagnostic performance, achieving an accuracy of 78.6% in a 28-question spot-diagnosis quiz, significantly surpassing the 62.7% accuracy of the most experienced human evaluators. AI models consistently outperformed human participants across various diagnostic categories, particularly in conditions with subtle visual features such as lymphedema, where distinguishing patterns can be challenging. The findings further indicate that while clinical experience enhances diagnostic accuracy, the improvements plateau after approximately 20 cases, suggesting inherent limitations in human diagnostic capabilities. This plateau may stem from the reliance of traditional medical training on history taking and physical examination, which are critical for clinicians to contextualize visual findings. However, this reliance presents a challenge in image-based diagnostics, where human evaluators, especially those with limited exposure, struggled to achieve high accuracy, reflecting the gap between conventional diagnostic methods and AI-driven image analysis. These results support that while human evaluators, particularly those with limited exposure, struggled with image interpretation, AI models have demonstrated the ability to extract subtle visual patterns with high precision.
Similar AI-based models have achieved high diagnostic performance in benchmark medical datasets, reinforcing the potential of deep learning for medical image classification. For instance, top-performing algorithms in the ISIC 2019 challenge for skin lesion classification have reported an AUC-ROC of up to 0.94 and accuracy exceeding 85%32. Likewise, models trained on the CheXpert dataset for chest X-ray classification have demonstrated an AUC-ROC of 0.93 and accuracy around 80% across multiple thoracic pathologies33,34. In the domain of diabetic retinopathy screening, Castilla et al. (2023)35 utilized a ResNet-18 architecture to detect referable diabetic retinopathy in large-scale public datasets, achieving an AUC-ROC of 0.955 and accuracy near 92% on a combined test set of 61,007 images. Given these precedents, our model’s of AUC-ROC of 0.96, along with an accuracy of 0.94 on the validation set and 0.786 on the 28-spot-diagnosis quiz, further supports its potential for clinical deployment in lower limb edema classification.
When diagnosing edema, physicians do not necessarily assess both legs simultaneously but rather focus on one limb first, identifying specific clinical features such as hyperpigmentation, swelling patterns, skin texture, and venous distension. Once these findings are noted, they compare the affected limb to the contralateral normal leg to confirm the presence and extent of edema. This structured approach allows clinicians to recognize subtle asymmetries and determine whether the observed features are consistent with conditions such as lymphedema, chronic venous insufficiency, or systemic edema.
Our AI model replicates these diagnostic processes by leveraging a specialized normal class, which acts as an internal reference for distinguishing abnormal features. Instead of requiring direct contralateral comparison, the AI has learned disease-specific patterns from thousands of annotated images, enabling it to identify characteristic signs of edema based on absolute and relative feature distributions.Even in the absence of a physical contralateral reference, the AI model can effectively distinguish between normal and pathological features by relying on its internal normal framework, similar to how an experienced clinician draws upon prior knowledge and mental imagery of typical limb anatomy.
To further understand how the model makes these distinctions, Grad-CAM heatmaps were utilized to analyze its decision-making process and enhance interpretability. These heatmaps provide visual insights into the specific anatomical features the model prioritizes during classification, helping to bridge the gap between AI-driven analysis and traditional clinical evaluation. Figure 6 further reveals that the network focuses on regions of swelling, textural irregularities, and discoloration, which are the same anatomical cues that clinicians routinely scrutinize when evaluating a single-leg image.
EfficientNetV2 effectively leveraged heatmap analyses to focus on clinically relevant regions, such as areas of swelling in lymphedema and hyperpigmentation in CVI and the overall leg shape in systemic disease demonstrating its precision in pattern recognition (Fig. 6). In contrast, human evaluators struggled with overlapping visual features, often misclassifying normal leg structures as pathological conditions. The plateau in human accuracy underscores the importance of AI augmentation, particularly in conditions where subtle differences in visual features make differentiation challenging.
