Abstract
The formation of the bilaterian anterior–posterior axis relies on deeply conserved patterning systems, yet how these evolved remains incompletely understood. As sister to or within Gnathifera, chaetognaths provide an informative phylogenetic context for investigating anterior–posterior patterning, yet their developmental genetics remain poorly studied. Here, we examine anterior patterning (otx, nk2.1, six3/6) and Hox gene expression in the chaetognath Spadella cephaloptera. Anterior patterning genes are expressed in cerebral neural regions as in other bilaterians. In contrast, Hox genes, including the previously undescribed med6, postC, and postD, show staggered expression along the nervous system, resembling the proposed ancestral bilaterian condition. Posterior Hox genes are also expressed in the postanal tail, a trait only present in chaetognaths and chordates, suggesting a link between Hox expansion and the independent emergence of this structure. Our results indicate that chaetognaths retain ancestral bilaterian features and provide insights into Hox-driven lineage-specific innovations within Spiralia.
Similar content being viewed by others
Introduction
The anterior–posterior (AP) axis is a fundamental organizational feature of bilaterian body plans, established by conserved transcription factors that establish regional identity during development. Anterior-specific genes (e.g., otx, nk2.1, six3/6, foxQ2) specify anterior neural and head structures, while Hox genes control positional identities along the AP axis (e.g.1,2,3,4). In many bilaterians, these genes also regulate neural regionalization through staggered expression in the developing central nervous system (CNS), indicating a conserved mechanism of CNS patterning across lineages5,6,7,8,9. While these mechanisms are well described in model systems such as vertebrates, arthropods, and annelids, our understanding of how these genes contribute to AP axis formation across the broader diversity of bilaterians remains incomplete.
Spiralians have historically been underrepresented in studies of Hox gene expression. However, recent research on annelids, molluscs, and other spiralian lineages has begun to uncover a variety of Hox expression profiles. Several lophotrochozoans, including some annelids and molluscs, exhibit canonical staggered Hox expression along the AP axis, particularly in the developing nervous system and non-neural ectoderm10,11,12,13,14. Others, however, display more restricted or non-staggered expression profiles, with Hox gene expression confined to particular tissues or developmental stages15,16,17,18,19. This variability highlights both conserved and divergent aspects of Hox function across Spiralia, but drawing broader evolutionary conclusions remains difficult due to limited taxon sampling, especially among underrepresented but phylogenetically informative lineages that appear to have maintained aspects of the ancestral body plan.
Chaetognaths (arrow worms) offer a particularly compelling model for comparative developmental studies. These marine invertebrates occupy a contentious phylogenetic position due to their mosaic of morphological and molecular traits, some of which resemble those of deuterostomes, others of protostomes, or suggest they form a distinct lineage altogether20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35. Recent phylogenomic analyses suggest that chaetognaths are either nested within Gnathifera or as a sister lineage to the group24,27. This phylogenetic positioning offers a unique opportunity to infer ancestral spiralian features and their roles in the evolution of bilaterian body plans.
Chaetognaths share several developmental and morphological features with rotifers (Monogononta, Bdelloidea, and Seisonidea), including the absence of spiral cleavage, the specification of primordial germ cells (PGCs) through preformation, a chitinous feeding apparatus, a trunk without external motile cilia, a postanal structure (tail/foot), and the central-post class Hox gene, medpost16. These shared traits support the proposed evolutionary link between chaetognaths and gnathiferans. Despite this phylogenetic proximity, chaetognaths retain a full complement of anterior, central, and posterior Hox genes31,36, in contrast to rotifers, which have a highly reduced Hox repertoire and entirely lack posterior-class genes16,37. Nevertheless, a comprehensive understanding of AP axis patterning in chaetognaths remains limited compared to other spiralians. Earlier chaetognath studies have examined the Hox complement of Spadella cephaloptera and Flaccisagitta enflata31,36, revealing a repertoire, although incomplete, that broadly resembles that of other bilaterians. Papillon et al.38 first reported expression of a central-class Hox gene in the ventral nerve center (VNC), suggesting a possible neural role. By comparison, in the rotifer Brachionus manjavacas, Hox gene expression is restricted to developing neural tissues and appears only partially staggered16. For Gnathifera, expression of anterior patterning genes nk2.1 and pax6 has been described in the brain of the juvenile rotifer Epiphanes senta39, and pax6 is also expressed in the anterolateral region of the rotifer Brachionus plicatilis during embryogenesis, although the exact tissue identity remains uncertain40. These differences, combined with limited data available, underscore the need for comparative developmental studies in chaetognaths to clarify whether AP patterning in this lineage reflects conserved spiralian and/or bilaterian mechanisms, or represents a distinct innovation within chaetognaths.
While this manuscript was under revision and already uploaded to BioRxiv41, a chromosome-level genome of Paraspadella gotoi was published, revealing an expanded, physically linked Hox cluster shaped by local duplication and partially staggered expression for a subset of genes in the early juvenile VNC42. This study provides a valuable genomic context for chaetognath Hox organization but leaves expression coverage incomplete. Here, we examine the expression profiles of the AP-patterning genes otx, nk2.1, and six3/6 in the chaetognath S. cephaloptera during embryonic and early post-hatch development, providing the most extensive spatiotemporal data to date on anterior-patterning genes (otx, nk2.1, six3/6) and broad coverage of the Hox repertoire in a chaetognath. We show that AP-patterning genes are expressed in the cerebral ganglion and developing head, consistent with their conserved role in brain and anterior tissues specification across Bilateria. We also describe an updated Hox gene complement for S. cephaloptera, including several previously undescribed members, and demonstrate staggered Hox expression along the AP axis of the VNC and postanal tail. These spatially organized expression patterns suggest that chaetognaths retain core bilaterian mechanisms of CNS patterning, which appear to have been secondarily lost in several other bilaterian lineages, despite their peculiar developmental traits and uncertain phylogenetic position.
Methods
Identification of gene homologs
We retrieved AP patterning (otx, six1/3, nk2.1, Hox) genes from the S. cephaloptera draft transcriptome43,44 using blastx searches45 against protein sequences from the NCBI GenBank non-redundant protein database. We also searched for gastrulation brain homeobox (gbx) and forkhead homeobox Q2 (foxQ2), two transcription factors involved in axial patterning and neural development in many bilaterians3,13,14, but found no gbx and foxQ2 homologs in the S. cephaloptera transcriptome. To determine gene orthology, phylogenetic analyses were conducted using reference protein sequences from a broad range of bilaterians (Supplementary Table 1). Additionally, Hox genes from selected chaetognath species, inferred from transcriptomic data, were also incorporated in the phylogenetic reconstruction. Briefly, raw RNA-seq reads from several chaetognath species generated by Marlétaz et al.27 were downloaded from the GenBank Sequence Read Archive. Raw reads of the rotifer Rotaria rotatoria were also included. Adapter removal, quality filtering, and trimming were performed using Trimmomatic (v0.39)46, and transcriptome assemblies were generated using rnaSPAdes (v4.1)47, both with default parameters. The resulting transcriptomes were screened for Hox genes using DIAMOND blastx (v2.1.11)48 against a custom database of protostome Hox gene sequences used in this study, applying an e-value threshold of 1e-20. SRA accession numbers used in the analysis are provided in Supplementary Table 2 and the resulting Hox sequences from the assemblies are presented in Supplementary Note.
Amino acid sequences were aligned using MAFFT (v7.490)49,50, implemented in Geneious Prime (v2023.0.1). The resulting multi-sequence alignments were then trimmed with ClipKit (v2.2.3)51. Maximum likelihood (ML) analyses were performed in IQTREE (v2.3.6), incorporating ModelFinder for amino-acid substitution model selection and conducting 1000 ultrafast bootstrap replicates52,53,54, along with SH-aLRT test replicates for Hox genes55. Substitution models used in phylogenetic analyses are indicated in Supplementary Table 2. Additionally, phylogenetic reconstruction for Hox genes was also performed using Bayesian inference in MrBayes (v3.2.7a)56, employing the same substitution model as the ML analysis (i.e., VT + G + F). The analysis was run for 50 million generations, sampling every 1000 generations across eight independent runs, each with four chains. Convergence was assessed using multiple diagnostics in MrBayes: Average Standard Deviation of Split Frequencies (ASDSF), Potential Scale Reduction Factor (PSRF), and Effective Sample Size (ESS). While ASDSF of 0.0206 is slightly above the recommended 0.01 threshold, ESS values exceeded 500 across parameters, and PSRF remained approximately 1.000 for all numeric parameters. This follows standard recommendations for Bayesian phylogenetic inference indicating robust convergence despite a minor ASDSF deviation57,58. The first 25% of sampled generations were discarded as burn-in prior to calculating the consensus tree.
Animal collection
Live adult specimens of S. cephaloptera (Busch, 1851) were collected in June 2024 during low tide from the intertidal zone off Roscoff, France (48°43'47.2“N 3°59'12.2“W). Specimens were obtained by sweeping hand nets and plankton nets through the shallow algal meadows and were subsequently housed in aquaria in the aquatic research facility of the University of Vienna. The aquaria contain a combination of natural and artificial seawater, with a temperature and salinity of 14°C and 35‰, respectively. Sexually matured individuals were transferred to plastic Petri dishes (14.1 cm radius), which were refreshed with natural sea water every day. The animals were fed with Artemia sp. nauplii until they laid eggs. Embryos and hatchlings (8–20 h post-hatching) were manually collected. Additionally, some hatchlings were allowed to develop to early juveniles (7–10 days post-hatching) before they were sampled.
All specimens were fixed in 4% paraformaldehyde in a buffer containing 0.1 M MOPS pH 7.4, 2 mM EGTA, 1 mM MgSO4, 2.5 M NaCl. The specimens used for whole-mount RNA Fluorescence in situ Hybridization (RNA-FISH) were fixed overnight (14–18 h) at 4 °C, while for whole-mount Hybridization Chain Reaction (HCR-FISH)59,60,61, they were fixed for only 1–1.5 h. Following fixation, the specimens were washed thrice for five minutes each in PBTw (1x PBS, pH 7.5 with 0.1% Tween-20), washed three times in ice-cold methanol for 10 minutes each, and then stored in 100% methanol at −20°C until use.
