Abstract
Some species of the Leucosphyrus Group of Anopheles mosquitoes in Southeast Asia are highly anthropophilic and efficient vectors of human malaria parasites, while others primarily feed on non-human primates (NHP) and transmit NHP malaria parasites. The evolutionary history of this group, particularly the origin of anthropophily, was studied using phylogenomic analysis of 2,657 high-confidence nuclear single-copy orthologous genes and 13 mitochondrial protein coding genes from 40 individuals of 11 species. Molecular dating and ancestral state reconstruction revealed that monkey-feeding is ancestral with speciation of monkey-feeding species dating to the Pliocene within Sundaland (Malay peninsula, Borneo, Sumatra and Java) which was covered in tropical rain forests during this period. Although less parsimonious alternatives cannot be excluded, molecular dating, ancestral state reconstruction and reticulation analysis indicated that anthropophily most likely evolved once, involving adaptive introgression, in the early Pleistocene in Sundaland, giving rise to multiple descendent anthropophilic species. Such early origination of anthropophily must necessarily have been in response to the arrival of early hominins (Homo erectus) rather than anatomically modern humans, likely associated with loss and fragmentation of rainforests during the early Pleistocene. The early origination of anthropophily also provides independent non-archaeological evidence supporting the limited fossil record of early hominin colonization in Southeast Asia around 1.8 Mya.
Similar content being viewed by others
Introduction
Mosquito-borne diseases present a significant burden on human health, with malaria alone causing an estimated 249 million cases and 608,000 deaths worldwide in 20221. The propensity of mosquitoes of a particular species to feed on humans (anthropophily) is the primary factor influencing their potential to spread pathogens that cause disease2,3,4,5,6,7. Although mosquitoes can be opportunistic in their host selection (e.g8,9.,, many species display varying degrees of host specificity10,11. Understanding the evolutionary origins of anthropophily and the circumstances that triggered its development can provide critical insights into mitigating the impacts of novel diseases due to mosquito-borne pathogens.
The Anopheles leucosphyrus group (hereafter, Leucosphyrus Group) comprises 20 recognized mosquito species in Southeast Asia (SE Asia)12,13,14,15. These species exhibit intrinsic differences in host preference, as demonstrated by host attraction experiments, blood-meal analysis, and variation in transmission of human and non-human primate (NHP) malarias (Supplementary Table S1)13,16,17,18,19. Notably, several species are highly anthropophilic and extremely efficient vectors of human malaria parasites. These include An. dirus, An. baimaii, and An. scanloni of the Dirus Complex found in mainland SE Asia, and An. balabacensis of the Leucosphyrus Complex from Borneo (Sabah and Kalimantan)17,20,21,22,23 (Supplementary Table S1). Conversely, species such as An. macarthuri, An. pujutensis, and An. hackeri blood-feed only in the forest canopy on NHP, including monkeys, gibbons, and orangutans, transmitting NHP malaria parasites (Fig. 1, Supplementary Table S1)24,25,26,27. Anopheles nemophilous (of the Dirus complex), An. latens, and An. introlatus (of the Leucosphyrus Complex) are less host-specific, feeding on both NHPs in the canopy and humans on the ground, apparently driven by host availability (Supplementary Table S1). As host choice experiments most often compared humans on the ground, to monkeys in the canopy, it is not possible to separate a tendency to seek hosts on the ground rather than in the canopy as a distinct trait23,28,29 (Supplementary Table SI).
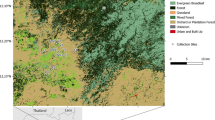
Map representing the distributions of specimens collected in Southeast Asia. Shading indicates the present-day distributions of forest types in mainland and insular Southeast Asia, adapted from Morley55. Black dots on the map represent collection sites. The number adjacent to the dots represents individual species collected from that site according to the species list on the left. The color of the numbers and species names indicate distinct blood-feeding behaviors; blue–NHP feeding, red–mixed-feeding, purple–human feeding, derived from published literature (listed in Supplementary Table S1). The brown outer line represents the outline of the exposed Sunda Shelf at the Last Glacial Maxima (currently 120 m below sea level)57. The two short brown lines represent biogeographic barriers: the Kangar-Pattani line in the south and the Isthmus of Kra in the north. The Isthmus of Kra and the W-H (Wallace-Huxley) line (grey line) in the east mark the boundaries of Sundaland. The map was created using QGIS v.3.28 (http://www.qgis.org).
The establishment of anthropophily in multiple species of the Leucosphyrus Group could be attributed to the trait evolving independently multiple times following the arrival of anatomically modern humans in SE Asia 76,000–63,000 years ago30,31. Alternatively, anthropophily may have evolved once in an ancestral species, possibly in response to the colonization of SE Asia by early hominins. Conservative estimates place Homo erectus in China at least 1.6–1.7 million years ago (Mya), and possibly as long ago as 2.4 Mya32. However, the timeline of hominin colonization southwards into SE Asia remains contentious. Recent reports suggest that hominins may have arrived in Java between 1.333 and 1.8 Mya34. Increased aridity during the Late Pliocene and Early Pleistocene, particularly during periodic glacial periods, is considered to have resulted in the formation of a north-to-south corridor of seasonal forests and grasslands35, that facilitated early hominin migration through SE Asia into Java36. We used phylogenomics and analyses of trait evolution in the Leucosphyrus Group to characterize the evolutionary history of these mosquitoes in relation to historical environmental changes and host preference. Our findings offer independent, non-archaeological evidence for the timing and location of the early hominin colonization of SE Asia, providing new perspectives on the co-evolution of mosquitoes and their hosts.
Results and discussion
Genome-scale phylogenies reveal reticulate evolution in the Leucosphyrus Group
To elucidate the evolutionary history of the Leucosphyrus Group, we sequenced 38 individual mosquitoes of 11 species: An. dirus (n = 5), An. baimaii (n = 6), An. scanloni (n = 6), An. cracens (n = 1), An. nemophilous (n = 2), An. introlatus (n = 1), An. balabacensis (n = 5), An. hackeri (n = 1), An. latens (n = 5), An. macarthuri (n = 3), and An. pujutensis (n = 3). These data were supplemented with the publicly available genomes of An. dirus37 and An. cracens38. Many of these species are particularly challenging to collect, for example, involving sampling larvae from animal wallows deep in the forest and from remote locations. Specimens of the 11 species studied here were accumulated over several years (from 1992 to 2020) and include all species of the Leucosphyrus Group from Sundaland and Indochina except, for logistical reasons, those restricted to Sumatra and the Philippines. The 11 species studied here include members of all three Subgroups (Leucosphyrus, Riparis and Hackeri) as originally proposed by Peyton and later adopted by Sallum et al.17. They also represent all three blood-feeding behaviors (human, NHP and mixed human-NHP) (Fig. 1, Supplementary Table S2). Orthology inference using An. dirus and an outgroup species, An. farauti, identified 2,657 high-confidence nuclear single-copy orthologous genes (nSCOs) across 40 genomes. Phylogenetic reconstructions were performed using coalescent-based summary analyses with ASTRAL39 and maximum-likelihood (IQ-TREE)40 analyses on 2,657 nuclear and 13 mitochondrial protein-coding DNA sequences (Fig. 2).
