Abstract
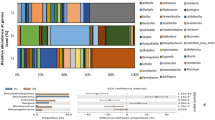
Phylum- and class-specific PCR primers were tested for the production of clone libraries and for denaturing gradient gel electrophoresis (DGGE) analysis of complex bacterial communities. Primers were designed to specifically amplify 16S rRNA gene fragments of the phyla Bacteroidetes, Planctomycetes and Firmicutes, of three classes of the phylum Proteobacteria, the Alphaproteobacteria, Betaproteobacteria and Gammaproteobacteria, and of the Cyanobacteria (including chloroplast 16S rRNA genes). The specificity of the seven primer pairs was tested by producing clone libraries from environmental DNA samples from mesotrophic (Norwegian coastal) and oligotrophic (Northern Atlantic Gyre) environments. Five of the seven primer pairs specifically amplified target 16S rRNA gene sequences. Exceptions were the Betaproteobacteria- and Firmicutes-specific primers, which were relatively successful with coastal water mesocosm samples but less so with the Northern Atlantic Gyre sample. Phylogenetic analysis of sequences from the Gammaproteobacteria clone library revealed that the coastal sample yielded a number of clones that clustered within clades that belong to the oligotrophic marine Gammaproteobacteria (OMG) group, indicating that this group is not confined exclusively to the oligotrophic environment. Comparison of the bacterial diversity of the environmental DNA sample from the coastal and the open ocean using a two- or three-step nested PCR-DGGE process revealed significant differences in the bacterial communities. The application of the group-specific primers provides a higher resolution genetic fingerprinting approach than existing DGGE primer sets.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
References
Amann RI, Ludwig W, Schleifer K-H . (1995). Phylogenetic identification of individual microbial cells without cultivation. Microbiol Rev 59: 143–169.
Ashelford KE, Chuzhanova NA, Fry JC, Jones AJ, Weightman AJ . (2005). At least 1 in 20 16S rRNA sequence records currently held in public repositories is estimated to contain substantial anomalies. Appl Environ Microbiol 71: 7724–7736.
Ashelford KE, Weightman AJ, Fry JC . (2002). PRIMROSE: a computer program for generating and estimating the phylogenetic range of 16S rRNA oligonucleotide probes and primers in conjunction with the RDP-II database. Nucleic Acids Res 30: 3481–3489.
Baker GC, Smith JJ, Cowan DA . (2003). Review and re-analysis of domain-specific 16S primers. J Microbiol Methods 55: 541–555.
Blackwood CB, Oaks A, Buyer JS . (2005). Phylum- and class-specific PCR primers for general microbial community analysis. Appl Environ Microbiol 71: 6193–6198.
Blümel M, Süling J, Imhoff JF . (2007). Depth-specific distribution of Bacteroidetes in the oligotrophic Eastern Mediterranean Sea. Aquat Microb Ecol 46: 209–224.
Brinkmeyer R, Knittel K, Jürgens J, Weyland H, Amann R, Helmke E . (2003). Diversity and structure of bacterial communities in Arctic and Antarctic pack ice. Appl Environ Microbiol 69: 6610–6619.
Calvo L, Vila X, Abella CA, Garcia-Gil LJ . (2004). Use of ammonia-oxidizing bacterial-specific phylogenetic probe Nso1225 as a primer for fingerprint analysis of ammonia-oxidizer communities. Appl Microbiol Biotechnol 63: 715–721.
Cho JC, Giovannoni SJ . (2004). Cultivation and growth characteristics of a diverse group of oligotrophic marine gammaproteobacteria. Appl Environ Microbiol 70: 432–440.
Chouari R, Le Paslier D, Daegelen P, Ginestet P, Weissenbach J, Sghir A . (2005). Novel predominant archaeal and bacterial groups revealed by molecular analysis of an anaerobic sludge digester. Environ Microbiol 7: 1104–1115.
Curtis TP, Sloan WT, Scannell JW . (2002). Estimating prokaryotic diversity and its limits. Proc Natl Acad Sci USA 99: 10494–10499.
Dar SA, Kuenen JG, Muyzer G . (2005). Nested PCR-denaturing gradient gel electrophoresis approach to determine the diversity of sulfate-reducing bacteria in complex microbial communities. Appl Environ Microbiol 71: 2325–2330.
Freitag TE, Prosser JI . (2003). Community structure of ammonia-oxidizing bacteria within anoxic marine sediments. Appl Environ Microbiol 69: 1359–1371.
Garrity GM, Bell JA, Lilburn TG . (2003). Taxonomic Outline of the Procaryotes. Bergey's Manual of Systematic Bacteriology, 2nd edn. Release 4.0. Springer-Verlag: New York. 395 pp.
