Abstract
Standardizing structural isomeric relationships and evaluating their distribution in chemical space remain major challenges in cheminformatics. Conventional molecular fingerprints and dimensionality reduction techniques are often sensitive to dataset size and structural complexity. Here, we introduce a molecular fingerprint, Structural Isomer Cumulative molecular fingerprint (SIC), that quantitatively captures relative structural differences among isomers with high precision. SIC consists of two variables: SICem, representing exact mass, and SICL, a cumulative descriptor derived from substructural differences. SICL enables calculation of relative structural distances within isomeric groups regardless of dataset size or molecular complexity. Using SIC, we successfully quantified structural differences across positional, skeletal, and functional group isomers, which were not adequately captured by existing descriptors. Furthermore, a scatter plot of SICem and SICL visualized metabolite distributions among cellular compartments, and nine endogenous metabolites were identified whose structural characteristics suggest potential toxicity.
Similar content being viewed by others
Introduction
Molecular fingerprints have been widely employed for compound similarity assessment, biological activity evaluation, and the construction of predictive models1,2,3, establishing themselves as core technologies in cheminformatics and drug discovery. Among widely used molecular fingerprints, the Atom Pair fingerprint (AP)4 represents descriptors based on interatomic distances across the entire molecule, enabling global structural evaluation. However, its sensitivity to subtle substructural differences is limited. In contrast, Extended-Connectivity Fingerprints (ECFP5) compute local environments around each atom, achieving high resolution for identifying substructures, but they struggle to capture holistic features such as overall molecular size and shape6. To address these limitations, MAP46 and MAP4C7 have been developed, which are less sensitive to molecular size and capable of more accurately encoding substructural information. Nonetheless, these approaches still face challenges in continuously and relatively evaluating distances between structural isomers. In particular, for compound groups comprising positional, skeletal, or functional group isomers, conventional fingerprint-based similarity metrics often fail to appropriately reflect their structural differences. Currently, no established method exists for continuously and relatively evaluating such fine-grained structural variations among isomers, which remains an important yet underexplored issue in many applied domains, including toxicity prediction, bioactive molecule screening, and metabolite identification. A numerical descriptor that enables the continuous and relative characterization of structural isomerism using a single variable is especially valuable for applications such as assessing the coverage of in-house compound libraries for unknown compound identification and conducting structure-based toxicity screening. In this study, we propose a novel molecular descriptor, the SIC fingerprint, which aims to quantify and standardize structural isomeric relationships. SIC consists of two components: SICem, representing the exact mass, and SICL, a cumulative structural distance metric derived from substructural differences. This framework enables visualization and comparison of chemical space within isomeric compound groups, independent of dataset size or molecular weight. The method was applied to various datasets containing endogenous and toxic compounds, including YMDB8,9, ECMDB10,11, HMDB12,13, T3DB14, and TOXRIC15. Compared to conventional fingerprints such as ECFP and Atom Pair, SIC demonstrated superior ability to evaluate structural isomeric relationships as relative structural distances. Furthermore, analysis based on SICL identified nine endogenous human metabolites whose structural features were located in close proximity to known toxic compounds in the SICL-defined chemical space, suggesting potential toxicological relevance.
Results
The Structural Isomer Cumulative molecular fingerprint (SIC) provides a standardized framework for representing datasets consisting of structurally isomeric compounds
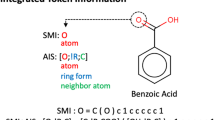
Conventional molecular fingerprints often exhibit substantial bias due to factors such as dataset size, molecular weight, and structural complexity. Furthermore, when chemical space is visualized using dimensionality reduction methods such as principal component analysis (PCA), it is often difficult to interpret the resulting distributions in terms of structural features. The SIC addresses these challenges by providing a structural distance metric that is relatively insensitive to molecular size and dataset scale. SIC enables continuous and relative quantification of structural differences, ranging from subtle substructural variations to large-scale scaffold-level differences. Moreover, when compounds are visualized in the two-dimensional space defined by SICL and SICem, the resulting chemical space exhibits high interpretability (Fig. 1). SIC consists of two variables: SICL, a structural distance metric, and SICem, the monoisotopic exact mass. An overview of the SICL calculation is shown in Fig. 2. To calculate SICL, compounds are first grouped by molecular formula to ensure relative comparisons within isomeric groups. For each compound, a list of substructures is defined based on atom and bond information (Supplementary Data 1). The molecular center was defined as the centroid of the 2D atomic coordinates of all atoms, generated using RDKit. The planar distance from this center (Lsub) was then computed for each substructure. Within each formula group, the median distance (Lmedian) was calculated for each substructure type and used as a reference. When the same substructure pattern occurred multiple times within a molecule, each occurrence was treated as an independent instance. Thus, multiple instances of the same functional group (e.g., R-NH₂) could differ in their classification, with some considered structurally divergent and others not, depending on their individual distances relative to the group median. This design allows SIC to capture positional differences of identical functional groups within a molecule. If the absolute difference between Lsub and Lmedian exceeds a predefined threshold (0.01), the substructure is considered structurally divergent. The product of its distance (Li) and molecular weight (Mi) is computed. The sum of these products across all such substructures is denoted as S, and the SICL value is obtained by normalizing this sum with the monoisotopic mass M:
The SIC fingerprint consists of two variables: SICL, which represents the cumulative structural distance between differing substructures, and SICem, which is the exact monoisotopic mass of the compound. This design enables explicit quantification of relative structural differences among isomeric compounds sharing the same molecular formula.