While EfficientNetV2 shows strong diagnostic potential, distinguishing between conditions with overlapping clinical features remains a challenge. The AI model’s most frequent misclassifications occurred between lymphedema and CVI, as both conditions share hyperpigmentation, fibrosis, and skin thickening. Additionally, systemic edema was sometimes misclassified as CVI, reflecting a common diagnostic difficulty where generalized swelling mimics localized venous insufficiency. Interestingly, these error patterns closely mirrored human evaluator misclassifications, suggesting that EfficientNetV2 reasoning aligns, at least in part, with the clinical decision-making process of physicians
Misclassifications have significant clinical implications. A misdiagnosis of lymphedema as CVI may lead to delays in lymphatic therapy, whereas mistaking CVI for lymphedema could result in unnecessary interventions focused on lymphatic dysfunction rather than venous pathology. Similarly, systemic edema being misclassified as CVI could obscure underlying conditions such as heart failure or renal dysfunction, delaying appropriate treatment. Minor variations in skin texture and swelling patterns occasionally led to normal cases being flagged as edema, prompting unwarranted clinical evaluations.
To mitigate these errors and enhance interpretability, AI explainability tools such as Grad-CAM heatmaps can serve as a guide, enabling clinicians to assess the model’s focus areas and ensure its predictions align with clinical reasoning. Grad-CAM can help clinicians better understand model predictions by highlighting key anatomical features the AI considers relevant, allowing physicians to visually assess whether the model’s decision aligns with clinical reasoning. By overlaying saliency maps on the original image, Grad-CAM clarifies which anatomical regions such as the medial calf, ankle region, or areas of skin discoloration are most indicative of edema, guiding clinicians to re-examine borderline features that may not be immediately apparent. This is particularly useful in ambiguous cases, such as differentiating lymphedema from CVI, where overlapping clinical signs can complicate diagnosis. Heatmaps can reveal subtle morphological or textural variations that might prompt further investigations, such as duplex ultrasound, to confirm a suspected condition.
Beyond improving diagnostic accuracy, Grad-CAM fosters AI-clinician synergy by acting as a visual second opinion. When heatmaps align with clinical findings, they reinforce clinician confidence in AI-driven classifications. In contrast, cases where AI focuses on seemingly irrelevant regions that prompt clinicians to question or override the model’s recommendation. This interpretability strengthens trust in AI tools, ensuring they function as decision-support systems rather than autonomous diagnostic entities. Additionally, heatmaps serve an educational role, guiding trainees on which morphological cues are most relevant for distinguishing different types of edema. By integrating Grad-CAM explanations into clinical workflows, AI-assisted diagnosis becomes more transparent and actionable, ensuring that interpretability tools not only explain model predictions but also meaningfully enhance clinical decision-making.
While deep learning models like EfficientNetV2 have shown promise in medical diagnostics, some studies highlight challenges and limitations that warrant consideration. For instance, a systematic review by Takahashi et al.36 critically assessed the design, reporting standards, and claims of studies comparing diagnostic deep learning algorithms with expert clinicians. The review found that many studies exhibited a high risk of bias and poor adherence to reporting standards, raising concerns about the robustness and generalizability of AI models in clinical settings. Additionally, a study by Nagendra et al.37discussed the limitations of AI in clinical diagnosis, emphasizing that while AI systems can process vast amounts of data, they may lack the nuanced understanding that human clinicians possess. The study cautioned against overreliance on AI, advocating for a balanced approach that combines AI tools with human expertise to ensure accurate and ethical medical decision-making. These perspectives suggest that despite the potential of AI models, their integration into medical diagnostics should be approached with careful validation and oversight to address inherent limitations and ensure patient safety.
Although there are some opposing views on the potential of AI, recent studies have shown promising results for AI, particularly CNN-based models, in performing medical diagnoses. Our findings align with and expand upon existing research. For instance, Goyal et al.38 demonstrated the effectiveness of DFUNet, a CNN architecture for diabetic foot ulcer classification, achieving an AUC score of 0.961. Their work highlighted the potential of CNNs to address specific medical challenges with cost-effective and scalable solutions. In lymphedema context, AI has demonstrated remarkable results in diagnosing lymphedema through image-based approaches. For instance, Nomura et al.39 developed a CAD system utilizing ResNet-34 to diagnose lower extremity lymphedema., achieving 92.9% accuracy and an AUC of 0.967 with fat-enhanced CT images. While Nomura et al. focused on CT imaging for lymphedema, this study applied CNNs to diverse conditions, demonstrating adaptability and robustness. Additionally, the use of heatmap analyses in both studies highlights the shared importance of interpretability, fostering clinical trust in AI systems.
