Abstract
As climate change threatens marine ecosystems, efforts to restore coral reefs using resilient corals are increasing. This conservation approach remains limited by our understanding of cellular mechanisms of resilience and trade-offs. Here, we demonstrate that downregulation of pa-BAK stabilizes the coral-algal endosymbiosis and slows bleaching during acute heat stress in Pocillopora acuta through coordinated expression of gene clusters. The improvement in thermal tolerance was closely related to the downregulation efficiency in individual corals. Oxidative DNA damage, a hallmark of thermal stress response, was prevented in corals with stabilized symbiosis, likely through a decrease in mitochondrial ROS release. We hypothesize that this manipulation causes a cascading molecular response, which may impact other traits such as oxidative mitochondrial damage, proving detrimental over the longer term. Developing our understanding of heat-stress defense mechanisms that promote stability in the coral-algal symbiosis is fundamental for effective modern coral reef restoration practices based on improving ecosystem resilience.
Similar content being viewed by others
Introduction
Ocean temperatures are virtually certain to increase due to anthropogenic emissions, leading to more frequent and intense warming events that result in the breakdown of the coral-algal symbiosis1,2. Reef corals are mixotrophic organisms, heavily relying on the photosynthetic activity of their endosymbionts, unicellular algae in the family Symbiodiniaceae, which meet metabolic carbon requirements in exchange for carbon dioxide and a tightly controlled transfer of nitrogen and phosphorus3,4,5. This symbiosis is the energetic foundation of tropical coral reefs, and its breakdown results in a phenomenon known as coral bleaching, which compromises the structure and function of 6corals and the reefs they build6. Tolerance to environmental stress varies both among and within species7,8, creating a mosaic of stress and mortality during environmental perturbation. Pocillopora acuta is widespread across the Indo-Pacific and in Kāne’ohe Bay, Hawaiʻi9, where it is among the most sensitive species to natural10 and experimental heat stress11. This sensitivity makes P. acuta an ideal model for investigating strategies to enhance coral resilience to thermal stress12,13.
The molecular and cellular mechanisms underpinning the initiation, maintenance, and disruption of the coral-algal symbiosis are still largely unknown. Multiple mechanisms have been proposed14, including programmed cell death pathways (PCD; apoptosis, and symbiophagy), which play an important role in natural conditions and laboratory-induced bleaching experiments12,15,16,17,18,19,20. These pathways involve the interaction of “pro-life” and “pro-death” proteins that inhibit or activate the execution of the pathway through post-translational modifications and physical protein-protein interactions21. BAK and BAX are two major apoptosis proteins that heterodimerize upon pro-death signaling, causing immediate permeabilization of the mitochondrial membrane, release of cytochrome c, fragmentation of mitochondria, and activation of caspases—proteases that degrade the content of the cell22. Each is also involved in the autophagy cascade; both pathways are closely interconnected and can compensate for each other in certain conditions17,22,23. In P. acuta corals pre-exposed to sublethal temperature stress, the expression of the pro-survival gene pa-Bcl-2 increases relative to pro-death genes pa-Bak and pa-Bax during the first 24 h of acute thermal stress, while the corals become more thermally resilient12. The inhibition of pa-Bcl-2 negates this beneficial phenotype, confirming the role of this protein in coral bleaching.
Since inhibition of pa-Bcl-2 increased the bleaching rate of more resilient corals, the impairment of BAK/BAX heterodimerization might prevent or delay the bleaching cascade in wild-type individuals. Vertebrate models show that BAK and/or BAX knockdown and knockout are successful strategies to prevent PCD, even upon increased pro-death signaling22,24,25, and targeting BAK may be a more successful strategy than targeting BAX to avert PCD26,27,28.
Many associated pathways have been proposed to play a role in coral bleaching and improved thermal tolerance12,13,15,17,29,30. This complex physiology leads to a range of cellular and molecular consequences, including the accumulation of (oxidative) DNA damage in heat-stressed corals, which can be exacerbated by thermal preconditioning or by the addition of external antioxidants13,31,32. The ROS-dependent bleaching theory postulates that during heat stress, symbionts are the main source of reactive oxygen species (ROS) which, unless scavenged by the host antioxidant defense, are detrimental to the host cell and trigger symbiosis disruption (reviewed in ref. 33). Increased DNA damage and the resultant compromised genome stability could be trade-offs to induced symbiosis stability in heat-stressed corals, such as those with manipulated gene expression.
In this work, we analyzed the connection between BAK manipulation, symbiosis stability, cellular oxidative stress, and genes involved in antioxidant defense, DNA repair, programmed cell death, and AMPK and mTOR signaling pathways. We show that coral thermal resilience can be improved by the downregulation of a single gene, eliciting a complex molecular response that mitigates oxidative DNA damage in heat-stressed corals. While the precise mechanisms behind this improved phenotype remain unknown, we propose a novel pathway (AMPK/Foxo3a/Nrf2) that affects multiple pro-life and antioxidant genes involved in coral response to heat stress.
Methods
Coral collection and experimental setup
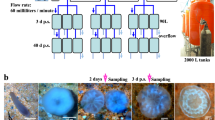
Twelve colonies of Pocillopora acuta (~ 15 to 20 cm in diameter) were collected in 2022 and 2023 at 3 different patch reefs in Kāneʻohe Bay, Hawaiʻi. Although clonality in this brooding species is common, it can be minimized by collecting from separate patch reefs34, so we opted to use spatially widespread sampling to maximize the likelihood of gathering distinct genotypes. Colonies were fragmented into ~7 cm nubbins and maintained in shallow flow-through seawater tanks under natural light until the start of the experiment. The experimental setup is depicted in Supplementary Fig. 1. An additional 6 colonies were collected in 2024 for the ROS analysis experiment.
siRNA-mediated knockdown
The experiment was conducted as described in ref. 13 with minor modifications. Freshly cut coral fragments (~3 cm) were placed into 20 ml cultivation vials, completely submerged in a flow-through tank with ambient-temperature seawater, and left to acclimatize overnight. This step is crucial for the siRNA transfection to be successful. The next morning, vials were moved into a temperature-controlled illuminated waterbath (280 series waterbath, Thermo Scientific with Hydra 64 HD LED Reef Light, AquaIllumination). Seawater was removed without disturbing the corals, and the siRNA transfection was carried out according to the manufacturerʻs instructions using a transfection mix consisting of 20 μl of 10 nM siRNA (siBAK targeting pa-BAK mRNA and siNTC designed as a negative control with no known target in the P. acuta mRNA sequence database. The duplexes were designed using the BLOCK-iT™ RNAi Designer online platform by Thermofisher. See Supplementary Table 1 for sequences) and 16 μl INTERFERin® reagent (Polyplus) in 400 μl of 0.2 μm filtered seawater pipetted directly onto the exposed coral. After 2 min, the coral fragment was covered in 5 ml of 0.2 μm filtered seawater and incubated at ambient temperature for 6 h. 30 minutes after the transfection, soft aquarium airline PVC tubing (4 mm in inside diameter, NACX) was inserted into each cultivation vial, and the seawater was aerated with approximately 1 bubble every 3 s to prevent hypoxic stress without disturbing the corals. After the 6-h incubation, coral fragments were removed from the transfection mixture, placed in a 6-well plate, and completely submerged in a flow-through seawater tank. 24 h after the beginning of the siRNA transfection, corals were exposed to acute heat stress ramping from ambient temperature (25.3–26.5 °C, depending on the day of the experiment) to 32 °C over 2.5 h and held at 32 °C. After 48 h in the stress treatment, we evaluated fragments using confocal microscopy (see below), and small subsamples were flash-frozen in liquid nitrogen and maintained at −80 °C until extractions for gene expression and DNA damage analyses. We re-evaluated the corals using confocal microscopy at 96 h in the stress treatment.