Structurally isomeric compounds are grouped by molecular formula, and predefined substructures are extracted. For each substructure, the planar distance from the geometric center of the molecule is calculated (Lsub) and compared to the median distance across all compounds in the group (Lmedian). If the deviation exceeds a defined threshold, the substructure is considered distinct, and its contribution to the cumulative structural difference is weighted by molecular weight (M). Red circle: geometric center of the molecule based on atomic coordinates. Red arrows: distances from substructures to the molecular center. Blue arrows: median distances (Lmedian) for each substructure across the group. Lsub: substructure-to-center distance in each compound. Lmedian: median substructure distance across all compounds. M: molecular weight of each substructure.
In this formulation, compounds with many substructures that are both heavy and distant from the group median contribute to a higher SICL value. The second axis, SICem, represents the monoisotopic mass. Therefore, in the two-dimensional SIC space, a wide spread along the SICL axis indicates the presence of diverse structural isomers. Conversely, compounds with similar SICL values tend to be structurally similar, often representing positional isomers with substructures situated at comparable distances.
Benchmarking the performance of chemical space visualization for structurally isomeric compounds
Chemical space visualizations based on conventional molecular fingerprints are often strongly influenced by dataset size and molecular weight. This effect becomes particularly pronounced in structurally isomeric compounds, which share the same molecular formula but differ in structure, making similarity evaluation difficult. In this study, we compared the visualization performance of several existing molecular fingerprints (MQN16, MAP4C, MHFP17, MACCS Key18, RDKit19, ECFP, and AP) with that of my newly developed fingerprint, the SIC, using principal component analysis (PCA). The evaluation was conducted on two isomer sets retrieved from PubChem: compounds with the formula C₆H₆O₂ (377 compounds) and C₄₈H₈₉NO₁₈ (31 compounds). Dimensionality reduction was performed using PCA, a linear method with high interpretability and reproducibility20. As a result, MQN, MAP4C, MHFP, MACCS Key, RDKit, and ECFP exhibited significantly biased distributions depending on dataset size and molecular weight (Fig. 3). In particular, AP showed a strong dependency on molecular weight. These fingerprints failed to produce chemically interpretable distributions. In contrast, SIC was minimally affected by dataset size or molecular weight. The first principal component (PC1) primarily reflected molecular weight, while the second (PC2) captured subtle structural differences. These results suggest that SIC is a highly effective method for visualizing and comparing structurally isomeric compounds.
Structurally isomeric compounds with the same molecular formula were retrieved from PubChem. Red circles represent compounds with the formula C₆H₆O₂ (n = 377), and blue circles represent compounds with C₄₈H₈₉NO₁₈ (n = 31). The PCA distribution shows that conventional fingerprints are strongly influenced by dataset size and molecular weight, leading to biased chemical space representations.
Performance evaluation of molecular fingerprints/Tanimoto similarity scores and scaled SICL
SICL inherently represents structural diversity within a compound set as relative distances between compounds, thus eliminating the need for pairwise comparisons such as those used in conventional Tanimoto similarity. However, to ensure a fair performance comparison, this study compared SICL with conventional molecular fingerprints and Tanimoto similarity-based structural similarity evaluations. SICL values were divided by their maximum value and scaled to the range 0–1. To enable pairwise comparison using SICL, the absolute difference between the normalized SICL values for each compound pair was calculated, and this difference was subtracted from one to derive a pairwise similarity score. This procedure reinterprets the numerical distance between SICL values as a measure of “similarity,” allowing direct comparison with traditional fingerprint-based methods evaluated by Tanimoto coefficients. The evaluation was performed using five sets of structural isomers obtained from PubChem: C₆H₆O₂ (377 compounds), C₁₂H₁₄O₇S (141 compounds), C₆H₁₆O₁₈P₄ (22 compounds), C₃₉H₇₉N₂O₆P (79 compounds), and C₄₈H₈₉NO₁₈ (31 compounds). The compounds selected for these evaluations were determined based on molecular formula and molecular weight, which fall within the structural evaluation range targeted by SIC.
