Abstract
Photobiomodulation (PBM) has been widely utilized in regenerative medicine, including dermatology, dentistry, and neurology. However, the optimal energy density of PBM for human umbilical cord mesenchymal stem cells (hUC-MSCs) remains underexplored, hindering its development and potential clinical application. This study aims to identify the optimal wavelength and irradiation fluence for promoting the proliferation of hUC-MSCs by comparing the effects of different wavelengths and irradiation fluences. Our results show that green light enhances the anti-inflammatory properties of hUC-MSCs, with the 76s being the most effective in inhibiting IL-6 and GM-CSF. Blue light with 38 s is more effective in promoting angiogenesis, significantly increasing the mRNA and protein secretion of VEGF, HGF, and FGF2 compared to the non-irradiated group. The peak secretion times varied, with VEGF and FGF2 peaking at 72 h and HGF at 24 h. RNA-Seq confirms the significant roles of blue and green light in inhibiting epithelial-mesenchymal transition and inflammation. In vitro co-culture models and conditioned media experiments validate these anti-inflammatory effects. These findings have important implications for accelerating the clinical application of stem cell therapies and provide new references for PBM use in hUC-MSCs.
Similar content being viewed by others
Introduction
Regenerative medicine is a discipline dedicated to repairing or replacing damaged tissues and organs, with the goal of restoring normal function and health of the human body1,2,3. Wound healing is a significant area within regenerative medicine, involving the synergistic action of various cells and molecules to achieve wound closure, infection prevention, and tissue reconstruction4,5. However, factors such as age, disease, and environment may interfere with or delay the wound healing process, leading to the formation of chronic wounds6. This not only causes pain and burden to the patient but also imposes significant economic pressure on the healthcare system. Therefore, it is urgently necessary to find effective methods to promote wound healing. Stem cells, with their self-renewal and multi-differentiation characteristics, are considered unique tools in regenerative medicine7. In adulthood, stem cells do not significantly self-renew under physiological conditions, but when cell damage requires regeneration, the potential of stem cells changes with these physiological environmental changes. The potential for stem cells to differentiate into different types of cells is referred to as potency. The tissue from which stem cells are extracted will determine their potency, as this will change the potential for differentiation. However, despite the many advantages of mesenchymal stem cells, there are some limitations in the application process, mainly including: maintaining biological activity, quantifying active substances, and conditions for preservation and transportation8. These issues limit stability and reliability and also increase the cost and risk of their application. Therefore, it is necessary to find a cell-free method with the same yield and efficacy.
In recent years, cell-free therapy, as an emerging strategy in regenerative medicine, has received increasing attention9,10,11. Cell-free therapy refers to the use of a group of factors or molecules secreted by cells (such as exosomes, cytokines, and other signaling molecules), rather than the cells themselves, to achieve tissue repair and regeneration. This therapy has gained significant attention in regenerative medicine because it can avoid the potential risks associated with traditional cell-based therapies, such as immune rejection and tumor formation. Analysis of the secretome highlights its influence on various biological processes, including angiogenesis, neurogenesis, tissue repair, immunomodulation, wound healing, as well as anti-fibrotic and anti-tumor activities, all of which contribute to tissue maintenance and regeneration11. These factors or molecules are collectively referred to as secretomes, including soluble proteins, free nucleic acids, and extracellular vesicles12. Extracellular vesicles are a class of nanoparticles wrapped by the cell membrane, which can carry information molecules such as proteins, nucleic acids, and lipids, and interact with target cells through fusion or receptor-mediated methods, thereby regulating the function of target cells13. Extracellular vesicles can be divided into apoptotic bodies, microparticles, and exosomes according to their sources and sizes. The secretomes of individual cells and tissues are specific and vary with fluctuations in physiological states or pathological conditions14. Compared with applications based on stem cells, the use of cell-free therapy (such as secretomes derived from mesenchymal stem cells) in regenerative medicine offers key advantages, including the following advantages of secretomes: they can simulate the biological activity of cells without the need for cell survival and proliferation; they can avoid the risks of immune rejection and tumor formation associated with cell transplantation15,16,17.
On the other hand, photobiomodulation (PBM) is a physical therapy method that uses low-intensity lasers to stimulate cells, as shown in Fig. 1a18,19. Some studies have shown that after photon absorption, the excited molecules influence biological effects by modulating intracellular metabolic pathways. Light exposure can either increase adenosine tri-phosphate (ATP) and cyclic AMP levels, leading to bio-stimulation, or damage cytochromes, resulting in inhibition, depending on the exposure. Both processes occur in the mitochondria. Cytochrome c oxidase (COX), the key photoreceptor, is a terminal enzyme in the respiratory chain, playing a critical role in oxidative phosphorylation. COX contains heme, copper, magnesium, and zinc, and transfers electrons from cytochrome c to oxygen, which drives proton pumps, ultimately generating ATP20. Due to its safety and non-invasive characteristics, it has received widespread attention from researchers21. Its principle is the photochemical reaction of specific molecules (such as cytochrome c oxidase, mitochondria, receptors, etc.) of living cells to photons (i.e., light energy), including the visible light and near-infrared bands in the electromagnetic spectrum (with the power density of 10 mW/cm2−500 mW/cm2) stimulating the irradiated cells22. This non-invasive and non-toxic light therapy method has a wide range of applications, including dermatology, oral medicine, neurology, ophthalmology, cardiovascular medicine, oncology, sports medicine, rehabilitation medicine, etc23. However, it is regrettable that although PBM is now widely used, its optimal energy density for human umbilical cord mesenchymal stem cells (hUC-MSCs) has not been thoroughly studied in clinical applications, which seriously hinders the development of PBM in wound repair treatment and may even have a negative impact on it (such as carcinogenesis, burns, etc.)24,25. hUC-MSCsare an ideal model for studying mesenchymal cells of the inner connective tissue because they are easy to obtain, cause no harm to either the mother or the infant, and maintain their biological characteristics after cryopreservation and thawing26,27,28. Moreover, compared to bone marrow-derived mesenchymal stem cells, hUC-MSCs have a higher cell yield, stronger proliferative capacity, and lower immunogenicity. These properties make hUC-MSCs an excellent model for studying mesenchymal cells of the inner connective tissue29,30. Therefore, it is very necessary to conduct in-depth research on the scientific dosage of PBM.