Bleaching rate assessment
We evaluated bleaching using confocal microscopy analysis (Zeiss LSM-710) following ref. 12. EC Plan-Neofluar 2.5X/0.075 objective with a total magnification of 25X was used for the imaging. Images were acquired in Zen Imaging Software (ZEN 2.3 SP1 FP3 black, v14.0.28.201) by Zeiss. Host cell fluorescence was excited with the 405 nm laser and collected in the emission range 454–621 nm with master gain set at 650, digital gain 1.0, laser power 28%, and pinhole 35.5. Symbiodinium fluorescence was excited with the 405 nm laser and collected in the emission range 655–718 nm with master gain set at 575, digital gain 1.0, laser power 28%, and pinhole 35.5.
The natural fluorescence of algal-derived chlorophyll a was normalized to the natural fluorescence signal of animal-derived GFP protein and used as a proxy for the symbiont density in the coral tissue12,35. Coral fragments were measured the day before siRNA transfection and 48 and 96 h post-heat stress. Three snaps of the top side of each coral were taken, which covered the entire fragment area. Ten circular regions of interest (ROI) were analyzed on the focal plane of each picture (Fiji v2.14.0/1.54f36,), avoiding the polyp mouth and tentacles that emit intense GFP-like signal. All ten ROIs were averaged, and one value per snap was used for the statistical analysis. Three snaps were analyzed for each coral per timepoint per treatment.
DNA damage assay
DNA was extracted from corals at the 48 h timepoint (72 h after the transfection) using Quick-DNA/RNATM Microprep Plus Kit (Zymo Research). The level of 8-Hydroxy-2’-deoxyguanosine (8-OhdG) – a proxy for oxidative DNA damage – was analyzed in corals using the DNA Damage Competitive ELISA Kit (Invitrogen) as described in ref. 13. 500 ng of total genomic DNA in 20 μl was denatured at 95 °C for 5 min, cooled down on ice, and digested to single nucleotides by 20 U of Nuclease P1 (NEB) at 37 °C for 30 min followed by dephosphorylation by 1 U of Shrimp Alkaline Phosphatase (NEB) for 30 min at 37 °C. All the enzymes were inactivated at 75 °C for 10 min. DNA was then diluted 1:5 in 1x Assay buffer and processed according to the kit instructions (DNA Damage Competitive ELISA Kit, Invitrogen). We used the MyCurveFit.com online platform (My Assays Ltd.) to model the standard curve and to predict result values, as shown in Supplementary data 1.
Gene expression analysis
Gene expression was analyzed as described in ref. 12 with minor changes. RNA was extracted from samples collected at the 48 h timepoint (72 h after the transfection) using Quick-DNA/RNATM Microprep Plus Kit (Zymo Research), including the DNase I step. This timepoint was a total of 72 h post-siRNA delivery (including the 24 h of transfection and recovery), which is generally considered the latest timepoint for mRNA level analysis because the induced mRNA downregulation loses efficiency over time (Interferrin™ protocol, Polyplus). We chose this timepoint as a compromise between the time at which we can still observe efficient downregulation and the onset of the phenotypic shift in the bleaching rate. 1000 ng of RNA was reverse transcribed with a High-Capacity cDNA reverse transcription kit (ThermoFisher Scientific) with the addition of 20 U of Murine Rnase Inhibitor (New England Biolabs). qPCR reactions were run in 12 μl with PowerUpTM SYBRTM Green Master Mix (Applied Biosystems) for 40 cycles according to the manufacturerʻs instructions. The efficiency of each primer pair was tested with a dilution curve followed by a melting curve analysis. Pa-EF-1 (elongation factor 1a) was used as a normalizing gene based on the optimization process published in ref. 12. Expression of target genes in siBAK-treated corals was calculated relative to the siNTC-treated corals with the ΔΔCt method.
Each analysis included one-sample per treatment for each colony.
We analyzed the expression of the following genes: pa-GR (Glutathione reductase, XM_027196629.1), pa-OGG1 (8-oxoguanine DNA glycosylase, XM_027192323.1), pa-FOXO3a (forkhead box O3, XM_027182315.1), pa-HO-1 (heme oxygenase, XP_027042110.1), pa-BI-1 (Bax inhibitor 1, XM_027189407), pa-Bcl-2 (B-cell lymphoma 2, XM_027196784.1), pa-BAK (Bcl-2 homologous antagonist/killer, XM_027189902.1), pa-BAX (Bcl-2 associated X protein, XM_027185654.1), pa-Cas3 (Caspase 3, XM_027199402.1), pa-GABARAP (GABA type A receptor-associated protein, XM_027194913.1), pa-LC3 (microtubule-associated proteins 1A/1B light chain 3B, XM_027182987.1), pa-AMPK (AMP-activated protein kinase, XM_027190931), and pa-mTOR (mechanistic target of rapamycin, XM_027197193). Sequences of all primers used in this study are listed in Supplementary Table 1.
ROS microscopy analysis
Coral fragments of 6 different colonies with successful BAK pa-BAK knockdown from a separate experiment were used for ROS analysis. At 6 h after heat stress (~ 27 h post-transfection), each coral was sampled for gene expression analysis (as described above). We chose this timepoint based on the previous experiments showing the expression of pa-BAK in heat-stressed P. acuta peaks around 6 h of heat stress12.
The tissue of corals with successful BAK knockdown was scraped with a small bottle brush into 5 ml of 0.45 μm filtered seawater and spun at 850 g for 4 min following ref. 37. Coral cells were resuspended in 100 μl of filtered seawater and let acclimate under artificial light for 15 min Hoechst 33342 (Invitrogen, 10 mg/ml) was diluted 2500x to stain DNA, and CellROXTM Orange Reagent (Invitrogen) was added at a final concentration of 5 μM to the cells and incubated for 15 min. Incubation time was adjusted from the recommended 30 min based on the optimizations. A shorter time prevents oversaturation of the CellRox dye, which is a cumulative dye. Stained tissue culture was then transferred to 10-well PTFE Printed Microscope slides (Electron Microscopy Sciences) and covered with a gold-seal coverglass (Electron Microscopy Sciences). Until this moment, all work took place at temperatures relevant to the treatment type –26 °C for ambient temperature controls and 32 °C for the heat-stressed siNTC and siBAK corals. Slides were then placed into a microscopy chamber preheated at 26 °C and scanned using a confocal microscope Zeiss LSM-710. Z-scans with a set number of slices (12) and a similar interval between them (~2.5 μm) were taken with the following channels: (i) GFP-like, ex 480/em 504–573 nm, (ii) chlorophyll a, ex 633/em 647-721 nm, (iii) Hoechst 33342, ex 405/em 410–504 nm, (iv) CellRox Orange, ex 561/em 572–632 nm. These spectra were determined using the Smart Setup function with the “Best compromise” setting in the microscopy software (ZEN 2.3 SP1 FP3 black, v14.0.28.201). This software interface allows for an advanced system hardware configuration for a chosen set of dyes.
Five scans per well in 4 different wells were taken, totaling 20 scans per colony per treatment. Each scan was chosen to contain at least two but preferably multiple symbiont cells, and they were used as limits for the Z-scan settings. Due to the inbuilt incompatibility of the ZEN Black software with the microscope bright field camera, we used a hand-held iPhone 12 mini camera to capture the cells in natural light. These photos were not used for quantitative analysis but served as a visual aid for categorizing cells based on morphological features. Scans of cells with no added dyes were taken to verify that coral cells do not have autofluorescence in the respective spectra.
