Abstract
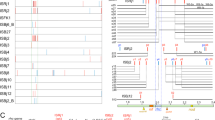
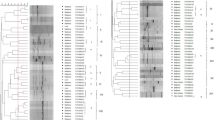
Comparative genomic hybridization (CGH) was performed with nine strains of Bradyrhizobium japonicum (a symbiotic nitrogen-fixing bacterium associated with soybean) and eight other members of the Bradyrhizobiaceae by DNA macroarray of B. japonicum USDA110. CGH clearly discriminated genomic variations in B. japonicum strains, but similar CGH patterns were observed in other members of the Bradyrhizobiaceae. The most variable regions were 14 genomic islands (4–97 kb) and low G+C regions on the USDA110 genome, some of which were missing in several strains of B. japonicum and other members of the Bradyrhizobiaceae. The CGH profiles of B. japonicum were classified into three genome types: 110, 122 and 6. Analysis of DNA sequences around the boundary regions showed that at least seven genomic islands were missing in genome type 122 as compared with type 110. Phylogenetic analysis for internal transcribed sequences revealed that strains belonging to genome types 110 and 122 formed separate clades. Thus genomic islands were horizontally inserted into the ancestor genome of type 110 after divergence of the type 110 and 122 strains. To search for functional relationships of variable genomic islands, we conducted linear models of the correlation between the existence of genomic regions and the parameters associated with symbiotic nitrogen fixation in soybean. Variable genomic regions including genomic islands were associated with the enhancement of symbiotic nitrogen fixation in B. japonicum USDA110.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
Accession codes
References
Akaike H . (1973). Information theory and an extension of the maximum likelihood principle. In: Petrov BN, Csaki F (eds). 2nd International Symposium on Information Theory. Akadimial Kiado: Budapest. pp 267–281.
Baginsky C, Brito B, Imperial J, Ruiz-Argüeso T, Palacios JM . (2005). Symbiotic hydrogenase activity in Bradyrhizobium sp. (Vigna) increases nitrogen content in Vigna unguiculata plants. Appl Environ Microbiol 71: 7536–7538.
Basit HA, Angle JS, Salem S, Gewaily EM, Kotob SI, van Berkum P . (1991). Phenotypic diversity among strains of Bradyrhizobium japonicum belonging to serogroup 110. Appl Environ Microbiol 57: 1570–1572.
Becker A, Berges H, Krol E, Bruand C, Ruberg S, Capela D et al. (2004). Global changes in gene expression in Sinorhizobium meliloti 1021 under microoxic and symbiotic conditions. Mol Plant Microbe Interact 17: 292–303.
Bontemps C, Golfier G, Gris-Liebe C, Carrere S, Talini L, Boivin-Masson C . (2005). Microarray-based detection and typing of the Rhizobium nodulation gene nodC: potential of DNA arrays to diagnose biological functions of interest. Appl Environ Microbiol 71: 8042–8048.
Brechenmacher L, Kim MY, Benitez M, Li M, Joshi T, Calla B et al. (2008). Transcription profiling of soybean nodulation by Bradyrhizobium japonicum. Mol Plant Microbe Interact 21: 631–645.
Brochet M, Couve E, Zouine M, Vallaeys T, Rusniok C, Lamy MC et al. (2006). Genomic diversity and evolution within the species Streptococcus agalactiae. Microbe Infect 8: 1227–1243.
Cardinale CJ, Washburn RS, Tadigotla VR, Brown LM, Gottesman ME, Nudler E . (2008). Termination factor Rho and its cofactors NusA and NusG silence foreign DNA in E. coli. Science 320: 935–938.
Carter B, Wu G, Woodward MJ, Anjum MF . (2008). A process for analysis of microarray comparative genomics hybridisation studies for bacterial genomes. BMC Genomics 9: 53.
Chang WS, Franck LW, Cytryn E, Jeong S, Joshi T, Emerich WD et al. (2007). An oligonucleotide microarray resource for transcriptional profiling of Bradyrhizobium japonicum. Mol Plant Microbe Interact 20: 1298–1307.
Ditta G, Virts E, Palomares A, Kin C . (1987). The nifA gene of Rhizobium meliloti is oxygen regulated. J Bacteriol 169: 3217–3223.
Dobrindt U, Agerer F, Michaelis K, Janka A, Buchrueser C, Samuelson M et al. (2003). Analysis of genome plasticity in pathogenic and commensal Escherichia coli isolates by use of DNA arrays. J Bacteriol 185: 1831–1840.