Sequencing, riboprobe synthesis, and whole-mount fluorescent in situ hybridization (FISH)
RNA extracted from hatchlings and adults using RNAqueous™-Micro Total RNA Isolation Kit (Invitrogen GmbH, Karlsruhe, Germany) was used to generate a cDNA library with First Strand cDNA Synthesis Kit for RT-PCR (AMV) (Roche Molecular Biochemicals, Vienna, Austria). Riboprobes were generated either directly from PCR amplicons or from plasmid inserts. For PCR-derived riboprobes, reverse primers carried a 5’ T7 promoter sequence and resulting products were purified with QIAquick PCR Purification Kit (Qiagen Vertriebs GmbH, Vienna, Austria). For cloning, gene fragments amplified with PCR were cloned into pGEM-T Easy vectors (Promega, Mannheim, Germany), transformed into competent Escherichia coli cells, and then purified with QIAprep® Miniprep (Qiagen Vertriebs GmbH, Vienna, Austria). The purified fragments from both methods were sent to Microsynth Austria for Sanger sequencing to validate their specificity. Primer sequences and PCR conditions are listed in Supplementary Table 2. Digoxigenin (DIG)-labeled sense riboprobes from the PCR product and linearized DNA were generated using the T7 polymerase and digoxigenin RNA labeling mix kit (Roche Diagnostics, Mannheim, Germany). Total riboprobe concentrations were estimated using NanoDrop 2000 spectrophotometer (ThermoFisher Scientific, Massachusetts, USA). Traditional whole-amount fluorescent in situ hybridization (RNA-FISH) experiments on hatchlings were conducted following the procedure described in refs. 43,62. HCR-FISH59,63 was carried out with genes (Sce-otx, Sce-nk2.1, Sce-hox2, Sce-medpost, Sce-postB, Sce-postC, and Sce-postD) for which riboprobes failed to work with RNA-FISH. These probes were also used in embryos, as RNA-FISH is not effective with early embryonic stages62. HCR Probes for each gene were designed using the program insitu_probe_generator.py (https://github.com/rwnull/insitu_probe_generator64) and were purchased from Integrated DNA Technologies (München, Germany). Hairpins were purchased from Molecular Instruments, Inc. (California, USA). HCR-FISH was performed following Choi et al.59, with additional modifications from Bruce et al.65. Further adjustments were made for embryos, with dextran sulfate concentration reduced from 10% to 2% to minimize severe morphological distortion. Information on the PCR primer sequences and HCR probes is provided in Supplementary Table 3 and Supplementary Data 1, respectively. For hatchlings and juveniles, 2–3 individuals were analyzed per run across two independent runs (total n = 4–6 per stage). For embryos, 1–2 individuals were analyzed per run across two independent runs (total n = 2–4). An expression pattern was considered supported if it was observed in ≥3 hatchling/juvenile individuals and in ≥2 embryos.
Post-embryonic stages were incubated in Vectashield® (Vector Laboratories, California, USA) for 20 minutes before mounting with 70% 2,2’-thiodiethanol (TDE) in PBS. To minimize embryo deformation, specimens were incubated in a series of Vectashield®-PBS solutions (10%, 30%, 60%, 80%, and 100%) for 8 minutes at each concentration. Clearing was then achieved by titrating 100% TDE into 2 µL Vectashield containing the embryo in 0.5 µL increments at 5-min intervals (6–8 additions) until optically cleared. The specimens were then mounted in the solution used for documentation. Specimens were scanned with Leica TCS SP5 (Leica Microsystems, Heidelberg, Germany) confocal laser scanning microscope. Brightness/contrast adjustments and Z-projections were performed with Fiji66. Figure assembly and schematic drawings were prepared in Inkscape (https://inkscape.org).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Results
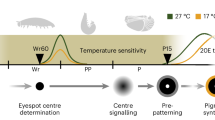
Staging overview for Spadella cephaloptera embryogenesis
Embryonic stages of S. cephaloptera were defined by morphology following John67, rather than time post-fertilization, because oviposition timing under laboratory conditions was unreliable (Fig. 1Ai-iv; Supplementary Fig. 1Ai–iv). Early gastrula: visible blastopore; no neuroectodermal thickening (Supplementary Fig. 1Ai). Mid gastrula: ventrolateral neuroectoderm emerges as a mitotically active thickening (Supplementary Fig. 1Aii). Late gastrula: axial elongation, intestinal invagination, blastopore closure, and further internalization of neuroectodermal cells (Supplementary Fig. 1Aiii). Early elongation: curvature along the AP axis, distinct head and tail buds, and aggregation of the neuroectodermal cells at the medioventral midline forming the nascent VNC (Supplementary Fig. 1Aiv). These landmarks frame the spatiotemporal gene expression analyses in the succeeding sections.
A Schematic representations of early (i), mid (ii), and late gastrula (iii) in ventral view, and early elongation stage (iv) in lateral view. B–G Expression patterns of Sce-otx (B), Sce-nk2.1 (C), Sce-med6 (D), Sce-medpost (E), Sce-postC (F), Sce-postD (G) across embryonic stages (i – iv). No expression data are available for Sce-postC at the mid gastrula stage. Purple outlines demarcate the neuroectoderm (ii, iii) and the developing ventral nerve center (iv). Scale bars: 50 μm. Encircled asterisk indicates the position of the blastopore. Orientation is indicated in the upper right corner of each panel. ect, ectoderm; eme, endomesoderm; hd, head bud; in, intestine; mes, mesoderm; nbs, neuroblast of the developing VNC; nec, neuroectoderm; tl, tail bud.
Gene orthology and expression patterns of anterior-patterning genes
We identified single-copy gene orthologs of otx, nk2.1, and six3/6 from the S. cephaloptera draft transcriptome by BLAST and ML phylogeny (Supplementary Figs. 2 and 3; see Ordoñez and Wollesen62 for Sce-nk2.1 orthology assignment).
Otx expression
In the early gastrula, Sce-otx is expressed in the anterior ectoderm and PGCs (Fig. 1Bi; Supplementary Fig. 1Bi). At the mid-gastrula stage, expression persists in the anterior ectoderm (Fig. 1Bii; Supplementary Fig. 1Bii). By the late gastrula stage, it extends to both the anterior and posterior regions of the neuroectoderm (Fig. 1Biii; Supplementary Fig. 1Biii). During early elongation, Sce-otx shows broad expression in the head bud and the polar tips of the lateral neuroblasts of the nascent VNC (Fig. 1Biv; Supplementary Fig. 1Biv).
In hatchlings, Sce-otx is broadly expressed in the head, with a strong signal concentrated in the anterior cerebral ganglion (Fig. 2B), presumptive ventral cephalic ganglia precursors (Fig. 2C), head epidermis (Fig. 2C–E), cephalic adhesive papillae (white arrowheads in Fig. 2B, D), and around the future mouth opening (asterisk in Fig. 2B, D). In the VNC, Sce-otx expression remains in the anterior and posterior tips of the lateral somata clusters and shows a bilateral streak pattern along the anterior inner cells (Fig. 2B, C). Early juveniles express Sce-otx throughout the head (Supplementary Fig. 4B–F), including the brain (dotted outline in Supplementary Fig. 4C), corona ciliata (dashed circle in Supplementary Fig. 4C), eyes (circles in Supplementary Fig. 4C), perioral epidermis (red arrowheads in Supplementary Fig. 4D, E), hood (Supplementary Fig. 4E), and cephalic epidermis (Supplementary Fig. 4B, C, E). In the trunk, Sce-otx is expressed in the lateral somata clusters (Supplementary Fig. 4B, D) and in the proliferating PGCs (red arrowheads in Supplementary Fig. 4F). PGC expression is detected at all stages except late gastrula and early elongation (Fig. 2E; red arrowheads in Supplementary Fig. 1Bi, ii and Supplementary Fig. 4F).
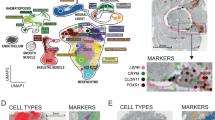
Gene transcripts (yellow) are visualized with AlexaFluor-647 (B–H) or AP-Fast Blue (I–L), and cell nuclei are counterstained with DAPI (purple). A, A’ General morphology of a hatchling shown in schematic drawing (A) and maximum intensity projection of DAPI-stained whole animal (A’). Top: dorsal view; bottom: lateral view. B–E Sce-otx expression pattern. B Dorsal and lateral maximum projections. C Dorsal section of the anterior body showing expression in the mid-anterior cerebral ganglion (cg) and presumptive ventral cephalic ganglia precursors (vcg). D Transverse section of the anterior head showing Sce-otx expression at the base of the head (red arrowheads) and in the cephalic adhesive papillae (ap, white arrowheads). E Higher magnification of primordial germ cells (pgc; red arrowheads), shown in DAPI only (left) and merged channels (right). F–H Sce-nk2.1 expression pattern. F Dorsal and lateral maximum projections. G Dorsal section of the anterior body showing expression in the cerebral ganglion and intestine (outlined). H Transverse section posterior to the cerebral ganglion showing expression in the anterior esophagus (es). I–L Sce-six3/6 expression pattern. I Dorsal and lateral maximum projections. J, L Dorsal sections of the anterior body showing expression in the cerebral ganglion and dorsal esophagus. K Dorsal section along the dorsal-most portion of the head showing Sce-six3/6+ cells in the eyes (solid circles) and inner corona ciliata (dashed circle). L Transverse section through the head along the esophagus, showing Sce-six3/6 expression in its dorsal domain. Scale bars: 50 μm, except panels B, F, and I (100 μm). Asterisk indicates the position of the mouth opening. Orientation is indicated in the top-right corner of each panel. ap cephalic adhesive papillae, cc corona cilata, cf caudal fin, cg cerebral ganglion, co ciliary tuft/fence organ, es esophagus, ey eye, in intestine, lms longitudinal muscle somata, lsc lateral somata clusters, mvs medioventral somata clusters, np neuropil of the ventral nerve center, pgc primordial germ cells, vcg ventral cephalic ganglia precursor.
Nk2.1 expression
Sce-nk2.1 expression is concentrated in the mid anterior cell population of the endomesoderm in early and mid gastrula (Fig. 1Ci, ii; Supplementary Fig. 1Ci, ii). In the late gastrula, these Sce-nk2.1+ cells extend posteriorly along the midline, a domain that will give rise to the future intestine (Fig. 1Ciii; Supplementary Fig. 1Ciii). A distinct anteroventral ectodermal domain also emerges, though its identity remains uncertain (inset iii’ in Supplementary Fig. 1Ciii). During early elongation stage, Sce-nk2.1 signal is detected in the mid-anterior head bud and in cells located in the developing tail, here referred to as tail bud (white arrowheads in Supplementary Fig. 1Civ).
At hatching, Sce-nk2.1 expression appears as a semi-arch, marking the anterior part of the developing brain (Fig. 2F – H) and outlines the digestive track, from the esophagus to the posterior gut, with the strongest signal intensity in the hindgut (Fig. 2F). In early juveniles, expression is observed in the anterior tip of the brain and in patches of neurons in its posterior domain (Supplementary Fig. 4G, H). Sce-nk2.1 remains active in the cells of the intestine at this stage (Supplementary Fig. 4G, I, J).