All the scans were analyzed using Fiji software (Fiji v2.14.0/1.54 f). Cells were divided into four categories based on the combination of specific fluorescence signals and bright light morphology. Symbiont cells showed a high chlorophyll-like signal and were brown under the bright light. To distinguish between symbionts still embedded in a host cell and free symbionts, we visualized the Hoechst 33342 signal. Hoechst 33342 is highly cell-permeable, allowing for living cell staining (Manufacturer’s information); however, it does not efficiently enter the plant cell wall without prior permeabilization/fixation. Therefore, symbiont cells with a visible blue spot inside or on the exterior were labeled as symbionts in a host cell. Two or three symbionts in a host cell were identified only when a connective membrane was visible in the bright light picture. Finally, host cells were first assessed in the bright light picture for size and shape. Only circular cells with no visible damage or abnormalities (indicating cellular death) and of at least approximately 3 μm in diameter were allowed for the analysis. Besides cells with a strong GFP signal (covering the whole cell surface or concentrated in one spot), we identified host cells with no GFP signal in agreement with previously published work38. One region with no cells per scan was analyzed as a background measurement to normalize for non-ROS-related oxidation of the CellRox Dye (during manipulations, etc.). Symbionts harbor a specific phase-separated organelle – the pyrenoid – that enhances photosynthetic carbon assimilation via the enzyme Rubisco39. Pyrenoids in symbiont cells were highly autofluorescent in the orange spectrum used to visualize CellRox Orange. This artifact was not present in all symbiotic algae, so we excluded this region of the cell in our analysis because it oversaturated the signal from other cells without pyrenoids (Supplementary Fig. 2).
Statistical analyses
All statistical analyses and visualizations were conducted in R, Version 2023.06.1 + 52440. We evaluated normality and heteroscedasticity for each dataset. We analyzed pa-BAK expression at 48 h post-heat stress and designated each trial a successful knockdown when the pa-BAK expression was less than 80% of siNTC-treated corals. The efficiency of siBAK knockdown was analyzed with a one-sample t-test, comparing the expression to 1. The Chl a/GFP ratio calculated from confocal microscopy was analyzed using a generalized linear mixed model in lme441 with treatment and time as main effects and colony as a random effect (ratio~treatment*time + (1|coral)).
Bleaching rate was calculated for each treatment as the slope of the fit between time (hours) and Chl a/GFP and used to calculate the tolerance gain (the change in bleaching after siBAK knockdown) using the following formula:
pa-BAK knockdown efficiency was calculated as the gene expression in siBAK normalized to siNTC via the ΔΔCt method, subtracted from one, and expressed in percentages using the following formula:
We calculated the tolerance gain for each coral and compared it to the pa-BAK knockdown efficiency using a Pearson correlation.
For hierarchical clustering and correlation analyses of gene expressions, we used the “raw gene expression” value calculated using the ΔCt method. It represents the relative abundance of target gene mRNA compared to that of the reference (normalizing) gene in each sample.
We analyzed the hierarchical clustering using the ‘complete’ method in the R package hclust to find relative relationships between genes and evaluated the pairwise correlation between all genes in each sample using a Pearson correlation. A one-sample t-test was used to evaluate if the relative expression of each gene was different from 1. The relationship between pa-BAK expression at 48 h and other genes was evaluated using a Pearson correlation.
The oxidative DNA damage marker 8-OhdG was compared between siNTC and siBAK using the Paired t-test, and we used a Pearson correlation to compare DNA damage to the tolerance gain.
The ROS signal was analyzed using a generalized linear mixed model in lme4 with treatment and celltype as main effects and colony and time of scan as a random effect (adjusted_signal~treatment*celltype + (1|coral) +(1|time)). The individual relationships were tested with the Tukey post hoc testing using the function lsmeans (package “emmeans”42); contrasts were pulled using the function cld (package “multcomp”43) with a significance threshold set as 0.05.
Figure preparation
Graphs in this manuscript were prepared in RStudio Version 2023.06.1 + 524 using packages ggplot244, ggtext45, cowplot46, and viridis47. Microscopy pictures were assembled in GIMP (2.10.38). All schematic figures in this manuscript were created from scratch by the authors using a combination of Microsoft PowerPoint for Mac (Version 16.95.4) and the Sketchbook app for iPad (Version 6.1.0). Illustrations were either hand-drawn or assembled using stock icons and shapes available within the Microsoft Office package.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Results
pa-BAK knockdown improves bleaching resilience of corals
We delivered siBAK to a total of 12 P. acuta colonies across multiple independent trials. After our first delivery attempt, we observed successful knockdown (defined as >20% reduction in expression compared to control) in 6 colonies (50%). For the remaining 6 colonies where knockdown was not initially successful, we performed a second delivery attempt; however, no further attempts were made. We successfully downregulated pa-BAK expression in 9 out of the 12 targeted colonies (75% of colonies, representing 56.3% of all samples, including repeated attempts of the same colony). Expression after knockdown averaged 57.5% ± 25.1% (mean ± SD, range: 2% to 79%, Fig. 1A) of control expression levels.
A Successful siRNA-mediated downregulation of pa-BAK in Pocillopora acuta. The expression is normalized to the control treatment (siNTC) by the ΔΔCT method. The error bar represents standard deviation (n = 9 colonies). B Bleaching rate in heat-stressed corals with successful pa-BAK knockdown (siBAK) compared to their controls (siNTC), points represent each microscopy snap. Three snaps were taken for each coral (n = 9). C Correlation between the tolerance gain (calculated as the percent difference between the bleaching rate of treated (siBAK) and control corals (siNTC) for each colony) and the magnitude of pa-BAK knockdown (calculated as percentual downregulation compared to the control), R stands for Pearson correlation coefficient (n = 9).
We then analyzed the bleaching rate in heat-stressed corals by comparing Chl a (symbiont-derived signal) to the GFP (host-derived signal) ratio over time12,35. Corals were scanned the day before the siRNA delivery, 48 h into the acute heat stress (i.e., 72 h post-siRNA delivery), and 96 h into the acute heat stress (i.e., 120 h post-siRNA delivery). Exposure to heat significantly bleached the corals, illustrated via decreased Chl a : GFP ratio (p(time) < 0.001, Fig. 1B). Corals with successful pa-BAK knockdown bleached 49.5% slower than controls (p(treatment:timepoint) = 0.004). When examining the signals separately, the GFP signal increased, in agreement with previously published data12, with a larger increase in siBAK corals compared to siNTC (Supplementary Fig. 3A). Thus, the difference in bleaching rate is explained by the decreasing Chl a signal corresponding to the loss of symbionts or chlorophyll degradation, eventually (Supplementary Fig. 3B), and cannot be explained by the GFP dynamics of treated corals. Corals in which BAK downregulation was not successful did not show any shift in bleaching rate compared to siNTC-treated corals (Supplementary Fig. 3C).