This research offers several strengths and contributions. By evaluating the performance of CNN models across multiple diagnostic categories, this study provides a comprehensive assessment of AI’s capabilities in medical imaging. Heatmap analyses added transparency to model decision-making, enabling clinicians to understand and trust the AI’s focus areas. The comparison of human and AI performance underscored the complementary roles of both, emphasizing that AI systems can augment clinicians’ expertise, particularly in complex diagnostic scenarios. These findings contribute significantly to advancing the role of AI in medical diagnostics especially in the AI-assisted decision-making context, allowing for improved clinical outcomes and stronger AI-human collaboration.
Conclusion
This research focused on developing an AI-based classification system to differentiate between various types of lower limb edema, with a particular emphasis on enhancing diagnostic tools for identifying lymphedema. The findings revealed that the EfficientNetV2 architecture exhibited the highest accuracy in classifying edema types, demonstrating its potential as a reliable diagnostic aid. By leveraging deep learning and utilizing appropriate preprocessing techniques, the AI model showed promising capability in accurately identifying lymphedema, venous insufficiency, and systemic edema of the legs.
The developed AI-based classification system can assist healthcare professionals in making accurate and timely diagnoses, potentially improving patient outcomes and streamlining treatment strategies. Moreover, automating the initial screening process for lower limb edema through deep learning could alleviate the burden on healthcare providers. Additionally, in the context of lower limb edema, Grad-CAM serves as a valuable visualization tool by highlighting key anatomical features such as skin texture, swelling patterns, or vascular signs, which are indicative of specific edema types. This interpretability not only enhances trust in AI-driven decisions but also allows clinicians to validate predictions with their expertise, fostering a more reliable diagnostic workflow.
As AI technology continues to evolve, integrating interpretability mechanisms like Grad-CAM into diagnostic systems can promote a more transparent, reliable, and collaborative approach between AI and healthcare providers. This will ensure that AI tools not only enhance diagnostic accuracy but also align with the clinical rationale guiding patient care. The developed AI-based classification system holds promise for revolutionizing lower limb edema diagnosis, serving as both a diagnostic aid and an educational resource for healthcare providers. Ultimately, the successful implementation of AI-driven diagnostic tools will depend on continuous collaboration between AI experts and clinicians to ensure their effectiveness in real-world clinical settings.
Limitations
This research has important limitations. One of the key challenges in AI-driven lower limb edema classification is the lack of publicly available datasets. Given that this field is still in its early stages, our study represents one of the first curated datasets specifically designed for this purpose. Data were collected from two independent medical institutions, incorporating a diverse range of clinical settings. However, despite this effort, our dataset primarily includes patients from a single geographic region, which may limit generalizability to broader populations. Expanding the dataset to include patients from different ethnic backgrounds and healthcare environments would enhance its representativeness and improve the robustness of AI predictions across diverse clinical settings. As AI-assisted edema diagnosis continues to develop, the availability of larger, multi-institutional datasets will be crucial for improving model performance.
Future work
To enhance the accuracy and applicability of the diagnostic tool, future research should focus on several key areas. Future studies should focus on validating AI models with independent, multi-center datasets beyond the current study population. Additionally, collaboration with other research institutions could support the establishment of a standardized dataset for AI-driven lower limb edema classification, ultimately facilitating model benchmarking and improving clinical translation. With these considerations, ongoing efforts will aim to solidify AI’s role as a reliable diagnostic tool while ensuring its performance remains consistent across various clinical contexts. The development of a more diverse and representative dataset will be essential in optimizing the real-world applicability of AI-assisted lower limb edema classification Additionally, validating the AI’s high accuracy in real clinical settings by integrating leg edema image classification with comprehensive clinical information would better mimic the typical diagnostic practices of healthcare professionals. This approach would ensure that the AI system not only excels in image analysis but also effectively supports clinicians in making accurate, well-rounded diagnoses in real-world scenarios.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Besharat, S. et al. Peripheral edema: A common and persistent health problem for older americans. PLoS ONE 16, e0260742 (2021).
Moffatt, C. J. et al. Lymphoedema: an underestimated health problem. QJM 96, 731–738 (2003).
Rockson, S. G. & Rivera, K. K. Estimating the population burden of lymphedema. Ann. N. Y. Acad. Sci. 1131, 147–154 (2008).