First, we compared the average similarity scores by molecular weight between the molecular fingerprints with average Tanimoto similarity scores from previous studies and the SICL with average pairwise similarity scores obtained in this study (Fig. 4a). The results showed that the average Tanimoto similarity scores of conventional molecular fingerprints tended toward 1.0 with increasing molecular weight. In contrast, the scaled SICL remained stable between ~0.5 and 0.8, indicating that it is independent of molecular weight. The average pairwise similarity score derived from the scaled SICL also remained stable, between ~0.8 and 0.9, similarly showing molecular weight-independent behavior. The higher average pairwise similarity score compared with the scaled SICL itself can be explained by the definition of SICL, which represents the relative differences in the mass and structural distances of substructures within each molecular formula group. Consequently, computing pairwise similarity from SICL values, already representing relative structural distances, leads to redundant evaluation of the same structural relationships.
a Comparison of average Tanimoto similarity scores and SICL across molecular formula groups. SICL_Norm represents SICL values normalized by the maximum, while SICL_Norm_pairwise denotes pairwise similarity scores calculated from normalized SICL values within each formula group. b Distribution of pairwise similarity scores for each molecular formula group (box plots).
Next, we compared the distribution of average pairwise similarity scores across different molecular weights (Fig. 4b). The results showed that for conventional molecular fingerprints with average Tanimoto similarity scores, the interquartile range (IQR) narrowed and tended toward 1.0 as molecular weight increased. In contrast, the scaled SICL exhibited a stable IQR between ~0.8 and 0.95, demonstrating molecular weight-independent behavior.
As similarity scores approach 1.0, they indicate higher structural similarity. Previous studies have discussed the limitations of conventional fingerprints in distinguishing substructural differences among high molecular-weight compounds6,21. The SIC developed in this study can evaluate structural similarity independently of molecular weight. Unlike conventional approaches, it does not require calculating pairwise similarity scores using molecular fingerprints. Instead, it directly evaluates the structural differences among compounds within the same molecular formula group as relative distance values.
Benchmarking chemical space visualization performance using public compound databases
To evaluate the practical utility of SIC, we conducted a comparative analysis using multiple compound databases spanning from prokaryotic to eukaryotic organisms. As reference methods, we selected AP and ECFP, which, although clearly influenced by dataset size and molecular weight, showed relatively less distortion in distribution compared to other fingerprints, as demonstrated in the results of Fig. 3. First, using the ECMDB, YMDB, and HMDB datasets (Supplementary Data 2), we performed chemical space visualizations. The results indicated that AP and ECFP tended to produce compressed distributions due to data size and molecular weight effects, resulting in limited coverage of chemical space (Fig. 5). In contrast, SIC demonstrated the broadest distribution, effectively capturing a greater range of structural diversity. This trend was consistent across HMDB and YMDB, which represent complex metabolic networks of eukaryotic organisms and encompass broader chemical diversity. By contrast, ECMDB showed a narrower spread, likely reflecting the simpler metabolic repertoire of E. coli, with its limited range of molecular weights and structural types. To assess whether this broader distribution holds biological relevance, we evaluated the structural diversity of compounds associated with different organelles, based on subcellular annotations provided in HMDB. The SIC-based visualization revealed that small-molecule metabolites were broadly distributed in the nucleus and mitochondria, whereas the cytosol exhibited a more diverse distribution, including higher-molecular-weight compounds. This trend may reflect functional differences in metabolism among subcellular compartments. To verify that these observed distributions were not simply due to differences in dataset size, we compared two databases with relatively similar numbers of compounds: HMDB (2933 endogenous metabolites) and T3DB (3457 toxic compounds). This analysis aimed to evaluate differences in chemical space coverage based on structural diversity alone. The results showed that with AP and ECFP, compound distributions from both databases largely overlapped, and no distinct clustering was observed. In contrast, SIC revealed clear distinctions: HMDB compounds exhibited broad diversity, including high-molecular-weight lipids, whereas T3DB compounds showed structural diversity biased toward low-molecular-weight compounds. Taken together, these findings suggest that SIC is a useful method for effectively evaluating structural diversity across a wide range of compounds, from endogenous metabolites to toxic xenobiotics.