In this study, we systematically evaluate the influence of different parameters of PBM on the healing potential of hUC-MSCs, as shown in Fig. 1b. First, we characterize the extracted hUC-MSCs by identifying the positive expression of CD73, CD90, and CD105, and the negative expression of Neg CKTL hematopoietic markers. We then assess the impact of different fluence of light intensity on the healing potential of hUC-MSCs, using variable fluence: 0/1/2/4/8/10 J/cm², output power density of 50 mW/cm² (see more details in Table 1 and Fig. S1) with blue light (450–455 nm) and green light (530–535 nm). The potential of hUC-MSCs to promote wound healing under different wavelengths and energy densities is compared from multiple angles, including cell viability assays, expression differences of proliferation and apoptosis-related genes, opsin expression, and gene expression and content of secretory factors. RNA-Seq sequencing reveals that the green light most significantly downregulate pathways related to inflammation and fibrosis. Starting from an inflammation cell model, we co-cultivate hUC-MSC conditioned medium with M1 macrophages to investigate the effects of light-treated hUC-MSCs on the polarization of M1 and M2 macrophages and the expression of inflammation-related genes. Finally, this work systematically analyzes and discusses the mechanisms by which the biological effects of light wavelengths and stem cell secretion effects contribute to wound healing. Additionally, with the advancement of regenerative medicine and tissue engineering, the findings of this study will aid in the development of stem cells or secretomes with specific bioregulatory properties, leading to the production of biotherapeutic products that improve healing quality and speed, thereby providing more effective clinical treatment options. Based on these results, further development of customized therapies can be pursued to offer patients more personalized and precise medical solutions, potentially providing valuable guidance for subsequent research in PBM and interdisciplinary medical engineering applications.
(a) Schematic diagram of the application of PBM in regenerative medicine. (b) Flow of this study. We explore the most effective PBM experimental parameters and investigate how the retinoid family responds to blue or green light PBM, examining the impact of blue or green light PBM on the repair potential of hUC-MSCs using RNA-Seq, and we reveal the underlying mechanism by which PBM regulates hUC-MSCs in macrophages through paracrine effects. Co-culture experiments and RNA sequencing provide insights into this intricate process. PBM, hUC-MSC denote photobiomodulation and human umbilical cord mesenchymal stem cells.
Results and discussion
Impact of PBM with different energies on hUC-MSC proliferation
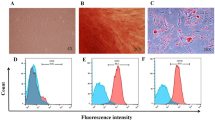
After seeding hUC-MSCs extracted from umbilical tissue in culture dishes for different days, adherent cells can be observed under a microscope. These cells exhibit a spindle-shaped or long spindle-shaped morphology. After 3 days of culture, the spindle-shaped cells elongate and occasionally intertwine in a network-like arrangement. Surface antigens of hUC-MSCs show positive expressions for CD90, CD105, and CD73, with expression rates of 100%, 99.89%, and 99.98%, respectively, and a negative expression rate of 0.52% for Neg CKTL. Flow cytometry data confirmed the mesenchymal stem cell characteristics of the isolated cells. MSC-specific markers are highly expressed, with CD73 at 98.7%, CD90 at 99.9%, and CD105 at 85.3%. Negative markers for hematopoietic stem cells, CD45 and CD34, including the macrophage marker CD11b, are not expressed (Fig. 2a). This validates that the isolated cells possessed MSC characteristics. To explore the effects of PBM on hUC-MSCs, we conducted comparative experiments using specific laser irradiation parameters. It is well known that laser parameters such as wavelength, power, and irradiance energy significantly impact cell proliferation31. While previous studies have shown that visible light can stimulate stem cell proliferation, few studies have focused on the shorter wavelengths within the blue-green spectrum, particularly for hUC-MSCs. Given the variability in irradiation parameters, methods, and cell types used, comparing results across studies is challenging. Therefore, we standardized the radiation source output power density at 50 mW/cm² and evaluate the effects of green light (500 nm) and blue light (455 nm) with different irradiance energies on hUC-MSCs.
The LED light device and control interface are shown in Fig. 2b, with the control center on the left adjusting the output power and irradiation time of each row. When cells are exposed to the longest set time (190 s), the post-irradiation temperature of the medium was recorded using an infrared thermal imaging system, showing no significant increase in medium temperature. The center wavelengths of the blue and green LED lights are depicted in the figure. To determine whether hUC-MSCs stimulate photoreceptors, we used qRT-PCR to explore the molecular expression basis of photopigments in hUC-MSCs. We aimed to identify photoreceptors responsive to blue and green light according to previous studies31,32. The results indicated that OPN4 is a gene with significant expression in response to blue and green light. We examined the gene expression levels of OPN4 and OPN1SW in hUC-MSCs at three time points (0 h, 24 h, and 48 h) following blue light (38 s; BLT) and green light (76 s; GLT) irradiation in Fig. 2c. Under green light, OPN4 expression increased significantly immediately after irradiation (p < 0.001) but decreased slightly at 24 h compared to 0 h. In contrast, OPN4 expression under blue light was generally lower than under green light (p < 0.001) and returned to control group levels by 48 h (p > 0.05). OPN1SW expression, however, increased significantly under blue light immediately (p < 0.001) but decreased at 24 and 48 h to levels similar to the control group. Additionally, it is important to note that statistical differences were observed in the expression of factors related to inflammatory pathways and photopigments. These differences may stem from variations in light exposure conditions (e.g., wavelength, intensity, duration) and subtle changes in the cell culture environment, which can influence cellular responses to light, leading to variations in expression levels.
We further assessed hUC-MSC proliferation over three days after exposure to blue and green light for different durations (19 s, 38 s, 76 s, 152 s, and 190 s)33,34,35. In the blue light group, the 38 s and 76 s groups treatments significantly increased cell proliferation at 24 h, 48 h, and 72 h (p < 0.01), with the 76 s group showing the highest activity. The 152 s and 190 s treatments reduced proliferation compared to the control group, while the 19 s treatment showed a slight difference (Fig. 2d). In the green light group, only the 152 s treatment significantly inhibited cell activity, while other durations showed no significant difference from the control. Due to the strong inhibitory effect observed with the 152 s treatment in both blue and green light groups (p < 0.001), this fluence was excluded from subsequent experiments. Optimal cell viability was observed at 72 h post-treatment (p< 0.01), and overall, monochromatic blue light was more effective than green light in promoting hUC-MSC proliferation. At 8 J/cm², both blue and green light inhibited proliferation, whereas intermediate fluences had more pronounced stimulatory effects (Fig. 2e)32,36,37.