Genome-scale phylogenies of the Leucosphyrus Group. Trees were constructed using concatenation-based (IQ-TREE) (right) and ASTRAL (left) approach. The nuclear trees (top) were constructed using 2,657 single copy orthologs. The mitochondrial trees (bottom) were constructed using 13 protein-coding genes. Numbers on the branches denote bootstrap values. Species names are color-coded according to feeding preferences as in Fig. 1 and table S1. Based on the morphological classification by Sallum et al.17, the Leucosphyrus Group comprises the Riparis, Hackeri and Leucosphyrus Subgroups. The Leucosphyrus Subgroup is further divided into the Dirus Complex (An. baimaii, An. dirus, An. scanloni, An. cracens and An. nemophilous) and the Leucosphyrus Complex (An. balabacensis, An. latens and An. introlatus)17. The morphological classification and phylogenetic classification do not agree therefore, based on the nuclear phylogeny, we classify An. dirus, An. baimaii and An. scanloni as the Dirus Clade (D Clade), An. balabacensis and An. cracens as the Balabacensis Clade (B Clade), and An. introlatus and An. nemophilous as the Nemophilous Clade (N Clade). Different population of An. latens and An. macarthuri are denoted by: Th–Thailand, B–Borneo and PM–Peninsular Malaysia. The color of the species names indicates distinct blood-feeding behaviors; blue–NHP feeding, red–mixed-feeding, purple–human feeding.
There were some inconsistencies between the resultant nuclear and mitochondrial phylogenies (Fig. 2). Anopheles dirus and An. baimaii, though distinct from each other in the nuclear phylogenies, are indistinguishable in mitochondrial phylogenies, consistent with mitochondrial introgression from An. baimaii to An. dirus as previously reported41. Anopheles pujutensis (Hackeri Subgroup) is placed within the Dirus Complex in the mitochondrial phylogenies which is very different from its placement with other NHP-feeding species in the nuclear phylogenies. We infer this to be the result of older mitochondrial introgression from a member of the Dirus Complex into An. pujutensis (Fig. 2). Incomplete lineage sorting cannot be an explanation in either of these two cases since there is no evidence of incomplete lineage sorting in the nuclear phylogeny with all individuals within a species forming distinct clades and given that lineage sorting would be even faster for mtDNA with its lower effective population size. Anopheles cracens is also differently placed between the mitochondrial and nuclear trees (Fig. 2). Due to this mitochondrial introgression, we focused subsequent analyses on the nuclear data.
The tree topologies for each genomic region (nuclear or mitochondrial) were consistent across phylogenetic methods, except for the placement of a midpoint root among An. macarthuri, An. hackeri, and An. pujutensis, indicating variation in branch length estimation between ASTRAL and IQ-TREE methods. Discordance between coalescent-based (ASTRAL) and concatenation-based (IQ-TREE) methods is commonly observed and can be attributed to multiple causes, including differing evolutionary histories among loci, rate variation among lineages, or model misspecification42,43. To investigate these discrepancies, we used PhyloNet44 to construct phylonetworks. As the number of reticulations increased from one to three, the likelihood of the phylonetworks increased (Supplementary Fig. S1). Although more than three reticulations might be likely, further exploration was limited by computational power. All networks with 1–3 reticulations indicated either substantial introgression and/or incomplete lineage sorting of nuclear loci between lineages leading to An. pujutensis, An. hackeri, and An. macarthuri and some introgression between these lineages and more recently derived species (Supplementary Fig. S1). The introgression or incomplete lineage sorting may significantly contribute to discrepancies in the placement of the midpoint root (Fig. 2).
Given the large number of nuclear loci used, general congruence of topology between different methods of phylogenetic reconstruction, and high levels of bootstrap support for most branches, the interpretations of evolutionary history below utilise the nuclear phylogenies that we consider to best represent the species history. In making these interpretations we consider how introgression and/or incomplete lineage sorting could contribute to uncertainty.
Evolutionary timeline and geographical origin of mosquito feeding preference
Overall, the nSCO phylogenies challenge the current morphology-based taxonomic classification17. While there is support for the monophyly of the Leucosphyrus Subgroup, the phylogenies do not support the monophyly of the Dirus and Leucosphyrus Complexes within this Subgroup (Fig. 2). Despite uncertainty in the order of basal branching due to introgression, the high topological concordance of the nuclear phylogenies provides a robust phylogenetic framework to study the evolution of host preference for species of the Leucosphyrus Group.
The interpretations below depend on the estimates of divergence times, so we have taken several steps to make the dating as reliable as possible. Incomplete linage sorting in rapidly diverging species and the large effective population sizes expected of mosquitoes, could lead to over estimation of divergence time due to coalescence times being much older than species divergence times45. Further, introgression as inferred here could also introduce error into dating. To address these issues, divergence times were estimated under the multi-species coalescent46. To minimize the issues of rate heterogeneity among genes and topological conflict of gene trees with the species tree, divergence times were estimated using a subset of 25 genes that exhibited the most clock-like behaviour (had the lowest root-to-tip variance) and that had a high degree of conformity to the species tree, evaluated using SortaDate47. Use of this reduced number also served to make the analysis computationally tractable. This approach of using reduced, clock-like datasets has been widely applied in molecular dating studies (e.g48,49,50.,. An advantage of this approach is that it enables a strict molecular clock (Fig. 3) to be used for dating which gives more precise estimates than a relaxed clock (Fig. S3). For this reason, these results are presented below, but divergence dating using a relaxed-clock model with the same 25 nuclear genes generated very similar divergence estimates (Fig. S2).
Chronogram of the Leucosphyrus Group using a Bayesian approach. The divergence dating analysis was carried out using 25 clock-like genes and the coalescent species tree estimation method. The estimated divergence times are indicated by the numbers at the nodes; the blue error bars denote 95% highest posterior densities, and the numbers in parentheses are confidence intervals. The green vertical shading represents the time interval proposed for the arrival of early hominins in Southeast Asia based on fossil data between 1.333 and 1.8 million years ago34. The vertical grey line represents the time of arrival of anatomically modern humans30. The color of the species names indicates distinct blood-feeding behaviors; blue–NHP feeding, red–mixed-feeding, purple–human feeding. (see Fig. 1 and Supplementary Table S1).
Due to lack of fossil or geological calibration points, divergence dating here relies on the use of a molecular clock. As no estimates of mutation rate are available for Anopheles, we applied the Drosophila melanogaster mutation rate of 2.8 × 10⁻⁹ mutations per site per generation51. While mutation rates differ between species, they are not expected to vary substantially between Drosophila and Anopheles as these reasonably closely related Dipteran taxa share many similar life history traits (including generation times and population sizes), metabolic rates, recombination rates and genomic architecture. Potential error associated with the application of the Drosophila mutation rate to Anopheles should not be forgotten in the discussion below, but it is expected to be minimal, as reflected by the widespread use of the Drosophila mutation rate for Anopheles species52,53,54. The mutation rate is expected to be very similar to the substitution rate (the rate at which new mutations are passed on to future generations) for non-coding regions where new mutations are largely expected to be neutral. By contrast, the substitution rate in coding data is expected to be significantly lower than the mutation rate and not accounting for this would lead to underestimation of divergence times55. Thawornwattana et al.52 have demonstrated in Anopheles gambiae that divergence rate in the coding regions is 0.524X lower than in non-coding regions, so a scaling factor of 0.524 has been applied here to the mutation rate to account for the use of only coding data (Fig. 3).