YMDB and ECMDB datasets included only metabolites that were both detected and quantified. For HMDB, only compounds categorized as “detected and quantified”, “endogenous”, and “organelle-associated” were included. Yellow circles indicate metabolites detected in the cytoplasm, green circles represent those detected in the nucleus, and red circles correspond to those detected in mitochondria. For the comparison between HMDB and T3DB, red circles denote metabolites from HMDB that were both detected and quantified, while blue circles represent compounds recorded in T3DB.
Evaluation of SIC utility based on structural distances between endogenous eukaryotic metabolites and toxic compounds
Among endogenous metabolites in eukaryotes, some compounds are listed in toxic compound databases such as T3DB and TOXRIC (Supplementary Data 3). Compounds structurally similar to known toxic substances are considered to carry a higher risk of toxicity22. Therefore, the accurate identification of metabolites structurally analogous to toxic compounds (even at the level of positional isomers) is considered critically important for risk assessment. The SIC enables the calculation of structural distances based on relative differences between structural isomers. To evaluate the performance and utility of SIC, we analyzed endogenous metabolites (from HMDB and YMDB) and toxic compounds (from T3DB and TOXRIC) to identify potential novel toxic candidates among endogenous metabolites. Using InChIKeys, we annotated endogenous metabolites that are already listed as toxic compounds in those databases. Then, Compounds were grouped based on molecular formulae derived from their canonical SMILES and calculated SIC values. For visualization, SICL and SICem were directly used as axes, allowing chemical space mapping based on structural similarity and substructural differences weighted by molecular mass (Fig. 6). Notably, several endogenous metabolites without known toxicity were found to cluster closely with known toxic compounds, particularly at the level of positional or functional group isomers. To further validate this, we manually examined the top nine endogenous metabolites that showed the smallest SICL differences to a toxic compound, sorted within each group of compounds sharing the same molecular formula (Fig. 7). The dataset used comprised pairs where one compound was a known toxicant and the other had no toxicity annotation despite being a structural isomer. The threshold of the top nine was selected because there were 94 compound pairs with SICL scores above 0.9, and manual evaluation of all cases was not practical (Fig. 8c). Thus, this analysis represents a pilot assessment to verify the plausibility of SIC-based toxic analog identification. Within the top nine, compounds such as 6-Hydroxynicotinic acid (HMDB0002658, C6H5NO3) and 2-Butanol (HMDB0011469, C4H10O) were judged to have low toxicity potential. 2-Hydroxyfluorene (HMDB0013163, C13H10O), while part of a dataset that included a toxic positional isomer, was matched as the most similar compound based on a functional group isomer, indicating a possible false positive in the SICL score. However, Methylsuccinic acid (HMDB0001844, C5H8O4) is a positional isomer of a known toxic compound and therefore may carry a significantly high toxicological risk, despite lacking current toxicity annotation. Ethyl hexanoate (YMDB01381, C8H16O2) was a compound for which the toxic form could be generated through ester hydrolysis via esterases (EC 3.1.1.x class). Pyruvaldehyde (HMDB0001167, C3H4O2) and Propanal (HMDB0003366, C3H6O), both bearing reactive aldehyde groups, were also found to be highly similar to toxic counterparts and may represent hazardous endogenous compounds, especially in the context of aging-related accumulation and abnormal chemical modifications23,24,25,26,27,28,29,30,31.
Red circles (Positive) represent compounds with reported toxicity, while blue circles (Negative) indicate endogenous metabolites with no known toxicity reports. Compounds are plotted using SICL (y-axis) and SICem (x-axis), which represent structural distance and exact monoisotopic mass, respectively.
For each toxic compound, the top nine endogenous metabolites with the smallest SICL distance from the toxic compound were selected from the HMDB and YMDB databases. Manual curation was performed to assess the potential toxicity or structural plausibility of each candidate. Examples include metabolites with functional groups or backbone structures closely resembling known toxic compounds, as well as cases judged unlikely to exhibit toxicity.
a SICL-based structural distance distribution for all possible pairs of toxic compounds and non-toxic endogenous metabolites (from YMDB and HMDB) sharing the same molecular formula. Plots indicate the number of pairs with absolute SICL differences below each threshold. b Comparison of similarity scores using AP, ECFP, and SICL fingerprints. For AP and ECFP, Tanimoto coefficients were used; for SICL, scores were normalized to a 0–1 scale by dividing by the maximum value. Blue circles indicate AP, red circles ECFP, and black circles SICL. c Evaluation of high-similarity pairs (similarity ≥ 0.9) between toxic compounds and non-toxic metabolites using each fingerprint. The number of positional isomer pairs and the number of falsely identified identical compounds (similarity = 1.0) were also assessed. Notably, SICL yielded no pairs with identical similarity scores (n = 0), whereas AP and ECFP produced 4 and 3 such cases, respectively.