(a) Identification of surface markers on hUC-MSCs by flow cytometry. (b) Illumination device and temperature assessment, and LED light source spectrum. (c) Relative gene expression of OPN4 and OPN1SW and the asterisks indicate statistical significance (*p < 0.05, **p < 0.01, ***p < 0.001, n = 3 per group). (d) Cell count under different conditions under varying wavelengths and irradiance fluence at different time points (24 h, 48 h, and 72 h, n = 3 per group). (e) Microscope images reveal the impact of PBM on hUC-MSCs. The conditions include control (no treatment), +BLT, +GLT, as well as combined blue and green light (+ BL + GLT). hUC-MSC, +BLT, and + GLT denote human umbilical cord mesenchymal stem cells, blue light and green light.
qRT-PCR analysis on the hUC-MSCs
Ki67, a nuclear antigen present in proliferating cells, is believed to be connected to chromatin and mitosis, making it one of the most widely used markers for proliferating cells33. PCNA is closely related to DNA synthesis and plays a crucial role in initiating cell proliferation, serving as an objective indicator of the proliferative state of cells38. Next, we assessed the expression of proliferation markers Ki67 and PCNA, which are indicative of cell proliferation39. In the blue light group, the 38 s treatment significantly increased Ki67 and PCNA expression compared to other fluences (p < 0.01), and the BCL2/BAX ratio indicated enhanced anti-apoptotic ability (p< 0.001)40,41. A partial increase in PCNA expression was also observed in the 19 s group (p < 0.05). In contrast, no significant changes in anti-apoptotic ability were observed in the green light group, but proliferation-related genes showed significant differences at various fluences, with the 76 s group showing the most notable increase (p < 0.01). We also examined angiogenic factors, including VEGF, HGF, and FGF2, using qRT-PCR (Fig. 3a-b). In the blue light group, VEGF expression increased significantly in the 38 s group, with levels 1.5 times higher than the control (***p < 0.0001). HGF expression was significantly elevated in the 19 s and 38 s groups at 48 h post-irradiation, with the highest increase observed in the 38 s group (p < 0.001). FGF2 expression peaked in the 38 s group, with no significant differences observed at other fluences. In the green light group, no significant differences were observed in angiogenesis-related gene expression.
ELISA analysis on the hUC-MSCs
To further investigate the impact of PBM on the paracrine properties of hUC-MSCs, we conducted ELISA on conditional media collected 24 h, 48 h, and 72 h post-irradiation and mock treatment in Fig. 4. The detection of VEGF revealed that overall, the secretion in the blue light 38 s and green light 76 s groups is significantly higher than that in the control group. The protein secretion of VEGF is more pronounced in the blue light 38 s group compared to the green light 76 s group (p < 0.01). Additionally, comparing − LLT and + BLT_38 groups and − LLT and + GLT_76 groups, the secretion of VEGF in the + BLT_38 group shows no significant difference from the control group at 24 h and 48 h but gradually exhibited significant differences at 72 h (p< 0.001). In contrast, the secretion of VEGF in the green light group showed no significant difference from the control group at 24 h but significantly increased at 48 h and 72 h in Fig. 4. This result indicated that the expression of pro-angiogenic factors induced by blue and green light may have a certain delay depending on the wavelength and fluence42,43. For HGF secretion, there was no significant difference between the B38 and G76 groups and the control group, whereas the green light 190 s group show a significant difference from the control group (p < 0.05). The secretion of FGF2 in the blue light 38 s group was significantly higher than that in the untreated group at 24 h, 48 h, and 72 h, while the green light 76 s group showed no significant differences at these time points, maintaining a relatively higher level compared to the control group. Overall, the secretion was only second to the blue light 38 s group (p < 0.01). The regulation of cytokines such as TIMP-1, GM-CSF, and IL-6 contributes to improve wound healing. ELISA analysis showed that the secretion of TIMP-1 by hUC-MSCs significantly increases in the blue light 76 s and 190 s groups compared to the untreated control group, with the 76 s group showing higher secretion than the 190 s group, and the differences are statistically significant. The secretion of GM-CSF and IL-6 decrease significantly after irradiation, especially in the green light 76 s and 190 s groups.
RNA-Seq study on the effects of PBM on the hUC-MSCs
Compared to control cells under no light conditions, RNA-Seq identifies global changes in gene expression of hUC-MSCs pre-treated with blue light at 2 J/cm² or green light at 4 J/cm². We conducted pairwise correlation analysis of the gene expression levels of the samples and plot a correlation heatmap to reflect the overall similarities and differences between the samples, as shown in Figs. 5a-b. Scatter plots of the PBM-treated groups versus the control samples are shown in Figs. 5c-d. Many genes are significantly upregulated or downregulated after PBM treatment with blue or green light compared to the blank control group. Gene Set Enrichment Analysis (GSEA) of normalized counts of all genes expressed under blue and green light conditions compared to the Hallmark database genes expressed under control conditions showed significant enrichment (FDR < 25%) of six gene sets in the control group. The categories most enriched under blue light included downregulated genes in EMT and NFkB-TNFA signaling. Under green light, the categories most enriched included downregulated genes in Angiogenesis, NFkB-TNFA signaling, TGF-β signaling, and IL-6-JAK-STAT3 signaling. Post-mRNA sequencing alignment analysis showed that blue light significantly downregulated the EMT pathway, indicating anti-fibrotic potential. Similarly, green light significantly downregulated vascularization-related genes and anti-inflammatory pathways, aligning with the previous ELISA and qRT-RNA results (see more details in Fig. S2-S3).
Transcriptome analysis of the effect of PBM on hUC-MSCs. Differential gene volcano plots of (a) blue light of 2 J/cm2 vs. non-irradiated group and (b) green light of 4 J/cm2 vs. non-irradiated group. Heat map of differential gene clustering with (c) blue light and (d) green light group. hUC-MSCs and PBM mean human umbilical cord mesenchymal stem cells and photobiomodulation.