To test the reliability of the nuclear molecular clock used above we also conducted divergence dating using the mitochondrial COI gene as the molecular clock rate of 2.3% divergence per million years has been well established for this gene by Brower56. Although direct comparisons are difficult due to the issue of mitochondrial introgression, the mitochondrial chronogram (Fig. S3 ) estimates divergence dates that are highly consistent with the nuclear chronogram. Notably, a key divergence date at node N0, which is associated with a transition from the ancestral state, has highly consistent dates and confidence intervals in the mitochondrial (2.98 Mya (95% CI: 3.4–2.5 Mya) (Fig. S3) and nuclear 3.1Mya (95% CI: 3.8–2.3 Mya) (Fig. 3) chronograms. The use of a carefully curated set of genes and the congruency between the mitochondrial and nuclear clocks give considerable confidence in the resulting chronogram, which was used to estimate divergence times (Fig. 3), and to reconstruct the ancestral states for host preference (Fig. 4a) and biogeographical distribution (Fig. 4b) using Reconstruction of Ancestral States (RASP) v457. Below, we integrate these lines of evidence, together with information on historical environmental change, to infer the evolutionary history of the group. The use of molecular clocks, rather than a fossil or geological calibration point, necessarily introduces some uncertainty into dating so interpretations of lineage-specific traits below are made with this caveat in mind.
Reconstruction of ancestral states for blood-feeding behavior and biogeography. (4a) Trait evolution analysis of host preference for blood-feeding. The node N0 represents a switch from feeding only on NHPs in the canopy to also feeding on humans on the ground. Node N1/N2 represents the earliest inferred switch to a preference for feeding on humans. Node N3 represents the ancestor of the Dirus Clade; N4–ancestor of the Nemophilous Clade, and N5–ancestor of the Balabacensis Clade. (4b) Reconstruction of the ancestral states for biogeography. This analysis shows dispersal at every node except for node N5 where “V” inside the pie denotes a vicariance event. The schematic maps on the right (i–iii) show dispersal within the Leucosphyrus Group for the corresponding part of the phylogeny, where (i) the orange color shows that peninsular Malaysia and Borneo is the ancestral range of monkey-feeding groups, (ii) green shows subsequent dispersal between Peninsula Malaysia, Sumatra, Borneo, and Java followed by (iii) shown in blue, dispersal of the Dirus Clade to and across the Indochinese region. Black arrows indicate direction of dispersal. The map was created using QGIS v.3.28 (http://www.qgis.org).
Ancestors of the leucosphyrus group were feeding on non-human primates in Sundaland
Extensive introgression and/or lineage sorting among An. macarthuri, An. hackeri, and An. pujutensis complicates the precise estimation of divergence times for these species. Despite uncertainty in the relationships between these three species we can infer with confidence that these NHP-feeding species are basal and diverged during the early Pliocene (5.3–3.6 Mya) (Fig. 3). Ancestral trait reconstruction for host preferences (Fig. 4a) corroborates the expectation that NHP-feeding is the ancestral state. Additionally, biogeographical reconstruction (Fig. 4b) indicates these three species originated in Sundaland (Fig. 1), encompassing present-day Borneo, peninsular Malaysia, peninsular Thailand below the Isthmus of Kra, Sumatra, and the currently submerged Sunda Shelf. During this period, the region was covered by extensive permanently humid (perhumid) rainforests58,59, providing ample opportunity for specialization in feeding on NHPs in the forest canopy.
Speciation associated with Plio-Pleistocene environmental change
The Pliocene and early Pleistocene were characterized by increasingly cooler and drier global climates60,61. It is during this period, characterized by extensive environmental change, that the Leucosphyrus Subgroup emerged, with the ancestral species An. latens diverging at node N0, around 3.1 Mya [95% CI: 3.8–2.3 Mya] (Fig. 3). Subsequently, four divergence events (nodes N1–N4, Fig. 3) occurred in the early to mid-Pleistocene (2.3–1.3 Mya) within Sundaland (Fig. 4b), with divergence at N1 giving rise to the Dirus Clade (comprising An. dirus, An. baimaii and An. scanloni) and divergence at N2 giving rise to the Nemophilous (An. nemophilous and An. introlatus) and Balabacensis (An. balabacensis and An. cracens) Clades (Figs. 2 and 3). It has long been viewed that eustatic sea-level changes were the major driver of diversification during the Pleistocene, with repeated splitting and reformation of the Sundaland landmass58,59,62. This view has changed with the recognition of subsidence of the Sunda Shelf during the Pleistocene, which means that it must have been exposed as a single landmass continuously until 400,000 years ago63. It is only after 400,000 years that Java, Sumatra, Borneo, and Indochina–Peninsular Malaysia were separated by elevated sea levels during interglacial periods (Fig. 1). Consequently, throughout the late Pliocene and most of the Pleistocene the balance of evidence favors a seasonal corridor that extended from Indochina southwards through central Sundaland, featuring more open and seasonal forests and potentially including grasslands particularly during drier glacial periods35,64,65,66,67,68,69,70. Therefore, we propose that the apparent burst of speciation occurring at nodes N0, N1–N4 within Sundaland at this time involved adaptation to novel forest types, hosts associated with these new habitats, or vicariance involving repeated fragmentation of perhumid rainforests. Since bi-directional dispersal was detected (Fig. 4b) between present-day peninsular Malaysia, Sumatra, Borneo, and Java, it indicates that there may have been limited periods of forest connectivity across Sundaland59. The recent divergence of An. cracens and An. balabacensis, which are restricted to peninsular Malaysia and Borneo, respectively, at node N5 around 0.5 Mya [95% CI: 0.7–0.4 Mya] (Figs. 3 and 4b), is attributed to vicariance by the RASP v4 analysis, likely due to the formation of separate landmasses during interglacial periods.
Crossing the barrier of the isthmus of Kra and origin of the Dirus clade
The Isthmus of Kra (ISK) marks a significant biogeographical boundary for many forest species, including birds67, due to the seasonally dry climates in lowlands further north of ISK, in Thailand59. While An. nemophilous is found both north and south of the ISK barrier, its distribution is confined to seasonal evergreen forests south of the ISK, and above the Kangar-Pattani line (K-PL) and patches found on the Thai-Cambodia border (Fig. 1)20. Speciation of the ancestor of the Dirus Clade likely involved adaptation to the seasonal forests north of ISK that enabled it to cross this biogeographical boundary. This ancestral species migrated northward, where it subsequently gave rise to at least three species (An. dirus, An. baimaii, and An. scanloni), with An. dirus and An. baimaii dispersing extensively eastwards and westwards, respectively (Fig. 4b and c(iii)), becoming major vectors of human malaria parasites across much of Indochina68.
Two-stage transition to feeding on humans in Sundaland
The shift away from strict canopy feeding on NHPs is inferred to have begun in the Late Pliocene (Fig. 3), with the emergence of the Leucosphyrus Subgroup (node N0, Figs. 3 and 4a). The late Pliocene is characterized by the transition from perhumid to seasonal and open forest types and increased savannah59,65,71. During this period a diverse assemblage of terrestrial mammals adapted to these novel habitats is likely to have inhabited forest-savannah mosaics in Sundaland based on fossil evidence from Early Pleistocene sites66. Unlike its canopy-feeding ancestors, the basal species of this Subgroup, An. latens, readily feeds on humans and other mammals on the ground as well as NHPs in the canopy24,26,65,66,67,68,69,70,71,72 (Supplementary Table S1). The increased abundance of ground dwelling host species during the late Pliocene could therefore have trigged an adaptive evolutionary innovation in host-seeking behavior of An. latens involving a willingness to seek hosts on the ground. This evolutionary transition could have acted as the bridge to human-feeding behaviour.