Six3/6 expression
Sce-six3/6 is mostly expressed in the head of hatchlings (Fig. 2I - L), including the cerebral ganglion, eyes, corona ciliata, anterodorsal epidermal cells, and the anterodorsal cells of the esophagus (Fig. 2J, L). The lateral somata clusters show a faint Sce-six3/6 expression (Fig. 2J). Sce-six3/6 expression in early juveniles is observed in the brain (Supplementary Fig. 4K, dotted outline in L), corona ciliata (dashed outline in Supplementary File 4 Fig. L), eyes (circles in Supplementary Fig. 4L), vestibular ganglia, esophageal ganglia (Supplementary Fig. 4M), and in the lateral somata clusters (Supplementary Fig. 4K, N).
The Spadella Hox gene complement
Chaetognaths are currently known to possess ten Hox genes representing five paralogous groups: the anterior class Hox gene hox1 (PG 1), hox3 (PG 3), the central class genes hox4–hox8 (PG 4-8), the central/posterior chimera medpost, and the posterior class genes postA and postB (PG 9-15)31,36. From the S. cephaloptera draft transcriptome, we recovered orthologs of previously reported Hox genes in chaetognaths using ML (Supplementary Fig. 5) and Bayesian phylogenetic analyses (Supplementary Fig. 6), while also identifying four previously undescribed Hox genes in chaetognaths: Sce-hox2, Sce-med6, Sce-postC, and Sce-postD. Surveying public chaetognath datasets reveal med6 and postC in multiple species (e.g. Paraspadella gotoi and Sagitta elegans), while hox2 and postD were not recovered (Fig. 3). Given that the majority of chaetognath Hox genes have already been characterized in earlier studies31,36, we focus here on the four newly identified genes recovered from our transcriptome survey, which expand the known Hox complement in this group.
A schematic phylogeny based on Marlétaz et al.27 illustrates the generalized Hox gene complement for Deuterostomia, Ecdysozoa, Lophotrochozoa, and Rotifera–Acanthocephala, alongside a species-level comparison within Chaetognatha. Boxes to the right of each group or species indicate the presence or undetected status of Hox genes across paralog groups (PG1–PG8 and posterior Hox genes, where applicable). Filled circles indicate identified Hox genes. Diamonds with question marks denote genes not retrieved from transcriptomic data in chaetognath species, though absence cannot be confirmed due to limitations such as low sequencing coverage or stage-specific expression.
Orthology analysis assigns Sce-hox2 to the hox2/proboscipedia (pb) cluster, where it contains five amino-acid residues conserved in PG2: R (position 2), L (4), N (10 and 23), and V (45) (Supplementary Fig. 7). All med6 sequences form a well-supported clade with chaetognath hox8, both nested within PG8 together with lox2, lox4, ubx, and abdA (Supplementary Figs. 5, 6). Med6 displays high similarity with central Hox genes, sharing all eleven diagnostic residues31: Q(6), T(7), R(10), LTR(R/K)RRI(26–32), and E(59) (Supplementary Fig. 8). Notably, med6 shows posterior-class diagnostic residue (Y at position 20).
Sce-postC and Sce-postD cluster with the post1/hox11 – 13 group of the posterior Hox class (Supplementary Figs. 5, 6). Posterior-class residues31,36 K(3) and R(18) are present in both, but V(21) is only found in Sce-postD (Supplementary Fig. 9). PostC also shares as a spiralian post1 signature36 Y(20) with postA and postB (Supplementary Fig. 9).
Expression patterns of Spadella Hox genes
Fluorescent in situ hybridization analyses revealed a staggered spatial and temporal expression pattern of Hox genes along the AP axis during embryonic and post-embryonic development in S. cephaloptera (Fig. 1D–G and Fig. 4; Supplementary Fig. 1D–G and Supplementary Fig. 11).
Gene transcripts (yellow) are visualized with AlexaFluor-647 (C, J, K, M–O) or AP-Fast Blue (B, D–I, L) and cell nuclei are counterstained with DAPI (purple). A General morphology of a hatchling in maximum intensity projection of DAPI-stained whole animal. The eyes are encircled and gut in dashed structure in between the lateral somata clusters (lsc). The position of the mouth and anal opening are indicated by an asterisk and a cross, respectively. (i) Dorsal view, (ii) lateral view, (iii, iv) transverse profile of the trunk (iii) and the tail (iv). B–O Expression of Hox genes in trunk structures (B–L, N, O) and the tail (J–L, N, O). F–I Sce-hox5 – Sce-hox8 are expressed in the medioventral somata clusters (mvs) (marked with red arrowheads). (Ii, ii) Expression of Sce-hox8 in the trunk longitudinal muscle is marked by white arrowheads. (Jii – Lii, Nii, Oii) Insets show expression of Hox genes in the epidermis and mesodermal derivatives of the tail. M Weak background staining is visible but does not represent specific Sce-postB expression. Scale bars: 100 μm, except panels Biii–Oiii (50 μm) and all insets (25 μm). Orientation is indicated in (A). cc corona ciliata, cf caudal fin, cg cerebral ganglion, co ciliary tuft/fence organ, dlm dorsal longitudinal muscle, epi epidermis, in intestine, lc lateral cells, lf lateral fin, lm longitudinal muscle, lsc lateral somata clusters, mc mesenterial cells, mvs medioventral somata clusters, vlm ventral longitudinal muscle.
Gastrula to early elongation stage
During early-mid gastrulation, Sce-med6 is expressed in the mid-posterior ectodermal domain (Fig. 1Di, ii; Supplementary Fig. 1Di, ii), expanding by late gastrula to pre-posterior ectoderm, posterior mesoderm, and posterior neuroectoderm (Fig. 1Diii; Supplementary Fig. 1Diii). Sce-medpost is expressed in ectodermal and endodermal regions slightly posterior to the embryonic midline in early-mid gastrula (Fig. 1Ei, ii; Supplementary Fig. 1Ei, ii), and by late gastrula in the posterior mesoderm, posterior neuroectoderm, and pre-posterior ectoderm (Fig. 1Eiii; Supplementary Fig. 1Eiii). The posterior Hox genes, Sce-postC and Sce-postD, are expressed around the blastopore in early-mid gastrula (Fig. 1Fi, Gi, Gii; Supplementary Fig. 1Fi, Gi, Gii), and later expression localizes to the posterior tip of the ectoderm, mesoderm, and neuroectoderm (Fig. 1Fiii, Giii; Supplementary Fig. 1Fiii, Giii). In the early elongation stage, Sce-med6 and Sce-medpost expression occupy mid-posterior trunk and the pre-posterior lateral neuroblasts (Fig. 1Div, Eiv; Supplementary Fig. 1Div, Eiv), whereas Sce-postC and Sce-postD are restricted to the posterior lateral neuroblasts and the tail bud region (Fig. 1Fiv, Giv; Supplementary Fig. 1Fiv, Giv). Sce-hox2 expression first appears at early elongation in the anterior and posterior regions of the lateral neuroblast of the nascent VNC (Supplementary Fig. 10).
Hatching to juvenile stage
In hatchlings and early juveniles, the lateral somata clusters of the VNC exhibit a staggered Hox patterning (Fig. 4, Supplementary Fig. 11). In the hatchling, Sce-Hox1 is expressed in the inner anterior regions and in the longitudinal muscles of the trunk (Fig. 4Bi–iii). Sce-hox2 is broadly expressed except at the anterior and posterior tips (Fig. 4Ci–iii). Sce-hox3 appears as two vertical bands (Fig. 4Di–iii) overlapping with Sce-hox2 in the mid-region but extending only slightly before its anterior and posterior boundaries. Sce-hox4, Sce-hox5, Sce-hox6, and Sce-hox7 expression patterns form horizontal bands in the mid-region with graded overlaps (Fig. 4E–H). Sce-hox8 is expressed broadly in the posterior lateral somata clusters (Fig. 4Ii–iii), with additional expression in posterior longitudinal muscles around the trunk-tail boundary (white arrowheads in Fig. 4Ii, inset in 4Iii). Sce-hox5–8 are also expressed in the medioventral somata clusters of the VNC (red arrowheads in Fig. 4Fi–Ii, Fiii–Iiii). In the posterior region, Sce-med6 is expressed as a horizontal band (Fig. 4Ji–iii), while Sce-medpost displays a triangular-shaped domain (Fig. 4Ki–iii). Expression of posterior-class genes (Sce-postA, Sce-postC, Sce-postD) is restricted to the posterior VNC, with the latter two extending slightly further toward the tip (Fig. 4Li–iii, Ni–iii, Oi–iii). Notably, Sce-postB is undetectable at all stages examined (Fig. 4Mi–iii). Early juvenile VNC patterns resemble those of hatchlings (Supplementary Fig. 11).
Sce-hox8, Sce-med6, Sce-postC, and Sce-postD are also expressed in the hindgut (Fig. 4Ii, Ji, Ni, Oi). In the tail, a sequential posterior Hox arrangement is observed in hatchlings and early juveniles (Fig. 4Jiii–Liii, Niii, Oiii; Supplementary Fig. 11J, K, M, N). Sce-med6 is expressed in the anterior tail (Fig. 4Ji, ii; Supplementary Fig. 11J), Sce-medpost in the anterior half (Fig. 4Ki, ii; Supplementary Fig. 11K), Sce-postA in the pre-posterior end (Fig. 4Li, ii), and both Sce-postC and Sce-postD in the posterior tail including the caudal fin (Fig. 4Ni, ii, Oi, ii; Supplementary Fig. 11M, N). No expression of Sce-postA was detected in the tail of early juveniles (Supplementary Fig. 11L). Hox expression in the tail is mainly observed in longitudinal muscles and faintly visible in epidermal, fin, and mesoderm-derived (e.g. lateral, mesenterial, and medial) cells (inset in Fig. 4Jii – Lii, Nii, Oii).
Discussion
Our study explores the molecular basis of anterior-posterior axis formation in chaetognaths, a group that is proposed to occupy a Gnathifera-affiliated position within Spiralia24,27. Despite their deep evolutionary history, chaetognaths have retained a remarkably conserved, streamlined, tripartite body plan that has undergone minimal morphological divergence over the past 500 million years68. Their comparatively conservative body architecture offers a unique opportunity to examine the deployment of axial patterning genes in a lineage with limited body plan divergence. Here, we characterized the expression profiles of AP-patterning genes across multiple, hitherto unexplored, developmental stages.
