Abstract
Oligodendrocytes are the myelinating cells within the central nervous system, but the mechanisms by which transcription factors (TFs) cooperate for gene regulation in oligodendrocytes remain unclear. We introduce coTF-reg, an analytical framework that integrates scRNA-seq and scATAC-seq data to identify cooperative TFs co-regulating the target gene (TG). First, we identify co-binding TF pairs in the same oligodendrocyte-specific regulatory regions. Next, we train a deep learning model to predict each TG expression using the co-binding TFs’ expressions. Shapley interaction scores reveal high interactions between co-binding TF pairs, such as SOX10-TCF12. Validation using oligodendrocyte eQTLs and their eGenes that are regulated by these cooperative TFs show potential regulatory roles for genetic variants. Experimental validation using ChIP-seq data confirms some cooperative TF pairs, such as SOX10-OLIG2. Prediction performance of our models is evaluated through holdout data and additional datasets, and an ablation study is also conducted. The results demonstrate stable and consistent performance.
Similar content being viewed by others
Introduction
Oligodendrocytes play key functional roles in the central nervous system (CNS) function, including that they are responsible for myelination1,2. Myelination is a complex neurodevelopmental process that begins during brain development in the third trimester of pregnancy and increases steadily during childhood, but it can also be dynamically regulated in the context of learning and diseases affecting the mature CNS3,4. Also, Oligodendrocyte dysfunction and myelin abnormalities have been reported in CNS disorders2,5,6. Multidirectional interactions between neuronal and glial cells are required for CNS function7, including interactions between oligodendrocytes and neurons through myelination8. Therefore, it is critical to better understand the functions and roles of oligodendrocytes and myelin.
Gene expression of oligodendrocyte development from oligodendrocyte progenitor cells (OPC) is governed by complex gene regulatory mechanisms involving transcription factors (TFs)3,4. TFs often work in a combinatorial fashion to regulate gene expression from regulatory elements9,10. For example, some TFs such as SOX10 and OLIG2 cooperate during the induction of genes for differentiation and myelin formation11,12,13,14. Enhancers can increase transcription levels from promoters and transcription start sites (TSS), and much of the regulatory code that drives cell type-specific gene expression resides in these distal regulatory elements. Especially, some active enhancers are associated with the gene expression that characterizes cell identity and functions15. Thus, it is important to identify active oligodendrocyte-specific enhancers as well as promoters and the co-binding TFs that are responsible for their activity.
Next-generation sequencing technologies, including single-cell RNA sequencing (scRNA-seq) and the assay for transposase-accessible chromatin sequencing (scATAC-seq), have provided important insights into cell-type-specific gene regulation. Recent functional genomic resources such as PsychENCODE216 and GTEx17, and emerging tools for integrating multi-omics data enable creating cell-type-level gene regulatory networks (GRNs) linking TFs and their binding sites (TFBS), regulatory elements to target genes (TGs). Those networks can reveal the cell-type-specific regulatory roles of TFs via regulatory elements. Moreover, additional bioinformatic tools such as SCENIC+18, Signac19, and scGRNom20 predict cell-type-specific gene regulatory networks to explain potential TF-TG relationships. However, most of these studies and tools focus on relationships between individual TFs and TGs instead of TF-TF interactions and their effects on TG expression. Consequently, due to the lack of tools, the mechanistic roles of cooperative TFs in establishing cell type-specific gene regulation remain uncharacterized.
To tackle these challenges, we introduce coTF-reg, an analytical framework that integrates scRNA-seq and scATAC-seq data to identify cooperative TFs co-regulating the TG. coTF-reg identifies cooperative co-binding TFs along with active regulatory elements for gene regulation as hallmarks of active oligodendrocyte-specific regulatory elements. First, it identifies co-binding TF pairs in these regulatory regions. Second, a deep learning model is trained to predict TG expression based on the expression profiles of co-binding TFs. Third, Shapley interaction scores are computed to evaluate the interactions between TF pairs. Our findings reveal high interactions between co-binding TF pairs, such as SOX10-TCF12. Validation using oligodendrocyte eQTLs and their eGenes that are regulated by these cooperative TFs showed potential regulatory roles for genetic variants. Experimental validation using ChIP-seq data confirmed some cooperative TF pairs, such as SOX10-OLIG2 and SOX10-NKX2.2. Prediction performance of our models was evaluated through holdout data and additional datasets, and an ablation study was also conducted. The results demonstrated stable and consistent performance. Overall, our results create an analytic framework in which co-binding TF pairs cooperatively activate the TG expression through oligodendrocyte-specific regulatory elements.
Results
Deep learning and single-cell multi-omics for identifying cooperative transcription factors in oligodendrocytes
In order to predict cooperative TFs involved in oligodendrocyte gene regulation, we designed coTF-reg, which integrates scRNA-seq and scATAC-seq data to identify the cooperative TFs that co-regulate the target gene (TG) expression in oligodendrocytes (Fig.1, Methods and Materials). Briefly, we first used scATAC-seq data with peak-to-gene links21. Second, among the regulatory regions for various cell types, we focused on those specific to oligodendrocytes. We then identified transcription factor binding sites (TFBSs) and co-binding TF pairs through motif co-occurrence and co-enrichment analyses. Third, we trained deep neural networks (DNNs) to predict the expression levels of the TGs and measure interaction effects between co-binding TFs on the expression levels of TGs using gene expression from scRNA-seq data22 and computed Shapley interaction (SI)23,24 scores for co-binding TF pairs and found cooperative TF pairs. Fourth, we built a gene regulatory network based on SI scores for co-binding TF pairs. Lastly, as an independent validation, to validate the cooperative TF pairs we found, we mapped oligodendrocyte eQTLs onto the regulatory regions where cooperative TF pairs exist, performed Liftover analysis and co-enrichment analysis using ChIP-seq data, and applied Boolean rules to characterize the cooperativity of regulatory factors. To evaluate the prediction performance of our models, we used other publicly available datasets and conducted an ablation study by generating random TF sets to predict TG expressions.
Inputs for the coTF-reg pipeline are scATAC-seq peak-gene links and scRNA-seq. It infers transcription factor binding sites (TFBSs) in regulatory regions and identifies co-binding TF pairs. Then, it measures cooperativity of co-binding TFs by predicting TF-TG relationships for the levels of expression using deep learning models and Shapley interaction scores. It outputs a gene regulatory network linking co-binding TF pairs with their TGs and regulatory variants on the regulatory regions where co-TFs have their binding sites.
Identification of the co-binding transcription factors in oligodendrocyte-specific regulatory regions
First, we identified a set of 787 oligodendrocyte-differentially accessible and oligodendrocyte-specific regulatory regions by comparison of oligodendrocyte scATAC-seq data to other brain cell types. In this set, we identified 958 motifs for inferred TFBSs using the JASPAR database. Second, we used co-occurrence analysis and co-enrichment analysis to identify 8101 co-binding TF pairs out of 458,403 possible TF pairs (‘Methods and Materials: Co-enrichment analysis’ for more details). We removed TF pairs from the same families and applied a cutoff (<0.1) for false discovery rate (FDR) yielding 8101 co-binding TF pairs. There were 206 TFs that have co-binding TFs linked to 445 TGs (Supplementary Data) that are oligodendrocyte specific in 643 regulatory regions (Fig. 2a). We annotated the regulatory regions to categorize them into promoters (32.5%) and enhancers (67.5%) (Fig. 2b).
a Summary statistics of transcription factor (TF)-target gene (TG) links. b Peak annotation. c Distributions of the numbers of co-binding TFs, TGs, and peaks for individual transcription factors. d Correlations between the numbers of co-binding transcription factors and target genes, as well as the numbers of co-binding TFs and peaks. e Distribution of the numbers of co-binding TF pairs, TGs, and peaks for transcription factors by families.