Dobrindt U, Hochhut B, Hentschel U, Hacker J . (2004). Genomic islands in pathogenic and environmental microorganisms. Nat Rev Microbiol 2: 414–424.
Dorman CJ . (2007). H-NS, the genome sentinel. Nat Rev Microbiol 5: 157–161.
Giraud E, Moulin L, Vallenet D, Barbe V, Cytryn E, Avarre J et al. (2007). Legumes symbioses: absence of nod genes in photosynthetic bradyrhizobia. Science 316: 1307–1312.
Giuntini E, Mengoni A, Filippo CD, Cavalieri D, Aubin-Horth N, Landry CR et al. (2005). Large-scale genetic variation of the symbiosis-requiered megaplasmid pSymA revealed by comparative genomic analysis of Sinorhizobium meliloti natural strains. BMC Genomics 6: 158.
Gupta SR, Mok A . (2007). Phylogenomics and signature proteins for the alpha proteobacteria and its main groups. BMC Microbiol 7: 106.
Hartmann A, Catroux G, Amarger N . (1992). Bradyrhizobium japonicum strains identification by RFLP analysis using the repeated sequence RSα. Lett Appl Microbiol 15: 15–19.
Hinchliffe SJ, Isherwood KE, Stabler RA, Prentice MB, Rakin A, Nichols RA et al. (2003). Application of DNA microarrays to study the evolutionary genomics of Yersinia pestis and Yersinia pseudotuberculosis. Genome Res 13: 2018–2029.
Ihaka R, Gentleman R . (1996). R: a language for data analysis and graphics. J Comput Graph Stat 5: 299–314.
Ishii S, Sadowsky MJ . (2008). Escherichia coli in the environment: implications for water quality and human health. Microbes Environ 23: 101–108.
Ishizuka J, Suemasu Y, Mizogami K . (1991). Preference of Rj-soybean cultivars for Bradyrhizobium japonicum for nodulation. Soil Sci Plant Nutr 37: 15–21.
Israel DW, Mathis JN, Barbour WM, Elkan GH . (1986). Symbiotic effectiveness and host-strain interactions of Rhizobium fredii USDA 191 on different soybean cultivars. Appl Environ Microbiol 51: 898–903.
Ito N, Itakura M, Eda S, Saeki K, Oomori H, Yokoyama T et al. (2006). Global gene expression in Bradyrhizobium japonicum cultured with vanillin, vanillate, 4-hydroxybenzoate and protocatechuate. Microbes Environ 21: 240–250.
Jordan DC . (1984). Rhizobiaceae. In: Krieg NR, Holt JG (eds). Bergey's Manual of Systematic Bacteriology. Williams & Wilkins: Baltimore. pp 235–244.
Jordan IK, Koonin EV . (2004). Horizontal gene transfer and prokaryotic genome evolution. In: Miller RV, Day MJ (eds). Microbial Evolution. Gene Establishment, Survival, and Exchange. ASM Press: Washington, DC, pp 319–338.
Kamagata Y, Fulthorpe RR, Tamura K, Takami H, Forney LJ, TieJe JM . (1997). Pristine environments harbor a new group of oligotrophic 2,4-dichlorophenoxyacetic acid-degrading bacteria. Appl Environ Microbiol 63: 2266–2272.
Kaneko T, Nakamura Y, Sato S, Minamisawa K, Uchiumi T, Sasamoto S et al. (2002). Complete genomic sequence of nitrogen-fixing symbiotic bacterium Bradyrhizobium japonicum USDA110. DNA Res 9: 225–256.
King G . (2007). Chemolithotrophic bacteria: distribution, functions and significance in volcanic environments. Microbes Environ 22: 309–319.
Koide T, Zaini PA, Moreira LM, Vencio RZN, Matsukuma AY, Durham AM et al. (2004). DNA microarray-based genome comparison of a pathogenic and a nonpathogenic strain of Xylella fastidiosa delineates genes important for bacterial virulence. J Bacteriol 186: 5442–5449.
Koyama O, Manome A, Okubo M, Yokomaku T, Tanaka H . (2007). Necrosis gene-based monitoring and control of potato scab disease. Microbes Environ 22: 123–127.
Larimer W, Chain FP, Hauser L, Lamerdin J, Malfatti S, Do L et al. (2004). Complete genome sequence of the metabolically versatile photosynthetic bacterium Rhodopseudomonas palustris. Nat Biotechnol 22: 55–61.
Leonard LT . (1943). A simple assembly for use in the testing of cultures of rhizobia. J Bacteriol 45: 523–525.