These data reveal spatially organized domains associated with axial and neural regionalization (Fig. 5), uncovering both ancestral bilaterian features and chaetognath-specific innovations. Our findings provide valuable insights into how chaetognath developmental programs relate to those of other spiralians, contributing to our understanding of conserved and lineage-specific mechanisms of AP patterning in Bilateria.
A–E Spatial expression domains of anterior-patterning and Hox genes at key developmental stages: (A) early gastrula, (B) mid gastrula, (C) late gastrula, (D) early elongation, and (E) hatchling (1 dph). Orientation is indicated in the top-right corner of each panel. Colored bars represent approximate gene expression domains, with bar color denoting germ layer origin: neuroectoderm (light purple), ectoderm (blue), mesoderm (dark gray), endoderm (red), and germ cells (dark yellow).
The expression patterns of the anterior patterning genes otx, six3/6, and nk2.1 in S. cephaloptera resemble those observed in the anterior-most regions of other bilaterians13,14,69,70,71,72,73 (Fig. 5).
Sce-otx is consistently expressed in the anterior embryonic domain from gastrula to early elongation. In hatchlings, both Sce-otx and Sce-six3/6 are broadly expressed in anterior regions, consistent with their conserved roles in specifying anterior structures across diverse bilaterian taxa69,72,73,74,75,76,77,78,79. Although embryonic data for Sce-six3/6 are unavailable, its post-embryonic expression remains in the anterior head, in line with its canonical pattern.
In many bilaterians, six3/6 expression occupies the anterior-most domain with otx expression positioned immediately posterior3,12,73,79,80,81. In Spadella, however, Sce-otx and Sce-six3/6 overlap at the anterior-most tip of the head, with Sce-otx occupying the anterior-most region across stages. This deviation from the common bilaterian arrangement suggests a derived reorganization of six3/6-otx spatial hierarchy and raises the possibility that the spatial relationship is more evolutionarily labile than previously recognized. Expression of both genes also extends into anterior structures beyond the CNS: Sce-six3/6 in the eyes, corona ciliata, and anterior epidermis, and Sce-otx in cephalic adhesive papillae, oral region, and ciliary sensory organs, reflecting deployment beyond neural territories as observed in many bilaterians77,79,81,82,83,84,85,86. Notably, Sce-otx is also expressed in PGCs, a divergence from the canonical bilaterian otx patterns that may reflect lineage-specific co-option to germline precursors in Spadella.
Sce-nk2.1 occupies the medial anterior domain of the cerebral ganglion, whereas Sce-pax6 marks the lateral regions62. This mediolateral partitioning resembles patterns known from vertebrates and annelids87,88,89, and is hypothesized as an ancestral bilaterian feature retained in several spiralians, including rotifers39 and brachiopods39,78, but absent in planarians90,91 and the nemertean Lineus ruber39,92. The mediolateral nk2.1–pax6 subdivision in S. cephaloptera, despite their compact hatchling brain, suggests that aspects of ancestral neural patterning can persist in lineages with derived morphologies. Beyond neural structures, Sce-nk2.1 appears in anterior cells of the endomesoderm of early-mid gastrula, later extending along the developing intestinal rudiment by late gastrula. At hatching, it spans the gut with stronger foregut and hindgut signal, as in annelids93,94. By contrast, ecdysozoan and molluscan patterns show foregut-restricted nk2.1/scarecrow (scro)/ceh-2495,96,97, whereas chordates express nk2.1 more broadly across the gut98,99,100. Overall, nk2.1 gut expression in S. cephaloptera is consistent with its roles in gut patterning39,93,95,96,99,100 and with its involvement, together with foxA and otx, in endomesoderm specification across Cnidaria and Bilateria72,93,101,102. Sce-foxA and Sce-nk2.1 are expressed in the developing gut62 but Sce-otx is notably absent, suggesting that S. cephaloptera has likely retained only a subset of the core gene regulatory mechanisms associated with endoderm specification and the development of the digestive tract.
Our findings also reveal that the chaetognath Spadella cephaloptera possesses a complete set of Hox genes with 14 genes from all paralogous groups, including new members of the central and posterior Hox groups (Fig. 3). By comparison, a recent genomic analysis of the related spadellid P. gotoi recovers 15 Hox genes, including additional central- and posterior-class genes, while reportedly lacking hox342. However, we detect hox3 transcripts in publicly available P. gotoi RNA-seq data, suggesting that the species likely possesses the full complement. Taken together, these findings represent the largest and most diverse Hox complement reported for protostomes without whole-genome duplication. Among metazoans, P. gotoi matches the number of canonical Hox genes (excluding medpost) found in the cephalochordate Branchiostoma (15 Hox genes)103. This stands in contrast to the extensive Hox expansions in many arthropods and vertebrates, which are often associated with genome duplication events104,105,106.
Previous studies identified up to ten chaetognath Hox genes31,36. We recovered orthologs of the genes and detected four previously undescribed genes in the group: Sce-hox2, Sce-med6, Sce-postC, and Sce-postD. Sce-hox2 was assigned to PG2 based on orthology inference and diagnostic residues, confirming a complete anterior Hox complement in chaetognaths. No hox2 and postD ortholog was found in other chaetognath transcriptomic datasets potentially due to stage-restricted expression or low sequencing depth (Supplementary Data 1).
The new central-class gene med6 clusters with hox8 inside PG8 and retains all the canonical central homeodomain signature. Similar to hox8, med6 also exhibits a divergent UbdA-like parapeptide motif (xxIxELN) typically found in ecdysozoan ubx/abdA and lophotrochozoan lox2/lox4 orthologs36 (Supplementary Fig. 8). Matus et al.36 hypothesized that the last common ancestor of protostomes possessed a central-class Hox with the UbdA parapeptide, which subsequently underwent independent duplications and divergence that occurred in Ecdysozoa and Lophotrochozoa, giving rise to ubx/abdA and lox2/lox4, respectively. In this framework, after splitting with lophotrochozoans, chaetognath hox8 and med6 may also have arisen from a shared ancestral UbdA-bearing central gene followed by lineage-specific modifications that resulted in partial UbdA signal.
We also identified two previously uncharacterized posterior class Hox genes, Sce-postC and Sce-postD. Both share several plesiomorphic features with ecdysozoan abdB, lophotrochozoan post1 and post2, and deuterostome hox9–15 (Supplementary Fig. 9), but lack clear chaetognath-specific motifs. Phylogenetic analyses place them variably: ML recovers postC/D as sister to post1/hox11–15, whereas Bayesian inference yields polytomy with postA and postB, complicating inferences about their evolutionary relationship with other bilaterian posterior Hox genes. The scattered distribution of chaetognath postA–D, rather than forming a single monophyletic group, suggests lineage-specific duplications of an ancestral posterior gene, followed by sequence divergence that has obscured shared molecular signatures. This pattern resembles Posterior Flexibility, a phenomenon described in deuterostomes where posterior Hox genes show elevated sequence divergence and poor phylogenetic resolution107. While Posterior Flexibility has not been formally reported in protostomes, the unresolved affinities of chaetognath postA–D may reflect similar evolutionary dynamics.
Sce-postB, a predicted paralog of Sce-postA, was recovered in transcriptomic data and confirmed by PCR, indicating transcriptional activity. However, no expression was detected by FISH at the stages examined, which may reflect low abundance, restricted expression timing, or technical sensitivity limits (e.g. probe inefficiency). Without additional temporal or spatial resolution, the expression dynamics and potential developmental role of Sce-postB remain unresolved.
Chaetognaths appear to have retained the ancestral Hox repertoire, supporting the hypothesis of an urbilaterian complement of at least seven Hox genes spanning the anterior, central, and posterior class genes108,109. Their Hox complement appears to have expanded through lineage-specific duplication and divergence, yielding one of the most extensive Hox repertoires among bilaterians (Fig. 3). We initially hypothesized that this expansion reflects small-scale events (tandem or segmental duplication) rather than whole-genome duplication (WGD), given the lack of widespread duplication in other transcription factors families (e.g., anterior patterning genes). The P. gotoi genome reveals a tandemly duplicated Hox cluster without WGD signatures42, reinforcing the view that the expansion of chaetognath Hox likely occurred through local duplication. While P. gotoi is not a direct substitute for a Spadella genome, its similar Hox content and cluster structure support inferences about expansion mechanisms in the lineage. In contrast, the Rotifera–Acanthocephala clade shows marked reduction with some lineages retaining only one or two anterior-class genes37 and despite an ancient WGD in Bdelloidea, they only retain a subset of ancestral Hox genes110. These divergent patterns underscore the distinctive chaetognath trajectory in Hox evolution, highlighting their importance for reconstructing the early diversification of bilaterians and contributing to a broader understanding of how Hox gene evolution may have shaped metazoan body plan complexity.
The earliest reported chaetognath Hox expression data came from SceMed4 (PG5), which was shown to be expressed as two bilateral domains in the VNC of Spadella38, suggesting a role of Hox genes in patterning the ventral CNS of chaetognaths38. Our data extended this view by revealing a staggered expression profile of multiple Hox genes along the AP axis of the VNC in S. cephaloptera.
During the late gastrula stage, a subset of central and posterior class Hox genes (Sce-med6, Sce-medpost, Sce-postC, and Sce-postD) are broadly expressed in the posterior neuroectoderm (Fig. 5C). By early elongation, these domains resolve into a staggered AP (Fig. 5D), with Sce-hox2 occupying a more anterior domain in the nascent VNC. This staggered pattern is maintained into hatchlings and juveniles, where 13 Hox genes show distinct expression along the VNC (Fig. 5E). These findings suggest a temporal and spatial continuity of Hox-mediated AP patterning in the developing central nervous system that begins during early neuroectoderm formation and persists through later development.
The AP-staggered pattern reflects a conserved mechanism observed across many bilaterians, in which Hox expression regionalizes the neuroectoderm and developing neural structures during specific developmental stages. In bilaterians, this pattern is evident in the nemertean Pantinonemertes californiensis larva75, in several molluscan groups11,111,112,113,114,115,116, and annelids10,117,118,119,120, though often coupled to segmental organization except in the leech Helobdella which shows a staggered Hox expression only in ventral neural tissues121,122. Hox genes in early development of annelids primarily function in defining overall segment organization and specifying segmental fate rather than being solely involved in patterning the CNS. In the rotifer Brachionus manjavacas16 and the brachiopod Terebratalia transversa19, only a partial resemblance of staggered patterning can be observed11. Outside Spiralia, staggered expression regionalizes, to varying degrees, the developing CNS in arthropods8,123,124, a hemichordate5, chordates9,125,126, and acoels127,128.