The density plots show the distributions of the number of co-binding TF pairs, the number of TGs, and the number of peaks for individual TFs that are co-bound to other TFs. Most of the TFs have 50 to 103 co-binding TFs (median = 78). The distribution of the number of TGs for TFs is right-skewed, and many TFs have 76 to 172 TGs linked. The distribution of the number of peaks for TFs is also right skewed, and the most frequent intervals were between 75 and 180 peaks (Fig. 2c). The distributions of the numbers of TGs and peaks for co-binding TF pairs are approximately normal. On average, co-binding TF pairs have 60 TGs linked and 59 peaks (Supplementary Fig. 1). Additionally, other density plots show the distributions of the number of TGs and the number of peaks for co-binding TF pairs and bar plots display the numbers of co-binding TFs, TGs, and peaks for individual TFs by their family categories (Fig. 2d). Co-binding TFs have 4 to 115 TGs (median = 59) and 4 to 123 peaks (median = 56) and the most frequent motifs are associated with TF families with C2H2 zinc finger (ZF), bZip, and bHLH DNA-binding domains (Fig. 2e). C2H2 ZF proteins are a large family and C2H2 ZF TFs (e.g., ZNF2425 and KLF9/1326) are known to play significant roles in the development and function of oligodendrocytes, which are the myelinating cells of the CNS. These TFs can regulate the expression of genes essential for oligodendrocyte differentiation, survival, and myelination processes25,27,28.
We computed Pearson correlation coefficient (r) to measure correlations between the number of co-binding TFs and the number of TGs and the number of co-binding TFs and the number of peaks for individual TFs (Fig. 2d). The number of co-binding TFs and the number of TGs for individual TFs are strongly positively correlated (r = 0.70). It suggests that TFs that are co-bound to other TFs tend to have more TGs linked to them. The number of co-binding TFs and the number of peaks for individual TFs are also strongly positively correlated (r = 0.67). It shows that co-binding TFs may exist in many different peaks.
In the following sections, we incorporate RNA-seq data to explore gene expression relationships between TFs and TGs, train deep learning models to predict TG expression using co-binding TFs, and compute TF interaction scores using the trained models.
Oligodendrocyte gene expression relationships between transcription factors and target genes
A single cell study identified the unique gene expression profile of oligodendrocytes compared to other brain cell types22, as shown by the two dimensional Uniform Manifold Approximation and Projection (UMAP) space after computing latent representations of the neighborhood graph (Fig. 3a). The UMAP embeddings reveal that oligodendrocytes exhibit a distinct expression profile compared to other cell types. This separation suggests that oligodendrocytes have unique transcriptional programs that differentiate them from neighboring cell types. The distinct clustering of oligodendrocytes in the UMAP space indicates specialized functional roles and may reflect their involvement in myelination and maintenance of neural integrity. In order to focus on oligodendrocyte-specific mechanisms of gene regulation, we conducted differential expression testing using 17,946 genes and 20,191 metacells and identified 4387 differentially expressed genes (DEGs) for oligodendrocytes. We found 445 TGs out of 507 TGs of oligodendrocyte-specific regulatory elements (88%) were DEGs for oligodendrocytes. Subsequently, we conducted enrichment analysis for these 445 TGs revealing their involvement in crucial biological processes for oligodendrocytes, such as oligodendrocyte development, oligodendrocyte differentiation, and myelination (Fig. 3b).
a UMAP for eighteen cell type in middle temporal gyrus region, b Enrichments analysis for target genes that are oligodendrocyte-specific, c Pairwise two-sided t-tests for the correlation (between the expression of the TFs and their TGs) comparison for three TF groups: 10 oligodendrocyte key transcription factors, 83 oligodendrocyte-specific non-key transcription factors, and 103 non-oligodendrocyte-specific transcription factors, and d Boxplots for the expression levels of the three categories in (c) (each column is an example for each category).
We categorized TFs into oligodendrocyte key TFs, oligodendrocyte-specific non-key TFs, and non-oligodendrocyte-specific TFs using oligodendrocyte expression level and the list of key TFs (see ‘Methods and Materials: Key TFs’ for more details). ‘Oligodendrocyte-specific key TFs’ are oligodendrocyte differentially expressed TFs and key TFs, ‘oligodendrocyte-specific non-key TFs’ are oligodendrocyte differentially expressed TFs but not key TFs, and ‘non-oligodendrocyte-specific TFs’ are neither oligodendrocyte differentially expressed TFs nor key TFs. The key oligodendrocyte TFs were defined based on mouse loss-of-function studies that have shown that specific TF’s are critical for oligodendrocyte differentiation. The key TF’s include SOX1029, SOX230,31, SOX832, MYRF33, OLIG134, OLIG235, TCF7L236,37, ZNF2425, NKX2.238, and NKX6.239.
Each of 206 TF, who have co-binding TFs, regulates a different set of TGs, and we computed correlations between the expression of the TFs and their TGs in the three categories. Pairwise two-sided t-tests show that correlations between TFs and TGs in oligodendrocyte key TF pairs and those in non-oligodendrocyte-specific TF pairs are significantly different (p < 0.001). It also indicates that correlations between TFs and TGs in oligodendrocyte-specific non-key TF pairs and those in non-oligodendrocyte-specific TF pairs are significantly different (p < 0.001) (Fig. 3c). The results for differential expression testing show that the six TFs in the two categories, oligodendrocyte key TF pairs and oligodendrocyte-specific non-key TF pairs are all significant and up-regulated (Fig. S8).
We color-coded the UMAP embeddings based on the expression level of the TFs (Fig. 3a) and selected three TFs as examples for each category. Oligodendrocyte-specific key TFs such as SOX10, MYRF, and OLIG2 are specifically highly expressed in oligodendrocytes. Oligodendrocyte-enriched non-key TFs, including RBPJ, JUND, and KLF7, are expressed in multiple cell types but are more highly expressed in oligodendrocytes. Non-oligodendrocyte-specific TFs, such as RUNX1, HLF, and CREB1, are not specifically expressed in oligodendrocytes (Fig. 3d).
Deep learning and Shapley interaction scores to measure cooperativity of co-binding transcription factors
To understand the complex relationships between TFs for predicting TGs, we built deep learning models. We trained a deep learning model for each of the 445 TG. Each model used the expression levels for the 206 TFs that have co-binding TFs to predict a TG expression level. We used seven hidden layers in each DNN (Fig. 4a). We excluded co-binding TF-TG pairs that exhibited high variability in their SI scores (coefficient of variance > 0.5). Using a trained model and a hold-out test dataset, we computed SI scores for TFs in each DNN. Additionally, we determined the percentile SI score for all co-binding TF pairs. Then, a two-sided t-test to compare the mean values for the percentile SI scores of key co-binding TF pairs and non-key co-binding TF pairs revealed a significant difference between the two groups (p < 0.0001) (Fig. 4b).
a Deep learning architecture, b Percentile distribution of interaction scores for 577 key-transcription factor pairs and 7029 non-key transcription factor pairs, c Top forty-eight SI interaction scores for key transcription factor pairs, and d Top forty-eight SI interaction scores for non-key transcription factor pairs.
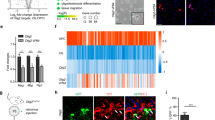
To emphasize the several important key co-binding TF pairs, we selected the top forty-eight interacting pairs for each key co-binding TF pair, such as SOX10, MYRF, OLIG1, OLIG2, NKX6.2, and TCF7L2, and generated a heatmap for their SI scores scaled from 0 to 1 (Fig. 4c). Similarly, we chose the top forty-eight interacting co-binding TF pairs for non-key TFs and created another heatmap for their SI scores scaled from 0 to 1 (Fig. 4d). We noticed that the SI scores for key-TF co-binding pairs have higher values than those for non-key co-binding TF pairs.