Co-culture of PBM-treated hUC-MSCs and M1 macrophages
Conditioned media (CM) represents a complete regenerative environment composed of cell-derived secretomes and vesicular molecules. The soluble components of secretomes can be separated and prepared through centrifugation, filtration, polymer-based precipitation methods, ion-exchange chromatography, and size-exclusion chromatography. Both components can independently trigger regeneration and repair, mediating the neogenesis of tissue-engineered organs in vitro. It has been demonstrated that CM derived from mesenchymal stem cells (MSCs) is sufficient to significantly improve multiple biomarkers of pathophysiology44. During the initial inflammatory response of wound healing, M1 macrophages are present in large numbers and produce a significant amount of pro-inflammatory factors, whereas M2 macrophages dominate during the resolution phase, primarily secreting anti-inflammatory and growth factors and exhibiting high phagocytic activity to clear necrotic and damaged cells from the wound surface. Therefore, the M2 phenotype and phagocytic function of macrophages are critical for wound repair. Based on RNA-Seq sequencing results, we further investigated the regulation of anti-inflammatory and anti-fibrotic capacities of hUC-MSCs under different light conditions. A co-culture model is constructed, dividing the cells into the upper layer (M1 macrophages) and the lower layer (hUC-MSCs) using cell chambers, as shown in Fig. 6a. The pore membrane of the chamber allows cytokines from both layers to pass through, simulating the effect of paracrine factors from hUC-MSCs on M1 macrophages in vivo. After 48 h of co-culture (n= 3 per group), the mRNA expression levels of M1-related pro-inflammatory factors (IL-1α, IL-6) and M2-related anti-inflammatory factor arginase (Arg1) are detected by qRT-PCR in the following groups: M1 (LPS + IFN-γ), M2 (IL-13 + IL-4), +BLT/+GLT (LPS + IFN-γ + PBM-CM), -LLT (LPS + IFN-γ + CM). The expression levels of pro-inflammatory cytokines IL-1α and IL-6 were both downregulated under the influence of secreted factors derived from PBM-treated hUC-MSCs, regardless of whether they were pretreated with blue or green light, as shown in Fig. 6b45,46,47,48. Specifically, for IL-1α, its expression was slightly reduced following treatment with secretomes from green light-preconditioned hUC-MSCs, and this reduction was marginally more pronounced compared to the blue light group. However, neither group demonstrated a statistically significant decrease in IL-1α expression. In contrast, IL-6 expression was markedly suppressed in both blue and green light preconditioned groups, with a more substantial reduction observed in the green light group compared to blue light, indicating a more prominent anti-inflammatory effect mediated by green light. Regarding the anti-inflammatory marker arginase-1 (Arg1), its expression was significantly higher in hUC-MSC co-culture groups compared to the M1 macrophage control. Notably, the expression level of Arg1 in the blue light group was comparable to the no-light group. However, in comparison, the green light group exhibited a significantly greater elevation in Arg1 expression, further supporting the enhanced M2 polarization effect induced by green light preconditioning.
(a) Schematic representation of the co-culture model of M1 macrophages with treated or non-light-treated hUC-MSCs cells. (b) Effect of PBM & hUC-MSCs on inflammation-related genes in M1 macrophages through paracrine effects (*p < 0.05; **p < 0.01). hUC-MSCs and PBM mean human umbilical cord mesenchymal stem cells and photobiomodulation.
Regulation of M1 macrophage polarization by conditional medium from PBM-Treated hUC-MSCs
To investigate whether photobiomodulation (PBM) preconditioning of hUC-MSCs modulates their immunoregulatory capacity, we evaluated the effects of PBM-conditioned medium on macrophage polarization. MSCs are known to influence immune responses through the secretion of paracrine factors, and macrophages, as key regulators of inflammation and repair, can adopt distinct phenotypes (M1 vs. M2) depending on microenvironmental signals49. Shifting macrophages toward the M2 phenotype is especially critical in promoting wound resolution and tissue regeneration50. After concentrating the PBM-conditioned medium fivefold, it was added to the basal medium of M1-polarized macrophages and incubated for 48 h.
Flow cytometry analysis revealed distinct expression patterns of M1 and M2 surface markers under green and blue light pretreatment conditions (Fig. 7). CD40 expression remained consistent across all groups, with nearly identical levels in the blue and green light groups51. In contrast, CD80 expression was slightly lower in the blue light group compared to the green light group. For M2 markers, more prominent differences were observed between the groups. CD206 expression was significantly higher in the green light group compared to both the blue light and untreated control groups. CD206 expression in the blue light group was slightly reduced relative to the control. Regarding CD209, both light-treated groups exhibited higher expression than the untreated group, with the green light group showing a moderately elevated level compared to the blue light group. These findings suggest that green-light-treated hUC-MSCs induce a more robust M2 polarization effect than blue-light-treated or untreated cells52,53,54. Our flow cytometry data confirmed that the number of CD206-positive macrophages in the green-light-treated hUC-MSC (GLT-MSC) group was significantly greater than in the blue-light-treated (BLT-MSC) or control (DMEM or S-MSC) groups. Conversely, CD86-positive macrophages were more abundant in the BLT and control groups, further supporting the enhanced anti-inflammatory transition induced by green light. This co-culture-based polarization model is widely used to simulate in vivo immune–stromal cell interactions and to assess the paracrine regulatory potential of stem cells. Numerous studies have demonstrated that MSC-derived cytokines and extracellular vesicles can drive macrophages toward the M2 phenotype, thereby limiting inflammation and promoting tissue regeneration50,55,56. M2 macrophages play a central role in the proliferative and remodeling phases of wound healing by secreting pro-regenerative growth factors such as VEGF, TGF-β, and IGF-1, which enhance fibroblast and endothelial cell proliferation, matrix remodeling, and angiogenesis57. Therefore, our results support the conclusion that green light PBM enhances the immunomodulatory secretome of hUC-MSCs, which in turn promotes macrophage M2 polarization58,59. This pathway may provide a mechanistic link between light-based preconditioning and improved regenerative outcomes in stem-cell-based therapies.
Limitations of the study
While this study demonstrates the immunomodulatory and regenerative potential of PBM-preconditioned hUC-MSCs through paracrine mechanisms, several limitations should be acknowledged. First, we did not perform direct measurements of oxidative stress, ATP production, or DNA synthesis, which are commonly associated with the intracellular effects of PBM. Second, the central scratch assay was not applied, as hUC-MSCs are less migratory than fibroblasts, and their therapeutic effects are predominantly mediated via secretome-driven modulation of immune responses. Instead, macrophage co-culture and polarization analysis were employed to capture these paracrine effects. Future studies will incorporate intracellular functional assays and wound closure models to provide a more comprehensive understanding of PBM effects.