In S. cephaloptera, the staggered Hox expression in the VNC may also contribute to regional specialization38, potentially influencing the differentiation of distinct neuronal subtypes along the AP axis, similar to how Hox genes regulate region-specific neuronal fates in Drosophila8,129, C. elegans130, and vertebrates131. Although direct evidence for a spatial correlation between neuronal diversity and Hox gene expression domains in chaetognaths is currently unavailable, the arrangement of RFamide-immunoreactive (ir) neurons along the AP axis of the VNC offers a potential framework for speculation. RFamine-ir neurons are serially arranged, with lateral cells (L-group) located anteriorly and dorsal cells (D-group) positioned centrally (D1 – D5) and posteriorly (D6, D7, and the giant X neuron found only in the Spadellidae)22,132,133,134,135. Conceptually mapping these onto Hox domains, it is plausible that L-group neurons nest within Sce-hox1-3 domains, midline G-groups with central-class domains, and posterior neurons with Sce-medpost and posterior-class domains. If such spatial alignment reflects underlying functional relationships, it would support the hypothesis that Hox genes contribute to neural subtype patterning in chaetognaths, as in other bilaterians. This underscores the need for future studies on chaetognath neuronal diversity and the developmental mechanisms guiding its specification.
P. gotoi exhibits an AP-staggered patterning for a subset of central- and posterior-class Hox genes in the VNC of early juveniles42, paralleling our findings in Spadella41. However, the reported expression patterns of some genes (e.g., Pgo-hox1 and Pgo-medpost) appear broader (VNC) or differ in tissue association (e.g., ciliary fence receptors), which may reflect species-specific differences, developmental staging, or technical variation (e.g., chromogenic ISH versus HCR-FISH).
The spatially staggered expression of Hox genes in the VNC of S. cephaloptera aligns chaetognaths with a diverse set of bilaterians exhibiting similar AP neural patterning, including annelids, molluscs, arthropods, and vertebrates. Given that chaetognaths represent one of the earliest-diverging lineages within Spiralia, these findings considerably broaden the phylogenetic scope of Hox-mediated nervous system regionalization. This supports the long-standing view that staggered Hox expression along the AP axis is an ancestral bilaterian trait, likely present in early spiralians. While Hox-mediated neural patterning is broadly conserved, the extent and precision of staggered patterns vary across lineages and developmental stages (e.g., partial or transient patterns in rotifers and brachiopods16,69 and departures from expected axial position in certain molluscs116). This variation may reflect lineage-specific modifications or reductions of an ancestral patterning mechanism, underscoring the significance of chaetognaths in preserving a complete, temporally sustained, and spatially ordered deployment of Hox genes during nervous system development. This conservation, along with the conserved bauplan since the early Cambrian, raises the possibility that chaetognaths retain a neural Hox expression pattern more reflective of the ancestral spiralian, or even bilaterian, condition. Conversely, if chaetognaths independently recruited Hox genes for nervous system patterning, it would represent a striking case of parallel evolution involving a deeply conserved genetic toolkit. Consequently, these findings position chaetognaths as a critical group for reconstructing early bilaterian nervous system evolution.
Hox genes, however, are not restricted to neural tissues and are also expressed in non-neural domains. From gastrula stages onward, Sce-med6, Sce-medpost, Sce-postC, and Sce-postD are expressed in the posterior mesoderm and ectoderm in spatially staggered domains largely confined to the posterior half of the embryo (Fig. 5A–D). In hatchlings, several Hox genes are expressed in non-neural structures of the trunk, including longitudinal musculature and the posterior region of the endoderm-derived intestine (Fig. 4), suggesting potential roles in mesodermal and endodermal regionalization.
In the trunk musculature, Sce-hox1 marks the mid-section longitudinal muscles (weakening posteriorly), while Sce-hox8 is confined to posterior longitudinal muscles. The absence of other Hox gene expression domains in the trunk musculature suggests a limited Hox involvement in mesodermal patterning at this stage. Since hox1/labial and hox8/lox2/lox4 orthologs are not typically implicated in mesodermal patterning in other spiralians, their expression here may reflect co-option for S. cephaloptera trunk muscle development. Although, it remains unclear whether this represents chaetognath-specific adaptation or an underexplored spiralian role.
Several Hox genes (Sce-hox8, Sce-med6, Sce-postC, and Sce-postD) are also expressed in the posterior intestine of the hatchling, overlapping with previously described zones of cell-type specialization: anterior light-granule cells (luminal digestion) and posterior dark-granule cells (absorption and intracellular digestion)136,137. The posterior localization of Sce-postC and Sce-postD, particularly at the rectal terminus, also suggests roles in hindgut regionalization and anus formation. Posterior Hox genes expression in the hindgut also occurs in Drosophila138, the tunicate Ciona intestinalis125, the sea cucumber Apostichopus japonicus139, and vertebrates (e.g.140,141). Hox genes such as hox3, hox6/lox5, hox7/antp are expressed in specific gut domains in the scaphopod Antalis entalis116 and in the annelid Capitella10. Taken together, these findings suggest that Hox genes may contribute to hindgut regionalization in S. cephaloptera, and that Hox recruitment to the gut may have repeatedly occurred across bilaterians. Whether this reflects shared ancestry or convergent co-option remains unresolved.
Such tissue-specific roles naturally prompt questions about the underlying genomic organization of chaetognath Hox genes. Colinearity in S. cephaloptera remains untested because the genomic Hox gene order is unknown. The genome of P. gotoi reveals a physically linked Hox cluster and support a spatial colinearity in early juvenile VNC42. This raises the possibility that a similar genomic organization underlies the VNC-wide AP-staggered deployment across most Hox loci observed in S. cephaloptera. Our embryonic expression data, however, point to a possible disruption of temporal colinearity. Sce-med6, Sce-medpost, Sce-postC, and Sce-postD are detected at the gastrula stage, while the anterior-class Hox gene Sce-hox2 only initiates during early elongation. Because this inference is based on a single anterior gene, temporal data for the full Hox set are needed to evaluate the broader pattern. Overall, these findings suggest that chaetognaths retain spatial aspects of ancestral Hox regulation while exhibiting modifications in their temporal deployment, consistent with partial decoupling of spatial and temporal colinearity observed in some bilaterians5,119,142,143.
Although S. cephaloptera has been suggested to belong to a morphologically derived benthic lineage (Spadellidae)144, its complete Hox complement and AP-staggered deployment offer a useful comparative baseline for inferring posterior patterning in Chaetognatha. Central and posterior Hox genes (Sce-med6, Sce-medpost, Sce-postA, Sce-postC, and Sce-postD) exhibit spatially staggered expression along the postanal tail (Fig. 5E), consistent with their involvement in caudal regionalization. During embryogenesis, these genes (except Sce-postA) are expressed in the posterior epidermal and mesodermal cells from gastrulation to early elongation stages, consistent with their later localization in the hatchling tail. Their dual localization in the posterior VNC and tail is more consistent with pleiotropy and tissue-specific readouts of a single AP axis rather than axis duplication, as no ectopic anterior markers (e.g., six3/6, otx, nk2.1) or locally replicated Hox series (full complement) are detected outside the VNC. Moreover, the restriction of extra non-neural domains to posterior-side genes is consistent with Posterior Flexibility, wherein posterior-class Hox genes are coopted into lineage-specific morphological elaborations, particularly in appendages and posterior structures145. However, whether these deployments are mechanistically decoupled remains to be investigated.
Postanal tails are rare among bilaterians and have only been described in chordates and chaetognaths. In chordates, postanal tail development involves posterior-class Hox genes (hox9–13), together with regulatory factors such as sonic hedgehog (shh) and brachyury (bra/tbxt)146,147,148. While this raises the possibility of functional parallels, no shh ortholog is detected in S. cephaloptera transcriptome, and bra expression in P. gotoi is restricted to anterior embryonic regions, particularly the stomodeum and presumptive mouth149. Combined with deep phylogenetic divergence and presumed differences in cell lineage, these observations favor independent origin of chaetognath and chordate postanal tails rather than a shared ancestral program.
Further support comes from comparisons with monogonont rotifers, the closest gnathiferan relatives for which Hox gene expression data are available. Across the Rotifera–Acanthocephala clade, posterior Hox genes are absent, and in monogononts, Hox expression is restricted to neural tissues and their postanal extension (i.e. foot) is not patterned by Hox genes16,37,110. Consequently, these comparisons suggest that postanal tails of chaetognaths arose via independent co-option of posterior Hox genes to pattern novel caudal structures.
Fossil data provide crucial context for understanding the evolutionary origins of the postanal tail in chaetognaths. Stem-group chaetognaths (e.g. Amiskwia sagittiformis and Timorbestia koprii), exhibit a subterminal anus with a short postanal extension (caudal fin)150,151, while early crown-group taxa from the early Cambrian (e.g. Protosagitta spinosa and Capinatator praetermissus) possess a more elongate tail posterior to the anus152,153, suggesting tail elongation emerged within the crown chaetognath lineage. Variation among modern taxa suggests ecological or functional pressures may have shaped tail morphology. The staggered deployment of posterior Hox genes in the tail, together with its persistence in both fossils and extant species, supports the hypothesis that this structure emerged through lineage-specific co-option of ancestral Hox-based patterning, contributing to a novel posterior anatomy.
Despite its derived ecology, S. cephaloptera retains a postanal tail and staggered posterior Hox expression, providing a comparative system for investigating Hox roles in posterior elaboration. The correspondence between morphological persistence and posterior-Hox deployment is compatible with the hypothesis of conserved developmental program, but comparative expression and functional tests are required to assess whether a shared mechanism underlies tail maintenance across Chaetognatha.
Conclusion
Our study provides a comprehensive analysis of anterior-patterning (otx, nk2.1, six3/6) and Hox gene expression in the chaetognath Spadella cephaloptera, providing new insights into the AP axis patterning in this lineage. We show that anterior patterning genes are expressed in the cerebral ganglion and associated head structures, supporting their conserved role in anterior CNS development across Bilateria. In addition, we document a staggered expression of Hox genes, including the newly described Sce-med6, Sce-postC, and Sce-postD, along the VNC, hindgut, and postanal tail. These patterns suggest that chaetognaths maintain core bilaterian AP patterning programs while also deploying lineage-specific modifications. The VNC regionalization parallels neural patterning observed in other bilaterians, highlighting shared developmental mechanisms despite divergent morphologies. As a lineage occupying a key phylogenetic position within Spiralia, chaetognaths provide a critical comparative perspective for reconstructing spiralian evolution. Their retention of a complete Hox cluster, in contrast to the reduced Rotifera-Acanthocephala repertoire, underscores the diversity of Hox gene utilization across Spiralia. By addressing a long-standing gap in spiralian evo-devo, this study establishes chaetognaths as an essential model for reconstructing ancestral spiralian developmental programs and for understanding how conserved genetic toolkits are independently co-opted and elaborated across animal evolution.