We also validated our model prediction performance for one TG, myelin basic protein (MBP), using additional data (Supplementary Fig. 2)40. We regressed the scaled actual values on the scaled predicted values. For our primary dataset, we obtained an R-squared of 0.81 and a r of 0.90 (Supplementary Fig. 2a). Furthermore, when analyzing another dataset, we observed an R-squared of 0.69 and a r of 0.83, affirming the predictive capability of our model architecture (Supplementary Fig. 2b).
Oligodendrocyte gene regulatory network analysis for cooperative TF pairs and transcription factor hierarchy
We chose one pair of co-binding TF with the highest interaction scores from six key co-binding TF pairs, including SOX10, MYRF, OLIG1, OLIG2, NKX6.2, and TCF7L2. We built a gene regulatory network (GRN) for these cooperative TF pairs and their TGs that are co-regulated by them (Fig. 5a). We found that a TG, CALD1, is co-regulated by three key cooperative TF pairs, SOX10-TCF12, RORA-OLIG2, and FOXP1-NKX6.2 and another TG, PPP1R16B, is co-regulated by three key cooperative TF pairs, RORA-OLIG2, FOXP1-NKX6.2, and FOXP1-OLIG1. There are other TGs, such as AMOTL2, BOK, CALD1, FA2H, and CPM, in the GRN that are co-regulated by two pairs of cooperative TFs.
a Gene regulatory network for six key cooperative transcription factor pairs with the highest interaction scores, b A plot of in-degree (I) vs out-degree (O) for the transcription factors that have I and O in the gene regulatory network, and b Transcription factor hierarchy. Each node depicts a transcription factor. In (b, c), the edges colored in orange are the top-level (master) regulators, the edges colored in yellow are the middle-level regulators, and the edges colored in white are the bottom-level regulators.
We computed in-degree and out-degree for eighteen TFs that can also be TGs at the same time since TF feed forward and feedback loops are common (Fig. 5b). Then, we conducted a TF hierarchy analysis and found eight top-level regulators (Fig. 5c), called ‘Master regulators’, including SOX10, SOX2, and SOX8, which are key TFs that are known to play critical roles in oligodendrocyte differentiation30,31,32. The other five master regulators, MEIS1, MEIS2, RBPJ, JUND, and ZNF281, are categorized as oligodendrocyte-specific non-key TFs in Fig. 3c. All TFs that are middle-level regulators and bottom-level regulators, except for MYRF, are categorized as oligodendrocyte-specific non-key TFs. MYRF is one of the key TFs which is specifically activated in myelinating oligodendrocytes. PROX1 has been identified as being important for oligodendrocyte differentiation41,42. Most of these eighteen TFs are expressed in both oligodendrocytes and OPCs (Supplementary Fig. 4). It provides evidence that oligodendrocyte differentiation is pre-set in OPCs43.
Independent validation for cooperative TFs
eQTL mapping
As an independent assessment of the regulatory regions we mapped oligodendrocyte eQTLs44 onto oligodendrocyte-specific regulatory regions to explain the causal relationships between the expression levels of the co-binding TF pairs we identified and their target genes (TGs). Using chromosome and position of eQTL SNPs (eSNPs) from oligodendrocyte eQTLs, eSNPs integrated with a total of 643 oligodendrocyte-specific regulatory regions (Fig. 2a). This integration facilitates the identification of potential regulatory connections between the eSNPs and the co-binding TFs in these regions, enhancing our understanding of how genetic variations influence the expression levels of the identified co-binding TF pairs and their corresponding TGs. Notably, it provides evidence of causation if the eQTL genes and TGs are identical where co-binding TF pairs occur, indicating that these co-binding TF pairs are co-regulating TG expressions.
First, among 4.8 million oligodendrocytes eQTLs, we filtered 2 million significant (FDR < 0.05) eQTLs. Second, we mapped these significant eSNPs onto oligodendrocyte-specific regulatory regions (Fig. 6a). In total, 383 eSNPs and 159 eGenes were mapped onto 188 regulatory regions. Among these, 373 eSNPs and 153 eGenes (and TGs) were found in 179 regulatory regions associated with key TF pairs. Enrichment analysis for TGs indicates their strong involvement in biological processes such as oligodendrocyte development, myelination, and oligodendrocyte differentiation. (Fig. 6b)
a eQTL mapping onto oligodendrocytes regulatory regions, b ChIP-seq peak for cooperative TF pairs that co-regulate MBP. The ChIPseq profile is for SOX10, and solid blocks indicate called peaks for the specified transcription factors. c Gene ontology enrichment analysis of target genes associated with oligodendrocyte eQTL’s, and d,e Heatmaps show distribution of SOX10 ChIP-seq reads centered on the previously defined OLIG2 and NKX2.2 sites.
Validation of cooperative TF pairs
The model generated from human epigenome and expression data predicted a number of enriched TF pairs within oligodendrocyte-specific TF regulatory elements. In order to test if the coordination occurs as predicted, we utilized rat oligodendrocyte ChIP-seq data that were available for selected transcription factors. One predicted pair was OLIG2/SOX10, which had previously been shown to be extensively colocalized in analyses of rat oligodendrocytes45. To visualize the preferential binding of SOX10 on a global scale, a read density plot for SOX10 ChIP-seq reads11 was generated centered on the previously defined OLIG2 peaks45 in oligodendrocytes (Fig. 6c). In line with previous analysis, the average read density of SOX10 is highly enriched over OLIG2 bound sites. A newly found pair predicted by the model was that of NKX2.2 and SOX10, and we generated a similar plot of SOX10 ChIP-seq reads over a defined set of NKX2.2 ChIP-seq peaks in oligodendrocytes46, and we found a similarly high enrichment of SOX10 binding on ~40% of NKX2.2 binding sites (Fig. 6d). An example of the colocalization is shown for the MBP gene, which MBP is a crucial TG in oligodendrocytes as a key component of the myelin sheath47,48. Expression of MBP is essential for the differentiation and maturation of oligodendrocytes49,50, and MBP maintains the structure and integrity of the myelin sheath51. As shown in Fig. 6e, there are at least 2 sites upstream of MBP where there is colocalization of SOX10 with NKX2.2 and OLIG2.
Boolean cooperativity of TF pairs
We applied a logic circuit to characterize Boolean cooperativity of TFs using Loregic52. A total of 206 TFs that form 8101 co-binding TF pairs were input. 6660 (82.2%) out of the 8101 co-binding TF pairs have consistent triplets—matching the same logic gate across all targets, demonstrating strong cooperation between the activities of the two TFs on the TGs. More than half of the TF1-TF2-TG pairs are categorized as “AND” indicating a positive correlation between TG expression and the expression of both TF1 and TF2 (Fig. S6a). We also achieved permutation scores to remove logic gates chosen by random. Still, 6092 TF pairs have consistent triplets and 64% of triplets are categorized as “AND” (Fig. S6b).
Independent validation for the prediction performance of the models
Model prediction validation and ablation study
Using Multi-omics scRNA-seq data21 from the same cells as the scATAC-seq data in the main analysis, we trained deep learning models and computed SI scores. Forty-eight SI scores for key-TF pairs in Fig. 4c were selected. The correlation between the SI scores computed from the main data and the Multi-omics data are shown in Fig. S5a. We also ran a two-sided t-test to compare the mean values for the percentile SI scores of key co-binding TF pairs and non-key co-binding TF pairs as we did for the main data (Fig. 4b). There was a significant difference between the two groups (p < 0.0001) (Fig. S5b).
Model performance was evaluated using the holdout data. Additionally, we included three more publicly available scRNA-seq datasets: Multi-omics, ROSMAP40, and Cross-disorder53, and validated the prediction performance of our model for each TG. Here, TGs were predicted using the trained models and the entire datasets. The holdout data, Multi-omics, and ROSMAP show consistently low normalized root mean squared error (NRMSE), while more than 75% of predictions in Cross-disorder also have low NRMSE. NRMSE can be compared across genes (Fig. S7a).