MacLean AM, Finan TM, Sadowsky MJ . (2007). Genomes of the symbiotic nitrogen-fixing bacteria of legumes. Plant Physiol 144: 615–622.
Maier RJ, Triplett EW . (1996). Toward more productive, efficient, and competitive nitrogen-fixing symbiotic bacteria. Crit Rev Plant Sci 15: 191–234.
Manning SD, Motiwala AS, Sprigman AC, Qi W, Lacher DW, Oullette LM et al. (2008). Variation in virulence among clades of Escherichia coli O157:H7 associated with disease outbreaks. Proc Natl Acad Sci USA 105: 4868–4873.
Masuda T, Fujita K, Kogure K, Ogata S . (1989). Effect of CO2 enrichment and nitrate application on vegetative growth and dinitrogen fixation of wild and cultivated soybean varieties. Soil Sci Plant Nutr 35: 357–366.
Mauchline TH, Fowler JE, East AK, Sartor AL, Zaheer R, Hosie AHF et al. (2006). Mapping the Sinorhizobium meliloti 1021 solute-binding protein-dependent transportome. Proc Natl Acad Sci USA 47: 17933–17938.
Minamisawa K, Itakura M, Suzuki M, Ichige K, Isawa T, Yuhashi K et al. (2002). Horizontal transfer of nodulation genes in soil and microcosms from Bradyrhizobium japonicum to B. elkanii. Microbes Environ 17: 82–90.
Minamisawa K, Nakatsuka Y, Isawa T . (1999). Diversity and field site variation of indigenous populations of soybean bradyrhizobia in Japan by fingerprints with repeated sequence RSα and RSβ. FEMS Microbiol Ecol 29: 171–178.
Molouba F, Lorquin J, Willems A, Hoste B, Giraud E, Dreyfus B et al. (1999). Photosynthetic bradyrhizobia from Aeschynomene spp. are specific to stem-nodulated species and form a separate 16S ribosomal DNA restriction fragment length polymorphism group. Appl Environ Microbiol 65: 3084–3094.
Nautiyal CS, Hegde SV, van Berkum P . (1988). Nodulation, nitrogen fixation, and hydrogen oxidation by pigeon pea Bradyrhizobium spp. in symbiotic association with pigeon pea, cowpea, and soybean. Appl Environ Microbiol 54: 94–97.
Nieuwkoop AJ, Banfalvi Z, Deshmane N, Garhold D, Schall MG, Sirotkin KM et al. (1987). A locus encoding host range is linked to the common nodulation genes of Bradyrhizobium japonicum. J Bacteriol 169: 2631–2638.
Obert C, Sublett J, Kaushal D, Hinojosa E, Barton T, Tuomanen EI et al. (2006). Identification of a candidate Streptococcus pneumoniae core genome and regions of diversity correlated with invasive pneumococcal disease. Infect Immun 74: 4766–4777.
Pearson BM, Pin C, Wright J, I'Anson K, Humphrey T, Wells JM . (2003). Comparative genome analysis of Campylobacter jejuni using whole genome microarrays. FEBS Lett 554: 224–230.
Pessi G, Ahrens HC, Rehrauer H, Lindemann A, Hauser F, Fischer HM et al. (2007). Genome-wide transcript analysis of Bradyrhizobium japonicum bacteroids in soybean root nodules. Mol Plant Microbe Interact 20: 1353–1363.
Sadowsky MJ, Cregan PB, Gottfert M, Sharma A, Gerhold D, Rodriguez-Quinones F et al. (1991). The Bradyrhizobium japonicum nolA gene and its involvement in the genotype-specific nodulation of soybeans. Proc Natl Acad Sci USA 88: 637–641.
Saeki Y, Aimi N, Tsukamoto S, Yamakawa T, Nagamoto Y, Akao S . (2006). Diversity and geographical distribution of indigenous soybean-nodulating bradyrhizobia in Japan. Soil Sci Plant Nutr 52: 418–426.
Saeki Y, Kaneko A, Hara T, Suzuki K, Yamakawa T, Nguyen MT et al. (2005). Phylogenetic analysis of soybean-nodulating rhizobia isolated from alkaline soils in Vietnam. Soil Sci Plant Nutr 51: 1043–1052.
Saito A, Kawahara M, Ikeda S, Ishimine M, Akao S, Minamisawa K . (2008). Broad distribution and phylogeny of anaerobic endophytes of cluster XIVa clostridia in plant species, including crops. Microbes Environ 23: 73–80.