Conclusion
This study investigated the effects of PBM on hUC-MSCs and their potential in wound healing. Our findings demonstrated that PBM, particularly with blue and green light, significantly enhances the expression of proliferative and angiogenic markers in hUC-MSCs. RNA-Seq analysis revealed that PBM modulates various gene pathways, promoting anti-inflammatory and anti-fibrotic effects. Additionally, co-culture experiments with M1 macrophages indicate that PBM-treated hUC-MSCs can effectively promote the polarization of M2 macrophages, enhancing their anti-inflammatory and tissue repair capabilities. These results suggest that PBM-treated hUC-MSCs hold significant promise for improving wound healing and regenerative therapies.
Materials and methods
Laser device
The light source used in this experiment was a secondary light-emitting diode (LED) laser (China Chengdu Huayuan Light Source Control Co., Ltd.) with 24 light-emitting heads, designed to fit a 24-well plate size, adhering to a “one head per well” configuration. The dual-frequency laser, equipped with multimode fibers, emitted wavelengths of 500 nm and 455 nm for cell irradiation. The laser was wrapped in plastic film to protect the light source from environmental influences. The 24-well plate (4 × 6 configuration) was mounted on the laser panel, with slight gaps and partitions to minimize interference between light sources and facilitate equipment cooling. Based on previous studies using similar power settings10,11,12,13, the laser power was set to 50–500 mW/cm². All LED emission heads were set to 50 mW/cm². The exposure times of 0 s, 19 s, 38 s, 76 s, 152 s, and 190 s, respectively, corresponding to fluences were 0, 1, 2, 4, 8, and 10 J/cm². hUC-MSCs were exposed to the LED light source at room temperature. At the same time points, control group cells were kept in a dark environment. After the exposure was completed, all groups of cells were kept in a CO₂ incubator. We use an infrared thermal imager (TESTO-865, Germany) to evaluate the temperature of the medium after light treatment47,48.
Light source and irradiation parameters
hUC-MSCs were purchased from Aoyinbio (Shanghai, China) and Oribiotech Co. from Shanghai, China, and isolated from Wharton’s jelly of the umbilical cord through physical dissection and enzymatic digestion using collagenase and hyaluronidase. The hUC-MSCs used in the experimental group were isolated and differentiated by Sichuan Cunde Therapeutics Co., Ltd (Chengdu, China), as shown in Table 1. The cells were cultured in a suspension medium designed for MSC growth, regularly monitored, and passaged upon reaching optimal density. Cell passaging was done by treating with trypsin, followed by centrifugation and re-seeding at a 1:3 ratio in T175 culture flasks. Before passaging, Cell dissociation was treated with trypsin, then collected and counted for subsequent experimental procedures. For the experiments, cells at passages 5–6 were used, seeded at a density of 1.5 × 104 cells/well in a 24-well plate, and prepared for photobiomodulation (PBM) treatment after 24 h of incubation. Characterization of hUC-MSCs was confirmed by flow cytometry, which verified positive expression of surface markers CD73, CD90, and CD105, while hematopoietic markers CD34 and CD45 showed negative expression. For antigen identification, cells were incubated with corresponding antibodies (CD90-FITC, CD105-PerCP-CY5.5, CD73-APC-A, and Neg CKTL-PE-A) and analyzed. The cells were then subjected to PBM treatment and incubated for 24 h, 48 h, and 72 h, with cell lysates and supernatants collected for further analysis.
CCK8 cell viability and proliferation assay
The enhanced CCK8 assay kit (CCK-8) from Beyotime (Shanghai, China) was used to assess cell viability. The CCK-8 assay measures the quantity of formazan dye produced due to reduced dehydrogenase activity within cells, which is proportional to the number of living cells. In brief, the fourth-generation cells from three different batches of hUC-MSCs were cultured in a 24-well plate at a density of 1.5 × 104 cells/well. Before irradiation, the cells were washed once with PBS. Fresh serum-free culture medium was added, and the cells were then exposed to LED light. After irradiation, hUC-MSCs were incubated for 24 h. Subsequently, CCK-8 solution was added to each well and incubated for 1 h. After incubation, the cell suspension was transferred to a 96-well plate with approximately 100 µL per well. Each sample was tested in triplicate. The absorbance at 450 nm was measured using a microplate reader (Männedorf). The cell number was calculated as the percentage of viable cells relative to untreated control cells (n = 3 per group).
Quantitative reverse transcription polymerase chain reaction (qRT-PCR)
Primers for qRT-PCR were designed using Primer-BLAST (NCBI) to ensure specificity and efficiency. The sequences for the target genes and the housekeeping gene GAPDH were obtained from the NCBI database. The following criteria were used for primer design:
VEGF/F: AGGGCAGAATCATCACGAAGT
VEGF/R: AGGGTCTCGATTGGATGGCA
HGF/F: GCTATCGGGGTAAAGACCTACA
HGF/R: CGTAGCGTACCTCTGGATTGC
FGF2/F: AGAAGAGCGACCCTCACATCA
FGF2/R: CGGTTAGCACACACTCCTTTG
PCNA/F: CCTGCTGGGATATTAGCTCCA
PCNA/R: CAGCGGTAGGTGTCGAAGC
Ki-67/F: ACGCCTGGTTACTATCAAAAGG
Ki-67/R: CAGACCCATTTACTTGTGTTGGA
BAX/F: CCCGAGAGGTCTTTTTCCGAG
BAX/R: CCAGCCCATGATGGTTCTGAT
BCL2/F: GGTGGGGTCATGTGTGTGG
BCL2/R: CGGTTCAGGTACTCAGTCATCC
OPN4/F: ACCGCTACCTGGTAATCACAC
OPN4/R: GCGTGAAGCTCATGTAGTCCC
OPN1SW/F: ATGGGCCTCAGTACCACATT
OPN1SW/R: GCAACTTTTTGTAGCGCAGT
CXCL12/F: ATTCTCAACACTCCAAACTGTGC
CXCL12/R: ACTTTAGCTTCGGGTCAATGC
CXCL4/F: ACTACACCGAGGAAATGGGCT
CXCL4/R: CCCACAATGCCAGTTAAGAAGA
GAPDH-F: GGAGCGAGATCCCTCCAAAAT
GAPGH-R: GGCTGTTGTCATACTTCTCATGG
RNA extraction
Total ribonucleic acid (RNA) was extracted from the cell culture supernatant using a commercial RNA purification kit (specify brand, catalog number if applicable) following the manufacturer’s instructions. Briefly, 200 µl of sample was lysed in Buffer DRL containing linear acrylamide, followed by the addition of isopropanol to precipitate nucleic acids. The lysate was applied to a DNA/RNA spin column, and sequential wash steps were performed using Buffer RW1 and Buffer RW2 to remove contaminants. Finally, RNA was eluted with 30–50 µl of RNase-free water preheated to 65 °C. All centrifugation steps were conducted at the specified speeds and durations recommended in the kit protocol. The concentration and purity of RNA were assessed using a NanoDrop spectrophotometer (Thermo Fisher Scientific, USA), and samples were stored at − 80 °C until further use.