We also conducted an ablation study to compare the prediction performance of our models. Another dataset for 206 random TFs that are neither co-binding nor cooperative was created and their prediction performance was compared to that of 206 co-binding TFs (Fig. S7b). The model prediction performance is much better overall when 206 TFs used for predicting TGs are either co-binding, cooperative, or both.
Discussion
With resources provided by advances in single-cell sequencing, some studies54,55,56,57 have elucidated the roles of several TFs, enabling the construction of cell type-specific gene regulatory networks to explain potential TF-TG relationships using bioinformatic tools. However, most of these studies and tools primarily focus on relationships between independent TFs and TGs.
This study introduces an analytical framework, coTF-reg, which identifies co-binding TFs and their TGs in oligodendrocyte-specific regulatory regions. Deep learning models predicted TG expression levels using the expression levels of co-binding TF pairs, and we computed TF SI scores to define highly interacting co-binding TF pairs as ‘cooperative’ TFs that co-regulate TG expression levels. We found that the key co-binding TF pairs tend to highly interact with each other compared to non-key co-binding TF pairs for predicting TG expression levels. Independent validation, such as mapping eQTLs onto the regulatory regions, provides evidence for causal relationships between co-binding TF pairs and TGs. Additionally, converting these regions to the rat genome assembly coordinates and measuring the density of ChIP-seq signals for key cooperative TFs show that many of these TF pairs are enriched in the regulatory regions, indicating their collaborative role in co-regulating TG expression levels. We defined specific key TFs and examined co-binding TF pairs containing them, along with their interactions in predicting TG expression levels. We then compared these results with those of non-key TF pairs. Overall, co-binding TF pairs with known regulators of oligodendrocyte development exhibit higher SI scores, suggesting that they not only regulate TG expression individually but also cooperatively. We identified several highly cooperative TF pairs, such as SOX10 and OLIG212,58, which are already known. Additionally, we discovered previously unreported cooperative pairs, such as SOX10 and NKX2.2.
Our study demonstrates several strengths. First, we concentrate on interactions between co-binding TF pairs and their impact on TG expression using deep learning approaches. Deep learning can elucidate complex TF relationships and their effects on TG expression levels. Second, the coTF-reg pipeline can be used by general users with any scATAC-seq and scRNA-seq data. The code for coTF-reg is openly available on GitHub, allowing users to input their scATAC-seq and scRNA-seq data for specific purposes. Third, we provide a comprehensive analytical framework that incorporates analyses utilizing co-bindings by motif and expression levels. We define ‘cooperative’ TF pairs as TF pairs significantly co-enriched across regulatory regions, exhibiting high SI scores in terms of expression when predicting TG expression. The term cooperativity has often been applied to co-bindings of TFs to nearby sites that facilitates stabilized binding due to protein-protein interactions, but in our model, we use TF pairs that can bind to sites in the same regulatory regions, since TF’s can coordinately activate enhancers without direct interactions.
Nevertheless, there are some limitations to our study. To begin with, it’s important to note that more than two TFs can co-regulate TG expression59,60. However, our current tool is limited to analyzing interactions between two co-binding TFs. In future research, developing or applying more sophisticated methods capable of handling clusters of TFs that co-regulate the same TG expression will be informative. Moreover, our method for identifying binding sites relies on the position frequency matrices in the motif database. While both SOX10 and MYRF are key TFs for oligodendrocytes, we encountered difficulty in obtaining sufficient binding sites for MYRF. Consequently, we had to supplement with a different motif for MYRF based on our prior knowledge. More generally, the definition of TF motifs relies on disparate methods, and limitations of motif generation and analysis have been noted previously. Nonetheless, our analysis provided TF-TF coordination that we could validate using data from previous studies. We predict that future analysis can be used to determine if the predicted TF pairing plays a role in oligodendrocyte differentiation, since reliance on single factor studies is not able to recapitulate the important combinatorial functions of TF’s in generating cell type-specific gene expression patterns. Lastly, there can be alternative methods for establishing cooperative relationships between TFs, such as Boolean rules61,62,63,64,65. Logic-based models are also powerful tools for understanding the complex interactions among regulatory TFs in gene regulation. Developing new tools that incorporate Boolean rules and machine learning approaches will help us effectively infer more intricate TF relationships, paving the way for future research aimed at unraveling the complexities of gene regulation.
Methods
coTF-reg pipeline workflow
First, published scATAC-seq data with peak-to-gene links21 is inputted into the coTF-reg pipeline. Second, transcription factor binding sites (TFBSs) and co-binding TF pairs in the oligodendrocyte regulatory regions are identified through motif co-occurrence and co-enrichment analyses. Third, deep neural networks (DNNs) to predict the expression levels of the TGs are trained and the interaction effects between co-binding TFs on the expression levels of TGs using gene expression from scRNA-seq data22 are measured by computing Shapley interaction (SI)23,24 scores. Fourth, a gene regulatory network is built based on SI scores for co-binding TF pairs. Fifth, a TF hierarchy analysis is used to define TFs as regulators in three categories. Lastly, as an independent validation, to validate the cooperative TF pairs: 1. The oligodendrocyte eQTLs are mapped onto the regulatory regions where cooperative TF pairs exist, 2. Liftover analysis and co-enrichment analysis using ChIP-seq data are conducted, 3. Boolean rules are applied to characterize the cooperativity of regulatory factors. To evaluate the prediction performance of our models: 1. Other publicly available datasets are used as validation data to predict TG expressions, 2. Ablation study is implemented by generating random TF sets to predict TG expressions.
Step 1: Infer transcription factor binding sites
We inferred transcription factor binding sites (TFBSs) in 787 scATAC-seq peak regions that have linkages with TGs.
-
a)
The R package GenomicRanges was used to format the ATAC-seq peaks into genomic ranges.
-
b)
Position frequency matrices (PFMs) for the 949 motifs in JASPAR2022 database66 were set in R, along with nine additional PFMs for the important modified motifs based on our prior knowledge.
-
c)
TFBSs in the scATAC-seq peak regions were inferred using a R package, motifmatchr67.
Step 2: Identify co-binding transcription factor pairs
We identified co-binding TF pairs using the inferred TFBSs in Step 1.
-
a)
All possible TF-TF pairs with binding sites in the scATAC-seq peak regions were considered.
-
b)
TF pairs from the same families were excluded.
-
c)
Co-enrichment analysis: Co-occurrence analysis was conducted to find TF pairs that have overlapping regions. We then conducted hypergeometric tests to find significantly enriched TF pairs in the same regions. We used multiple testing corrections via FDR and applied FDR < 0.1 cutoff. We define the TF pairs that are co-enriched (FDR < 0.1) as ‘co-binding’ TF pairs.
-
d)
Gene regulatory networks (GRNs) were constructed for TG-co-TF pair-peak links and matched TGs and co-TF pairs to the scRNA-seq data.
-
e)
Lowly expressed TGs and TFs were removed from the GRNs by applying a cutoff, median expression level > 1; more than half of the cells are expressed, from the GRNs.
-
f)
Differential expression testing was implemented using Seurat68 and selected TGs that are oligodendrocyte specific in the GRNs.
-
g)
Peaks were annotated as promoters or enhancers using annotatr69.
Step 3: Measure cooperativity of co-binding transcription factors
Gene expression levels of the co-binding TF pairs from scRNA-seq data were incorporated into deep learning models to predict the expression levels of the TGs and measure interaction effects between co-binding TFs on the expression levels of TGs using Shapley interaction (SI) scores.
-
a)
Metacells for the cells in scRNA-seq data were projected using a Python package, metacells70.
-
b)
Expression levels of TFs that have co-binding TFs and TGs were used to construct deep learning models for each TG using PyTorch71 in Python.