Gasparis, A. P., Kim, P. S., Dean, S. M., Khilnani, N. M. & Labropoulos, N. Diagnostic approach to lower limb edema. Phlebology 35, 650–655 (2020).
Yodrabum, N., Khaogate, K., Chaikangwan, I., Aojanepong, C. & Taweepraditpol, S. Lymphaticovenular anastomosis for patients with lymphedema of the upper extremity at siriraj hospital: A quantitative analysis study. J. Medical Assoc. Thailand104 (2021).
Vignes, S. et al. Primary lymphedema french national diagnosis and care protocol (pnds; protocole national de diagnostic et de soins). Orphanet J. Rare Dis. 16, 1–12 (2021).
Grada, A. A. & Phillips, T. J. Lymphedema: Pathophysiology and clinical manifestations. J. Am. Acad. Dermatol. 77, 1009–1020 (2017).
Orhurhu, V. et al. Management of lower extremity pain from chronic venous insufficiency: A comprehensive review. Cardiology Therapy 10, 111–140 (2021).
de Moraes Silva, M. A., Nakano, L. C., Cisneros, L. L. & Miranda Jr, F. Balneotherapy for chronic venous insufficiency. Cochrane Database Syst. Rev. (2023).
Sterns, R. H., Emmett, M. & Forman, J. P. Pathophysiology and etiology of edema in adults. UpToDate. Waltham, MA: UpToDate (2013).
Fu, M. R. et al. Machine learning for detection of lymphedema among breast cancer survivors. Mhealth4 (2018).
Wei, X. et al. Developing and validating a prediction model for lymphedema detection in breast cancer survivors. Eur. J. Oncol. Nurs. 54, 102023 (2021).
Ozmen, B. B. & Schwarz, G. Predicting breast cancer related lymphedema after immediate lymphatic reconstruction: an artificial intelligence approach with synthetic data. Plastic Reconstructive Surgery-Global Open 12, 65 (2024).
Krizhevsky, A., Sutskever, I. & Hinton, G. E. Imagenet classification with deep convolutional neural networks. Adv. Neural Inform. Process. Syst.25 (2012).
Simonyan, K. & Zisserman, A. Very deep convolutional networks for large-scale image recognition. arXiv preprint arXiv:1409.1556 (2014).
Huang, G., Liu, Z., Van Der Maaten, L. & Weinberger, K. Q. Densely connected convolutional networks. In: Proceedings of the IEEE conference on computer vision and pattern recognition, 4700–4708 (2017).
Szegedy, C. et al. Going deeper with convolutions. In: Proceedings of the IEEE conference on computer vision and pattern recognition, 1–9 (2015).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In: Proceedings of the IEEE conference on computer vision and pattern recognition, 770–778 (2016).
Iandola, F. N. Squeezenet: Alexnet-level accuracy with 50x fewer parameters and\(<\) 0.5 mb model size. arXiv preprint arXiv:1602.07360 (2016).
Howard, A. et al. Searching for mobilenetv3. In Proceedings of the IEEE/CVF international conference on computer vision, 1314–1324 (2019).
Tan, M. & Le, Q. Efficientnetv2: Smaller models and faster training. In: International conference on machine learning, 10096–10106 (PMLR, 2021).
Dosovitskiy, A. et al. An image is worth 16x16 words: Transformers for image recognition at scale (2021). arXiv:2010.11929.
Han, K. et al. Transformer in transformer. In: Neural Information Processing Systems (2021).
Liu, Z. et al. Swin transformer: Hierarchical vision transformer using shifted windows. In: 2021 IEEE/CVF International Conference on Computer Vision (ICCV), 9992–10002, https://doi.org/10.1109/ICCV48922.2021.00986 (2021).
Wu, H. et al. Cvt: Introducing convolutions to vision transformers. In: 2021 IEEE/CVF International Conference on Computer Vision (ICCV), 22–31, https://doi.org/10.1109/ICCV48922.2021.00009 (2021).
Heo, B. et al. Rethinking spatial dimensions of vision transformers. In: 2021 IEEE/CVF International Conference on Computer Vision (ICCV), 11916–11925, https://doi.org/10.1109/ICCV48922.2021.01172 (2021).