Evolution of vectors of malaria parasites in response to early hominin colonization of Southeast Asia
Mosquitoes employ multiple senses to track their hosts, but evolutionary changes in olfactory genes, particularly those involving the fine tuning of olfactory receptors through modification of their expression and specificity, are crucial for developing a preference for human body odor73. Large numbers of odorants and olfactory genes are involved in host specificity and genomic studies in Ae. aegypti74, An. farauti75, Culex pipiens76 and An. gambiae77 reveal that multiple genetic changes at these and other genes are required for the evolution of anthropophily, i.e. a strong, evolved preference for human blood. It is not surprising therefore that anthropophily is uncommon amongst the ~ 3500 known mosquito species78. It is therefore more parsimonious to consider that anthropophily within a taxon has a single origin. Accordingly, it is improbable that there were multiple independent switches to anthropophily in the human-preferring species of the Dirus and Balabacensis Clades, which diverged around 1.3–0.5 Mya (Fig. 3). Even taking into account some uncertainty in the molecular clock used for these divergence estimates these clades far predate the arrival of anatomically modern humans in SE Asia 76,000–63,000 years ago30,31. We therefore reject with confidence the hypothesis that anthropophily in the Leucosphyrus Subgroup evolved in response to the arrival of modern humans in SE Asia.
Using the same molecular clock rate as applied in this study and a species tree, anthropophily would be inferred to have evolved ~ 509,000–61,000 years ago in the lineage leading to the major African malaria vectors, An. gambiae and An. coluzzii52. This would date it well before the development of agriculture several thousand years ago as has been previously suggested79. Since An. gambiae originates in West African forests80 the switch to anthropophilly may instead have occurred in response to modern humans entering this forested region ~ 150,000 years ago81. The emergence of anthropophily in the domestic form of Aedes aegypti74 and the molestus ecotype of Culex pipiens7, both date to within the last 10,000 years, apparently in response to growing human populations and environmental change. Together these studies indicate that the abundance of a novel host source is a key requisite for triggering the evolution of anthropophily. The Fig. 3 indicates that anthropophily in the Leucosphyrus Subgroup emerged much earlier than in other anthropophilic mosquito species. If a strictly bifurcating tree were assumed, anthropophily could have evolved either: by the time of node N1 [95% CI: 2.9–1.8 Mya] and subsequently been lost in the lineage leading to An. nemophilous and An. introlatus; or evolved twice along the lineages leading to the Dirus Clade (N3, 1.6 Mya [95% CI: 2.0–1.2 Mya]) and the Balabacensis Clade (N5, 0.5 Mya [95% CI: 0.7–0.4 Mya]). However, the short internode distance between N1 and N2, their highly overlapping confidence intervals for divergence times, and the low level of bootstrap support for N2 in the ASTRAL phylogenetic tree (Fig. 2) indicate that this period of speciation history more likely corresponds to a phylogenetic network rather than a bifurcating tree. A network would result from introgression among incipient species and/or incomplete lineage sorting between N1 and N2. This is further supported by both the detection of reticulation from our reticulation analysis (Fig. S1) and mitochondrial introgression (Fig. 2).
Numerous genomic studies have shown the importance of introgression in promoting species adaptation to novel environments i.e. adaptive introgression82, including during speciation within the An. gambiae complex79. We therefore consider the most parsimonious argument for the evolution of anthropophily in the Leucosphyrus Subgroup to be that it evolved once only through the process of adaptive introgression at nodes N1/N2 as this accommodates the multiplicity of genes underlying the trait and negates any need to invoke loss of this trait. According to this hypothesis, anthropophily would have evolved between the extremes of the N1 and N2 confidence intervals i.e. between 2.9 and 1.6 mya (Fig. 3) when all the lineages were in Sundaland, and prior to the divergence of the Dirus Clade (node N3, 1.6 Mya [95% CI: 2.0–1.2 Mya]) further north in Indochina. This hypothesis could be tested against the above alternatives by identifying the genes underlying anthropophily and characterizing their evolutionary history.
Dating of the evolution of anthropophily in the Leucosphyrus Group to 2.9–1.6 mya overlaps with the earliest proposed date for the arrival of early hominins (Homo erectus) into Sundaland at 1.8 Mya34, but not with the more recent proposed date of 1.3 Mya33. Our findings suggest that anthropophily in the Leucosphyrus Group emerged in Sundaland in the early Pleistocene in response to the arrival of early hominins who must have not only been present in this region by this time but must have been in substantial numbers to drive adaptation to human host preference. This supports the hypothesis of Husson et al.34 that early hominins were present and abundant in Sundaland ~ 1.8 Mya, prior to their dispersal via land bridges to Java. Middle Pleistocene fossils of Homo erectus indicate their prolonged occupation on the exposed Sundaland landmass, likely associated with extensive river systems83. In the context of the very fragmentary nature of the fossil record in tropical SE Asia our findings contribute an important piece of evidence to the broader puzzle of the colonization of hominins in insular Southeast Asia.
Materials and methods
Fieldwork and sampling/Taxon sampling
Mosquito specimens (n = 38) were used to represent 11 of the 20 species in the Leucosphyrus Group, including species belonging to each Subgroup and representing all blood-feeding behaviours (Supplementary Table S2). Except for colony material of An. cracens, all the specimens were obtained as either larvae or adults from field collections between 1995 and 2020 (Supplementary Table S2). Most adult mosquitoes were collected using human landing catch, while An. pujutensis, An. macarthuri, An. hackeri, An. introlatus and one An. balabacensis and one An. baimaii were collected as larvae (Supplementary TableS2 ). The larvae were reared to adults for morphological identification to species or as belonging to the An. dirus complex using keys by Reid13 and E L Peyton17. Members of the An. dirus complex were identified to species based on ITS2 sequence data84.
DNA extraction, library Preparation and sequencing
DNA extractions of whole mosquitoes were carried out following a phenol-chloroform protocol. Total genomic DNA extraction for a few mosquito specimens was performed with DNeasy Blood and Tissue Kits (Qiagen®), digested overnight with proteinase K following the manufacturer’s protocol, and eluted with elution buffer to either 50–100 µl. Extracted genomic DNA was quantified with a Qubit 4.0 fluorometer (Invitrogen, Carlsbad, CA, USA) using the manufacturer’s protocol and 20 µl of DNA (10-100ng/µl) per sample was sequenced at the Earlham Institute (Norwich, UK). Low Input, Transposase Enabled (LITE) libraries were constructed and sequenced on two lanes of the NovaSeq 6000 SP flow cell with 150 bp paired-end reads.
Quality control of Raw reads
With a sequencing depth of ~ 30X, we were able to generate 3 million paired-end DNA sequence reads and roughly 10 GB of data per sample. To minimise sequencing errors several quality control steps were taken to filter out low-quality sequencing reads. Raw FASTQ reads were filtered using the software TrimGalore v0.6.7 developed by Babraham Bioinformatics85 by trimming adapters and low-quality bases (Phred quality ≥ 30) and cleaned sequences were reanalysed using FastQC86 and MultiQC (Babraham Bioinformatics).