-
c)
SI scores for TF pairs were computed in each deep learning model.
-
d)
Interaction matrices for the SI scores were generated in deep learning models and the mean interaction scores for co-binding TF pairs were calculated.
-
e)
Coefficients of variation (CV)72 of the interaction scores for each co-binding TF pair were computed and the pairs with CV values higher than 0.5 were removed.
Step 4: Gene regulatory network and TF hierarchy analysis
A gene regulatory network was built for six key cooperative TFs.
-
a)
One cooperative TF pair for each of the six key TFs was selected based on the top interaction scores.
-
b)
A gene regulatory network was bulit linking cooperative TF pairs to TGs.
-
c)
TGs co-regulated by cooperative TF pairs were selected.
-
d)
A network plot was generated using Cytoscape73.
Step 5: TF hierarchy analysis
TFs that can be TGs were chosen, and we implemented hierarchy analysis74 for those TFs.
-
a)
In-degree (I) and out-degree (O) for the TFs were calculated.
-
b)
Hierarchy height metrics for the TFs were computed.
-
c)
TFs were classified as top-regulator, middle-regulator, or bottom-regulator.
Step 6: Independent validation
We implemented eQTL mapping, ChIP-seq enrichment analysis, and Boolean cooperativity analysis for validating cooperative TF pairs and model prediction validation and an ablation study for validating the prediction performance of the models.
Validation of cooperative TF pairs
eQTL mapping
We mapped the significant (FDR < 0.05) oligodendrocyte eQTLs onto the scATAC-seq peak regions.
-
a)
Publicly available oligodendrocyte eQTL data44 were downloaded and the significant (FDR < 0.05) eQTLs were extracted.
-
b)
The significant eQTLs were mapped to the scATAC-seq peak regions in the GRNs.
-
c)
The results were verified by comparing the number of eQTLs mapped onto the peak regions for key-TF pairs and non-key-TF pairs.
ChIP-seq enrichment analysis
We performed the LiftOver analysis to convert genome coordinates for rat to human hg38 assembly using UCSC Genome Browser75.
-
a)
Genome coordinates for human hg38 assembly were converted to the rn5 rat genome coordinates for human (hg19) assembly.
-
b)
Overlapping genome coordinates between conserved (from hg38 to rn5) assembly and the regulatory regions in the GRN were identified.
-
c)
Cooperative TF pairs in the overlapping regions identified, along with the TGs they co-regulate.
Using the results from the LiftOver analysis, we tried to find signals in co-enriched binding sites for cooperative key TF pairs in rat oligodendrocyte ChIP-seq data. Heatmaps were created via EAseq76. ChIP-seq tracks were visualized using UCSC genome browser. Previous ChIP-seq datasets for SOX10, OLIG2, and NKX2.2 are available at GEO accession numbers: GSE64703, GSE42447 and GSM1906296.
Boolean cooperativity of TF pairs
We applied a logic circuit to characterize Boolean cooperativity of TFs using Loregic52. Loregic is a computational tool, integrating gene expression and regulatory network, to characterize the cooperativity of regulatory factors. It uses 16 possible two-input-one-output logic gates (e.g. AND) to describe triplets of two factors regulating a common target. The GRN was inputted including co-binding TFs-TG links. Then, we binarized the gene expression levels to Boolean values 1 and 0 to represent high and low gene expression, respectively, using BoolNet77. BoolNet assigned Boolean values to expression data on the basis of modular co-expression patterns by K-means clustering across inputted samples and therefore accounts for differences in the dynamic ranges of expression among genes in the input data. The triplet gene expression data was extracted and matched to all possible logic gates. We selected consistent logic gates. We also ran 100 permutation tests to find significant logic gates.
Validation of the prediction performance
Model prediction validation
To verify the performance of deep learning model architectures, we trained a deep learning model for predicting a TG, MBP using another data40. The trained model was used to predict the expression level of MBP and compared the results with the model for MBP using the main data.
Using Multi-omics scRNA-seq data21 from the same cells as the scATAC-seq data in the main analysis, we trained deep learning models and computed SI scores, following the same processes we did in coTF-reg pipeline for identifying cooperative TFs in oligodendrocyte gene regulation (‘Step 2 Measure cooperativity of co-binding TFs’) for the main scRNA-seq data.
Model performance was evaluated using the SEA-AD22 holdout data. We also include three more publicly available scRNA-seq datasets: Multi-omics21, ROSMAP40, and Cross-disorder53, and validate the prediction performance of our model for each TG. Here, TGs were predicted using the trained models and the entire datasets. Normalized root mean squared error (NRMSE) is used to compare the performance across different datasets.
Ablation study
It is important to assess whether the 206 co-binding TFs effectively predict their TGs. Another dataset with 206 random TFs that are neither co-binding nor cooperative was generated to evaluate the prediction performance of our models. We used our trained models to predict holdout data for random TFs and compared their prediction performance to that of 206 co-binding TFs.
Single-cell ATAC-seq data
Chromatin accessibility data21 was used for the main analyses. Brain samples were selected and eight thousand nuclei from each sample were subjected to the Chromium Next GEM Single-Cell Multiome ATAC-seq. We filtered oligodendrocyte-specific peak-gene links for our analyses. 930 peaks and 606 genes were initially chosen.
Single-cell RNA-seq data
SEA-AD (Main analysis)
The data for the whole taxonomy collected from dorsolateral prefrontal cortex (1,395,601 cells) were downloaded through the Open Data Registry on AWS as AnnData objects (h5ad format)22. The cells for disease were excluded and only the controls were retained. Then, we projected metacells for the whole taxonomy and found 2004 metacells and 17,946 genes for oligodendrocytes.
Multi-omics
The normalized and quality controlled data was gained from the CELLxGENE (RRID:SCR_021059) portal. Brain samples were selected and eight thousand nuclei from each sample were subjected to the Gene Expression protocol (10x Genomics). We filtered 5459 cells for oligodendrocyte.
ROSMAP
The processed count matrix for oligodendrocyte was downloaded from a supplementary website for ‘Single-cell atlas reveals correlates of high cognitive function, dementia, and resilience to Alzheimer’s disease pathology’40. We projected metacells for the controls only and found 7072 metacells and 16,707 genes.
Cross-disorder
Post quality control filtered data was obtained from the CELLxGENE portal. We projected metacells for oligodendrocyte controls and found 1004 metacells and 21,248 genes for oligodendrocytes.
Uniform manifold approximation and projection for dimension reduction
We gained scRNA-seq data for the whole taxonomy collected from dorsolateral prefrontal cortex through the Open Data Registry on AWS as AnnData objects (h5ad format)22. There were 1,395,601 cells across 18 sub-cell types. A total of 18,431 hg38 protein-coding genes, obtained via BioMart78, were selected from 36,517 genes. We normalized the data to a depth of 10,000 and log1 transformed it using Scanpy79 in Python. Then, the highly variable genes (HVGs) were identified using dispersion-based methods80 to normalize dispersion, obtained by scaling with the mean and standard deviation of the dispersions for genes falling into a given bin for mean expression of genes. The cutoffs for the mean dispersions for genes were a minimum of 0.0125 and a maximum of 3, and for the minimum dispersion was 0.5. We identified 3032 HVGs and scaled each gene to unit variance to clip values exceeding standard deviation of 10. To reduce the dimensionality of the data, we ran principal component analysi and used top 30 PCs to compute the neighborhood graph of the cells. Finally, we embedded the neighborhood graph with 20 neighbors in two dimensions using Uniform Manifold Approximation and Projection for Dimension Reduction (UMAP)81.
Differential expression testing
We inputted metacells for all cell types to identify oligodendrocyte-specific genes using Seurat v4 in R. We used the Poisson likelihood ratio test in FindMarkers function assuming that gene expression follows the negative binomial distribution. Oligodendrocytes and oligodendrocyte precursor cells (OPCs), and astrocytes that are known as major cell types among glia in the CNS were grouped and the other fifteen cell types were compared. We used a cutoff, FDR (<0.05) to select differentially expressed genes in the oligodendrocyte group.