Data availability
The assembled Spadella cephaloptera transcriptome is available on Zenodo under the accession https://doi.org/10.5281/zenodo.760296044. Newly obtained S. cephaloptera sequence data that support the findings of this study have been deposited to GenBank with accession numbers: PX219847-PX219863. All other data supporting the findings of this study are available from the corresponding authors upon reasonable request.
Code availability
All scripts used to download, assemble, and annotate transcriptomes for Hox gene mining are available at GitHub (https://github.com/jfordonez/ChaetognathAPPatterning.git). The exact version of the code used in this study has been archived on Zenodo under the https://doi.org/10.5281/zenodo.17705168154.
References
Arenas-Mena, C., Cameron, A. R. & Davidson, E. H. Spatial expression of Hox cluster genes in the ontogeny of a sea urchin. Development 127, 4631–4643 (2000).
Feuda, R. & Peter, I. S. Homologous gene regulatory networks control development of apical organs and brains in Bilateria. Sci. Adv. 8, eabo2416 (2022).
Holland, L. Z. et al. Evolution of bilaterian central nervous systems: a single origin?. EvoDevo 4, 27 (2013).
Lowe, C. J. et al. Anteroposterior Patterning in Hemichordates and the Origins of the Chordate Nervous System. Cell 113, 853–865 (2003).
Aronowicz, J. & Lowe, C. J. Hox gene expression in the hemichordate Saccoglossus kowalevskii and the evolution of deuterostome nervous systems. Integr. Comp. Biol. 46, 890–901 (2006).
Faltine-Gonzalez, D., Havrilak, J. & Layden, M. J. The brain regulatory program predates central nervous system evolution. Sci. Rep. 13, 8626 (2023).
Hejnol, A. & Lowe, C. J. Embracing the comparative approach: how robust phylogenies and broader developmental sampling impacts the understanding of nervous system evolution. Philos. Trans. R. Soc. Lond. B: Biol. Sci. 370, 20150045 (2015).
Jarvis, E., Bruce, H. S. & Patel, N. H. Evolving specialization of the arthropod nervous system. Proc. Natl. Acad. Sci. 109, 10634–10639 (2012).
Philippidou, P. & Dasen, J. S. Hox genes: choreographers in neural development, architects of circuit organization. Neuron 80, 12–34 (2013).
Fröbius, A. C., Matus, D. Q. & Seaver, E. C. Genomic organization and expression demonstrate spatial and temporal hox gene colinearity in the lophotrochozoan Capitella sp. I. PLoS ONE 3, e4004 (2008).
Huan, P., Wang, Q., Tan, S. & Liu, B. Dorsoventral decoupling of Hox gene expression underpins the diversification of molluscs. Proc. Natl. Acad. Sci. 117, 503–512 (2020).
Marlow, H. et al. Larval body patterning and apical organs are conserved in animal evolution. BMC Biol 12, 7 (2014).
Steinmetz, P. R. H., Kostyuchenko, R. P., Fischer, A. & Arendt, D. The segmental pattern of otx, gbx, and Hox genes in the annelid Platynereis dumerilii: The segmental pattern of otx, gbx, and Hox genes. Evol. Dev. 13, 72–79 (2011).
Wollesen, T. et al. Brain regionalization genes are co-opted into shell field patterning in Mollusca. Sci. Rep. 7, 5486 (2017).
Currie, K. W. et al. HOX gene complement and expression in the planarian Schmidtea mediterranea. EvoDevo 7, 7 (2016).
Fröbius, A. C. & Funch, P. Rotiferan Hox genes give new insights into the evolution of metazoan bodyplans. Nat. Commun. 8, 9 (2017).
Gąsiorowski, L. & Hejnol, A. Hox gene expression during development of the phoronid Phoronopsis harmeri. EvoDevo 11, 2 (2020).
Salamanca-Díaz, D. A., Calcino, A. D., De Oliveira, A. L. & Wanninger, A. Non-collinear Hox gene expression in bivalves and the evolution of morphological novelties in mollusks. Sci. Rep. 11, 3575 (2021).
Schiemann, S. M. et al. Clustered brachiopod Hox genes are not expressed collinearly and are associated with lophotrochozoan novelties. Proc. Natl. Acad. Sci. 114 (2017).
Dunn, C. W. et al. Broad phylogenomic sampling improves resolution of the animal tree of life. Nature 452, 745–749 (2008).
Giribet, G., Distel, D. L., Polz, M., Sterrer, W. & Wheeler, W. C. Triploblastic Relationships with emphasis on the acoelomates and the position of Gnathostomulida, Cycliophora, Plathelminthes, and Chaetognatha: a combined approach of 18S rDNA sequences and morphology. Syst. Biol. 49, 539–562 (2000).
Harzsch, S. & Müller, C. H. A new look at the ventral nerve centre of Sagitta: implications for the phylogenetic position of Chaetognatha (arrow worms) and the evolution of the bilaterian nervous system. Front. Zool. 4, 14 (2007).
Helfenbein, K. G., Fourcade, H. M., Vanjani, R. G. & Boore, J. L. The mitochondrial genome of Paraspadella gotoi is highly reduced and reveals that chaetognaths are a sister group to protostomes. Proc. Natl. Acad. Sci. 101, 10639–10643 (2004).
Laumer, C. E. et al. Revisiting metazoan phylogeny with genomic sampling of all phyla. Proc. R. Soc. B: Biol. Sci. 286, 20190831 (2019).
Littlewood, D. T. J., Telford, M. J., Clough, K. A. & Rohde, K. Gnathostomulida—an enigmatic metazoan phylum from both morphological and molecular perspectives. Mol. Phylogenet. Evol. 9, 72–79 (1998).
Mallatt, J. & Winchell, C. J. Testing the new animal phylogeny: First use of combined large-subunit and small-subunit rRNAgene sequences to classify the protostomes. Mol. Biol. Evol. 19, 289–301 (2002).
Marlétaz, F., Peijnenburg, K. T. C. A., Goto, T., Satoh, N. & Rokhsar, D. S. A new Spiralian Phylogeny places the enigmatic arrow worms among Gnathiferans. Curr. Biol. 29, 312–318.e3 (2019).
Marlétaz, F. et al. Chaetognath phylogenomics: a protostome with deuterostome-like development. Curr. Biol. 16, R577–R578 (2006).
Matus, D. Q. et al. Broad taxon and gene sampling indicate that chaetognaths are protostomes. Curr. Biol. 16, R575–R576 (2006).
Papillon, D., Perez, Y., Caubit, X. & Le Parco, Y. Identification of Chaetognaths as Protostomes Is Supported By The Analysis Of Their Mitochondrial Genome. Mol. Biol. Evol. 21, 2122–2129 (2004).
Papillon, D., Perez, Y., Fasano, L., Le Parco, Y. & Caubit, X. Hox gene survey in the chaetognath Spadella cephaloptera: evolutionary implications. Dev. Genes Evol. 213, 142–148 (2003).
Paps, J., Baguñà, J. & Riutort, M. Lophotrochozoa internal phylogeny: new insights from an up-to-date analysis of nuclear ribosomal genes. Proc. R. Soc. B Biol. Sci. 276, 1245–1254 (2009).
Peterson, K. J. & Eernisse, D. J. Animal phylogeny and the ancestry of bilaterians: inferences from morphology and 18S rDNA gene sequences. Evol. Dev. 3, 170–205 (2001).
Philippe, H. et al. Acoelomorph flatworms are deuterostomes related to Xenoturbella. Nature 470, 255–258 (2011).
Telford, M. J. & Holland, P. W. The phylogenetic affinities of the chaetognaths: a molecular analysis. Mol. Biol. Evol. 10, 660–676 (1993).
Matus, D. Q., Halanych, K. M. & Martindale, M. Q. The Hox gene complement of a pelagic chaetognath, Flaccisagitta enflata. Integr. Comp. Biol. 47, 854–864 (2007).
Mauer, K. M. et al. Genomics and transcriptomics of epizoic Seisonidea (Rotifera, syn. Syndermata) reveal strain formation and gradual gene loss with growing ties to the host. BMC Genomics 22, 604 (2021).
Papillon, D., Perez, Y., Fasano, L., Le Parco, Y. & Caubit, X. Restricted expression of a median Hox gene in the central nervous system of chaetognaths. Dev. Genes Evol. 215, 369–373 (2005).
Martín-Durán, J. M. et al. Convergent evolution of bilaterian nerve cords. Nature 553, 45–50 (2018).
Boell, L. A. & Bucher, G. Whole-mount in situ hybridization in the Rotifer Brachionus plicatilis representing a basal branch of lophotrochozoans. Dev. Genes Evol. 218, 445–451 (2008).
Ordoñez, J. F. & Wollesen, T. Chaetognaths exhibit the most extensive repertoire of Hox genes among protostomes. https://doi.org/10.1101/2025.01.31.635887 (2025).
Piovani, L. et al. The genomic origin of the unique chaetognath body plan. Nature https://doi.org/10.1038/s41586-025-09403-2 (2025).
Wollesen, T., Rodriguez Monje, S. V, Oel, A. P. & Arendt, D. Characterization of eyes, photoreceptors, and opsins in developmental stages of the arrow worm Spadella cephaloptera (Chaetognatha). J. Exp. Zoolog. B: Mol. Dev. Evol. https://doi.org/10.1002/jez.b.23193 (2023).
Wollesen, T. Characterization of eyes, photoreceptors and opsins in developmental stages of the arrow worm Spadella cephaloptera (Chaetognatha). Zenodo https://doi.org/10.5281/ZENODO.7602960 (2023).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Bushmanova, E., Antipov, D., Lapidus, A. & Prjibelski, A. D. rnaSPAdes: a de novo transcriptome assembler and its application to RNA-Seq data. GigaScience 8, giz100 (2019).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 12, 59–60 (2015).
Katoh, K. MAFFT version 5: improvement in accuracy of multiple sequence alignment. Nucleic Acids Res. 33, 511–518 (2005).
Katoh, K. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 30, 3059–3066 (2002).
Steenwyk, J. L., Buida, T. J., Li, Y., Shen, X.-X. & Rokas, A. ClipKIT: A multiple sequence alignment trimming software for accurate phylogenomic inference. PLOS Biol. 18, e3001007 (2020).
Hoang, D. T., Chernomor, O., Von Haeseler, A., Minh, B. Q. & Vinh, L. S. UFBoot2: Improving the Ultrafast Bootstrap Approximation. Mol. Biol. Evol. 35, 518–522 (2018).