The degree of improvement in bleaching tolerance – the tolerance gain – was positively correlated with the magnitude of pa-BAK downregulation (R = 0.68, p = 0.043, Fig. 1C); corals with more pronounced pa-BAK knockdown bleached at a slower rate (normalized to their individual controls) during acute heat stress. This relationship was driven by the siRNA-mediated knockdown; the tolerance gain did not correlate with pa-BAK expression in heat-stressed corals in which BAK was not successfully downregulated (R = 0.3, p = 0.56, Supplementary Fig. 3D). However, this correlation seems to be primarily driven by the one coral with the most successful gene knockdown that also showed the highest tolerance gain. The use of a ranked correlation method (Kendall’s Tau or Spearman’s Rank Correlation test) did not find a significant correlation between the two traits (R = 0.22, p = 0.48, and R = 0.3, p = 0.44, respectively)
Genes connected to programmed cell death and oxidative stress response are correlated in heat-stressed corals upon BAK downregulation
To better understand which cellular and molecular pathways are affected during the phenotypic change associated with BAK downregulation, we examined the pairwise relationships of the expression of a variety of genes. Hierarchical clustering suggested that BAX and LC3 had relatively discrete expression profiles, while the rest of the genes were in a single node of the dendrogram (Fig. 2A).
A Hierarchical clustering of gene expression using the “complete” method in heat-stressed corals with downregulated pa-BAK. B Gene expression in heat-stressed corals with downregulated pa-BAK, dashed line is no change (n = 9 colonies). C Heat map of gene expression correlations. Numbers represent Pearson correlation values, significant (p < 0.05) in black, and non-significant (p > 0.05) in light purple (n = 9). * represents p-values lower than 0.05, ‡ represents p-values between 0.05 and 0.1.
To eliminate the possibility that pa-BAX compensated for the loss of pa-BAK, we examined the expression of pa-BAX in siBAK-treated corals (Fig. 2B). While slightly higher, pa-BAX expression was not statistically different in pa-BAK downregulated corals (p = 0.205) compared to the siNTC treatment, and the expression of pa-BAX was not correlated with pa-BAK (R = −0.11, p = 0.731, Fig. 2C). pa-OGG1 and pa-AMPK expression were also significantly downregulated in pa-BAK downregulated corals (p < 0.001 and p = 0.017, respectively). We observed a trend towards pa-LC3 upregulation, which was not statistically significant (p = 0.07; Fig. 2B).
For the Pairwise correlation analysis, raw gene expression values—defined as the difference in expression between siBAK and control coral—were compared across all coral colonies for each gene in the dataset, allowing us to identify genes with similar response patterns. This analysis revealed two major modules of broadly correlated genes (Fig. 2C and Supplementary Table 2). The first module consisted of pa-GAB, pa-mTOR, pa-GR, and pa-BI-1 (Pearson R > 0.86, p < 0.006), while the second group consisted of pa-AMPK, pa-foxo3, pa-HO-1, and pa-Bcl-2 (Pearson R > 0.77, p < 0.013; with the exception of pa-AMPK and pa-Bcl-2, which did not show significant correlation with each other). These gene groupings correspond to the global clustering patterns observed in Fig. 2A.
pa-Cas3 was strongly correlated with pa-LC3 (R = 0.84, p = 0.009) and pa-OGG1 was strongly correlated with pa-HO-1 (R = 0.91, p = 0.002). No gene expression besides pa-BAK was correlated with tolerance gain (Supplementary Fig. 4A). Due to technical issues, we did not obtain expression data for one colony for pa-GABARAP, pa-mTOR, pa-GR, and pa-BAX.
Markers of oxidative DNA damage decrease in corals after pa-BAK knockdown
Corals with effective pa-BAK knockdown showed significantly decreased levels of oxidative DNA damage after 48 h in heat stress, 72 h post-siRNA delivery (paired t-test, p = 0.011, Fig. 3A), and all colonies tested showed lower DNA damage in the siRNA treatment (n = 7). Additionally, the level of oxidative DNA damage negatively correlated with the tolerance gain (R = -0.83, p = 0.02, Fig. 3B) but not with the extent of the pa-BAK downregulation (R = 0.51, p = 0.25, Supplementary Fig. 3E). Expression of pa-foxo3 was positively correlated with the level of oxidative DNA damage marker in heat-stressed corals (R = 0.77, p = 0.044, Supplementary Fig. 4B). DNA extraction of two samples failed to produce enough material for the ELISA DNA damage analysis input requirements.
A The level of oxidative DNA damage marker (8-OHdG) in DNA of heat-stressed corals with successful pa-BAK knockdown (siBAK) and controls of the same colony (siNTC; n = 7). B Correlation between the tolerance gain (calculated as the percent difference between bleaching rates of treated (siBAK) and control corals (siNTC)) and the relative decrease of oxidative DNA damage marker in treated corals (n = 7). C The level of ROS in coral host cells, calculated as the mean fluorescent signal normalized by background signal. AT stands for ambient temperature; siBAK and siNTC underwent heat stress. D Example of microscopy images of each cell type captured in each channel. Hoechst 33342 stains DNA. CellRox becomes fluorescent after oxidation and is used as a proxy for reactive oxygen species (ROS) accumulation over time. Host cells (right side of the panel) are in three different categories based on the GFP signal.
pa-BAK knockdown mitigates ROS release in heat-stressed coral host cells
To examine whether the observed decrease in oxidative DNA damage was connected to a decrease in cellular oxidative stress, we conducted a second siRNA experiment and analyzed ROS release. pa-BAK was successfully inhibited in 6 different corals; gene expression after knockdown averaged 46.8% ± 16.1% (average ± SD) of control.
Based on cell morphology and presence/absence of autofluorescent signal in GFP and chlorophyll a spectra, we divided cells into different categories—Symbiont, Symbiont in host cell, 2+ Symbionts in host cell, and Host cell, and we measured the ROS-sensitive dye CellRox Orange-derived signal as a proxy for the level of intracellular oxidative stress (Fig. 3C, D and Supplementary information Fig. 5). This dye emits fluorescence when oxidized by ROS and is an accumulative dye. To eliminate the possibility of this artifact impacting our results, we normalized the fluorescence mean of each cell by the fluorescence mean of the scan background area. We also included the time of scan as a random effect in our statistical analysis.
In ambient conditions (AT), host cells and symbiotic cells (host cells harboring a symbiont cell) had a higher ROS signal than single symbiont cells (p < 0.001, Supplementary Fig. 5B). Heat stress increased intracellular oxidative stress in host cells in both siRNA treatments (siBAK and siNTC) compared to the ambient control (p < 0.001); however, the siBAK treatment had significantly less ROS signal than the siNTC treatment, indicating that pa-BAK knockdown efficiently alleviated it (Fig. 3C). ROS signal also increased in heat-stressed single symbiont cells (p < 0.001) but with no apparent effect of siRNA treatment (p = 0.97, Supplementary Fig. 5B). We observed no effect of temperature or treatment in symbiotic cells (p > 0.53, Supplementary Fig. 5B).
Discussion
Coral thermal tolerance is a complex trait driven by the interaction between cnidarian host and photosynthetic symbionts, making it especially difficult to describe and manipulate. As coral reefs face an increasingly stressful future of warming oceans driven by anthropogenic change, developing tools and a basic understanding of these mechanisms is critical for conservation and management. Here we show that the manipulation of a single gene slows coral bleaching, decreases oxidative DNA damage, and creates an extensive cellular cascade with potentially transformative effects.
The use of a single siRNA construct to target a gene of interest has known limitations, including the potential for off-target silencing48. Although there is a possibility that off-target interactions could influence some of the observed effects, this experiment was designed as a hypothesis-driven study rather than an exploratory multi-omic analysis where such interactions would present significant challenges to interpretation. We anticipated the phenotypic consequences of pa-BAK knockdown based on known function, and our results aligned with previous results. The observed effects support our measurements, demonstrating that the knockdown was effective and that the phenotypic changes are likely attributable to pa-BAK suppression rather than unintended siRNA interactions.