Position Frequency Matrices
Position frequency matrices (PFMs) for the 949 motifs in JASPAR2022 were used to infer TF binding sites. We added PFMs for MYRF, SP7, and OLIG2 that are one of the key TFs from another study82, Mus musculus in JASPAR202266, and HOCOMOCO v1283, respectively. We also included shorter motifs for other key TFs, such as SOX10, MYRF, ZNF24, NKX2.2, and SP7, considering their importance in oligodendrocytes (Supplementary Fig. 3).
Co-enrichment analysis
We used a hypergeometric test to assess whether a number of overlaps in the binding sites for two TFs follows a hypergeometric distribution. Specifically, given that a random variable \(X\) represents the possible outcomes of a hypergeometric process, the probability of getting k or more overlapping binding sites between two TFs inside a particular chosen set, as a hypergeometric random process, is
where \({N}\) is the total number of transcription binding sites for all TFs, \(m\) is the number of binding sites for TF1, n is the number of binding sites for TF2, and \(x\) is the number of overlapping binding (co-occurrence) sites between TF1 and TF2. We applied an FDR adjusted p-value as a cutoff (<0.1) for all possible TF pairs and chose co-binding TF pairs.
Key transcription factors
We defined ten key TFs that are oligodendrocyte marker genes based on mouse loss-of-function studies that have shown that specific TF’s are critical for oligodendrocyte differentiation. This includes SOX1029, SOX230,31, SOX832, MYRF33, OLIG134, OLIG235, TCF7L236,37, ZNF2425, NKX2.238, and NKX6.239 were chosen as key TFs. Ten ‘Oligodendrocyte-specific key TFs’ are oligodendrocyte differentially expressed TFs and key TFs, eighty-three ‘oligodendrocyte-specific non-key TFs’ are oligodendrocyte differentially expressed TFs but not key TFs, and a hundred-thirteen ‘non-oligodendrocyte-specific TFs’ are neither oligodendrocyte differentially expressed TFs nor key TFs. Especially, ten ‘Oligodendrocyte-specific key TFs’ play crucial roles in the development and differentiation of oligodendrocytes. They regulate various stages of oligodendrocyte maturation and promote the expression of myelin genes; essentially, they are key players in the process of myelination within the CNS.
Deep learning models
We inputted expression levels of TFs that have co-binding pairs into the deep neural network (DNN) models to predict TG expression levels. 2004 metacells (samples), 206 TFs (features), and a TG expression level (label) were used in the DNN models. A DNN for each TG was built to predict a TG expression level. The mean squared error (MSE) between predicted TG expression and actual TG expression was used as the loss function in DNN models. We cross-validated the training dataset (80% of the input samples) with 5-fold cross-validation and validated the best trained model on the 20% of hold-out validation dataset for the best use of data and to achieve reliable model performance. We used an early stopping function with patience 10 and determined the number of epochs and we set the batch size to 32. Adam with a learning rate 0.001 was used for training the models. The structure of our neural network model can be written as
where \(X\) denotes the input data and \(f\) represents the activation function, specifically the LeakyReLU function. TF expression levels serve as the input data, while Zi represents the output of the ith hidden layer. The final output of the model is the predicted TG expression level, and \({W}_{i}\) and \({b}_{i}\) are the weight matrix and bias vector for the \(i\)th layer, respectively.
To evaluate the performance of our neural network model, we utilize the Mean Squared Error (MSE) loss function. The MSE quantifies the average squared difference between the predicted outputs of the model, Z and the true labels in our regression task. Mathematically, we can express the MSE as follows:
where \({Z}_{i}\) represents the predicted output for the ith sample, and \({Y}_{{{true}}_{i}}\) denotes the true label corresponding to the ith sample.
Shapley interaction scores
We denote the set of all TFs by F, a feature i∈F, and a feature set S⊆F. We define the interaction effect between TF i and j, with feature set S, of a neural network f at a data point \({X}_{k}\) to be
where \(f\)\(({X}_{k}{;S})\) is the prediction at \({X}_{k}\) when only TFs in S are used, which often requires retraining the NN multiple times. A common approximation is to replace the absent features (i.e., F\S) by the corresponding values in a baseline CF\S, such that
The baseline is set as the empirical mean of each feature. The Shapley interaction score \({{SI}}_{{ij}}^{f}({X}_{k})\) is the expectation of \({\delta }_{{ij}}^{f}({X}_{k}{;S})\),
over a uniformly random chosen feature set \(S\) from \(F\). We use Monte-Carlo procedure84 to approximate \({{SI}}_{{ij}}^{f}({X}_{k})\) by a small number of samples of \(S\). To aggregate the local interaction effect at different data points into a global interaction effect, we take the expectation \(\left|{{SI}}_{{ij}}^{f}({X}_{k})\,\right|\) of w.r.t. the empirical data distribution \(p(X)\), such that
For our deep ensemble of deep learning models, we utilize a posterior distribution of functions \(q( \, f)\) induced by the ensemble distribution of the weights \(q(w)\), as outlined in Eq. (2). This ensemble approach involves training multiple instances of the model, each initialized with different random weights to promote diverse learning paths.
The weights \(w\) are drawn from a Gaussian prior, reflecting our initial uncertainty about their values. After training, we apply Bayesian inference techniques to update our beliefs about these weights and compute the posterior distribution \(q(w)\). This posterior captures the uncertainty in the model parameters, providing a more comprehensive understanding of the model’s behavior.
The function \(q( \, f)\) represents the expected output of the model across this ensemble of weights. To compute the interaction score, we take the expectation of the interaction score \({{SI}}_{{ij}}\) with respect to \(q(f)\). This is estimated by averaging \({N}_{f}\) samples drawn from the ensemble:
We compute Shapley interaction scores23,24 for the co-binding TF pairs, TF \(i\) and TF \(j\) using the trained DNN models and validation datasets. We calculate mean values for co-binding TF pairs using interaction matrices. We rank them by percentile and scaled them to 0 and 1 for easier interpretation.
Coefficient of variance
The coefficient of variation (CV) is a statistical measure of the dispersion of data points in a data series around the mean. The CV represents the ratio of the standard deviation to the mean, and it is a useful statistic for comparing the degree of variation from one data series to another, even if the means are drastically different from one another. The CV is defined as the ratio of standard deviation to the mean as follows:
Hierarchy analysis
We computed connectivity statistics, out-degree (O) and in-degree (I), for individual TFs to get a ‘hierarchy height’ metric (h), a normalized value of the difference between O and I for each TF. The \(h\) is calculated as
We defined TFs as top-regulator (h > 0.33), middle-regulator (−0.33 < h < 0.33), and bottom-regulator (h < -0.33) by their h values.
Statistics and reproducibility
Data manipulation and analyses were performed using Python 3.10.14 and R 4.3.1. All relevant information including the sample sizes in the groups for statistical tests are included in the figure legends. The plots in this study are generated by Scanpy79 (v1.10.3), and seaborn (v0.13.2) in Python and ggplot2 (v3.5.1) in R.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data supporting the results are included in Supplementary Data 1~5 and are publicly available on GitHub (https://github.com/daifengwanglab/coTF-reg). For the main analyses, scATAC-seq data were obtained from Supplementary Materials of the Multi-omics study21 and scRNA-seq data was sourced from SEA-AD: Seattle Alzheimer’s Disease Brain Cell Atlas (https://cellxgene.cziscience.com/collections/1ca90a2d-2943-483d-b678-b809bf464c30). For the independent validation, scRNA-seq data were acquired from the following websites: (1) Multiome (https://cellxgene.cziscience.com/collections/ceb895f4-ff9f-403a-b7c3-187a9657ac2c); (2) ROSMAP (https://compbio.mit.edu/ad_aging_brain/#loading-the-raw-data); (3) Cross-disorder (https://cellxgene.cziscience.com/collections/c53573b2-eff4-4c5e-9ad0-b24d422dfd9b).