Single-copy ortholog identification and filtering
The assembled and annotated proteomes of the reference species An. dirus s.s. (ENA accession-GCA_000349145.11.1) from Thailand and an outgroup species An. farauti (ENA accession-GCA_000473445.2) from Papua New Guinea were used to define groups of orthologous sequences using OrthoFinder287. Only single-copy orthologs (SCOs) that were present in all individuals and had a minimum length of 300-bp were chosen for downstream analysis, which resulted in 5,867 nuclear SCO protein-coding genes.
Assembly, alignment and filtering of nuclear SCOs
We used aTRAM 2.088, an iterative assembler that executes reference-guided local de novo assemblies, to assemble 5,867 SCOs from 38 genomes we sequenced. To do this, we used trimmed sequence reads that were first converted to a Blast database using the atram_preprocessor.py command of aTRAM v2.3.4. The amino acid sequences of 5,867 genes were used in tblastn along with the SPAdes assembler89, with five iterations, to create the aTRAM assemblies (atram.py script). The exon sequences from aTRAM assemblies were stitched together in the correct frame using the find_orthologs.py wrapper script90. Two publicly available genomes, one for An. dirus s.s. (GCA 000349145.1, Thailand)37 and one for An. cracens (GCA 002091845.1, peninsular Malaysia)38, were used in addition to the 38 novel genomes for phylogenetic analysis. From the 5,867 genes that were assembled by aTRAM we selected 2,928 that were present in all the samples for phylogenetic analysis. Each gene was aligned using the – auto flag in MAFFT91 and individual gene alignments were trimmed using the -automated1 flag in trimAl v. 1.4 0.192. Gene alignment of length > 500 bp was chosen for phylogenetic analysis. This filtering resulted in 2,657 SCOs.
Mitochondrial genome assembly and alignment
The mitochondrial genome was assembled using MITObim v1.9.1 program93. First, trimmed paired-end reads were interleaved using NGmerge v0.394. The complete mitochondrial genome from An. dirus s.s. (GenBank accession NC_036263) was used as a reference seed, and the assemblies were created with 30 iterations and --quick option using the Mitobim.pl script. The FASTA reads retrieved in the final iteration were annotated using default parameters in the MITOS web server95 (http://mitos.bioinf.uni-leipzig.de/). The GFF files generated by MITOS were imported to Geneious Prime v11.0.4. and 13 mitochondrial protein coding genes (PCGs) were extracted from mitogenomes. Two published mitogenomes of An. dirus s.s. NC_036263 (Hainan, China) and An. cracens, NC_020768 (Thailand), were also used along with 38 samples. Geneious v11.0.4’s MUSCLE aligner was used to align PCGs from 40 individuals, and trimming was performed so that all genes were the same length.
Sample validation
To validate the species identity and identify any potential contamination in our assembled sequences, we used the NCBI BLAST web interface to compare our ITS2 sequences assembled in aTRAM against the GenBank database. We also verified the COI sequences assembled using MITObim against BOLD and GenBank databases. Specifically, rDNA ITS2 was used to distinguish An. dirus from An. baimaii as they cannot be separated using mtDNA COI41. Morphological identification for An. hackeri, An. pujutensis, An. macarthuri, An. balabacensis was performed in the field by an expert taxonomist (Ralph E. Harbach).
Phylogenetic analysis
Two datasets were used to reconstruct the phylogeny of eleven species in the Leucosphyrus Group: mitochondrial PCGs and nuclear SCOs of 38 individuals and two reference genomes. Both concatenation and coalescent-based methods were used. We first concatenated all 13 PCGs in Geneious to make a bigger alignment file consisting of 9,900 bp. The nuclear dataset was made by concatenating 2,657 SCOs using the script concatenate.rb90 to make a supermatrix of 4,929,412 bp. Maximum likelihood phylogenetic reconstruction using the concatenated dataset was performed in IQ-TREE v2.1.240. We ran MODELFINDER96 in IQ-TREE with the –m MFP option to find the best model. The best-fit model: TIM2 + F + I + G4F was chosen for mitochondrial PCGs and GTR + F + R10 was chosen for the nuclear dataset according to Bayesian Information Criterion. Incorporating these models in the respective datasets, 100 ultrafast bootstraps were performed in IQ-TREE v2.1.2. For the coalescent-based tree reconstruction, we first generated gene trees for each gene using 100 rapid bootstrap replicates in RAxML v8.297 using a GTRGAMMA model. The gene trees for both the mitochondrial and nuclear datasets were then summarised using ASTRAL v5.7.8. Both the trees were visualised using iTOL98.
Divergence time Estimation
Chronograms for nSCOs were generated using StarBeast3 which infers species trees from gene trees under the multispecies coalescent (MSC) model taking into account incomplete lineage sorting and/or introgression46,99. However, due to the computational demands of the Bayesian approach, we were unable to make use of the entire nuclear dataset. We used gene selection in SortaDate47 to create a reduced dataset that could be processed with the available computing power and time. In a study by Jarvis et al.100, “clock-like” genes were found to evolve at a steady rate and reduce errors caused by model misspecification47. The SortaDate filtering procedure required two primary inputs: a species tree topology and rooted gene trees. The species tree was constructed using all the loci with RAxML97, serving as a reference for comparison. Individual gene trees were rooted using the outgroup An. macarthuri. This species was selected as the outgroup based on its position in phylogenetic trees when a much more distant outgroup, An. farauti, was used (data not shown). SortaDate evaluates each gene tree based on three criteria; (i) clock-likeness assessed using a root-to-tip variance statistic, indicating how consistent the gene’s evolutionary rate is across lineages, (ii) topological similarity/bipartition support measured by how closely the gene tree’s topology matches the provided species tree, and (iii) tree length evaluated by the total branch length of the gene tree, reflecting the amount of evolutionary information it contains. The genes were then ranked based on bipartition support, root-to-tip variance and tree length. Prioritizing bipartition support first ensures that the selected genes are topologically congruent with the species tree, minimizing topological conflicts. Following this with root-to-tip variance emphasizes the selection of genes that exhibit clock-like behaviour, which is crucial for accurate molecular dating. Tree length is considered last, and it helps to avoid overemphasis on genes with high evolutionary rates that may not be suitable for divergence-time estimation47. Using this approach we selected 25 clock-like nSCO nucleotide alignments to create a dataset for divergence dating and trait analysis.
Divergence times were estimated using the mutation rate derived from spontaneous mutations in whole genomes of Drosophila—2.8 × 10− 9 mutations per site per generation with 11 generations per year51,52,53,54. To account for only protein coding regions being used here, this mutation rate was scaled by 0.524 based on the relative mutation rate of coding to non-coding autosomal regions in Anopheles52 to yield a rate 1.47 × 10− 9 mutations per site per generation for the coding data used here.
The optimal substitution model, yielding GTR + 4 gamma count categories, was inferred using bModelTest101 in BEAST v2.6.7. Additionally,.xml configuration files with a strict clock and relaxed clock were generated using the template StarBeast3 in BEAUti v2.6.7. Two independent MCMC runs were carried out, each with a chain length of 100,000,000 and logging every 50,000 generations, until the effective sample size reached above 200. Divergence time were also estimated using mitochondrial CO1gene. The.xml configuration file was generated using relaxed clock in standard Beast template with GTR (+ I and + G) model. The MCMC runs were carried out with a chain length of 10, 000,000 and logging every 1000 generation until the effective sample size reached above 200. To determine whether each run converged, we used Tracer v1.7.2 to visually evaluate traces and effective sample sizes for posteriors, and likelihoods. We used the TreeAnnotator application in BEAST v2.6.746 to generate a maximum clade credibility tree. Trees were visualised in FigTree v1.4.4102.