ELISA analysis
To analyze the secretion of paracrine factors from hUC-MSCs, we used ELISA kits for human VEGF, human HGF, human FGF2, human TIMP-1, human IL-6, and human GM-CSF according to the manufacturer’s instructions. Conditioned media were collected after 48 and 72 h of treatment with blue-green OLED. The optical density (OD) values of each well were measured at 450 nm using a microplate reader.
RNA-Seq and gene set enrichment analysis (GSEA)
After 48 h of PBM intervention, total RNA was collected from hUC-MSCs in the PBM intervention groups (blue light group and green light group) and the blank control group using the TRIzol reagent kit (Takara). The RNA quality was assessed using gel electrophoresis and a Nanodrop spectrophotometer (Thermo, Waltham, MA, USA). mRNA enrichment, fragmentation, reverse transcription, and library construction were performed by Hengyuan BioTech Co., Ltd. (Chengdu). We analyzed the transcriptional abundance from a ranked list of all available genes (whole genome ranked list without cutoff values) using Gene Set Enrichment Analysis (GSEA) software version 4.1.0 to identify functionally related gene sets with statistically significant enrichment.
Co-culture of hUC-MSCs and macrophages
M0 macrophages (1 × 10⁶ cells) were treated with 20 ng/mL lipopolysaccharide (LPS) and interferon-γ (IFN-γ) in Texmacs medium at 37 °C and 5% CO₂ for 24 h to induce the M1 phenotype. Minimum essential medium-alpha (mem-alpha), with l-glutamine, ribonucleosides, and deoxyribonucleosides, without ascorbic acid is purchased from Oribiotech Co. from Shanghai. Mem-alpha contains no proteins, lipids, or growth factors. After passaging the adherent cells, the medium was switched from a-MEM + 10% FBS to the serum-free basal medium (a-MEM) before photobiomodulation (PBM) treatment. The cells were then cultured for an additional 48 h post-irradiation. Therefore, the factors detected by ELISA represent those secreted by hUC-MSCs after light exposure. Flow cytometry analysis confirmed the induction of the M1 phenotype in macrophages. As a control, M1 macrophages were treated with IL-13 (20 ng/mL) and IL-4 (20 ng/mL) to induce the M2 phenotype. The phenotype changes induced by LPS and IFN-γ were identified using M1 markers CD40 and CD86, and M2 markers CD206 and CD209. M1 macrophages (1 × 10⁵ cells) were seeded into 24 mm polyester membrane Transwell® inserts (0.4 μm pore size, Corning Inc.), and the lower chamber was seeded with untreated hUC-MSCs (1 × 10⁵ cells) and hUC-MSCs treated with blue or green light PBM. The co-culture was maintained at 37 °C and 5% CO₂ for 48 h. The next day, macrophages were collected for RNA extraction. For M2 macrophage polarization: After culturing with the supernatant from each group for 24 h, flow cytometry is used to detect specific proteins of M2 macrophage markers: CD206, CD163, Arg-1, and IL-10, to evaluate the anti-inflammatory activity of macrophages.
Statistical analysis
To ensure the scientific validity and reliability of the analysis results, data are presented as mean ± standard deviation. All tests were performed two-sided, with a significance level of *p < 0.05 considered statistically significant (*p < 0.05, **p < 0.01, ***p < 0.001, and ****p < 0.0001). GraphPad Prism was used for all statistical analyses. All qRT-PCR, CCK-8 assays, ELISA, and other tests were representative of three independent experiments. Experimental data were analyzed using GraphPad Prism 9.0 (GraphPad Software Inc., San Diego, CA, USA). Independent sample t-tests were used for comparisons between two independent samples, and one-way ANOVA was used for comparisons among different groups. Data are expressed as mean ± standard deviation (X ± S), with P < 0.05 indicating statistically significant differences.
Data availability
The data that support the plots within this paper and other finding of this study are available from the [github] 6aa8596 (https://github.com/YANHONGSCU/HongYangroup) or corresponding authors.
References
Tatullo, M. et al. Stem cells and regenerative medicine. In Stephen T. Sonis and Allessandro Villa (Eds.), Translational Systems Medicine and Oral Disease (387–407). Academic. (2020).
Bhaskar, B. & Nagarjuna, V. Biomaterials, Tissue Engineering, and Regenerative Medicine: A Brief Outline. Biomaterials in Tissue Engineering and Regenerative Medicine: From Basic Concepts to State of the Art Approaches, 3–17. (2021).
Santana, M. H. A. & Huber, S. C. History and evolution of regenerative medicine. In Maria H. Santana, Eliana B. Souto, Ranjita Shegokar (Eds.), Nanotechnology and Regenerative Medicine (23–44). Academic. (2023).
Kaur, G., Narayanan, G., Garg, D., Sachdev, A. & Matai, I. Biomaterials-based regenerative strategies for skin tissue wound healing. ACS Appl. Bio Mater. 5 (5), 2069–2106 (2022).
Rani Raju, N. et al. Multifunctional and smart wound Dressings—A review on recent research advancements in skin regenerative medicine .Pharmaceutics 14 (8), 1574 (2022).
Falanga, V. et al. Chronic wounds. Nat. Reviews Disease Primers. 8 (1), 50 (2022).
Gupta, S., Sharma, A. & Verma, R. S. Mesenchymal stem cells for cardiac regeneration: from differentiation to cell delivery. Stem Cell. Reviews Rep. 17 (5), 1666–1694 (2021).
Benoit, C., Virginie, C. & Boris, V. The use of NADES to support innovation in the cosmetic industry. In Lemonnier, N. & Antoniotti, S. (Eds.), Advances in Botanical Research (Vol. 97, 309–332). Academic. (2021).