Finally, we compared SICL scores with Tanimoto coefficients calculated using AP and ECFP fingerprints, scaling SICL to a 0–1 range. All pairs of toxicants and structurally related nontoxic endogenous metabolites were analyzed (Fig. 8). The majority of absolute SICL differences were below 1.0 (Fig. 8a). Each score was scaled by its maximum value and compared to AP and ECFP (Fig. 8b). SICL produced unique scores for every compound pair, sharply distinguishing structural differences with minimal redundancy (Supplementary Data 4). To evaluate the characteristics of high-similarity pairs, we analyzed those with scores ≥0.9. The focus of this comparison was not the number of pairs per se, but the proportion of structural isomers and the occurrence of false-perfect matches (score = 1.0 between distinct structures). SICL identified 94 such pairs, with a higher proportion of structural isomers and no false-perfect matches, whereas AP and ECFP identified 26 and 27 pairs, respectively, including several false-perfect matches (Fig. 8c). Furthermore, the proportion of positional isomeric pairs among high-similarity matches (score ≥ 0.9) was 22.3% for SICL, compared to 0% and 2.85% for ECFP and AP, respectively. Collectively, these findings demonstrate that SICL exhibits superior discriminative power in evaluating fine structural similarity between isomeric compounds, particularly in identifying endogenous metabolites closely resembling toxic compounds. Additionally, SICL provides a novel approach for capturing structural proximity beyond the reach of traditional molecular fingerprints evaluated using Tanimoto similarity, making it a promising tool for toxicological prediction and safety assessment.
Discussion
Conventional molecular fingerprint-based similarity evaluations are often influenced by dataset size and molecular complexity, and have continued to face challenges in the continuous and relative assessment of structurally isomeric compound groups. To address this issue, we developed a method that accumulates structural differences within compound groups sharing the same molecular formula, based on substructural differences and their associated molecular weights. Traditional approaches assess chemical space using overall molecular structure information, and therefore have limited capacity to distinguish small differences. In contrast, the SIC method computes structural distances using variables derived from differing substructures and molecular weight. This design minimizes the impact of dataset size and molecular weight, allowing for relative evaluation of structural distances even at the level of positional isomers.
To evaluate the utility of SIC, we compared its performance with that of existing molecular fingerprints. The comparison used isomer datasets with differing molecular weights and dataset sizes (C₆H₆O₂: 377 compounds; C₄₈H₈₉NO₁₈: 31 compounds) and employed principal component analysis (PCA) to visualize chemical space. we focused primarily on small molecules with molecular weights under 1000 Da. To ensure a fair performance comparison, this study evaluated SICL alongside conventional molecular fingerprints and their Tanimoto similarity scores. The evaluation was conducted using five sets of structural isomers obtained from PubChem: C₆H₆O₂ (377 compounds), C₁₂H₁₄O₇S (141 compounds), C₆H₁₆O₁₈P₄ (22 compounds), C₃₉H₇₉N₂O₆P (79 compounds), and C₄₈H₈₉NO₁₈ (31 compounds). Comparisons with larger or extremely small molecules were not conducted, and therefore future studies will be needed to explore SIC’s applicability across a broader range. In addition, we assessed the generalizability of SIC using diverse metabolite databases derived from prokaryotic and eukaryotic organisms (HMDB, YMDB, ECMDB), as well as toxic compound datasets (T3DB). The results showed that SICem and SICL explicitly represented structural distributions based on molecular weight and diversity. We also plan to evaluate how increasing dataset size affects chemical space visualization to assess its effectiveness for mapping the diversity of natural products. Furthermore, in toxicity assessments using SIC, we were able to identify numerous endogenous metabolites with structures highly similar to known toxic compounds, which had not been detected using previous methods. However, manual curation was labor-intensive, highlighting the need for standardized annotation criteria in the future.