Giovannoni SJ, Rappé MS, Vergin KL, Adair NL . (1996). 16S rRNA genes reveal stratified open ocean bacterioplankton populations related to the green non-sulfur bacteria. Proc Natl Acad Sci USA 93: 7979–7984.
Glöckner FO, Fuchs BM, Amann R . (1999). Bacterioplankton compositions of lakes and oceans: a first comparison based on fluorescence in situ hybridization. Appl Environ Microbiol 65: 3721–3726.
Hamasaki K, Taniguchi A, Tada Y, Long RA, Azam F . (2007). Actively growing bacteria in the inland sea of Japan, identified by combined bromodeoxyuridine immunocapture and denaturing gradient gel electrophoresis. Appl Environ Microbiol 73: 2787–2798.
Hicks RE, Amann RI, Stahl DA . (1992). Dual staining of natural bacterioplankton with 4′,6-diamidino-2-phenylindole and fluorescent oligonucleotide probes targeting kingdom-level 16S rRNA sequences. Appl Environ Microbiol 58: 2158–2163.
Holben WE, Feris KP, Kettunen A, Apajalahti JHA . (2004). GC fractionation enhances microbial community diversity assessment and detection of minority populations of bacteria by denaturing gradient gel electrophoresis. Appl Environ Microbiol 70: 2263–2270.
Jameson E, Joint I, Mann NH, Mühling M . (2007). Application of a novel rpoC1-RFLP approach reveals that marine Prochlorococcus populations in the Atlantic gyres are composed of greater microdiversity than previously described. Microb Ecol (in print). Published online, doi:10.1007/s00248-007-9259-5.
Lee SH, Malone C, Kemp PF . (1993). Use of 16S rRNA-targeted fluorescent probes to increase signal strength and measure cellular RNA from natural planktonic bacteria. Mar Ecol Prog Ser 101: 193–201.
Ludwig W, Strunk O, Westram R, Richter L, Meier H, Yadhukumar B et al. (2004). ARB: a software environment for sequence data. Nucleic Acid Res 32: 1363–1371.
Marchesi JR, Sato T, Weightman AJ, Martin TA, Fry JC, Hiom SJ et al. (1998). Design and evaluation of useful bacterium-specific PCR primers that amplify genes coding for bacterial 16S rRNA. Appl Environ Microbiol 64: 795–799.
Martín-Cuadrado A-B, López-García P, Alba J-C, Moreira D, Monticelli L, Strittmatter A et al. (2007). Metagenomics of the deep Mediterranean, a warm bathypelagic habitat. PLoS ONE 2: e914. Published online doi:10.1371/journal.pone.0000914.
McCaig AE, Embley TM, Prosser JI . (1994). Molecular analysis of enrichment cultures of marine ammonia oxidisers. FEMS Microbiol Lett 120: 363–367.
Mehling A, Wehmeier UF, Piepersberg W . (1995). Nucleotide sequences of streptomycete 16S ribosomal DNA—towards a specific identification system for streptomycetes using PCR. Microbiology 141: 2139–2147.
Miyamoto H, Yamamoto H, Arima K, Fujii J, Maruta K, Izu K et al. (1997). Development of a new semi-nested PCR method for detection of Legionella species and its application to surveillance of legionellae in hospital cooling tower water. Appl Environ Microbiol 63: 2489–2494.
Muyzer G, Brinkhoff T, Nübel U, Santegoeds C, Schäfer H, Waver C . (1998). Denaturing gradient gel electrophoresis (DGGE) in microbial ecology. In: Akkermans ADL, van Elsas JD, de Bruijn FJ (eds). Molecular Microbial Ecology Manual. Kluwer Academic Publishers: Dordrecht, The Netherlands. pp 1–27.
Muyzer G, De Waal EC, Uitterlinden AG . (1993). Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microbiol 59: 695–700.
Nübel U, Garcia-Pichel F, Muyzer G . (1997). PCR primers to amplify 16S rRNA genes from cyanobacteria. Appl Environ Microbiol 63: 3327–3332.
Pernthaler J, Amann R . (2005). Fate of heterotrophic microbes in pelagic habitats: focus on populations. Microbiol Mol Biol Rev 69: 440–461.
Rappé MS, Vergin K, Giovannoni SJ . (2000). Phylogenetic comparisons of a coastal bacterioplankton community with its counterparts in open ocean and freshwater systems. FEMS Microbiol Ecol 33: 219–232.