Da Silva, D., Crous, A. & Abrahamse, H. Photobiomodulation: an effective approach to enhance proliferation and differentiation of adipose-derived stem cells into osteoblasts. Stem Cells International, 2021. (2021).
Gomzikova, M. O. & Rizvanov, A. A. Current trends in regenerative medicine: from cell to cell-free therapy. Bionanoscience 7, 240–245 (2017).
Kumar, P. et al. The mesenchymal stem cell secretome: A new paradigm towards cell-free therapeutic mode in regenerative medicine. Cytokine Growth Factor Rev. 46, 1–9 (2019).
Steffens, R. C. & Wagner, E. Directing the way—receptor and chemical targeting strategies for nucleic acid delivery. Pharm. Res. 40 (1), 47–76 (2023).
Ayaz-Guner, S. et al. A comparative study on normal and obese mice indicates that the secretome of mesenchymal stromal cells is influenced by tissue environment and physiopathological conditions. Cell Communication and Signaling 18(1), 1–19 (2020).
Ritchie, S., Reed, D. A., Pereira, B. A. & Timpson, P. The cancer Cell Secretome Drives Cooperative Manipulation of the Tumour Microenvironment To Accelerate Tumourigenesis 10 (Faculty Reviews, 2021).
Daneshmandi, L. et al. Emergence of the stem cell secretome in regenerative engineering. Trends in Biotechnology 38(12), 1373–1384 (2020).
Arai, A. J., Haider, K. H. & S., & Researches and Applications of Stem Cell Secretome 191–223 (From Potential to Promise, 2021).
Liebert, A. et al. Photophysical mechanisms of photobiomodulation therapy as precision medicine. Biomedicines 11 (2), 237 (2023).
Ma, F. et al. Effects of Photobiomodulation and Low-Intensity Stretching on Delayed-Onset Muscle Soreness: A Randomized Control Trial. Photobiomodulation, Photomedicine, and Laser Surgery 40(12), 810–817 (2022).
González-Muñoz, A. et al. Efficacy of Photobiomodulation Therapy in the Treatment of Pain and Inflammation: A Literature Review. In Healthcare https://doi.org/10.3390/healthcare11070938 (2023).
Hammes, F., Goldschmidt, F., Vital, M., Wang, Y. & Egli, T. Measurement and interpretation of microbial adenosine tri-phosphate (ATP) in aquatic environments .Water Res. 44 (13), 3915–3923 (2010).
Paramaguru Rajesh, S. Analysis of Functional Outcome of Various Conservative Modalities in Treatment of Plantar Fasciitis: A Prospective study (Doctoral dissertation, Madurai Medical College, Madurai). (2020).
Andrabi, M., Upton, B. A., Lang, R. A. & Vemaraju, S. An expanding role for nonvisual Opsins in extraocular light sensing physiology. Annual Rev. Vis. Sci. 9 (1), 245–267 (2023).
Del Vecchio, A. et al. Laser photobiomodulation (PBM)—a possible new frontier for the treatment of oral cancer: A review of in vitro and in vivo studies. In Healthcare https://doi.org/10.3390/healthcare9020134 (2021).
Horton, L., Brady, J., Kincaid, C. M., Torres, A. E. & Lim, H. W. The effects of infrared radiation on the human skin. PhotoDermatol. PhotoImmunol. PhotoMed. 39 (6), 549–555 (2023).
Mansano, B. S. D. M. et al. do Nascimento, Enhancing the therapeutic potential of mesenchymal stem cells with light-emitting diode: implications and molecular mechanisms. Oxidative Medicine and Cellular Longevity, 2021. (2021).
Kim, K. et al. Photobiomodulation enhances the angiogenic effect of mesenchymal stem cells to mitigate radiation-induced enteropathy. Int J Mol Sci. https://doi.org/10.3390/ijms20051131 (2019).
Liu, J. et al. Numerical study on extinction performance of ag nanoparticles@ SiO2 ellipsoid. J. Mater. Res. Technol. 9 (3), 6723–6732 (2020).
Cases-Perera, O. et al. Development of secretome-based strategies to improve cell culture protocols in tissue engineering. Scientific reports https://doi.org/10.1038/s41598-022-14115-y (2022).
Hu, H. et al. Laser irradiation of electrode materials for energy storage and conversion. Matter 3(1), 95–126 (2020).
Malthiery, E. et al. Effects of green light photobiomodulation on Dental Pulp Stem Cells: enhanced proliferation and improved wound healing by cytoskeleton reorganization and cell softening. Lasers Med Sci. 36(2), 437–445. https://doi.org/10.1007/s10103-020-03092-1 (2021).
Cios, A. et al. Effect of different wavelengths of laser irradiation on the skin cells. Int J Mol Sci. https://doi.org/10.3390/ijms22052437 (2021).
Liu, C. et al. Mesenchymal stem cells pretreated with proinflammatory cytokines accelerate skin wound healing by promoting macrophages migration and M2 polarization. Regen Ther. https://doi.org/10.1016/j.reth.2022.06.009 (2022).
Fang, S. et al. Umbilical cord-derived mesenchymal stem cell-derived exosomal microRNAs suppress myofibroblast differentiation by inhibiting the transforming growth factor-β/SMAD2 pathway during wound healing. Stem Cells Transl Med. 5(10), 1425–1439 (2016).
Lv, Y. et al. Evaluation of the red & blue LED effects on cutaneous refractory wound healing in male Sprague-Dawley rat using 3 different multi-drug resistant bacteria. Lasers Surg Med. 54(5), 725–736 (2022).
Li, J. et al. Comparison of the biological characteristics of human mesenchymal stem cells derived from exfoliated deciduous teeth, bone marrow, gingival tissue, and umbilical cord corrigendum in/10.3892/mmr. 2022.12919 corrigendum in/10.3892/mmr. 2022.12920. Mol. Med. Rep. 18 (6), 4969–4977 (2018).
Manouras, T., Koufakis, E., Vasilaki, E., Peraki, I. & Vamvakaki, M. Antimicrobial hybrid coatings combining enhanced biocidal activity under visible-light irradiation with stimuli-renewable properties. ACS Appl. Mater. Interfaces. 13 (15), 17183–17195 (2021).