The benchmarking conducted in this study addressed several key issues in prior methods, including sensitivity to molecular weight and dataset size, limited resolution of substructural differences, and general applicability. As a result, we demonstrated that SIC can effectively minimize the influence of dataset size and molecular weight, and enable relative evaluation of structural distances at the level of positional isomers. Compared to existing molecular fingerprints, SIC showed greater clarity in capturing differences among structural isomers while reducing dependency on dataset size and molecular weight. Conventional visualization techniques for chemical space struggle to represent structural diversity explicitly. In contrast, SIC, comprising two fingerprint variables (SICL and SICem), enabled clear visualization of structural differences as relative distances. As shown in Fig. 5, the compound distribution was well organized, with SICem capturing molecular weight variation, while SICL reflected substructural differences. SIC also demonstrated high discriminative power by sensitively extracting endogenous metabolites structurally close to toxic compounds from databases such as T3DB and TOXRIC. Unlike previous methods, SIC avoided issues such as redundant scores or score saturation, allowing for the assignment of unique structural distances.
SIC minimizes the influence of dataset size and molecular weight and effectively captures differences among structural isomers. However, as it quantifies relative structural distances within groups sharing the same molecular formula, its application becomes unstable when the number of compounds in a group is insufficient, making practical use difficult in such cases. Therefore, identifying the minimum number of required comparison compounds for reliable SIC evaluation remains an important issue. On the other hand, SIC does not require the construction of predictive models and enables explicit evaluation of structural distances, making it a fast and simple tool for structural assessment, without the need for prior domain validation or large datasets typically required for machine learning approaches. Looking ahead, SIC may be extended to evaluate stereoisomers; however, this would require overcoming computational challenges such as incorporating quantum chemical calculations to assess structural stability.
A limitation of the present SIC implementation is that it relies on a predefined list of substructures. In this study, the evaluation was centered on endogenous metabolites and toxic compounds, and therefore the atomic types included were restricted to those most relevant to this context. From the standpoint of general chemical research, the omission of elements such as B or Si may appear unusual, and this limitation should be acknowledged. In addition, the current approach may yield false positives when substructures with nearly identical molecular weights are cumulatively evaluated, leading to compounds with different scaffolds being placed in closer proximity than expected along the SIC axis. This reflects the fact that SIC quantifies structural differences using 2D atomic coordinates and substructure molecular weights, which may not fully capture subtle electronic effects. In future work, the SMARTS list can be expanded to incorporate additional elements, and integration of 3D structural descriptors or quantum chemical information such as electron density distributions could provide a more robust evaluation of structural diversity. In addition, the dataset dependence of SICL has both advantages and drawbacks. For metabolomic applications, variability across datasets is useful for comparing isomeric repertoires between organisms, whereas in contexts such as drug discovery, dataset dependence may hinder evaluation of candidate chemical spaces. In such cases, fixed reference values for each substructure type would be preferable.
SIC offers a novel approach to the explicit quantification of structural differences among isomers and opens new possibilities in the evaluation of chemical diversity and toxicological screening. Future work will include expanding its application to stereochemical isomers and quantitatively validating its relationship with biological similarity. In addition, integration with clustering or visualization methods based on structural distance may enhance the accuracy and interpretability of compound classification and discovery pipelines.
Methods
Method for calculating the structural isomer cumulative molecular fingerprint
The calculation of the Structural Isomer Cumulative molecular fingerprint (SIC) was performed as follows:
1. Grouping by molecular formula: Canonical SMILES strings were used to calculate molecular formulas, and compounds with identical formulas were grouped together.
2. Substructure distance calculation: For each compound, a predefined list of substructures—each defined by a single atom and its bond types—was used to compute the 2D atomic coordinates of atoms (generated using RDKit). The molecular center was defined as the centroid of these coordinates, and the distance to this center (Lsub) was calculated for each substructure. This list was systematically constructed to comprehensively cover covalent bonding patterns of the atomic types analyzed in this study.
3. Median distance determination: For each substructure type across the group, the median distance Lmedian was computed.
4. Cumulative distance calculation: For each substructure where | Lsub - Lmedian |≧0.01 and substructure molecular weight Mi were recorded. The cumulative structural score S was then calculated as:
where i is the index of the substructure and n is the number of substructures meeting the threshold condition. The threshold of 0.01 was determined empirically through benchmarking analyses across datasets with different molecular weights and sample sizes. Thresholds ranging from 1 to 0.0001 were tested, and 0.01 was found to yield the most stable and consistent distributions (Supplementary Data 5).
5. Normalization and fingerprint output: The final SIC values consisted of two variables: (1) SICL, defined as the normalized cumulative score S/M, where M is the monoisotopic exact mass of the compound, and (2) SICem, the exact mass M itself (rounded to six significant digits).