Programmed cell death pathways (PCD) are one of the common mechanisms of coral-algal symbiosis disruption12,15,17,19,20,49. There is a well-documented cross-talk between apoptosis and autophagy, and several proteins such as Bcl-2, BAK, or BAX are described to have the power to activate/prevent both pathways, depending on particular cellular conditions21,50. The expression ratio of pro-life Bcl-2 to pro-death BAK and BAX increases during acute heat stress in P. acuta and Acropora millepora and is significantly higher in corals with higher thermal resilience12,15. Inhibition of pa-Bcl2 leads to the loss of the beneficial phenotype12. Since the disruption of symbiosis in corals is the major response mechanism to various stress conditions51, it is more meaningful—and also more challenging—to search for gene manipulations that can enhance, rather than disrupt, the stability of coral-algal symbiosis under thermal stress. Experiments in vertebrates have shown that BAK or/and BAX downregulation is a successful strategy to prevent PCD in cells22,24,26, and our results demonstrate that pa-BAK knockdown is sufficient to slow down but not fully prevent heat-induced coral bleaching (Fig. 1B). This may be due to the incomplete nature of expression decline using siRNA-mediated knockdown, leaving enough pa-BAK to heterodimerize with pa-BAX and activate the PCD. However, since we observed a weak correlation between pa-BAK expression and bleaching rate, mostly driven by one coral, it is possible that other mechanisms partially compensate for the loss of the pro-death PCD signaling.
Heterodimerization of BAK and BAX and the subsequent permeabilization of the mitochondrial membrane is considered a non-reversible point in the apoptotic cascade22: it triggers a self-destroying cellular mechanism based on the expression and activation of multiple executioner proteases of the family of caspases21. The detailed mechanism of BAK/BAX’s role in the activation of autophagy is not yet well understood, but it is also presumed that they are the main trigger before the execution of the pathway21,22,50. Several studies showed that depletion of both proteins is necessary to efficiently block the execution of the programmed cell death pathway25,52, potentially because BAK can be upregulated in BAX-depleted cells, efficiently compensating for the loss of function26. In our experiment, we did not observe a compensatory relationship between BAX and BAK upon knockdown (Fig. 2), suggesting that BAK has a unique, BAX-independent role in the disruption of the coral host-algal symbiosis. BAK’s distinctive BAX-independent role has also been described in model organisms27,28.
Coral thermal tolerance and stress response are intricate traits that involve myriad cell pathways and affect coral physiology and metabolism12,13,29,30,53,54,55. The complexity of these processes means there are likely to be off-target consequences of ‘forced’ thermal tolerance, so the evaluation of other potentially detrimental traits is critical from a basic and applied perspective. Thermal stress causes the accumulation of oxidative DNA damage markers in coral tissue, which can be prevented through preconditioning13. This oxidative mutation is usually recognized and repaired by DNA glycosylases, most commonly OGG1 (8-oxoguanine DNA glycosylase), via the process of base excision repair56. In alignment with the observed decrease in oxidative DNA damage (Fig. 3A), the expression of pa-OGG1 also significantly decreased in pa-BAK downregulated heat-stressed corals (Fig. 2B), suggesting DNA damage is prevented, not repaired, in pa-BAK knockdown corals. This is further corroborated by the analysis of ROS release in coral cells; both host cells and symbionts increase the oxidative environment in heat stress, but a part of it is exacerbated by the loss of pa-BAK in host cells, not symbionts, indicating that the siRNA treatment was affecting host cells only (Fig. 3C and Supplementary Fig. 5B). Interestingly, we did not observe an increase in ROS release in symbiotic cells during heat stress, suggesting they may have an improved antioxidant defense system at work. Our results also show a higher ROS-derived signal in host cells (including symbiotic cells) compared to symbiont cells. It may indicate that animal cells are the major source of ROS in the symbiosis, but these intercellular differences can also be explained by the differential permeability of a cell wall and a cellular membrane between an animal and a plant cell or by the difference in size between the cells (the ROS signal is normalized by the area).
We previously showed that BI-1-mediated upregulation of glutathione reductase, a ubiquitous cellular antioxidant, improves oxidative DNA damage defense, possibly via the Nrf2/ARE pathway in preconditioned (thus thermally more resilient) P. acuta13. Similarly, here, we also found a strikingly high correlation between pa-BI-1 and pa-GR expressions in heat stress (Fig. 2C and Supplementary Table 2), confirming this previously established relationship. However, neither of the antioxidant genes analyzed in this study, Glutathione reductase (GR) and Heme oxygenase 1 (HO-1), were differentially expressed in BAK-downregulated corals (Fig. 2B). We hypothesize that decreased oxidative stress in the cytoplasm of host cells is not caused by improved ROS scavenging but rather by reduced release of ROS from mitochondria.
Many proteins of the Bcl family (including BAK and BAX) have different non-apoptotic functions, often related to mitochondrial structure and functioning57. In normal conditions, BAK – unlike BAX – is localized inside mitochondria with its C-terminal region inserted in the lipid bilayer58. The observed phenotypic advantage of pa-BAK downregulation could be due to the changes in the pool of “available” single BAK protein inserted in the mitochondrial membrane, which is necessary for the formation of mitochondrial permeability transition pores (mPTP)59. Their regular openings help maintain healthy mitochondrial homeostasis through control of the release of excessive ROS to the cytoplasm, where it affects stress signaling and oxidative damage response pathways59,60.
In some cases, released ROS triggers the formation of mPTP and subsequent ROS release from neighboring mitochondria, activating a positive feedback loop called ROS-induced ROS release (RIRR), which plays a strategic role in extensive signaling and may lead to oxidative cellular damage and cell death via a PCD pathway59. The loss of BAK and BAX causes changes in the mPTP permeability, prevents mitochondrial swelling, and the cells resist cell death, but the inner membrane still undergoes stress-related reorganization60. We hypothesize that with less mPTP in mitochondria, cells in pa-BAK downregulate corals that accumulate less cellular ROS due to the impaired mitochondrial ROS homeostasis mechanism. It is thus important to further study possible long-term outcomes of such forced survival on coral mitochondrial and cellular health. Besides energy production, mitochondria play a vital role in intra- and extracellular communication, and their impaired function is connected to a multitude of symptoms and degenerative diseases (reviewed in ref. 61).
We found the expression of pa-foxo3 positively correlated with pa-AMPK, pa-HO-1, and pa-Bcl-2 (Fig. 2C and Supplementary Table 2). AMPK is a serine/threonine kinase that plays a major role in promoting mitochondrial health and is involved in various aspects of mitochondrial homeostasis (reviewed in ref. 62). It can be (probably indirectly) activated by mitochondria-derived ROS63. Foxo3a, a potent transcription factor, is linked to the modulation of stress responses upon oxidative stress, DNA damage, or nutrient shortage (reviewed in ref. 64). Its activity is primarily controlled by post-translational modifications, which determine subcellular localization. Heme oxygenase 1 (HO-1) is an inducible form of heme oxygenase enzyme that catalyzes heme breakdown, with byproducts that possess anti-inflammatory, anti-apoptotic, and antioxidant properties and a truncated form of HO-1 with low enzymatic activity acts as a transcriptional factor in the nucleus during stress response (reviewed in ref. 65). Bcl-2 expression is often controlled by the Nrf2 pathway66, but overexpression of Heme oxygenase 1 alone is enough to induce the expression of Bcl-2 in a rat heart, suggesting HO-1 might be an intermediary in the regulation67.