Kalyaanamoorthy, S., Minh, B. Q., Wong, T. K. F., Von Haeseler, A. & Jermiin, L. S. ModelFinder: fast model selection for accurate phylogenetic estimates. Nat. Methods 14, 587–589 (2017).
Minh, B. Q. et al. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Guindon, S. et al. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst. Biol. 59, 307–321 (2010).
Huelsenbeck, J. P. & Ronquist, F. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17, 754–755 (2001).
Rambaut, A., Drummond, A. J., Xie, D., Baele, G. & Suchard, M. A. Posterior summarization in Bayesian phylogenetics using Tracer 1.7. Syst. Biol. 67, 901–904 (2018).
Ronquist, F. et al. MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 61, 539–542 (2012).
Choi, H. M. T. et al. Third-generation in situ hybridization chain reaction: multiplexed, quantitative, sensitive, versatile, robust. https://doi.org/10.1101/285213 (2018).
Wang, Y. et al. Multiplexed in situ protein imaging using DNA-barcoded antibodies with extended hybridization chain reactions. Nucleic Acids Res. gkae592 https://doi.org/10.1093/nar/gkae592 (2024).
Tsuneoka, Y. & Funato, H. Modified in situ hybridization chain reaction using short hairpin DNAs. Front. Mol. Neurosci. 13, 75 (2020).
Ordoñez, J. F. & Wollesen, T. Unfolding the ventral nerve center of chaetognaths. Neural Develop 19, 5 (2024).
Choi, H. M. T., Beck, V. A. & Pierce, N. A. Next-generation in situ hybridization chain reaction: higher gain, lower cost, greater durability. ACS Nano 8, 4284–4294 (2014).
Kuehn, E. et al. Segment number threshold determines juvenile onset of germline cluster expansion in Platynereis dumerilii. J. Exp. Zoolog. B: Mol. Dev. Evol. 338, 225–240 (2022).
Bruce, H. S. et al. Hybridization chain reaction (HCR) in situ protocol v1. https://doi.org/10.17504/protocols.io.bunznvf6 (2021).
Schindelin, J. et al. Fiji: An open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
John, C. C. Habits, structure, and development of Spadella Cephaloptera. J. Cell Sci. S2-75, 625–696 (1933).
Telford, M. J. Evolution: arrow worms find their place on the tree of life. Curr. Biol. 29, R152–R154 (2019).
Andrikou, C., Passamaneck, Y. J., Lowe, C. J., Martindale, M. Q. & Hejnol, A. Molecular patterning during the development of Phoronopsis harmeri reveals similarities to rhynchonelliform brachiopods. EvoDevo 10, 33 (2019).
Eriksson, B. J., Samadi, L. & Schmid, A. The expression pattern of the genes engrailed, pax6, otd and six3 with special respect to head and eye development in Euperipatoides kanangrensis Reid 1996 (Onychophora: Peripatopsidae). Dev. Genes Evol. 223, 237–246 (2013).
Martín-Durán, J. M., Vellutini, B. C. & Hejnol, A. Evolution and development of the adelphophagic, intracapsular Schmidt’s larva of the nemertean Lineus ruber. EvoDevo 6, 28 (2015).
Omori, A., Akasaka, K., Kurokawa, D. & Amemiya, S. Gene expression analysis of Six3, Pax6, and Otx in the early development of the stalked crinoid Metacrinus rotundus. Gene Expr. Patterns 11, 48–56 (2011).
Redl, E., Scherholz, M., Wollesen, T., Todt, C. & Wanninger, A. Expression of six3 and otx in Solenogastres (Mollusca) supports an ancestral role in bilaterian anterior-posterior axis patterning. Evol. Dev. 20, 17–28 (2018).
Gonzalez, P., Uhlinger, K. R. & Lowe, C. J. The adult body plan of indirect developing hemichordates develops by adding a hox-patterned trunk to an anterior larval territory. Curr. Biol. 27, 87–95 (2017).
Hiebert, L. S. & Maslakova, S. A. Expression of Hox, Cdx, and Six3/6 genes in the hoplonemertean Pantinonemertes californiensis offers insight into the evolution of maximally indirect development in the phylum Nemertea. EvoDevo 6, 26 (2015).
Luo, Y.-J. et al. Nemertean and phoronid genomes reveal lophotrochozoan evolution and the origin of bilaterian heads. Nat. Ecol. Evol. 2, 141–151 (2017).
Oliver, G. et al. Six3, a murine homologue of the sine oculis gene, demarcates the most anterior border of the developing neural plate and is expressed during eye development. Development 121, 4045–4055 (1995).
Santagata, S., Resh, C., Hejnol, A., Martindale, M. Q. & Passamaneck, Y. J. Development of the larval anterior neurogenic domains of Terebratalia transversa (Brachiopoda) provides insights into the diversification of larval apical organs and the spiralian nervous system. EvoDevo 3, 3 (2012).
Steinmetz, P. R. et al. Six3 demarcates the anterior-most developing brain region in bilaterian animals. EvoDevo 1, 14 (2010).
Kaul-Strehlow, S., Urata, M., Praher, D. & Wanninger, A. Neuronal patterning of the tubular collar cord is highly conserved among enteropneusts but dissimilar to the chordate neural tube. Sci. Rep. 7, 7003 (2017).
Vöcking, O., Kourtesis, I. & Hausen, H. Posterior eyespots in larval chitons have a molecular identity similar to anterior cerebral eyes in other bilaterians. EvoDevo 6, 40 (2015).
Bruce, A. E. E. & Shankland, M. Expression of the Head Gene Lox22-Otx in the Leech Helobdella and the Origin of the Bilaterian Body Plan. Dev. Biol. 201, 101–112 (1998).
Carl, M., Loosli, F. & Wittbrodt, J. Six3 inactivation reveals its essential role for the formation and patterning of the vertebrate eye. Development 129, 4057–4063 (2002).
Hinman, V. F., Nguyen, A. T. & Davidson, E. H. Expression and function of a starfish Otx ortholog, AmOtx: a conserved role for Otx proteins in endoderm development that predates divergence of the eleutherozoa. Mech. Dev. 120, 1165–1176 (2003).
Loosli, F., Köster, R. W., Carl, M., Krone, A. & Wittbrodt, J. Six3, a medaka homologue of the Drosophila homeobox gene sine oculis is expressed in the anterior embryonic shield and the developing eye. Mech. Dev. 74, 159–164 (1998).
Ogura, A. et al. Loss of the six3/6 controlling pathways might have resulted in pinhole-eye evolution in Nautilus. Sci. Rep. 3, 1432 (2013).
Corbin, J. G., Rutlin, M., Gaiano, N. & Fishell, G. Combinatorial function of the homeodomain proteins Nkx2.1 and Gsh2 in ventral telencephalic patterning. Development 130, 4895–4906 (2003).
Murakami, Y. et al. Identification and expression of the lamprey Pax6 gene: evolutionary origin of the segmented brain of vertebrates. Development 128, 3521–3531 (2001).
Tessmar-Raible, K. et al. Conserved sensory-neurosecretory cell types in annelid and fish forebrain: Insights into hypothalamus evolution. Cell 129, 1389–1400 (2007).
Currie, K. W., Molinaro, A. M. & Pearson, B. J. Neuronal sources of hedgehog modulate neurogenesis in the adult planarian brain. eLife 5, e19735 (2016).
Scimone, M. L., Kravarik, K. M., Lapan, S. W. & Reddien, P. W. Neoblast specialization in regeneration of the planarian Schmidtea mediterranea. Stem Cell Rep 3, 339–352 (2014).
Gąsiorowski, L., Børve, A., Cherneva, I. A., Orús-Alcalde, A. & Hejnol, A. Molecular and morphological analysis of the developing nemertean brain indicates convergent evolution of complex brains in Spiralia. BMC Biol 19, 175 (2021).
Boyle, M. J., Yamaguchi, E. & Seaver, E. C. Molecular conservation of metazoan gut formation: evidence from expression of endomesoderm genes in Capitella teleta (Annelida). EvoDevo 5, 39 (2014).
Kerbl, A., Martín-Durán, J. M., Worsaae, K. & Hejnol, A. Molecular regionalization in the compact brain of the meiofaunal annelid Dinophilus gyrociliatus (Dinophilidae). EvoDevo 7, 20 (2016).
Zaffran, S., Das, G. & Frasch, M. The NK-2 homeobox gene scarecrow (scro) is expressed in pharynx, ventral nerve cord and brain of Drosophila embryos. Mech. Dev. 94, 237–241 (2000).
Harfe, B. D. & Fire, A. Muscle and nerve-specific regulation of a novel NK-2 class homeodomain factor in Caenorhabditis elegans. Development 125, 421–429 (1998).
Dunn, E. F. et al. Molecular paleoecology: using gene regulatory analysis to address the origins of complex life cycles in the late Precambrian. Evol. Dev. 9, 10–24 (2007).
Ristoratore, F. et al. Expression and functional analysis of Cititf1, an ascidian NK-2 class gene, suggest its role in endoderm development. Development 126, 5149–5159 (1999).
Takacs, C. M., Moy, V. N. & Peterson, K. J. Testing putative hemichordate homologues of the chordate dorsal nervous system and endostyle: expression of NK2.1 (TTF-1) in the acorn worm Ptychodera flava (Hemichordata, Ptychoderidae). Evol. Dev. 4, 405–417 (2002).
Venkatesh, T. V., Holland, N. D., Holland, L. Z., Su, M.-T. & Bodmer, R. Sequence and developmental expression of amphioxus AmphiNk2-1: insights into the evolutionary origin of the vertebrate thyroid gland and forebrain. Dev. Genes Evol. 209, 254–259 (1999).
Hejnol, A. & Martindale, M. Q. Acoel development indicates the independent evolution of the bilaterian mouth and anus. Nature 456, 382–386 (2008).
Perry, K. J. et al. Deployment of regulatory genes during gastrulation and germ layer specification in a model spiralian mollusc Crepidula. Dev. Dyn. 244, 1215–1248 (2015).
Pascual-Anaya, J. et al. Broken colinearity of the amphioxus Hox cluster. EvoDevo 3, 28 (2012).
Nong, W. et al. Horseshoe crab genomes reveal the evolution of genes and microRNAs after three rounds of whole genome duplication. Commun. Biol. 4, 83 (2021).
Ontano, A. Z. et al. Taxonomic Sampling and Rare Genomic Changes Overcome Long-Branch Attraction in the Phylogenetic Placement of Pseudoscorpions. Mol. Biol. Evol. 38, 2446–2467 (2021).