Robinson C, Poulton AJ, Holligan PM, Baker AR, Foster G, Gist N et al. (2006). The Atlantic Meridional Transect Programme (AMT): a contextual view 1995–2005. Deep Sea Res II 53: 1485–1515.
Scanlan DJ, West NJ . (2002). Molecular ecology of the marine cyanobacterial genera Prochlorococcus and Synechococcus. FEMS Microbiol Ecol 40: 1–12.
Sogin ML, Morrison HG, Huber JA, Welch DM, Huse SM, Neal PR et al. (2006). Microbial diversity in the deep sea and the underexplored ‘rare biosphere’. Proc Natl Acad Sci USA 103: 12115–12120.
Stach JEM, Maldonado LA, Ward AC, Goodfellow M, Bull AT . (2003). New primers for the class Actinobacteria: application to marine and terrestrial environments. Environ Microbiol 5: 828–841.
Stingl U, Desiderio RA, Cho J-C, Vergin KL, Giovannoni SJ . (2007). The SAR92 clade: an abundant coastal clade of culturable marine bacteria possessing proteorhodopsin. Appl Environ Microbiol 73: 2290–2296.
Stubner S . (2004). Quantification of Gram-negative sulphate-reducing bacteria in rice field soil by 16S rRNA gene-targeted real-time PCR. J Microbiol Methods 57: 219–230.
Venter JC, Remington K, Heidelberg JF, Halpern AL, Rusch D, Eisen JA et al. (2004). Environmental genome shotgun sequencing of the Sargasso Sea. Science 304: 66–74.
Vergin KL, Urbach E, Stein JL, DeLong EF, Lanoil BD, Giovannoni SJ . (1998). Screening of a fosmid library of marine environmental genomic DNA fragments reveals four clones related to members of the order Planctomycetales. Appl Environ Microbiol 64: 3075–3078.
Weisburg WG, Barns SM, Pelletier DA, Lane DJ . (1991). 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173: 697–703.
Widmer F, Seidler RJ, Gillevet PM, Watrud LS, Di Giovanni GD . (1998). A highly selective PCR protocol for detecting 16S rRNA genes of the genus Pseudomonas (sensu stricto) in environmental samples. Appl Environ Microbiol 64: 2545–2553.
Zaballos M, Lopez-Lopez A, Ovreas L, Bartual SG, D'Auria G, Alba JC et al. (2006). Comparison of prokaryotic diversity at offshore oceanic locations reveals a different microbiota in the Mediterranean Sea. FEMS Microbiol Ecol 56: 389–405.
Acknowledgements
This work is a core research activity of the Plymouth Marine Laboratory, a Collaborative Centre of the Natural Environment Research Council. Additional financial support came from the EU project ‘Microbial Marine Communities Diversity: from Culture to Function’ (MIRACLE) (EVK3-CT-2002-00087) and from an Natural Environment Research Council (NERC)-funded studentship (NER/S/A/2004/12171) allocated to JAW-A. We are grateful to Dr Hendrik Schäfer (Warwick University, UK) for useful discussion during this project. We also thank Ellie Jameson (PML) for the gift of the environmental DNA from the Northern Atlantic Gyre collected during the Atlantic Meridional Transect (AMT-15) cruise. The DNA sample from the mesocosm was collected during the ‘Pelagic Ecosystem CO2 Enrichment Study’ (PeECE; http://peece.ifm-geomar.de/) organized by Professor Ulf Riebesell (IFM-GEOMAR, Kiel, Germany) and carried out at the EU Large-Scale-Facility, University of Bergen, Norway.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website (http://www.nature.com/ismej)
Rights and permissions
About this article
Cite this article
Mühling, M., Woolven-Allen, J., Murrell, J. et al. Improved group-specific PCR primers for denaturing gradient gel electrophoresis analysis of the genetic diversity of complex microbial communities. ISME J 2, 379–392 (2008). https://doi.org/10.1038/ismej.2007.97
Received:
Revised:
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1038/ismej.2007.97
Keywords
This article is cited by
-
A bacterial formula with native strains as alternative to chemical fertiliser for tomato crop
Plant Growth Regulation (2024)
-
Exploring the status of global terrestrial and aquatic microbial diversity through ‘Biodiversity Informatics’
Environment, Development and Sustainability (2023)
-
Identification and characterization of chitinase-producing bacteria from gut of pleurostict scarab beetle grubs (Coleoptera: Scarabaeidae)
International Journal of Tropical Insect Science (2023)
-
Gut bacterial profile in Indian children of varying nutritional status: a comparative pilot study
European Journal of Nutrition (2021)
-
Bioactive carbon improves nitrogen fertiliser efficiency and ecological sustainability
Scientific Reports (2020)