AMPK activates Foxo3a mostly as a response to nutrient stress, leading to autophagy-related gene transcription64. Lower expression of pa-AMPK (Fig. 2B) could lead to decreased pa-foxo3 function in pa-BAK-knockdown corals, which would lead to a lower expression of autophagy/symbiophagy-related genes and delayed bleaching. We previously identified symbiophagy rather than apoptosis as the probable pathway involved in thermal resilience in P. acuta12. AMPK has also been proposed to boost the Nrf2-mediated HO-1 expression68, and Foxo3a putatively regulates the keap1/Nrf2 pathway, activating or repressing it based on specific conditions69. Foxo3a overexpression promotes the expression of HO-1 and other antioxidant enzymes in human lymphocytes70.
Additionally, pa-HO-1 strongly correlated with pa-OGG1 gene expression (Fig. 2C). Besides its role in DNA damage repair, OGG1 can also induce gene transcription, although the mechanisms remain largely unclear56. It is thus possible that one protein regulates the expression of the other gene or that they are both controlled by a third party. HO-1 knockdown leads to downregulation of OGG1 in irradiated mice lungs71; however, studies also show that both HO-1 and OGG1 are positively and directly regulated by the Nrf2 factor72,73. Furthermore, HO-1 can regulate the expression of Nrf2 in what appears to be a positive feedback loop65, which points to a complex regulatory network surrounding the Nrf2 stress–response pathway. Nrf2 is a pleiotropic factor, and it may thus play a major role in acclimatization to environmental stress in corals with improved symbiosis stability, impacting multiple important defense mechanisms in the organism, a hypothesis we already proposed in our previous research12.
Algal symbionts co-exist within the host cell in symbiosomes, which are believed to be late lysosomes in the state of arrested phagocytosis74. LC3 is often analyzed as a proxy for the amount and/or size of autophagosomes in the cell, as it is incorporated into the membrane of newly emerging phagosomes during the initialization phase75. Here, we observed a non-significant trend of pa-LC3 overexpression in pa-BAK-knockdown corals, which may indicate increased symbiont stability or enhanced symbiont turnover during acute heat stress75, but this hypothesis requires further investigation.
pa-GABARAP, another marker of phagosomes, did not respond to the treatment, but its expression was strongly correlated with pa-BI-1 and pa-mTOR. Research in model organisms suggests that GABARAP directly regulates lysosomal and autophagosomal biogenesis by inhibiting the AKT/mTOR pathway, which is in contradiction to our observations76. However, this regulation acts on the protein level through post-translational modifications and cellular translocation machinery, while we analyzed the mRNA levels. We did not find any literature discussing the functional interaction between GABARAP and BI-1, and the observed correlation may be coincidental.
Given the results of this study, we propose that the main pro-life and antioxidant genes (pa-GR, pa-BI-1, pa-Bcl-2, and pa-HO-1) are regulated through the AMPK/Foxo3a/Nrf2 signaling in heat-stressed Pocillopora acuta. In our model (Fig. 4), heat-stressed cells release higher mitochondrial ROS, which activate AMPK and trigger the AMPK/Foxo3a/Nrf2 stress signaling. The AMPK/Foxo3a pathway is ambiguous. It may activate or inhibit the same downstream pathways based on the particular conditions in the cell, for example, the level of mitochondrial ROS release69,77. It could also boost the antioxidant defense system as a response to short-term acute stress while triggering programmed cell death pathways during long-term stress, leading to delayed PCD pathways. Since BAK-depleted host cells decrease mitochondrial ROS release due to the possible loss of mPTP, AMPK activation and subsequent signaling for PCD pathways are weaker, leading to delayed bleaching. The possible downside of this apparent beneficial phenotype may be permanent dysregulation of mitochondrial homeostasis and accumulation of mitochondrial damage. These additional questions must be resolved to determine if the siBAK phenotype can be truly deemed beneficial for corals in the long-term.
pa-BAK is inserted into the mitochondrial membrane, where it contributes to the formation of the mitochondrial permeability transition pore (mPTP). This pore allows reactive oxygen species (ROS) to exit into the cytoplasm, helping to prevent mitochondrial damage. Under heat stress (A), the resulting excess of ROS activates the ROS-sensitive AMPK/Foxo3a signaling pathway, which initially triggers the expression of antioxidant genes via the Nrf2/ARE pathway. However, prolonged activation may shift the response toward programmed cell death and increased organismal stress, ultimately contributing to coral bleaching through the loss of symbiotic algae. In contrast, corals with downregulated pa-BAK (B) exhibit reduced mPTP formation in the mitochondrial membrane, limiting the release of ROS into the cytoplasm. This altered ROS signaling results in a different pattern of AMPK/Foxo3a pathway activation and appears to delay or reduce the bleaching response under heat stress.
In our experiment, we did not observe treatment-related changes in ROS accumulation in cells distinguished as “single symbiont in a host cell.” This may be due to the size of these cells compared to non-symbiotic cells; the difference in mitochondrial ROS release in BAK-depleted cells may be lost in the overall signal-to-area ratio used to quantify the microscopy signal. It is also possible that ROS homeostasis in symbiotic cells is altered, but stress response in the neighboring coral host (non-symbiotic) tissue and the animal as a whole makes the holobiont metabolic environment hostile for algal survival, and symbionts leave the coral by themselves, independently of the host, as discussed in ref. 5.
Additionally, we cannot exclude that the AMPK/Foxo3a regulation is independent of the putative Nrf2-mediated control of pa-GR, pa-BI-1, pa-Bcl-2, and pa-HO-1. Therefore, their correlation would not be causal but would exist in parallel, and the two major signaling pathways—Nrf2 and AMPK would act independently in heat-stressed corals. This hypothetical ambiguity needs to be further examined using the tools of functional genetics to overcome the limitations of describing fine-tuned signaling of cellular response pathways that are primarily regulated via post-translational modifications rather than gene expression.
Conclusion
This work demonstrates that corals can become more resilient during acute heat stress via targeted manipulation of a single gene. It also shows that corals modulate various pathways as a response to single gene manipulation, probably to cope with possible harmful trade-offs of a prolonged symbiosis in heat stress. Our experiment focused on a single timepoint, thus, the correlation analysis is unable to capture the dynamic nature of signaling pathways, and while some correlations may be spurious, we may also be missing other interactions that occur outside of our chosen timeframe.
These data shed light on the network of regulatory pathways involved in coral response to acute heat stress. Our results are crucial for assisted evolution approaches to conservation and restoration that target more resilient individuals and use them to support the health of coral reefs facing anthropogenic climate change8,78.
Data availability
All data needed to evaluate the conclusions in the paper are present in the paper and/or supplied as Supplementary Data.
References
Mauritsen, T. & Pincus, R. Committed warming inferred from observations. Nat. Clim. Change 7, 652–655 (2017).
Brown, B. E. Coral bleaching: causes and consequences. Coral Reefs 16, S129–S138 (1997).
Fox, M. D. et al. Gradients in primary production predict trophic strategies of mixotrophic corals across spatial scales. Curr. Biol. 28, 3355–3363.e4 (2018).
Roth, M. S. The engine of the reef: photobiology of the coral–algal symbiosis. Front. Microbiol. 5, 422 (2014).
Matz, M. V. Not-so-mutually beneficial coral symbiosis. Curr. Biol. 34, R798–R801 (2024).
Knowlton, N. et al. Rebuilding Coral Reefs: A Decadal Grand Challenge. https://coralreefs.org/publications/rebuilding_coral_reefs/ (ICRS, 2021).
Drury, C. Resilience in reef-building corals: the ecological and evolutionary importance of the host response to thermal stress. Mol. Ecol. 29, 448–465 (2020).
Caruso, C., Hughes, K. & Drury, C. Selecting heat-tolerant corals for proactive reef restoration. Front. Mar. Sci. 8, 632027 (2021).