Garcia-Fernàndez, J. & Holland, P. W. H. Archetypal organization of the amphioxus Hox gene cluster. Nature 370, 563–566 (1994).
Ferrier, D. E. K., Minguillón, C., Holland, P. W. H. & Garcia-Fernàndez, J. The amphioxus Hox cluster: deuterostome posterior flexibility and Hox14. Evol. Dev 2, 284–293 (2000).
Balavoine, G., De Rosa, R. & Adoutte, A. Hox clusters and bilaterian phylogeny. Mol. Phylogenet. Evol. 24, 366–373 (2002).
De Rosa, R. et al. Hox genes in brachiopods and priapulids and protostome evolution. Nature 399, 772–776 (1999).
Flot, J.-F. et al. Genomic evidence for ameiotic evolution in the bdelloid rotifer Adineta vaga. Nature 500, 453–457 (2013).
Barrera Grijalba, C. C., Rodríguez Monje, S. V., Gestal, C. & Wollesen, T. Octopod Hox genes and cephalopod plesiomorphies. Sci. Rep. 13, 15492 (2023).
Fritsch, M., Wollesen, T., De Oliveira, A. L. & Wanninger, A. Unexpected co-linearity of Hox gene expression in an aculiferan mollusk. BMC Evol. Biol. 15, 151 (2015).
Fritsch, M., Wollesen, T. & Wanninger, A. Hox and ParaHox gene expression in early body plan patterning of polyplacophoran mollusks. J. Exp. Zoolog. B: Mol. Dev. Evol. 326, 89–104 (2016).
Hinman, V. F., O’Brien, E. K., Richards, G. S. & Degnan, B. M. Expression of anterior Hox genes during larval development of the gastropod Haliotis asinina. Evol. Dev. 5, 508–521 (2003).
Samadi, L. & Steiner, G. Expression of Hox genes during the larval development of the snail, Gibbula varia (L.)—further evidence of non-colinearity in molluscs. Dev. Genes Evol. 220, 161–172 (2010).
Wollesen, T., Rodríguez Monje, S. V., Luiz De Oliveira, A. & Wanninger, A. Staggered Hox expression is more widespread among molluscs than previously appreciated. Proc. R. Soc. B: Biol. Sci. 285, 20181513 (2018).
Bakalenko, N. I., Novikova, E. L., Nesterenko, A. Y. & Kulakova, M. A. Hox gene expression during postlarval development of the polychaete Alitta virens. EvoDevo 4, 13 (2013).
Irvine, S. Q. & Martindale, M. Q. Expression patterns of anterior Hox Genes in the Polychaete Chaetopterus: correlation with morphological boundaries. Dev. Biol. 217, 333–351 (2000).
Kulakova, M. et al. Hox gene expression in larval development of the polychaetes Nereis virens and Platynereis dumerilii (Annelida, Lophotrochozoa). Dev. Genes Evol. 217, 39–54 (2007).
Wei, M. et al. Echiuran Hox genes provide new insights into the correspondence between Hox subcluster organization and collinearity pattern. Proc. R. Soc. B: Biol. Sci. 289, 20220705 (2022).
Kourakis, M. J. et al. Conserved anterior boundaries of hox gene expression in the central nervous system of the leech Helobdella. Dev. Biol. 190, 284–300 (1997).
Nardelli-Haefliger, D., Bruce, A. E. E. & Shankland, M. An axial domain of HOM/Hox gene expression is formed by morphogenetic alignment of independently specified cell lineages in the leech Helobdella. Development 120, 1839–1849 (1994).
Technau, G. M., Berger, C. & Urbach, R. Generation of cell diversity and segmental pattern in the embryonic central nervous system of Drosophila. Dev. Dyn. 235, 861–869 (2006).
Serano, J. M. et al. Comprehensive analysis of hox gene expression in the amphipod crustacean Parhyale hawaiensis. Dev. Biol. 409, 297–309 (2016).
Ikuta, T., Yoshida, N., Satoh, N. & Saiga, H. Ciona intestinalis Hox gene cluster: its dispersed structure and residual colinear expression in development. Proc. Natl. Acad. Sci. 101, 15118–15123 (2004).
Wada, H., Garcia-Fernàndez, J. & Holland, P. W. H. Colinear and segmental expression of Amphioxus Hox genes. Dev. Biol. 213, 131–141 (1999).
Hejnol, A. & Martindale, M. Q. Coordinated spatial and temporal expression of Hox genes during embryogenesis in the acoel Convolutriloba longifissura. BMC Biol 7, 65 (2009).
Moreno, E., Nadal, M., Baguñà, J. & Martínez, P. Tracking the origins of the bilaterian Hox patterning system: insights from the acoel flatworm Symsagittifera roscoffensis. Evol. Dev. 11, 574–581 (2009).
Suska, A., Miguel-Aliaga, I. & Thor, S. Segment-specific generation of Drosophila Capability neuropeptide neurons by multi-faceted Hox cues. Dev. Biol. 353, 72–80 (2011).
Zheng, C., Diaz-Cuadros, M. & Chalfie, M. Hox Genes promote neuronal subtype diversification through posterior induction in Caenorhabditis elegans. Neuron 88, 514–527 (2015).
Dasen, J. S., Liu, J.-P. & Jessell, T. M. Motor neuron columnar fate imposed by sequential phases of Hox-c activity. Nature 425, 926–933 (2003).
Goto, T., Katayama-Kumoi, Y., Tohyama, M. & Yoshida, M. Distribution and development of the serotonin-and RFamide-like immunoreactive neurons in the arrowworm, Paraspadella gotoi (Chaetognatha). Cell Tissue Res. 267, 215–222 (1992).
Harzsch, S. et al. Fine structure of the ventral nerve centre and interspecific identification of individual neurons in the enigmatic Chaetognatha. Zoomorphology 128, 53–73 (2009).
Harzsch, S. & Wanninger, A. Evolution of invertebrate nervous systems: the Chaetognatha as a case study. Acta Zool 91, 35–43 (2010).
Rieger, V. et al. Development of the nervous system in hatchlings of Spadella cephaloptera (Chaetognatha), and implications for nervous system evolution in Bilateria: Nervous system development in S. cephaloptera. Dev. Growth Differ. 53, 740–759 (2011).
Arnaud, J., Brunet, M., Casanova, J.-P., Mazza, J. & Pasqualini, V. Morphology and ultrastructure of the gut in Spadella cephaloptera (Chaetognatha). J. Morphol. 228, 27–44 (1996).
Perez, Y., Arnaud, J., Brunet, M., Casanova, J.-P. & Mazza, J. Morphological study of the gut in Sagitta setosa, S. serratodentata and S. pacifica (Chaetognatha). Functional implications in digestive processes. J. Mar. Biol. Assoc. U.K. 79, 1097–1109 (1999).
Bienz, M. Homeotic genes and positional signalling in the Drosophila viscera. Trends Genet. 10, 22–26 (1994).
Kikuchi, M., Omori, A., Kurokawa, D. & Akasaka, K. Patterning of anteroposterior body axis displayed in the expression of Hox genes in sea cucumber Apostichopus japonicus. Dev. Genes Evol. 225, 275–286 (2015).
Beck, F., Tata, F. & Chawengsaksophak, K. Homeobox genes and gut development. BioEssays 22, 431–441 (2000).
Yokouchi, Y., Sakiyama, J.-I. & Kuroiwa, A. Coordinated expression of Abd-B subfamily genes of the HoxA cluster in the developing digestive tract of chick embryo. Dev. Biol. 169, 76–89 (1995).
Seo, H.-C. et al. Hox cluster disintegration with persistent anteroposterior order of expression in Oikopleura dioica. Nature 431, 67–71 (2004).
Monteiro, A. S. & Ferrier, D. E. K. Hox genes are not always Colinear. Int. J. Biol. Sci. 95–103 https://doi.org/10.7150/ijbs.2.95 (2006).
Gasmi, S. et al. Evolutionary history of Chaetognatha inferred from molecular and morphological data: a case study for body plan simplification. Front. Zool. 11, 84 (2014).
Pascual-Anaya, J., D’Aniello, S., Kuratani, S. & Garcia-Fernàndez, J. Evolution of Hoxgene clusters in deuterostomes. BMC Dev. Biol. 13, 26 (2013).
Fodor, A. C. A. et al. The degenerate tale of Ascidian tails. Integr. Comp. Biol. 61, 358–369 (2021).
Herrmann, B. G. The mouse Brachyury (T) gene. Semin. Dev. Biol. 6, 385–394 (1995).
Schwaner, M. J. et al. Future tail tales: a forward-looking, integrative perspective on tail research. Integr. Comp. Biol. 61, 521–537 (2021).
Takada, N., Goto, T. & Satoh, N. Expression pattern of the Brachyury gene in the arrow worm paraspadella gotoi (chaetognatha). Genesis 32, 240–245 (2002).
Park, T.-Y. S. et al. A giant stem-group chaetognath. Sci. Adv. 10, eadi6678 (2024).
Walcott, C. D. Cambrian Geology and Paleontology II, No.5—Middle Cambrian Annelids. (Smithsonian Institution, 1911).
Briggs, D. E. G. & Caron, J.-B. A large Cambrian Chaetognath with supernumerary grasping spines. Curr. Biol. 27, 2536–2543.e1 (2017).
Vannier, J., Steiner, M., Renvoisé, E., Hu, S.-X. & Casanova, J.-P. Early Cambrian origin of modern food webs: evidence from predator arrow worms. Proc. R. Soc. B: Biol. Sci. 274, 627–633 (2007).
Ordoñez, J. F. & Wollesen, T. Anterior–posterior patterning in the chaetognath Spadella cephaloptera: insights into the evolution of the bilaterian nervous system and postanal tail. Zenodo https://doi.org/10.5281/ZENODO.17705168 (2025).
Acknowledgements
The authors are grateful for financial support from the Faculty of Life Sciences of the University of Vienna. This research was funded in whole, or in part, by the Austrian Science Fund (FWF) [P34665].
Author information
Authors and Affiliations
Contributions
J.F.O. designed and performed the experiments, performed the analyses, and wrote the manuscript. T.W. designed the experiments, designed and synthesized some riboprobes, secured funding, supervised, and contributed to analyzing the results and to writing the manuscript. Both authors have read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: Edwina McGlinn and Michele Repetto.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ordoñez, J.F., Wollesen, T. Anterior-posterior patterning in the chaetognath Spadella cephaloptera informs bilaterian nervous system and tail evolution. Commun Biol 9, 122 (2026). https://doi.org/10.1038/s42003-025-09398-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-09398-6