Code availability
The code for the analyses and figures is available at https://github.com/daifengwanglab/coTF-reg.
References
Bercury, K. K. & Macklin, W. B. Dynamics and mechanisms of CNS myelination. Dev. Cell 32, 447–458 (2015).
Nasrabady, S. E., Rizvi, B., Goldman, J. E. & Brickman, A. M. White matter changes in Alzheimer’s disease: a focus on myelin and oligodendrocytes. Acta Neuropathol. Commun. 6, 22 (2018).
Singh, D. K., Ling, E. & Kaur, C. Hypoxia and myelination deficits in the developing brain. Int. J. Dev. Neurosci. 70, 3–11 (2018).
Emery, B. & Lu, Q. R. Transcriptional and epigenetic regulation of oligodendrocyte development and myelination in the central nervous system. Cold Spring Harb. Perspect. Biol. 7, a020461 (2015).
Valdés-Tovar, M. et al. Insights into myelin dysfunction in schizophrenia and bipolar disorder. WJP 12, 264–285 (2022).
Maitre, M. et al. Myelin in Alzheimer’s disease: culprit or bystander? Acta Neuropathol. Commun. 11, 56 (2023).
Quan, L., Uyeda, A. & Muramatsu, R. Central nervous system regeneration: the roles of glial cells in the potential molecular mechanism underlying remyelination. Inflamm. Regen. 42, 7 (2022).
Simons, M. & Nave, K.-A. Oligodendrocytes: myelination and axonal support. Cold Spring Harb. Perspect. Biol. 8, a020479 (2016).
Elbaz, B. & Popko, B. Molecular control of oligodendrocyte development. Trends Neurosci. 42, 263–277 (2019).
Ibarra, I. L. et al. Mechanistic insights into transcription factor cooperativity and its impact on protein-phenotype interactions. Nat. Commun. 11, 124 (2020).
Lopez-Anido, C. et al. Differential Sox10 genomic occupancy in myelinating glia. Glia 63, 1897–1914 (2015).
Sock, E. & Wegner, M. Using the lineage determinants Olig2 and Sox10 to explore transcriptional regulation of oligodendrocyte development. Dev. Neurobiol. 81, 892–901 (2021).
Bujalka, H. et al. MYRF is a membrane-associated transcription factor that autoproteolytically cleaves to directly activate myelin genes. PLoS Biol. 11, e1001625 (2013).
Weider, M. et al. Nfat/calcineurin signaling promotes oligodendrocyte differentiation and myelination by transcription factor network tuning. Nat. Commun. 9, 899 (2018).
Heinz, S., Romanoski, C. E., Benner, C. & Glass, C. K. The selection and function of cell type-specific enhancers. Nat. Rev. Mol. Cell Biol. 16, 144–154 (2015).
Emani, P. S. et al. Single-cell genomics and regulatory networks for 388 human brains. Science 384, eadi5199 (2024).
Lonsdale, J. et al. The Genotype-Tissue Expression (GTEx) project. Nat. Genet 45, 580–585 (2013).
Bravo González-Blas, C. et al. SCENIC+: single-cell multiomic inference of enhancers and gene regulatory networks. Nat. Methods 20, 1355–1367 (2023).
Stuart, T., Srivastava, A., Madad, S., Lareau, C. A. & Satija, R. Single-cell chromatin state analysis with Signac. Nat. Methods 18, 1333–1341 (2021).
Jin, T. et al. scGRNom: a computational pipeline of integrative multi-omics analyses for predicting cell-type disease genes and regulatory networks. Genome Med. 13, 95 (2021).
Zhu, K. et al. Multi-omic profiling of the developing human cerebral cortex at the single-cell level. Sci. Adv. 9, eadg3754 (2023).
Gabitto, M. I., Travaglini, K. J., Rachleff, V. M. et al. Integrated multimodal cell atlas of Alzheimer’s disease. Nat. Neurosci. 27, 2366–2383 (2024).
Cui, T. et al. Gene–gene interaction detection with deep learning. Commun. Biol. 5, 1238 (2022).
Dhamdhere, K., Agarwal, A. & Sundararajan, M. The Shapley Taylor Interaction Index. https://doi.org/10.48550/ARXIV.1902.05622. (2019).
Elbaz, B. et al. Phosphorylation sState of ZFP24 controls oligodendrocyte differentiation. Cell Rep. 23, 2254–2263 (2018).
Bernhardt, C. et al. KLF9 and KLF13 transcription factors boost myelin gene expression in oligodendrocytes as partners of SOX10 and MYRF. Nucleic Acids Res. 50, 11509–11528 (2022).
Howng, S. Y. B. et al. ZFP191 is required by oligodendrocytes for CNS myelination. Genes Dev. 24, 301–311 (2010).
Al-Naama, N., Mackeh, R. & Kino, T. C2H2-Type Zinc finger proteins in brain development, neurodevelopmental, and other neuropsychiatric disorders: systematic literature-based analysis. Front. Neurol. 11, 32 (2020).
Aprato, J. et al. Myrf guides target gene selection of transcription factor Sox10 during oligodendroglial development. Nucleic Acids Res. 48, 1254–1270 (2020).
Zhao, C. et al. Sox2 sustains recruitment of oligodendrocyte progenitor cells following CNS demyelination and primes them for differentiation during remyelination. J. Neurosci. 35, 11482–11499 (2015).
Zhang, S. et al. Sox2 is essential for oligodendroglial proliferation and differentiation during postnatal brain myelination and CNS remyelination. J. Neurosci. 38, 1802–1820 (2018).
Turnescu, T. et al. Sox8 and Sox10 jointly maintain myelin gene expression in oligodendrocytes. Glia 66, 279–294 (2018).
Garnai, S. J. et al. Variants in myelin regulatory factor (MYRF) cause autosomal dominant and syndromic nanophthalmos in humans and retinal degeneration in mice. PLoS Genet. 15, e1008130 (2019).
Chen, Y. et al. The oligodendrocyte-specific G protein–coupled receptor GPR17 is a cell-intrinsic timer of myelination. Nat. Neurosci. 12, 1398–1406 (2009).
Wang, J. et al. Olig2 ablation in immature oligodendrocytes does not enhance CNS myelination and remyelination. J. Neurosci. 42, 8542–8555 (2022).
Zhao, C. et al. Dual regulatory switch through interactions of Tcf7l2/Tcf4 with stage-specific partners propels oligodendroglial maturation. Nat. Commun. 7, 10883 (2016).
Ye, F. et al. HDAC1 and HDAC2 regulate oligodendrocyte differentiation by disrupting the beta-catenin-TCF interaction. Nat. Neurosci. 12, 829–838 (2009).
Qi, Y. et al. Control of oligodendrocyte differentiation by the Nkx2.2 homeodomain transcription factor. Development 128, 2723–2733 (2001).
Southwood, C. et al. CNS myelin paranodes require Nkx6-2 homeoprotein transcriptional activity for normal structure. J. Neurosci. 24, 11215–11225 (2004).
Mathys, H. et al. Single-cell atlas reveals correlates of high cognitive function, dementia, and resilience to Alzheimer’s disease pathology. Cell 186, 4365–4385.e27 (2023).
Bunk, E. C. et al. Prox1 is required for oligodendrocyte cell identity in adult neural stem cells of the subventricular zone. Stem Cells 34, 2115–2129 (2016).
Kato, K. et al. Prox1 inhibits proliferation and is required for differentiation of the oligodendrocyte cell lineage in the mouse. PLoS ONE10, e0145334 (2015).