Ritson-Williams, R. & Gates, R. D. Coral community resilience to successive years of bleaching in Kāne’ohe Bay, Hawai’i. Coral Reefs 39, 757–769 (2020).
Yadav, S. et al. Fine-scale variability in coral bleaching and mortality during a marine heatwave. Front. Mar. Sci. 10, 1108365 (2023).
Jury, C. P. et al. Experimental coral reef communities transform yet persist under mitigated future ocean warming and acidification. Proc. Natl Acad. Sci. USA 121, e2407112121 (2024).
Majerova, E., Carey, F. C., Drury, C. & Gates, R. D. Preconditioning improves bleaching tolerance in the reef-building coral Pocillopora acuta through modulations in the programmed cell death pathways. Mol. Ecol. 30, 3560–3574 (2021).
Majerová, E. & Drury, C. Thermal preconditioning in a reef-building coral alleviates oxidative damage through a BI-1-mediated antioxidant response. Front. Mar. Sci. 9, 971332 (2022).
Helgoe, J., Davy, S. K., Weis, V. M. & Rodriguez-Lanetty, M. Triggers, cascades, and endpoints: connecting the dots of coral bleaching mechanisms. Biol. Rev. 99, 715–752 (2024).
Pernice, M. et al. Regulation of apoptotic mediators reveals dynamic responses to thermal stress in the reef building coral Acropora millepora. PLoS ONE 6, e16095 (2011).
Downs, C. A. et al. Symbiophagy as a cellular mechanism for coral bleaching. Autophagy 5, 211–216 (2009).
Dunn, S. R., Schnitzler, C. E. & Weis, V. M. Apoptosis and autophagy as mechanisms of dinoflagellate symbiont release during cnidarian bleaching: every which way you lose. Proc. R. Soc. B Biol. Sci. 274, 3079–3085 (2007).
Fujise, L., Yamashita, H., Suzuki, G. & Koike, K. Expulsion of zooxanthellae (Symbiodinium) from several species of scleractinian corals: comparison under non-stress conditions and thermal stress conditions. Galaxea J. Coral Reef. Stud. 15, 29–36 (2013).
Kvitt, H., Rosenfeld, H. & Tchernov, D. The regulation of thermal stress induced apoptosis in corals reveals high similarities in gene expression and function to higher animals. Sci. Rep. 6, 30359 (2016).
Tchernov, D. et al. Apoptosis and the selective survival of host animals following thermal bleaching in zooxanthellate corals. Proc. Natl Acad. Sci. 108, 9905–9909 (2011).
Galluzzi, L. et al. Molecular mechanisms of cell death: recommendations of the Nomenclature Committee on Cell Death 2018. Cell Death Differ. 25, 486–541 (2018).
Karch, J. et al. Autophagic cell death is dependent on lysosomal membrane permeability through Bax and Bak. eLife 6, e30543 (2017).
Denton, D. & Kumar, S. Autophagy-dependent cell death. Cell Death Differ. 26, 605–616 (2019).
Ieranò, C. et al. Loss of the proteins Bak and Bax prevents apoptosis mediated by histone deacetylase inhibitors. Cell Cycle 12, 2829–2838 (2013).
Jang, H.-S. & Padanilam, B. J. Simultaneous deletion of Bax and Bak is required to prevent apoptosis and interstitial fibrosis in obstructive nephropathy. Am. J. Physiol. Ren. Physiol. 309, F540–F550 (2015).
Hemmati, P. G. et al. Bak functionally complements for loss of Bax during p14ARF-induced mitochondrial apoptosis in human cancer cells. Oncogene 25, 6582–6594 (2006).
Iyer, S. et al. Robust autoactivation for apoptosis by BAK but not BAX highlights BAK as an important therapeutic target. Cell Death Dis. 11, 268 (2020).
Brooks, C. et al. Bak regulates mitochondrial morphology and pathology during apoptosis by interacting with mitofusins. Proc. Natl Acad. Sci. USA 104, 11649–11654 (2007).
Bay, R. A. & Palumbi, S. R. Rapid acclimation ability mediated by transcriptome changes in reef-building corals. Genome Biol. Evol. 7, 1602–1612 (2015).
Bellantuono, A. J., Granados-Cifuentes, C., Miller, D. J., Hoegh-Guldberg, O. & Rodriguez-Lanetty, M. Coral thermal tolerance: tuning gene expression to resist thermal stress. PLoS ONE. 7, e50685 (2012).
Nesa, B., Hidaka, M. & editors. Thermal stress increases oxidative DNA damage in coral cell aggregates. Proceedings of the 11th International Coral Reef Symposium, Ft. Lauderdale, Florida, 7–11 (2008).
Yakovleva, I. et al. Algal symbionts increase oxidative damage and death in coral larvae at high temperatures. Mar. Ecol. Prog. Ser. 378, 105–112 (2009).
Szabó, M., Larkum, A. W. D. & Vass, I. in Photosynthesis in Algae: Biochemical and Physiological Mechanisms (eds. Larkum, A. W. D., Grossman, A. R. & Raven, J. A.) Vol. 45, 459–488 (Springer International Publishing, Cham, 2020).
Huffmyer, A. S., Drury, C., Majerová, E., Lemus, J. D. & Gates, R. D. Tissue fusion and enhanced genotypic diversity support the survival of Pocillopora acuta coral recruits under thermal stress. Coral Reefs 40, 447–458 (2021).
Huffmyer, A. S., Matsuda, S. B., Eggers, A. R., Lemus, J. D. & Gates, R. D. Evaluation of laser scanning confocal microscopy as a method for characterizing reef-building coral tissue thickness and Symbiodiniaceae fluorescence. J. Exp. Biol. 223, jeb220335 (2020).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Allen-Waller, L. R., Jones, K. G., Martynek, M. P., Brown, K. T. & Barott, K. L. Comparative physiology reveals heat stress disrupts acid-base homeostasis independent of symbiotic state in the model Cnidarian Exaiptasia diaphana. J. Exp. Biol. 227, jeb246222 (2024).
Roger, L. M. et al. Evaluation of fluorescence-based viability stains in cells dissociated from scleractinian coral Pocillopora damicornis. Sci. Rep. 12, 15297 (2022).
He, S., Crans, V. L. & Jonikas, M. C. The pyrenoid: the eukaryotic CO2-concentrating organelle. Plant Cell 35, 3236–3259 (2023).
R Core Team. R: A Language and Environment for Statistical Computing. (R Foundation for Statistical Computing, 2020).
Bates, D., Mächler, M., Bolker, B. & Walker, S. Fitting linear mixed-effects models using lme4. J. Stat. Softw. 67, 1–48 (2015).
Lenth, R. V. emmeans: Estimated Marginal Means, Aka Least-squares Means. 1.10.5 https://doi.org/10.32614/CRAN.package.emmeans (2017).
Hothorn, T., Bretz, F. & Westfall, P. multcomp: Simultaneous Inference in General Parametric Models. 1.4–26 https://doi.org/10.32614/CRAN.package.multcomp (2002).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis. (Springer International Publishing: Imprint: Springer, Cham, 2016).
Wilke, C. O. & Wiernik, B. M. ggtext: Improved Text Rendering Support for ‘ggplot2’. The R Foundation https://doi.org/10.32614/cran.package.ggtext (2020).
Wilke, C. O. cowplot: Streamlined Plot Theme and Plot Annotations for ‘ggplot2’. The R Foundation https://doi.org/10.32614/cran.package.cowplot (2015).
Garnier, S. viridis: Colorblind-friendly Color Maps for R. The R Foundation https://doi.org/10.32614/cran.package.viridis (2015).