Saito A, Mitsui H, Hattori R, Minamisawa K, Hattori T . (1998). Slow-growing and oligtrophic soil bacteria phylogenetically close to Bradyrhizobium japonicum. FEMS Microbiol Ecol 25: 277–286.
Sameshima R, Isawa T, Sadowsky MJ, Hamada T, Kasai H, Shutsrirung A et al. (2003). Phylogeny and distribution of extra-slow-growing Bradyrhizobium japonicum harboring high copy numbers of RSα, RSβ and IS1631. FEMS Microbiol Ecol 44: 191–202.
Sameshima-Saito R, Chiba K, Minamisawa K . (2006). Correlation of denitrifying capability with the existence of nap, nir, nor and nos genes in diverse strains of soybean bradyrhizobia. Microbes Environ 21: 174–184.
Starkenburg RS, Chain SGP, Sayavedra-Soto AL, Hauser L, Land LM, Larimer WF et al. (2006). Genome sequence of the chemolithoautotrophic nitrite-oxidizing bacterium Nitrobacter winogradskyi Nb-255. Appl Environ Microbiol 72: 2050–2063.
Trung BC, Yoshida S . (1983). Improvement of Leonard jar assembly for screening of effective Rhizobium. Soil Sci Plant Nutr 29: 97–100.
Uchiumi T, Ohwada T, Itakura M, Mitsui H, Nukui N, Dawadi P et al. (2004). Expression islands clustered on the symbiosis island of the Mesorhizobium loti genome. J Bacteriol 186: 2439–2448.
van Berkum P . (1990). Evidence for a third uptake hydrogenase phenotype among the soybean bradyrhizobia. Appl Environ Microbiol 56: 3835–3841.
van Berkum P, Fuhrmann JJ . (2000). Evolutionary relationships among the soybean bradyrhizobia reconstructed from 16S rRNA gene and internally transcribed spacer region sequence divergence. Int J Syst Evol Microbiol 50: 2165–2172.
Wei M, Yokoyama T, Minamisawa K, Mitsui H, Itakura M, Kaneko T et al. (2008). Soybean seed extracts preferentially express genomic loci of Bradyrhizobium japonicum in the initial interaction with soybean Glycine max (L.) Merr. DNA Research advance online publication 29 May 2008. DNA Research advance online publication 29 May 2008; doi:10.1093/dnares/dsn012.
Zhou D, Han Y, Song Y, Tong Z, Wang J, Guo Z et al. (2004). DNA microarray analysis of genome dynamics in Yersinia pestis: insights into bacterial genome microevolution and niche adaptation. J Bacteriol 186: 5138–5146.
Acknowledgements
This study was supported in part by a grant-in-aid for Scientific Research on Priority Area ‘Comparative Genomics’, by grants-in-aid for Scientific Research (no. 17380046), by a grant from PROBRAIN, by Special Coordination Funds for Promoting Science and Technology and by a grant from the Tokachi Federation of Agricultural Cooperatives. We thank H Kouchi (National Institute of Agrobiological Sciences), C Harwood (The University of Iowa) and E Giraud (French National Institute for Agricultural Research) for array spotting, provision of R. palustris CGA009 and provision of Bradyrhizobium sp. strains ORS278 and BTAi1, respectively.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website (http://www.nature.com/ismej)
Rights and permissions
About this article
Cite this article
Itakura, M., Saeki, K., Omori, H. et al. Genomic comparison of Bradyrhizobium japonicum strains with different symbiotic nitrogen-fixing capabilities and other Bradyrhizobiaceae members. ISME J 3, 326–339 (2009). https://doi.org/10.1038/ismej.2008.88
Received:
Revised:
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1038/ismej.2008.88
Keywords
This article is cited by
-
Genome mining, antimicrobial and plant growth-promoting potentials of halotolerant Bacillus paralicheniformis ES-1 isolated from salt mine
Molecular Genetics and Genomics (2023)
-
Evolution of rhizobial symbiosis islands through insertion sequence-mediated deletion and duplication
The ISME Journal (2022)
-
Transposon sequencing analysis of Bradyrhizobium diazoefficiens 110spc4
Scientific Reports (2021)
-
Bradyrhizobium japonicum, B. elkanii and B. diazoefficiens Interact with Rice (Oryza sativa), Promote Growth and Increase Yield
Current Microbiology (2021)
-
Tapping into the maize root microbiome to identify bacteria that promote growth under chilling conditions
Microbiome (2020)