Ancestral state reconstruction
To infer historical biogeography of the Leucosphyrus Group including the relative roles of vicariance and dispersal, the ancestral states were reconstructed on a phylogenetic tree with the RASP (Reconstruct Ancestral State in Phylogenies) v4 software57 using Bayesian Binary MCMC (BBM) analysis. The Bayesian tree generated for divergence dating estimation in BEAST was used for this analysis. The geographical distributions of the Leucosphyrus Group species (Supplementary Table S3 ) were based on the literature17 (Supplementary Table S1) with each species assigned to one or more of the following eight geographical areas that capture biogeographical and landmass transitions: (A) Borneo, (B) Peninsular Malaysia + south Thailand below the Kangar-Pattani Line (K-PL), (C) from K-PL northwards to the Isthmus of Kra (ISK), (D) from ISK to northwestern Thailand along its border with Myanmar, (E) remaining area of Thailand + Laos + Vietnam + Cambodia, (F) Myanmar + northeast India, (G) Java, and (H) Sumatra. RASP analysis was also performed to track evolution of the trait for host preference in the Leucosphyrus Group. For this, three categories for trait state were used: (A) non-human primate feeders, (B) mostly anthropophilic, with a third category (AB) indicating feeding on both humans and NHPs without a strong preference, based on evidence from the literature outlined in Supplementary Table S1. For both historical biogeography inference and host preference trait analysis, the MCMC chains of the BBM analysis were run for one million generations, with a sampling frequency of every 100 generations and a 10% burn-in. A fixed JC + G (Jukes-Cantor + Gamma) model was used for the BBM analysis.
Data availability
All the trimmed genome sequence data are deposited in the NCBI SRA, BioProject ID [PRJNA1148154](https:/dataview.ncbi.nlm.nih.gov/object/PRJNA1148154?reviewer=8qu6nucnqaj8ihg744ttf0b07d). All the data files, scripts, and codes used in this study are deposited in Figshare [https://figshare.com/s/e82d3503a201e019eca7](https:/figshare.com/s/e82d3503a201e019eca7).
References
World Health Organization. World malaria report 2022. https://www.who.int/teams/global-malaria-programme/reports/world-malaria-report-2022 (accessed on 8th August 2024).
Lyimo, I. N. & Ferguson, H. M. Ecological and evolutionary determinants of host species choice in mosquito vectors. Trends Parasitol. 25, 189–196 (2009).
Takken, W. & Verhulst, N. O. Host preferences of blood-feeding mosquitoes. Ann. Rev. Entomol. 58, 433–453 (2013).
Smith, D. L. et al. Ross, macdonald, and a theory for the dynamics and control of mosquito-transmitted pathogens. PLoS Pathog. 8, e1002588 (2012).
Chaves, L. F., Harrington, L. C., Keogh, C. L., Nguyen, A. M. & Kitron, U. D. Blood feeding patterns of mosquitoes: random or structured? Front. Zool. 7, 1–11 (2010).
Stone, C. & Gross, K. Evolution of host preference in anthropophilic mosquitoes. Malar. J. 17, 1–11 (2018).
Haba, Y. et al. Ancient origin of an urban underground mosquito. Science 390 (6771), eady4515 (2025).
Kilpatrick, A. M., Kramer, L. D., Jones, M. J., Marra, P. P. & Daszak, P. West nile virus epidemics in North America are driven by shifts in mosquito feeding behavior. PLoS Biol. 4, e82 (2006).
Simpson, J. E. et al. Vector host-feeding preferences drive transmission of multi-host pathogens: West nile virus as a model system. Proc. Royal Soc. B: Biol. Sci. 279, 925–933 (2012).
Yan, J., Gangoso, L., Ruiz, S., Soriguer, R. & Figuerola, J. Martínez-de La Puente, Understanding host utilization by mosquitoes: determinants, challenges and future directions. Biol. Rev. 96, 1367–1385 (2021).
Wilkerson, R. C., Linton, Y. M. & Strickman, D. Mosquitoes of the World (Johns Hopkins University, 2021).
Colless, D. H. The Anopheles leucosphyrus group. Trans. Royal Entomol. Soc. Lond. 108, 37–116 (1956).
Reid, J. A. Anopheline mosquitoes of Malaya and Borneo. Stud. Inst. Med. Res. Malaysia. 31, 520 (1968).
Hii, J. & Rueda, L. M. Malaria vectors in the greater Mekong subregion: overview of malaria vectors and remaining challenges. Southeast. Asian J. Trop. Med. Public. Health. 44, 73–165 (2013).
Sinka, M. E. et al. The dominant Anopheles vectors of human malaria in the AsiaPacific region: occurrence data, distribution maps and bionomic précis. Parasites Vectors. 4, 1–46 (2011).
Obsomer, V., Defourny, P. & Coosemans, M. The Anopheles Dirus complex: Spatial distribution and environmental drivers. Malar. J. 6, 1–16 (2007).
Sallum, M. A., Peyton, E. L., Harrison, B. A. & Wilkerson, R. C. Revision of the leucosphyrus group of Anopheles (Cellia) (Diptera, Culicidae). Revista Brasileira De Entomol. 49, 1–152 (2005).
Carnevale, P. & Manguin, S. Review of issues on residual malaria transmission. J. Infect. Dis. 223, S61–80 (2021).
Van de Straat, B. et al. Burkot. Zoonotic malaria transmission and land use change in Southeast asia: what is known about the vectors. Malar. J. 21, 109 (2022).
Baimai, V., Harbach, R. E. & Kijchalao, U. Cytogenetic evidence for a fifth species within the taxon Anopheles Dirus in Thailand. J. Am. Mosq. Control Assoc. 4, 333–338 (1988a).
Peyton, E. L. A new classification for the leucosphyrus group of Anopheles (Cellia). Mosq. Syst. 21, 197–205 (1989).
Hii, J. L. et al. Transmission dynamics and estimates of malaria vectorial capacity for Anopheles balabacensis and An. flavirostris (Diptera: Culicidae) on Banggi island, Sabah, Malaysia. Ann. Trop. Med. Parasitol. 88, 91–101 (1988).
Harbach, R. E., Baimai, V. & Sukowati, S. Some observations on sympatric populations of the malaria vectors Anopheles leucosphyrus and Anopheles balabacensis in a village-forest setting in South Kalimantan. Southeast Asian J. Trop. Med. Public Health. 18, 241–247 (1987).
Wharton, R. H. & Eyles, D. E. Anopheles hackeri, a vector of Plasmodium Knowlesi in Malaya. Science 134, 279–280 (1961).
Warren, M. & Wharton, R. H. Symposium on Simian malaria. The vectors of Simian malaria: identity, biology, and geographical distribution. J. Parasitol. 49, 892–904 (1963).
Warren, M., Cheong, W. H., Fredericks, H. K. & Coatney, G. R. Cycles of jungle malaria in West Malaysia. Am. J. Trop. Med. Hyg. 19, 383–393 (1970).
Reid, J. A. & Weitz, B. Anopheline mosquitoes as vectors of animal malaria in Malaya. Ann. Trop. Med. Parasitol. 55, 180–186 (1961).
Brant, H. L. et al. Vertical stratification of adult mosquitoes (Diptera: Culicidae) within a tropical rainforest in Sabah, Malaysia. Malar. J. 15, 1–9 (2016).