Suzuki, N. et al. Differentiation of oligodendrocyte precursor cells from Sox10-venus mice to oligodendrocytes and astrocytes. Sci. Rep. 7, 14133 (2017).
Bryois, J. et al. Cell-type-specific cis-eQTLs in eight human brain cell types identify novel risk genes for psychiatric and neurological disorders. Nat. Neurosci. 25, 1104–1112 (2022).
Yu, Y. et al. Olig2 targets chromatin remodelers to enhancers to initiate oligodendrocyte differentiation. Cell 152, 248–261 (2013).
Aguado, L. C. et al. microRNA function is limited to cytokine control in the acute response to virus infection. Cell Host Microbe 18, 714–722 (2015).
Galiano, M. R. et al. Myelin basic protein functions as a microtubule stabilizing protein in differentiated oligodendrocytes. J. Neurosci. Res. 84, 534–541 (2006).
Aber, E. R. et al. Oligodendroglial macroautophagy is essential for myelin sheath turnover to prevent neurodegeneration and death. Cell Rep. 41, 111480 (2022).
Ehrlich, M. et al. Rapid and efficient generation of oligodendrocytes from human induced pluripotent stem cells using transcription factors. Proc. Natl Acad. Sci. USA 114, E2243–E2252 (2017).
Smirnova, E. V. et al. Comprehensive Atlas of the myelin basic protein interaction landscape. Biomolecules 11, 1628 (2021).
Snaidero, N. et al. Antagonistic Functions of MBP and CNP establish cytosolic channels in CNS Myelin. Cell Rep. 18, 314–323 (2017).
Wang, D. et al. Loregic: a method to characterize the cooperative logic of regulatory factors. PLoS Comput. Biol. 11, e1004132 (2015).
Rexach, J. E. et al. Cross-disorder and disease-specific pathways in dementia revealed by single-cell genomics. Cell 187, 5753–5774.e28 (2024).
Yashar, W. M. et al. Predicting transcription factor activity using prior biological information. iScience 27, 109124 (2024).
Duren, Z. et al. Sc-compReg enables the comparison of gene regulatory networks between conditions using single-cell data. Nat. Commun. 12, 4763 (2021).
Ferrari, C., Manosalva Pérez, N. & Vandepoele, K. MINI-EX: integrative inference of single-cell gene regulatory networks in plants. Mol. Plant 15, 1807–1824 (2022).
Duren, Z., Chen, X., Jiang, R., Wang, Y. & Wong, W. H. Modeling gene regulation from paired expression and chromatin accessibility data. Proc. Natl Acad. Sci. USA 114, E4914–E4923 (2017).
Liu, Z. et al. Induction of oligodendrocyte differentiation by Olig2 and Sox10: Evidence for reciprocal interactions and dosage-dependent mechanisms. Dev. Biol. 302, 683–693 (2007).
Kim, J. et al. The co-regulation mechanism of transcription factors in the human gene regulatory network. Nucleic Acids Res. 40, 8849–8861 (2012).
Fuda, N. J., Ardehali, M. B. & Lis, J. T. Defining mechanisms that regulate RNA polymerase II transcription in vivo. Nature 461, 186–192 (2009).
Buchler, N. E., Gerland, U. & Hwa, T. On schemes of combinatorial transcription logic. Proc. Natl Acad. Sci. USA100, 5136–5141 (2003).
Silva-Rocha, R. & De Lorenzo, V. Mining logic gates in prokaryotic transcriptional regulation networks. FEBS Lett. 582, 1237–1244 (2008).
Krumsiek, J., Marr, C., Schroeder, T. & Theis, F. J. Hierarchical differentiation of myeloid progenitors is encoded in the transcription factor network. PLoS ONE 6, e22649 (2011).
Malekpour, S. A., Shahdoust, M., Aghdam, R. & Sadeghi, M. wpLogicNet: logic gate and structure inference in gene regulatory networks. Bioinformatics 39, btad072 (2023).
Malekpour, S. A., Haghverdi, L. & Sadeghi, M. Single-cell multi-omics analysis identifies context-specific gene regulatory gates and mechanisms. Brief. Bioinform. 25, bbae180 (2024).
Castro-Mondragon, J. A. et al. JASPAR 2022: the 9th release of the open-access database of transcription factor binding profiles. Nucleic Acids Res. 50, D165–D173 (2022).
Alicia, S. motifmatchr: Fast Motif Matching in R. R package version 1.28.0. https://doi.org/10.18129/B9.bioc.motifmatchr (2024).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e29 (2021).
Cavalcante, R. G. & Sartor, M. A. annotatr: genomic regions in context. Bioinformatics 33, 2381–2383 (2017).
Ben-Kiki, O., Bercovich, A., Lifshitz, A. & Tanay, A. Metacell-2: a divide-and-conquer metacell algorithm for scalable scRNA-seq analysis. Genome Biol. 23, 100 (2022).
Paszke, A. et al. PyTorch: An imperative style, high-performance deep learning library. https://doi.org/10.48550/ARXIV.1912.01703. (2019).
Koopmans, L. H., Owen, D. B. & Rosenblatt, J. I. Confidence intervals for the coefficient of variation for the normal and log normal distributions. Biometrika 51, 25–32 (1964).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Gerstein, M. B. et al. Architecture of the human regulatory network derived from ENCODE data. Nature 489, 91–100 (2012).
Kent, W. J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
Lerdrup, M., Johansen, J. V., Agrawal-Singh, S. & Hansen, K. An interactive environment for agile analysis and visualization of ChIP-sequencing data. Nat. Struct. Mol. Biol. 23, 349–357 (2016).
Müssel, C., Hopfensitz, M. & Kestler, H. A. BoolNet—an R package for generation, reconstruction and analysis of Boolean networks. Bioinformatics 26, 1378–1380 (2010).
Smedley, D. et al. BioMart – biological queries made easy. BMC Genom.10, 22 (2009).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
McInnes, L., Healy, J. & Melville, J. UMAP: Uniform Manifold Approximation and Projection for Dimension Reduction. https://doi.org/10.48550/ARXIV.1802.03426. (2018).
Kim, D. et al. Homo-trimerization is essential for the transcription factor function of Myrf for oligodendrocyte differentiation. Nucleic Acids Res. 45, 5112–5125 (2017).
Vorontsov, I. E. et al. HOCOMOCO in 2024: a rebuild of the curated collection of binding models for human and mouse transcription factors. Nucleic Acids Res. 52, D154–D163 (2024).
Štrumbelj, E. & Kononenko, I. Explaining prediction models and individual predictions with feature contributions. Knowl. Inf. Syst. 41, 647–665 (2014).
Acknowledgements
This work was supported by National Institutes of Health grants, R21 NS128761, RF1MH128695, R01AG067025, R21NS128761, and National Science Foundation Career Award 2144475, and a core grant to the Waisman Center from NICHD (P50 HD105353).
Author information
Authors and Affiliations
Contributions
Conceptualization, J.S. and D.W.; Methodology, J.C., J.S., and D.W.; Formal Analysis, J.C.; Investigation, J.C., J.S., and D.W.; Writing – Original Draft, J.C.; Writing – Review & Editing; J.C., J.S., and D.W.; Supervision, J.S., and D.W.; Funding Acquisition, J.S. and D.W.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Seyed Malekpour and Ping-Han Hsieh for their contribution to the peer review of this work. Primary Handling Editors:Chien-Yu Chen and Mengtan Xing.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Choi, J.J., Svaren, J. & Wang, D. CoTF-reg reveals cooperative transcription factors in oligodendrocyte gene regulation using single-cell multi-omics. Commun Biol 8, 181 (2025). https://doi.org/10.1038/s42003-025-07570-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-07570-6
This article is cited by
-
Cooperative multi-view integration with a scalable and interpretable model explainer
Nature Machine Intelligence (2025)