Liu, Y. et al. Human umbilical cord-derived mesenchymal stem cells not only ameliorate blood glucose but also protect vascular endothelium from diabetic damage through a paracrine mechanism mediated by MAPK/ERK signaling. Stem Cell Res Ther. https://doi.org/10.1186/s13287-022-02927-8 (2022).
Ma, Y. et al. Photobiomodulation attenuates neurotoxic polarization of macrophages by inhibiting the notch1-HIF-1α/NF-κB signalling pathway in mice with spinal cord injury. Front Immunol. https://doi.org/10.3389/fimmu.2022.816952 (2022).
Han, X. et al. Protective effect of Gastrodin on myocardial ischemia–reperfusion injury and the expression of Bax and Bcl–2. Experimental Therapeutic Med. 17 (6), 4389–4394 (2019).
Kim, S. W. et al. Phototoxicity-free blue light for enhancing therapeutic angiogenic efficacy of stem cells. Cell Biol Toxicol. https://doi.org/10.1007/s10565-021-09656-3 (2021).
Dehghani Soltani, S. et al. Different effects of energy dependent irradiation of red and green lights on proliferation of human umbilical cord matrix-derived mesenchymal cells. Lasers Med. Sci. 31, 255–261 (2016).
Nieves, A. A. B. Effect of Low Wavelength Blue Light and Ultraviolet Light in Optically Active Human Umbilical Cord Mesenchymal Stem Cells and the Effect of Spinal Cord Injury at the Level of C3 to C5 in the Requirements of Mechanical Ventilation (Master’s thesis, College of Medicine-Mayo Clinic). (2024).
Moradi, A., Ghaffari Novin, M. & Bayat, M. A Comprehensive Systematic Review of the Effects of Photobiomodulation Therapy in Different Light Wavelength Ranges (Blue, Green, Red, and Near-Infrared) on Sperm Cell Characteristics in Vitro and in Vivo 1–28 (Reproductive Sciences, 2024).
Vilaça-Faria, H., Salgado, A. J. & Teixeira, F. G. Mesenchymal stem cells-derived exosomes: a new possible therapeutic strategy for Parkinson’s disease? Cells 8 (2), 118 (2019).
Serrage, H. et al. Under the spotlight: mechanisms of photobiomodulation concentrating on blue and green light. Photochem. Photobiol. Sci. 18 (8), 1877–1909 (2019).
Serrage, H. et al. Under the spotlight: mechanisms of photobiomodulation concentrating on blue and green light. Photochem. Photobiol. Sci. 18 (8), 1877–1909 (2019).
Dou, C. et al. Au-functionalized wrinkle graphene biosensor for ultrasensitive detection of Interleukin-6. Carbon 216, 118556 (2024).
Liu, J. et al. Inverse design a patternless solar energy absorber for maximizing absorption. Sol. Energy Mater. Sol. Cells. 244, 111822 (2022).
Lu, D. et al. Mesenchymal stem cells influence monocyte/macrophage phenotype: regulatory mode and potential clinical applications. Biomed. Pharmacother. 165, 115042 (2023).
Arabpour, M., Saghazadeh, A. & Nima Rezaei. Anti-inflammatory and M2 macrophage polarization-promoting effect of mesenchymal stem cell-derived exosomes. Int. Immunopharmacol. 97, 107823 (2021).
De Santa et al. The role of metabolic remodeling in macrophage polarization and its effect on skeletal muscle regeneration. Antioxid. Redox. Signal. 30 (12), 1553–1598 (2019).
Mao, J. et al. Advanced biomaterials for regulating polarization of macrophages in wound healing. Adv. Funct. Mater. 32 (12), 2111003 (2022).
Sharifiaghdam, M. et al. Macrophages as a therapeutic target to promote diabetic wound healing. Mol. Ther. 30 (9), 2891–2908 (2022).
Yan, L. et al. Macrophage plasticity: signaling pathways, tissue repair, and regeneration. MedComm. https://doi.org/10.1002/mco2.658 (2024).
Soufihasanabad, S. et al. In vivo polarization of M2 macrophages by mesenchymal stem cell-derived extracellular vesicles: A novel approach to macrophage polarization and its potential in treating inflammatory diseases. Med. Hypotheses. 187, 111353 (2024).
Chen, Y., Yang, L. & Li, X. Advances in mesenchymal stem cells regulating macrophage polarization and treatment of sepsis-induced liver injury. Front. Immunol. 14, 1238972 (2023).
Nazari, M. et al. Therapeutic Properties of M2 Macrophages in Chronic Wounds: An Innovative Area of Biomaterial-Assisted M2 Macrophage Targeted Therapy. Stem Cell Rev Rep. https://doi.org/10.1007/s12015-024-10806-3 (2024).
Ginani, F. et al. Effect of low-level laser therapy on mesenchymal stem cell proliferation: a systematic review. Lasers Med. Sci. 30, 2189–2194 (2015).
Eroglu, B. et al. Photobiomodulation has rejuvenating effects on aged bone marrow mesenchymal stem cells. Sci. Rep. 11 (1), 13067 (2021).
Funding
This work was supported by Luzhou Southwest Medical University Cooperative Climbing Program (2021) [2021LZXNYD-D06].
Author information
Authors and Affiliations
Contributions
ZS: Conceptualization, Investigation, Writing–original draft, Writing–review and editing, Methodology, Project administration, Resources, Software, Validation, Visualization. SL: Writing–review and editing. WC: Writing–review and editing. HY: Conceptualization, Resources, Supervision, Writing–review and editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
In accordance with the “Good Clinical Practice for Pharmaceutical Trials” issued by the National Medical Products Administration of the People’s Republic of China and the ethical principles of the “Declaration of Helsinki,” our research protocol has been approved by the ethics review committee of Puyang Renji Hospital composed of 15 experts and the approval number is (PYRJYY-20220701).
Consent for publication
All authors give consent for its publication in Scientific Reports.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visithttp://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Shi, Z., Li, S., Chen, W. et al. The effect of blue and green light on human umbilical cord mesenchymal stem cells for promoting proliferation and wound healing. Sci Rep 15, 14787 (2025). https://doi.org/10.1038/s41598-025-99083-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-99083-9
Keywords
This article is cited by
-
Photobiomodulation stimulates mitochondrial function and cell proliferation in meniscus-derived stem cells (MeSCs) via activation of TRPV1 channel
Scientific Reports (2025)
-
FTIR spectroscopic characterization reveals short-term macromolecular responses to photobiomodulation in mesenchymal stem cells
Scientific Reports (2025)