Eyles, D. E., Wharton, R. H., Cheong, W. H. & Warren, M. Studies on malaria and Anopheles balabacensis in Cambodia. Bull. World Health Organ. 30, 7–21 (1964).
Westaway, K. E. et al. An early modern human presence in Sumatra 73,000–63,000 years ago. Nature 548, 322–325 (2017).
Freidline, S. E. et al. Early presence of Homo sapiens in Southeast Asia by 86–68 Kyr at Tam Pà Ling, Northern Laos. Nat. Commun. 14, 3193 (2023).
Sawafuji, R., Tsutaya, T., Takahata, N., Pedersen, M. W. & Ishida, H. East and Southeast Asian hominin dispersal and evolution: a review. Q. Sci. Rev. 333, 108669 (2024).
Matsu’ura, S. et al. Age control of the first appearance datum for Javanese Homo erectus in the Sangiran area. Science 367, 210–214 (2020).
Husson, L. et al. Javanese Homo erectus on the move in SE Asia circa 1.8 Ma. Scientific Reports 12, 19012 (2022).
Heaney, L. R. A synopsis of Climatic and vegetational change in Southeast Asia. Trop. Forests Clim. 19, 53–61 (1991).
Roberts, P. & Amano, N. Plastic pioneers: hominin biogeography East of the Movius line during the pleistocene. Archaeol. Res. Asia. 17, 181–192 (2019).
Neafsey, D. E. et al. Highly evolvable malaria vectors: the genomes of 16 Anopheles mosquitoes. Science 346, 1258522 (2015).
Lau, Y. L. et al. Draft genomes of Anopheles cracens and Anopheles maculatus: comparison of Simian malaria and human malaria vectors in Peninsular Malaysia. PLoS One. 11, e0157893 (2016).
Mirarab, S. et al. ASTRAL: genome-scale coalescent-based species tree Estimation. Bioinformatics 30, 541–548 (2014).
Minh, B. Q. et al. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Walton, C. et al. Genetic population structure and introgression in Anopheles Dirus mosquitoes in South-east Asia. Mol. Ecol. 10, 569–580 (2001).
Shen, X. X., Steenwyk, J. L. & Rokas, A. Dissecting incongruence between concatenation- and quartet-based approaches in phylogenomic data. Syst. Biol. 70, 997–1014 (2021).
Vankan, M., Ho, S. Y. W. & Duchêne, D. A. Evolutionary rate variation among lineages in gene trees has a negative impact on species-tree inference. Syst. Biol. 71, 490–500 (2022).
Wen, D., Yu, Y., Zhu, J. & Nakhleh, L. Inferring phylogenetic networks using phylonet. Syst. Biol. 67, 735–740 (2018).
Tiley, G. P., Poelstra, J. W., Dos Reis, M., Yang, Z. & Yoder, A. D. Molecular clocks without rocks: new solutions for old problems. Trends Genet. 36 (11), 845–856 (2020).
Bouckaert, R. et al. BEAST 2.5: an advanced software platform for bayesian evolutionary analysis. PLoS Comput. Biol. 15, e1006650 (2019).
Smith, S. A., Brown, J. W. & Walker, J. F. So many genes, so little time: a practical approach to divergence-time Estimation in the genomic era. PloS One. 13, e0197433 (2018).
Zhou, B. F. et al. Phylogenomic analyses highlight innovation and introgression in the continental radiations of fagaceae across the Northern hemisphere. Nat. Commun. 13, (1), 1320 (2022).
Salles, M. M. et al. Ancient introgression explains mitochondrial genome capture and mitonuclear discordance among South American collared tropidurus lizards. Molecular Ecology, 34, e70130 (2025).
Aardema, M. L., Stiassny, M. L. & Alter, S. E. Genomic analysis of the only blind cichlid reveals extensive inactivation in eye and pigment formation genes. Genome Biol. Evol. 12, 1392–1406 (2020).
Keightley, P. D., Ness, R. W., Halligan, D. L. & Haddrill, P. R. Estimation of the spontaneous mutation rate per nucleotide site in a drosophila melanogaster full-sib family. Genetics 196, 313–320 (2014).
Thawornwattana, Y., Dalquen, D. & Yang, Z. Coalescent analysis of phylogenomic data confidently resolves the species relationships in the Anopheles Gambiae species complex. Mol. Biol. Evol. 35, 2512–2527 (2018).
Anopheles gambiae 1000 Genomes Consortium. Genetic diversity of the African malaria vector Anopheles Gambiae. Nature 552 (7683), 96 (2017).
Small, S. T. et al. Radiation with reticulation marks the origin of a major malaria vector. Proceedings of the National Academy of Sciences 117(50):31583-90 (2020).
Ho, S. Y. The changing face of the molecular evolutionary clock. Trends Ecol. Evol. 29 (9), 496–503 (2014).
Brower, A. V. Rapid morphological radiation and convergence among races of the butterfly Heliconius Erato inferred from patterns of mitochondrial DNA evolution. Proc. Natl. Acad. Sci. 91, 6491–6495 (1994).
Yu, Y., Blair, C. & He, X. RASP 4: ancestral state reconstruction tool for multiple genes and characters. Mol. Biol. Evol. 37, 604–606 (2020).
Lohman, D. J. et al. Biogeography of the Indo-Australian Archipelago. Annu. Rev. Ecol. Evol. Syst. 42, 205–226 (2011).
Morley, R. J. Assembly and division of the South and South-East Asian flora in relation to tectonics and climate change. J. Trop. Ecol. 34, 209–234 (2018).
Filippelli, G. M. & Flores, J. A. From the warm pliocene to the cold pleistocene: A Tale of two oceans. Geology 37, 959–960 (2009).
deMenocal, P. B. Climate and human evolution. Science 331, 540–542 (2011).
Voris, H. K. Maps of pleistocene sea levels in Southeast asia: shorelines, river systems and time durations. J. Biogeogr. 27, 1153–1167 (2000).
Husson, L., Boucher, F. C., Sarr, A. C., Sepulchre, P. & Cahyarini, S. Y. Evidence of sundaland’s subsidence requires revisiting its biogeography. J. Biogeogr. 47, 843–853 (2020).
Hamilton, R. et al. Forest mosaics, not savanna corridors, dominated in Southeast Asia during the last glacial maximum. Proc. Natl. Acad. Sci. U.S.A. 121, e2311280120 (2024).
Bird, M. I., Taylor, D. & Hunt, C. Palaeoenvironments of insular Southeast Asia during the last glacial period: a savanna corridor in sundaland? Q. Sci. Rev. 24, 228–2242 (2005).
Louys, J. & Meijaard, E. Palaeoecology of Southeast Asian megafauna-bearing sites from the pleistocene and a review of environmental changes in the region. J. Biogeogr. 37, 1432–1449 (2010).
Woodruff, D. S. The location of the Indochinese-Sundaic biogeographic transition in plants and birds. Nat. History Bull. Siam Soc. 51, 97–108 (2003).
Morgan, K. et al. Comparative phylogeography reveals a shared impact of pleistocene environmental change in shaping genetic diversity within nine Anopheles mosquito species across the Indo-Burma biodiversity hotspot. Mol. Ecol. 20, 4533–4549 (2011).
Colbourne, M. J., Huehne, W. H. & LaChance, F. S. The Sarawak anti-malaria project. Sarawak Museum J. 9, 215–248 (1959).