The SIC fingerprint is designed such that SICL increases when the substructures of a compound have both larger spatial deviation (SICL) from the median and higher molecular weights (Mi) within a group of structural isomers. By normalizing S with respect to M, variation due to overall molecular size is minimized. The substructure list was generated using SMART notation, covering atoms of C, N, P, O, S, H, Cl, Br, I, F, and Hg, with classification based on bonding type and coordination. Molecular weights Mi for substructures included the atomic mass of the central atom and its attached hydrogens. The threshold value of 0.01 for | Lsub - Lmedian | was selected based on performance evaluation, as it maximized the ability to distinguish positional isomers in compounds with molecular weights below 1000. All computations were implemented using Python 3.11.5, with RDKit 2023.3.2, pandas 1.3.5, SciPy 1.7.3, and NumPy 1.21.6. The SIC tool developed in this study is available at BioChemCalc (BCC, https://biochemcalc.com/).
Chemical space visualization
Molecular fingerprints were calculated using RDKit (version 2023.3.2) and mapchiral (version 0.0.7). For the comparison with existing fingerprints, the following specifications were used: Atom Pair (count fingerprint, 2048 bits), Morgan (binary, radius = 2, 2048 bits; corresponding to ECFP4), MACCS Keys (binary, 165 bits), MAP4C (binary, 2048 bits), MHFP (MinHash fingerprint, 128 dimensions), MQN (count, 42 dimensions), and RDKit (binary, 2048 bits). Prior to principal component analysis (PCA), all fingerprint vectors were standardized by Z-score normalization so that each feature had a mean of 0 and a standard deviation of 1. PCA was performed with scikit-learn (version 1.0.2), and the resulting scores were exported to Microsoft Excel for graphical visualization.
Use of public databases
Datasets of endogenous metabolites from both prokaryotic and eukaryotic organisms were retrieved from YMDB8,9 (Detected and quantified), ECMDB10,11, and HMDB12,13 (Detected and quantified, Endogenous). Toxic compounds were obtained from T3DB14 and TOXRIC15 (Toxicity Category: Carcinogenicity). Since the datasets included stereoisomeric information, InChIKeys were computed using RDKit from canonical SMILES. These InChIKeys were then used to merge stereoisomers into structural isomers. Additionally, metabolites from YMDB and HMDB that matched InChiKeys in T3DB and TOXRIC were extracted and classified as toxic compounds.
Data availability
The source data supporting the figures and graphs are provided in the Supplementary Data file. Additional data generated or analyzed during this study are available from the corresponding author upon reasonable request.
Code availability
The tools and source code developed are openly available and freely accessible under an open-access model at https://biochemcalc.com. They may be used without restriction for academic purposes. The source code implementing the SIC algorithm is openly available at https://github.com/TaiheiTorigoe/SIC and archived on Zenodo (DOI: 10.5281/zenodo.17446622) under the MIT License32.
References
Gomez-Bombarelli, R. et al. Automatic chemical design using a data-driven continuous representation of molecules. ACS Cent. Sci. 4, 268–276 (2018).
Wu, Z. et al. MoleculeNet: a benchmark for molecular machine learning. Chem. Sci. 9, 513–530 (2018).
Awale, M. & Reymond, J. L. The polypharmacology browser: a web-based multi-fingerprint target prediction tool using ChEMBL bioactivity data. J. Cheminform. 9, 11 (2017).
Carhart, R. E., Smith, D. H. & Venkataraghavan, R. Atom pairs as molecular features in structure-activity studies: definition and applications. J. Chem. Inf. Comput. Sci. 25, 64–73 (1985).
Rogers, D. & Hahn, M. Extended-connectivity fingerprints. J. Chem. Inf. Model 50, 742–754 (2010).
Capecchi, A., Probst, D. & Reymond, J. L. One molecular fingerprint to rule them all: drugs, biomolecules, and the metabolome. J. Cheminform. 12, 43 (2020).
Orsi, M. & Reymond, J. L. One chiral fingerprint to find them all. J. Cheminform. 16, 53 (2024).
Ramirez-Gaona, M. et al. YMDB 2.0: a significantly expanded version of the yeast metabolome database. Nucleic Acids Res. 45, D440–D445 (2017).
Jewison, T. et al. YMDB: the yeast metabolome database. Nucleic Acids Res. 40, D815–D820 (2012).
Sajed, T. et al. ECMDB 2.0: a richer resource for understanding the biochemistry of E. coli. Nucleic Acids Res. 44, D495–D501 (2016).
Guo, A. C. et al. ECMDB: the E. coli metabolome database. Nucleic Acids Res. 41, D625–D630 (2013).
Wishart, D. S. et al. HMDB: the human metabolome database. Nucleic Acids Res. 35, D521–D526 (2007).
Wishart, D. S. et al. HMDB 5.0: the human metabolome database for 2022. Nucleic Acids Res. 50, D622–D631 (2022).