Hassani, A. et al. Escaping the big data paradigm with compact transformers. ArXivabs/2104.05704 (2021).
Tu, Z. et al. Maxvit: Multi-axis vision transformer. In: European Conference on Computer Vision (2022).
Ding, M. et al. Davit: Dual attention vision transformers. In: Avidan, S., Brostow, G., Cissé, M., Farinella, G. M. & Hassner, T. (eds.) Computer Vision – ECCV 2022, 74–92 (Springer Nature Switzerland, 2022).
Borman, P. Lymphedema diagnosis, treatment, and follow-up from the view point of physical medicine and rehabilitation specialists. Turkish J. Phys. Med. Rehabilitation 64, 179 (2018).
Krishnan, S. & Nicholls, S. C. Chronic venous insufficiency: clinical assessment and patient selection. In: Seminars in Interventional Radiology, vol. 22, 169–177 (Copyright 2005 by Thieme Medical Publishers, Inc., 333 Seventh Avenue, New ?, 2005).
Yu, L., Chen, H., Dou, Q., Qin, J. & Heng, P.-A. Automated melanoma recognition in dermoscopy images via very deep residual networks. IEEE Trans. Med. Imaging 36, 994–1004 (2016).
Irvin, J. et al. Chexpert: A large chest radiograph dataset with uncertainty labels and expert comparison. In: Proceedings of the AAAI conference on artificial intelligence 33, 590–597 (2019).
Rajpurkar, P. et al. Chexpedition: Investigating generalization challenges for translation of chest x-ray algorithms to the clinical setting. arXiv preprint arXiv:2002.11379 (2020).
Castilla, T., Martínez, M. S., Leguía, M., Larrabide, I. & Orlando, J. I. A resnet is all you need: modeling a strong baseline for detecting referable diabetic retinopathy in fundus images. In: 18th International Symposium on Medical Information Processing and Analysis, vol. 12567, 212–221 (SPIE, 2023).
Takahashi, S. et al. Comparison of vision transformers and convolutional neural networks in medical image analysis: A systematic review. J. Med. Syst. 48, 1–22 (2024).
Nagendran, M. et al. Artificial intelligence versus clinicians: systematic review of design, reporting standards, and claims of deep learning studies. bmj368 (2020).
Goyal, M. et al. Dfunet: Convolutional neural networks for diabetic foot ulcer classification. IEEE Trans. Emerging Topics Comput. Intell. 4, 728–739. https://doi.org/10.1109/TETCI.2018.2866254 (2020).
Nomura, Y. et al. Computer-aided diagnosis for screening of lower extremity lymphedema in pelvic computed tomography images using deep learning. Sci. Rep. 13, 16214 (2023).
Acknowledgements
The authors are grateful to Ploypan Seesun and Pattarawadee Prakobphol from the Research Department, Faculty of Medicine Siriraj Hospital, Mahidol University, for data analysis. This research project was supported by Faculty of Medicine Siriraj Hospital, Mahidol University, Grant Number (IO) R016631050.
Author information
Authors and Affiliations
Contributions
T.L.: Data collection, Data analysis and interpretation of data. T.T.: Conceptualization, Methodology, Software, Validation, Formal analysis, Writing - original draft, Visualization, Supervision. S.A.: Data analysis interpretation of data and writing manuscript. A.U.: Software, Data analysis and interpretation of data. V.S.: Data Collection. N.S.: Data Collection. S.B.: Data collection. T.P.: Data Collection. W.C.: Data collection. N.Y.: Conceptualization, Methodology, Data analysis and interpretation of data, Validation, Writing - original draft. All authors reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare no competing interests.
Generative AI and AI-assisted technologies in writing process
During the preparation of this work, the authors used ChatGPT to improve readability and grammar. After using this, the authors reviewed and edited the content as needed and took full responsibility for the publication’s content.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lewsirirat, T., Titijaroonroj, T., Apichonbancha, S. et al. Deep learning-based classification of lymphedema and other lower limb edema diseases using clinical images. Sci Rep 15, 12453 (2025). https://doi.org/10.1038/s41598-025-97564-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-97564-5
This article is cited by
-
Decision-Making for Lymphedema Diagnostic and Surveillance Measures: A Case-Based Approach
Current Physical Medicine and Rehabilitation Reports (2025)