Wharton, R. H., Eyles, D. E., Warren, M. & Cheong, W. H. Studies to determine the vectors of monkey malaria in Malaya. Ann. Trop. Med. Parasitol. 58, 56–77 (1964).
van den Bergh, G. D., de Vos, J. & Sondaar, P. Y. The late quaternary palaeogeography of mammal evolution in the Indonesian Archipelago. Palaeogeogr., Palaeoclimatol. Palaeoecol. 171, 385–408 (2001).
Jeyaprakasam, N. K. et al. High transmission efficiency of the Simian malaria vectors and population expansion of their parasites Plasmodium cynomolgi and Plasmodium Inui. PLoS Negl. Trop. Dis. 17, e0011438 (2023).
McBride, C. S. Genes and odors underlying the recent evolution of mosquito preference for humans. Curr. Biol. 26, 41–46 (2016).
Rose, N. H. et al. Dating the origin and spread of specialization on human hosts in Aedes aegypti mosquitoes. Elife 12, e83524 (2023).
Ambrose, L. et al. Comparisons of chemosensory gene repertoires in human and non-human feeding Anopheles mosquitoes link olfactory genes to anthropophily. Iscience 25.7 (2022).
Bell, K. L. et al. Genetic and behavioral differences between above and below ground culex pipiens bioforms. Heredity 132 (5), 221–231 (2024).
Popkin-Hall, Z. R. & Slotman, M. A. Molecular evolution of gustatory receptors in the Anopheles Gambiae complex. BMC Ecol. Evol. 25, 22 (2025).
Hall, M. & Tamïr, D. Mosquitopia: The place of pests in a healthy world pp16-31(2022).
White B. J. Ecological Genomics of the Malaria Mosquito Anopheles Gambiae. University of Notre Dame; https://doi.org/10.7274/7p88cf97j1j(2010).
Schmidt, H. et al. Transcontinental dispersal of Anopheles gambiae occurred from West African origin via serial founder events. Communications biology 2.1, 473 : (2019).
Ben Arous, E. et al. Humans in africa’s wet tropical forests 150 thousand years ago. Nature 640, 402–407 (2025).
Horta, P. et al. Adaptive introgression as an evolutionary force: A Meta-Analysis of knowledge trends. Evol. Appl. 18.6, e70103 (2025).
Berghuis, H. W. K. et al. The late middle pleistocene homo erectus of the Madura Strait, first hominin fossils from submerged Sundaland. Quaternary Environ. Humans. 3, 100068 (2025).
Walton, C. et al. Identification of five species of the Anopheles Dirus complex from Thailand, using allele-specific polymerase chain reaction. Med. Vet. Entomol. 13 (1), 24–32 (1999).
F. Krueger, F. James, P. Ewels, E. Afyounian, B. Schuster-Boeckler, FelixKrueger/TrimGalore: v0.6.7. https://doi.org/10.5281/zenodo.5127899 (2021).
Andrews, S. S, FastQC: a quality control tool for high throughput sequence data. Available online at: (2010). http://www.bioinformatics.babraham.ac.uk/projects/fastqc
Emms, D. M. & Kelly, S. OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol. 20, 1–4 (2019).
Allen, J. M., LaFrance, R., Folk, R. A., Johnson, K. P. & Guralnick, R. P. aTRAM 2.0: an improved, flexible locus assembler for NGS data. Evolutionary Bioinf. 14, 1–4 (2018).
Bankevich, A. et al. Pyshkin, spades: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Malmstrøm, M. et al. Evolution of the immune system influences speciation rates in teleost fishes. Nat. Genet. 48, 1204–1210 (2016).
Katoh, K., Misawa, K., Kuma, I. & Miyata, T. MAFFT: a novel method for rapid multiple sequence alignment based on fast fourier transform. Nucleic Acids Res. 30, 3059–3066 (2002).
Capella-Gutiérrez, S., Silla-Martínez, J. M. & Gabaldón, T. TrimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Hahn, C., Bachmann, L. & Chevreux, B. Reconstructing mitochondrial genomes directly from genomic next-generation sequencing reads—a baiting and iterative mapping approach. Nucleic Acids Res. 41, e129 (2013).
Gaspar, J. M. NGmerge: merging paired-end reads via novel empirically derived models of sequencing errors. BMC Bioinform. 19, 1–9 (2018).
Bernt, M. et al. MITOS: improved de Novo metazoan mitochondrial genome annotation. Mol. Phylogenet. Evol. 69, 313–319 (2013).
Kalyaanamoorthy, S., Minh, B. Q., Wong, T. K., Von Haeseler, A. & Jermiin, L. S. ModelFinder: fast model selection for accurate phylogenetic estimates. Nat. Methods. 14, 587–589 (2017).
Stamatakis, A. The RAxML v8. 2. X manual. Heidleberg Inst. Theoretical Studies, 845, 1-61 (2016).
Letunic, I. & Bork, P. Interactive tree of life (iTOL) v4: recent updates and new developments. Nucleic Acids Res. 47, 256–259 (2019).
Douglas, J., Jiménez-Silva, C. L. & Bouckaert, R. StarBeast3: adaptive parallelized bayesian inference under the multispecies coalescent. Syst. Biol. 71, 901–916 (2022).
Jarvis, E. D. et al. Whole-genome analyses resolve early branches in the tree of life of modern birds. Science 346, 1320–1331 (2014).
Bouckaert, R. R. & Drummond, A. J. bModelTest: bayesian phylogenetic site model averaging and model comparison. BMC Evol. Biol. 17, 1–11 (2017).
Rambaut, A. FigTree v1.4.4, a graphical viewer of phylogenetic trees. 127–128. (2018). Available at http://tree.bio.ed.ac.uk/software/figtree/
Acknowledgements
We would like to thank the CSCMi entomology team from northeast India, Simone Nambanya from Lao PDR, and Thaung Hlaing from Myanmar for providing mosquito samples. We are grateful to Dr. Michael Matschiner from the University of Oslo for his guidance on data analysis. Additionally, we appreciate the support from Research IT and the use of the Computational Shared Facility at The University of Manchester.
Funding
Wellcome Trust grant 089229/Z/09/Z (CW). Wellcome Trust grant 097820/Z/11/A (CW). Dean’s Doctoral Scholarship from the University of Manchester (USS). U.S. National Institute of Allergy and Infectious Diseases, National Institutes of Health (NIH) Award Number U19AI089676 (JMC, AD, SA). BBSRC doctoral scholarship to Ben S. Broomfield
Author information
Authors and Affiliations
Contributions
Conceptualization: USS, CWMethodology: USS, REH, CWInvestigation: USS, CW, KM, REH, JH, MSC, PS, AP, DS, Visualization: USS, CWFunding acquisition: CW, USS, JMC, ADProject administration: USS, CWSupervision: CWWriting – original draft: USS, CWWriting – review & editing: USS, REH, JH, MSC, PS, AP, DS, BB, KM, SA, AD, YM-L, JMC, CW.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Disclaimer
The opinions or assertions contained herein are the private views of the authors, and are not to be construed as official, or as reflecting true views of the US National Institutes of Health, US Department of the Army, US Department of Defense, or the US Government. The material in this manuscript has been reviewed by the Walter Reed Army Institute of Research. There is no objection to its presentation and/or publication.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Singh, U.S., Harbach, R.E., Hii, J. et al. Early hominin arrival in Southeast Asia triggered the evolution of major human malaria vectors. Sci Rep 16, 6973 (2026). https://doi.org/10.1038/s41598-026-35456-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-35456-y