Lim, E. et al. T3DB: a comprehensively annotated database of common toxins and their targets. Nucleic Acids Res. 38, D781–D786 (2010).
Wu, L. et al. TOXRIC: a comprehensive database of toxicological data and benchmarks. Nucleic Acids Res. 51, D1432–D1445 (2023).
Awale, M., van Deursen, R. & Reymond, J. L. MQN-mapplet: visualization of chemical space with interactive maps of DrugBank, ChEMBL, PubChem, GDB-11, and GDB-13. J. Chem. Inf. Model 53, 509–518 (2013).
Probst, D. & Reymond, J. L. A probabilistic molecular fingerprint for big data settings. J. Cheminform. 10, 66 (2018).
Durant, J. L., Leland, B. A., Henry, D. R. & Nourse, J. G. Reoptimization of MDL keys for use in drug discovery. J. Chem. Inf. Comput Sci. 42, 1273–1280 (2002).
Landrum, G. Fingerprints in the RDKit. https://www.rdkit.org/UGM/2012/Landrum_RDKit_UGM.Fingerprints.Final.pptx.pdf (2012).
Jolliffe, I. T. & Cadima, J. Principal component analysis: a review and recent developments. Philos. Trans. A Math. Phys. Eng. Sci. 374, 20150202 (2016).
Bajusz, D., Racz, A. & Heberger, K. Why is Tanimoto index an appropriate choice for fingerprint-based similarity calculations?. J. Cheminform. 7, 20 (2015).
Wassenaar, P. N. H., Rorije, E., Vijver, M. G. & Peijnenburg, W. Evaluating chemical similarity as a measure to identify potential substances of very high concern. Regul. Toxicol. Pharm. 119, 104834 (2021).
Banerjee, S. Methylglyoxal-induced modification of myoglobin: an insight into glycation mediated protein aggregation. Vitam. Horm. 125, 31–46 (2024).
Zhu, Z. et al. Acrolein, an endogenous aldehyde induces synaptic dysfunction in vitro and in vivo: involvement of RhoA/ROCK2 pathway. Aging Cell 21, e13587 (2022).
Li, Y. et al. Oxidative stress and 4-hydroxy-2-nonenal (4-HNE): implications in the pathogenesis and treatment of aging-related diseases. J. Immunol. Res. 2022, 2233906 (2022).
Demir, E. & Marcos, R. Assessing the genotoxic effects of two lipid peroxidation products (4-oxo-2-nonenal and 4-hydroxy-hexenal) in haemocytes and midgut cells of drosophila melanogaster larvae. Food Chem. Toxicol. 105, 1–7 (2017).
Islam, U. L., Moinuddin, B., Mahmood, R. & Ali, A. Genotoxicity and immunogenicity of crotonaldehyde modified human DNA. Int J. Biol. Macromol. 65, 471–478 (2014).
Tong, Z. et al. Accumulated hippocampal formaldehyde induces age-dependent memory decline. Age (Dordr.) 35, 583–596 (2013).
Perluigi, M., Coccia, R. & Butterfield, D. A. 4-Hydroxy-2-nonenal, a reactive product of lipid peroxidation, and neurodegenerative diseases: a toxic combination illuminated by redox proteomics studies. Antioxid. Redox Sig. 17, 1590–1609 (2012).
Garaycoechea, J. I. et al. Genotoxic consequences of endogenous aldehydes on mouse haematopoietic stem cell function. Nature 489, 571–575 (2012).
Stein, S., Lao, Y., Yang, I. Y., Hecht, S. S. & Moriya, M. Genotoxicity of acetaldehyde- and crotonaldehyde-induced 1,N2-propanodeoxyguanosine DNA adducts in human cells. Mutat. Res. 608, 1–7 (2006).
Torigoe, T. SIC_ver_1. Zenodo https://doi.org/10.5281/zenodo.17446622 (2025).
Acknowledgements
I thank Ryohei Torigoe for his generous personal financial support that enabled this research. I also appreciate the valuable feedback and English editing assistance provided by Omidreza Heravizadeh, M.Sc., during the preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
Taihei Torigoe conceived the study, developed the SIC algorithm, performed all computational analyses, created figures and tables, and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The author declares no competing interests.
Peer review
Peer review information
Communications Chemistry thanks Yannick Djoumbou-Feunang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Torigoe, T. Structural Isomer Cumulative molecular fingerprinting method (SIC) for standardizing structural isomeric relationships. Commun Chem 8, 406 (2025). https://doi.org/10.1038/s42004-025-01798-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42004-025-01798-3