Jackson, A. L. et al. Expression profiling reveals off-target gene regulation by RNAi. Nat. Biotechnol. 21, 635–637 (2003).
Weis, V. M. Cellular mechanisms of Cnidarian bleaching: stress causes the collapse of symbiosis. J. Exp. Biol. 211, 3059–3066 (2008).
Lindqvist, L. M. et al. Autophagy induced during apoptosis degrades mitochondria and inhibits type I interferon secretion. Cell Death Differ. 25, 784–796 (2018).
Baker, A. C. & Cunning, R. in Diseases of Coral (eds. Woodley, C. M., Downs, C. A., Bruckner, A. W., Porter, J. W. & Galloway, S. B.) 396–409 (John Wiley & Sons, Inc, Hoboken, N. J, 2015).
Wei, M. C. et al. Proapoptotic BAX and BAK: a requisite gateway to mitochondrial dysfunction and death. Science 292, 727–730 (2001).
Palumbi, S. R., Barshis, D. J., Traylor-Knowles, N. & Bay, R. A. Mechanisms of reef coral resistance to future climate change. Science 344, 895–898 (2014).
Thomas, L. et al. Mechanisms of thermal tolerance in reef-building corals across a fine-grained environmental mosaic: lessons from Ofu, American Samoa. Front. Mar. Sci. 4, 434 (2018).
Drury, C., Dilworth, J., Majerová, E., Caruso, C. & Greer, J. B. Expression plasticity regulates intraspecific variation in the acclimatization potential of a reef-building coral. Nat. Commun. 13, 4790 (2022).
Wang, R., Hao, W., Pan, L., Boldogh, I. & Ba, X. The roles of base excision repair enzyme OGG1 in gene expression. Cell. Mol. Life Sci. 75, 3741–3750 (2018).
Gross, A. & Katz, S. G. Non-apoptotic functions of BCL-2 family proteins. Cell Death Differ. 24, 1348–1358 (2017).
Peña-Blanco, A. & García-Sáez, A. J. Bax, Bak and beyond—mitochondrial performance in apoptosis. FEBS J. 285, 416–431 (2018).
Zorov, D. B., Juhaszova, M. & Sollott, S. J. Mitochondrial reactive oxygen species (ROS) and ROS-induced ROS release. Physiol. Rev. 94, 909–950 (2014).
Karch, J. et al. Bax and Bak function as the outer membrane component of the mitochondrial permeability pore in regulating necrotic cell death in mice. eLife 2, e00772 (2013).
Shen, K. et al. Mitochondria as cellular and organismal signaling hubs. Annu. Rev. Cell Dev. Biol. 38, 179–218 (2022).
Herzig, S. & Shaw, R. J. AMPK: guardian of metabolism and mitochondrial homeostasis. Nat. Rev. Mol. Cell Biol. 19, 121–135 (2018).
Hinchy, E. C. et al. Mitochondria-derived ROS activate AMP-activated protein kinase (AMPK) indirectly. J. Biol. Chem. 293, 17208–17217 (2018).
Fasano, C., Disciglio, V., Bertora, S., Lepore Signorile, M. & Simone, C. FOXO3a from the nucleus to the mitochondria: a round trip in cellular stress response. Cells 8, 1110 (2019).
Costa Silva, R. C. M. & Correa, L. H. T. Heme oxygenase 1 in vertebrates: friend and foe. Cell Biochem. Biophys. 80, 97–113 (2022).
Niture, S. K. & Jaiswal, A. K. Nrf2 protein up-regulates antiapoptotic protein Bcl-2 and prevents cellular apoptosis. J. Biol. Chem. 287, 9873–9886 (2012).
Katori, M. et al. Heme oxygenase-1 overexpression protects rat hearts from cold ischemia/reperfusion injury via an antiapoptotic pathway1. Transplantation 73, 287–292 (2002).
Zimmermann, K. et al. Activated AMPK boosts the Nrf2/HO-1 signaling axis—a role for the unfolded protein response. Free Radic. Biol. Med. 88, 417–426 (2015).
Klotz, L.-O. & Steinbrenner, H. Cellular adaptation to xenobiotics: Interplay between xenosensors, reactive oxygen species and FOXO transcription factors. Redox Biol. 13, 646–654 (2017).
Xu, S. et al. FOXO3a alleviates the inflammation and oxidative stress via regulating TGF-β and HO-1 in ankylosing spondylitis. Front. Immunol. 13, 935534 (2022).
Yang, G. et al. Heme oxygenase-1 regulates postnatal lung repair after hyperoxia: role of β-catenin/hnRNPK signaling. Redox Biol. 1, 234–243 (2013).
Loboda, A., Damulewicz, M., Pyza, E., Jozkowicz, A. & Dulak, J. Role of Nrf2/HO-1 system in development, oxidative stress response and diseases: an evolutionarily conserved mechanism. Cell. Mol. Life Sci. 73, 3221–3247 (2016).
Shang, Q. et al. Nuclear factor Nrf2 promotes glycosidase OGG1 expression by activating the AKT pathway to enhance leukemia cell resistance to cytarabine. J. Biol. Chem. 299, 102798 (2023).
Davy, S. K., Allemand, D. & Weis, V. M. Cell biology of Cnidarian-Dinoflagellate symbiosis. Microbiol. Mol. Biol. Rev. 76, 229–261 (2012).
Klionsky, D. J. et al. Guidelines for the use and interpretation of assays for monitoring autophagy (3rd edition). Autophagy 12, 1–222 (2016).
Kumar, S. et al. Mammalian Atg8-family proteins are upstream regulators of the lysosomalsystem by controlling MTOR and TFEB. Autophagy 16, 2305–2306 (2020).
Agostini, F., Bisaglia, M. & Plotegher, N. Linking ROS levels to autophagy: the key role of AMPK. Antioxidants 12, 1406 (2023).
van Oppen, M. J. H., Oliver, J. K., Putnam, H. M. & Gates, R. D. Building coral reef resilience through assisted evolution. Proc. Natl Acad. Sci. 112, 2307–2313 (2015).
Gregg, T. M., Mead, L., Burns, J. H. R. & Takabayashi, M. in Ethnobiology of Corals and Coral Reefs (eds. Narchi, N. & Price, L. L.) 103–115 (Springer International Publishing, Cham, 2015).
Acknowledgements
We acknowledge the profound connection between Native Hawaiian culture and coral reefs79, including those found in Kāneʻohe Bay, Oʻahu, Hawaiʻi, USA, where this research was conducted. We hope to honor the unique relationship between local communities and place and the practice of mālama ʻāina (caring for and protecting the land). We would like to thank Kira Hughes, Khalil Smith, and Joshua Hancock for assistance with coral husbandry, experimental setup and project administration and the Coral Resilience Lab for helping with coral collection and maintenance. This work was funded by the National Science Foundation (IOS-2041401) and the Paul G. Allen Family Foundation. Corals were collected under permit SAP 2023-31. This is HIMB contribution #1997 and SOEST contribution #11959.
Author information
Authors and Affiliations
Contributions
E.M. conceived the experiments, conducted research, analyzed and visualized data and wrote the manuscript. C.S. conducted research, visualized data, and wrote the manuscript. C.D. analyzed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Pratima Chennuri, Lyuba Chechik and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary handling editor: Michele Repetto. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Majerová, E., Steinle, C. & Drury, C. BAK knockdown delays bleaching and alleviates oxidative DNA damage in a reef-building coral. Commun Biol 8, 1216 (2025). https://doi.org/10.1038/s42003-025-08671-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-08671-y